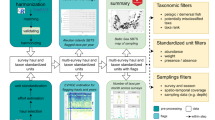
Abstract
Recognition of the threats to biodiversity and its importance to society has led to calls for globally coordinated sampling of trends in marine ecosystems. As a step to defining such efforts, we review current methods of collecting and managing marine biodiversity data. A fundamental component of marine biodiversity is knowing what, where, and when species are present. However, monitoring methods are invariably biased in what taxa, ecological guilds, and body sizes they collect. In addition, the data need to be placed, and/or mapped, into an environmental context. Thus a suite of methods will be needed to encompass representative components of biodiversity in an ecosystem. Some sampling methods can damage habitat and kill species, including unnecessary bycatch. Less destructive alternatives are preferable, especially in conservation areas, such as photography, hydrophones, tagging, acoustics, artificial substrata, light-traps, hook and line, and live-traps. Here we highlight examples of operational international sampling programmes and data management infrastructures, notably the Continuous Plankton Recorder, Reef Life Survey, and detection of Harmful Algal Blooms and MarineGEO. Data management infrastructures include the World Register of Marine Species for species nomenclature and attributes, the Ocean Biogeographic Information System for distribution data, Marine Regions for maps, and Global Marine Environmental Datasets for global environmental data. Existing national sampling programmes, such as fishery trawl surveys and intertidal surveys, may provide a global perspective if their data can be integrated to provide useful information. Less utilised and emerging sampling methods, such as artificial substrata, light-traps, microfossils and eDNA also hold promise for sampling the less studied components of biodiversity. All of these initiatives need to develop international standards and protocols, and long-term plans for their governance and support.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
6.1 Introduction
Current concerns about the Earth’s ecosystems and the loss of biodiversity drives the need to measure spatial and temporal variation in biodiversity from local to global scales (Costello 2001; Andréfouët et al. 2008a; Ash et al. 2009). In the ocean, over-fishing and other threats to species’ populations reduce resources for society, have altered ecosystems, and put many mammals, birds, reptiles, and fish in danger of extinction (e.g., Costello and Baker 2011; Hiscock 2014; Costello 2015; Webb and Mindel 2015). Global and regional scale assessments need data that are either collected by similar methods and procedures, or produce variables that can be integrated for analyses (Pereira et al. 2013). For example the EU Marine Strategy Framework Directive (MSFD) requires extensive measures of biodiversity and ecosystem functioning to monitor the health of European marine waters and to guide measures that ensure that they achieve a Good Environmental Status by 2021 (Boero et al. 2015). The World Ocean Assessment will emphasise the need for more standardised reporting of information (Inniss et al. 2016). To that end, variables that are ‘essential’ for the monitoring of biodiversity and understanding ecosystem change are being developed (Box 6.1). As yet, how to measure these variables, and manage and analyse the data, has not been elaborated. Here, we review methods used for field observations and sampling marine biodiversity, provide examples of methods and operational global monitoring programmes, and how data systems have emerged to assist in data publication and analysis. It cannot be assumed that established or popular methods are the most cost-effective and suitable for monitoring biodiversity. Thus we outline the potential of less prominent methods as well as those considered more conventional. This synthesis thus provides an introduction to how marine biodiversity may be monitored and assessed into the future.
Box 6.1. Essential Ocean Variables (EOVs)
Under the leadership of the Intergovernmental Oceanographic Commission (IOC) of UNESCO, the Global Ocean Observing System (GOOS) has proposed to develop an integrated framework for sustained ocean observing based on Essential Ocean Variables (EOVs). An EOV, should have by definition, a high impact in responding to scientific and societal issues and a high feasibility of sustained observation. These will include biogeochemical and biological variables (ecosystem EOVs), to help understand marine ecosystems, in addition to the existing physical ocean variables. At the same time GEO BON has been developing the Essential Biodiversity Variables (Pereira et al. 2013). GOOS in collaboration with GEO BON, has established the GOOS Panel on Biology and Ecosystems (GOOS BioEco), which is responsible for the development and assessment of ecosystem EOVs. This includes documentation, best practice, readiness, implementation strategies, coordination of activities, and fitness-of-purpose of data and information streams resulting from observations to improve their recommendations to policy-making. GOOS BioEco is also considering societal needs and human pressures affecting marine biodiversity and ecosystems to identify the EOVs. The first GOOS Biology technical expert workshop in Townsville, Australia in November 2013, resulted in a preliminary list of 42 candidate ecosystem EOVs. From these, 10 were selected for high impact and feasibility within four major areas identified as key for a healthy and productive ocean: (1) Productivity, (2) Biodiversity, (3) Ecosystem Services, and (4) Human activities and pressures. Some of the candidate EOVs that meet these requirements were chlorophyll, harmful algal blooms (HAB), zooplankton biomass and abundance, and the extent and live cover of marine communities such as coral reefs, mangroves, seagrasses, and salt marshes.
6.2 Sampling Methods
An impressive variety of methods have been used to sample marine species, including observations, nets, hooks, traps, grabs, sediment collection, sound, chemicals and electricity (Table 6.1) (e.g., Santhanam and Srinivasan 1994; Kingsford and Battershill 1998; Tait and Dipper 1998; Elliott and Hemingway 2002; Eleftheriou 2013; Hiscock 2014). All methods are selective, at least for body size by excluding smaller and/or larger organisms. Such bias should be explicitly recognised in the design and interpretation of field data. Because of methodological biases a comprehensive sampling of marine biodiversity across habitats, body sizes and trophic levels would need to use a variety of complementary methods. Such a suite of methods can produce an inventory of species present that reflect the environment, habitats, and ecology of an area.
A species inventory provides the evidence of what species are present and an estimate of species richness. Knowing which species are present is essential to distinguish those that are of socio-economic or ecological importance, endemic, threatened with extinction, introduced, or considered pests (McGeoch et al. 2016). Indeed, species richness is by far the most common measure of ‘diversity’ used in science and conservation management (Gotelli and Colwell 2001; Costello et al. 2004).
For microbes, species identification can be impractical and so ad hoc ‘metagenomic’ and ‘barcoding’ style guidance on molecular ‘Operational Taxonomic Units’ (OTU) are used as indicators of diversity. However, OTU are not standardised between studies, and values vary due to different resolution of the genes analysed for different taxa. For some taxa they may indicate genus level and others population level differences. For bacteria, the species concept used for eukaryotes is doubtfully applicable, and while they have high genetic diversity, the number of formally named ‘species’ is relatively low (Costello et al. 2013a, b). Thus, while an indicator of genetic diversity, OTU should not be equated with ‘species’.
Various methods and metrics have been used to characterise the relative abundance of species, including numbers of individuals, areal cover, and/or biomass within samples (Hiscock 2014). Assessments of measures of biodiversity thus need to consider that every sampling method is biased, and that different methods are required to sample different components of biodiversity. Thus quantitative sampling is best focused on measuring dynamics of particular species populations rather than measuring biodiversity across species. Instead, the relative abundance of species may be compared on semi-quantitative (e.g., log 10) abundance scales (e.g., Davies et al. 2001; Haegeman et al. 2013; Hiscock 2014). The more abundant and/or conspicuous species define communities and biotopes and indicate how an ecosystem functions in terms of habitat, productivity, and food-webs. Changes in the identity of the dominant species can indicate changes in the community present in space and time, and thus changes to the ecosystem. However, often the ecosystem effects of species are unrelated to their abundance or body size. For example, top-predators are typically low in abundance and density but large in body size. Thus a range of species of different guilds and body sizes should be sampled to monitor ecosystems.
In addition to the bias of how samples are taken, results will depend on when and where sampling takes place. The design of field surveys thus needs to be clear which habitats, body sizes and taxa it has focused on, and what has been excluded; i.e., how it has ‘stratified’ sampling. Perhaps the most effective way to place the data into an environmental context is to map the geographic distribution of environmental variables (e.g., depth, salinity, temperature, substratum, topography) and habitats (Costello 1992; Costello and Emblow 2005; Costello et al. 2005, 2010a; Hiscock 2014). These environmental variables can be mapped through ‘remote sensing’ from satellites, aircraft and ships (Andréfouët et al. 2008b, 2011) and can include: seabed depth, topography, and roughness; surface water colour (an estimate of phytoplankton biomass and dominance) and temperature; depth-profiles of density (salinity) and temperature; acoustic signatures of zooplankton and pelagic megafauna; and the distribution and extent of intertidal and shallow-water habitats such as coral reefs, kelp and seagrass beds, mangrove forests, and salt-marshes. As the technology improves and cost reduces, it is likely that ‘remotely operated’ and ‘autonomous’ vehicles (ROV, AUV) will become more commonly used for underwater and aerial surveillance. The potential of sound signatures in the marine environment as indicators of biodiversity is also being researched (Harris et al. 2015). Although sensors borne on satellites and aircraft may have limited ability to identify species they provide an invaluable environmental context for biodiversity, and may indicate global large-scale patterns in biodiversity (De Monte et al. 2013). They thus complement in situ observations and enable mapping of habitats and biotopes (e.g., Neilson and Costello 1999; Connor et al. 2006; Leleu et al. 2012; Remy-Zephir et al. 2012; Hiscock 2014). Other methods may identify species from images, such as video and still photography (Table 6.1). Techniques for unsupervised image processing continue to improve and may lead to an increased use of automated image systems for large and microscopic species. Crowd-sourcing is also increasingly assisting the digitisation of large ecological image libraries (Edgar et al. 2016).
6.2.1 Bottom Trawl Surveys
In many countries bottom trawl surveys are used for monitoring commercially important fish stocks. Although originally designed to provide fisheries independent information for fish stock assessment and management, they are now increasingly being used to analyse trends in the abundance, distribution and diversity of both commercial and non-commercial species of fish and epibenthos (e.g., Bianchi et al. 2000; Shackell and Frank 2003; Daan et al. 2005; Perry et al. 2005; Atkinson et al. 2011).
Bottom trawls come in different designs suited for catching fish on different types of seabed. Beam trawls use a horizontal metal beam to keep the mouth of the trawl open, and target flatfish and other near-bottom species. They sometimes have ‘tickler chains’ attached to the front bottom part of the gear to scare shrimps or flatfish up from the seabed and into the net. Otter trawls use otter boards (trawl doors) attached to the trawl net by wires to keep the mouth of the net horizontally open. Over fine grained sediments the otter boards generate clouds of suspended material on each side of the trawl net which helps to herd the fish into the mouth of the trawl. Often wings of netting are attached to both sides of the trawl mouth to increase the herding effect further. Vertically the mouth of an otter trawl is held open by floats and by a footrope to which weights, rollers or bobbins are attached. These vary from small rubber discs used on sandy or muddy bottoms to large metal balls that can roll over rocks or larger stones and prevent the footrope from becoming snagged on rougher and harder grounds. The body of the trawl is funnel-shaped and narrows from the mouth towards the cod end where the fish accumulate during the tow. It is the mesh size of the cod end that determines the size of the fish that are retained. In commercial trawl gears minimum mesh size regulations are often used to reduce the catch of juvenile undersized fish. However, in research surveys the mesh size in the cod end is usually small enough to ensure that the smaller species and individuals are retained. Pelagic trawls target fish such as anchovies, mackerels, and sardines in the water column.
The catch efficiency of a bottom trawl is defined as the proportion of the fish in the area swept by the gear that is retained in the cod end. The area swept equals the length of the tow multiplied by the width of the gear, where the latter often is assumed to correspond either to the spread of the wings or to the distance between the otter boards during fishing to account for the herding effect of the boards and bridles. However, the catch efficiency is influenced by a multitude of factors including the escape behaviour of the fish species, properties of the gear, and the fishing operation (Benoít and Swain 2003; Fraser et al. 2007, 2008; Queirolo et al. 2012; Weinberg and Kotwicki 2008; Winger et al. 2010; Sistiaga et al. 2015). Fish may escape by burrowing in the seabed, by swimming under the footrope, by escaping over the head-rope of the gear, or by passing through the meshes in the front part of the trawl. The size of the vertical and horizontal opening is often monitored during the tow by sensors attached to the gear and has been found to depend on the warp length and towing speed as well as the weight of the catch accumulating in the cod end. During fishing, fish accumulate in the mouth of the trawl where they try to keep pace with the gear. As individuals tire they fall back towards the cod end. How fast a fish will get tired, and whether it can outswim the gear is species and size dependent. The amount caught per area swept may also depend on the time of day because this can influence how close to the seabed the fish are found (Kotwicki et al. 2009). To ensure that catch rates can be compared across years, much is therefore done to standardise the trawling operation, the gear and the procedures for sampling and for analysing the catch (e.g., Miller 2013).
In some parts of the world standardised bottom trawl surveys have now been conducted for more than 50 years and some of the resulting data are publicly available or available upon request. ICES provides online access to a database with trawl survey data from the north eastern Atlantic (www.ices.dk/marine-data/data-portals/Pages/DATRAS.aspx) and similar databases are available for other areas such as the Eastern Bering Sea and Gulf of Alaska (www.afsc.noaa.gov/RACE/groundfish/survey_data). Additional data can be downloaded from international data portals such as OBIS (Table 6.2), but much data still reside in the custody of national fisheries research institutions. These data constitute a so far underutilised source of information on the distribution, abundance, and diversity of marine fishes on the world’s continental shelves.
6.2.2 Light Traps
Light traps are commonly used for collecting insects as a means for monitoring pest species. The American Center for Disease Control has had a standardised light trap for mosquito monitoring for over 50 years (Sudia and Chamberlain 1988). Moths, beetles and other crop pests are also commonly surveyed this way (Szentkiralyi 2002). However, light traps have a shorter history of use in the aquatic environment. The earliest uses were in freshwater for capturing insects and they were soon found to be excellent for collecting young fish (Hungerford et al. 1955) and zooplankton (Meekan et al. 2001; Øresland 2007), but also collect many benthic species that emerge from the benthos at night. They have been used extensively around coral reefs where the structural complexity of the reef system makes other methods susceptible to damage (Doherty 1987). There can be species, gender and ontogenetic specific responses to light traps making them more useful for some organisms than others. Species may vary in their abundance at different times of the night and lunar cycle. A benefit of light-trapping is that the animals are not harmed during collection, and have thus proved useful for sampling of museum specimens and laboratory animals (Doherty 1987; Holmes and O’Connor 1988). However, light trap catches may not work well in areas of high current or excessive turbidity. The potential of light traps for monitoring mobile benthic and demersal organisms, mostly crustaceans, has yet to be adequately explored. This ‘fish food’ component of biodiversity forms an important trophic link in many ecosystems, and has been overlooked in marine biodiversity monitoring.
6.2.3 Artificial Substrata
A problem in sampling the natural environment is that it is variable at every spatial scale, and thus the abundance of species sampled varies because of micro-habitat variation as well as changes in species abundance in space and over time. Advantages of artificial substrata are that they provide a standard replicable physical habitat and thus low variation between replicate samples. In addition their use avoids damage to natural habitat, and they can be low cost, amenable to experimental manipulation, easily deployed and retrieved, and rapidly processed (reviewed in Costello and Thrush 1991). Because the date and duration of deployment of artificial substrata is known their community can also be standardised for successional age. They can be hard panels, balls of plastic mesh, sediment trays, and made of a variety of materials. They can also capture species otherwise difficult to sample, such as mobile epifaunal macroinvertebrates that nestle into plastic mesh. Species composition and community structure has been found to be similar and comparable to natural substrata (Costello and Myers 1996). Artificial substrata have been long used in freshwater environments as a standard method of monitoring biodiversity, especially in large rivers and lakes where other methods may be difficult (APHA et al. 2007). They have had widespread use in experiments in the marine environment, such as looking at colonisation, succession, competition, and community stability on plastic mesh (e.g., Costello and Myers 1996) and flat panels (e.g., Atalah et al. 2007a, b; Wahl et al. 2011). Recently, hundreds of Autonomous Reef Monitoring Structures (ARMS) have become deployed on coral reefs and other habitats around the world (e.g., http://www.pifsc.noaa.gov/cred/arms.php; Leray and Knowlton 2015). ARMS are a stack of hard plastic plates that capture crevice living invertebrates otherwise difficult to collect without damaging reefs. Artificial substrata merit wider use in marine biodiversity monitoring considering their benefits of standardisation and lack of damage to natural habitat.
6.2.4 Microfossils
Microfossils are microscopic sized organisms that have hard parts with high fossilisation potential (e.g., calcareous or siliceous shells), including foraminiferans, ostracods, diatoms, radiolarians and coccolithophores, or are microscopic sized hard parts of larger organisms, including ichthyoliths. Microfossils can be a proxy for biodiversity patterns across a broader range of organisms, because they have excellent fossil records, occupy a wide range of ecological niches, and are abundant even in a small amount of sediment. Marine sediment cores available from almost the entire ocean through national and international drilling projects (e.g., International Ocean Discovery Program; IODP) include abundant microfossils and provide long-term continuous time-series sedimentary records at decadal, centennial, millennial, and multi-millennial time scales covering the entire Cenozoic Era. Thus microfossils in sediment cores are an archive that enables reconstruction of long-term time-series beyond the temporal coverage of recent biological monitoring (Yasuhara et al. 2015).
Sample procedures involve physical and chemical treatments of sediment subsamples to disaggregate consolidated sediment, clean up microfossils, concentrate specimens and remove extraneous material, for example, by freeze drying, hydrogen peroxide treatment, wet sieving, centrifugation, and acid treatment. The resulting sample can be mounted on a glass slide (e.g., for diatoms, radiolarians and coccolithophores) or manually picked from treated material onto a paper slide (e.g., for foraminiferans, ostracods and ichthyoliths) for counting under stereo and compound microscopes respectively.
For example, North Atlantic deep-sea ostracod diversity has been found to track global climate change for the last 500,000 years, being less during glacial and high during interglacial periods (Yasuhara et al. 2009). Climatic control of deep-sea ostracod diversity has also been shown for shorter, decadal-centennial time scales (Yasuhara et al. 2008). Latitudinal species diversity gradients of deep-sea ostracods in the North Atlantic Ocean were distinct during interglacials (including present day) but indistinct or collapsed during glacials (Yasuhara et al. 2009). These deep-sea diversity patterns in space and time in North Atlantic microfossil records are explained by temperature control of deep-sea biodiversity (Hunt et al. 2005; Yasuhara and Cronin 2008; Yasuhara et al. 2009, 2014). Further applications of microfossils as a model system for biodiversity research are found in Yasuhara et al. (2015).
6.2.5 Molecular Observations of Microbial Communities
Genomic analysis of marine microbes has become common both at the local (marine stations, localised cruises) and at the global scale. After the Global Ocean Sampling expedition (Venter et al. 2004) proved that high-throughput molecular approaches were able to reveal an unprecedented diversity of bacterial sequences, several other programs have quantified the molecular diversity and biogeography of planktonic communities. The Tara Oceans missions (http://oceans.taraexpeditions.org/en) have sampled coastal and open oceans worldwide (Bork et al. 2015), including eddies, upwellings, oxygen-minimum zones, coral reefs, regions of natural iron fertilisation, and lately the Arctic and Mediterranean regions. These missions are uncovering marine planktonic communities from viruses to protists, up to metazoan larvae. The Malaspina project (http://scientific.expedicionmalaspina.es) complements these observations with samples of the deep seas at the global scale, and ‘Ocean Sampling Day’ with about 150 stations globally sampled on the same day (Kopf et al. 2015). An increasing number of cruises include molecular high-throughput analyses of genes, transcripts, and metabolites of planktonic organisms, together with environmental variables such as physical and biochemical parameters (e.g., Atlantic Meridional Transect http://www.amt-uk.org).
6.3 Case Studies
6.3.1 The Continuous Plankton Recorder (CPR)
The Continuous Plankton Reorder (CPR) survey is the longest sustained and geographically most extensive marine biological survey in the world, covering ~1000 taxa over multi-decadal periods since 1931 (Edwards et al. 2010). It samples phytoplankton and zooplankton in oceans and shelf seas using ships of opportunity from ~30 different shipping companies, at monthly intervals on ~50 trans-ocean routes. In this way the survey autonomously collects biological and physical data from ships covering ~20,000 km of the ocean per month, ranging from the Arctic to the Southern Ocean. The survey is operated by the Sir Alister Hardy Foundation for Ocean Science (SAHFOS), an internationally funded charity with a wide consortium of stakeholders. Since the first tow of a CPR more than 6 million nautical miles of sea have been sampled and over 100 million data entries have been recorded. Plankton are collected on a band of silk and subsequently visually identified by experts. Additionally, over the last decade the CPRs have been equipped with modern chemical and physical sensors as well as molecular probes. The database and sample archive together provide a resource that can be utilised in a wide range of environmental, ecological and fisheries related research, e.g., molecular analyses of marine pathogens, modelling for forecasting and data for incorporation in new approaches to ecosystem and fishery management.
In 2011 SAHFOS, along with 12 other research organisations using the CPR from around the world formed a Global Alliance of CPR surveys (GACs) with the aim of developing new surveys and a global database, and producing a global ocean status report (Edwards et al. 2012). This global network of CPR surveys now routinely monitors the North Sea, North Atlantic, Arctic, North Pacific and Southern Ocean. New surveys are underway in Australian, New Zealand, Japanese and South African waters with a Brazilian and an Indian Ocean survey under development. These surveys provide coverage of large parts of the world’s oceans but many gaps still exist particularly in the South Atlantic, Indian and Pacific Oceans. This global network also brings together the expertise of approximately 60 plankton specialists, scientists and technicians from 14 laboratories around the world. Working together, centralising the database and working in close partnership with the maritime shipping industry, this global network of CPR surveys with its low costs and new technologies makes the CPR an ideal tool for an expanded and comprehensive marine biological sampling programme.
6.3.2 Tropical Coral Reefs
Monitoring of tropical shallow reefs is conducted with near-global coverage using methods described by English et al. (1994). Considerable effort has been invested in comparing the accuracy and agreement among different methods (e.g., Leujak and Ormond 2007; Facon et al. 2016). The emerging consensus is to focus on the output variables from monitoring, rather than the methods: e.g., proportional cover for sessile taxa, abundance or density per unit area for mobile taxa and biomass, particularly for fishes. This is consistent with emerging guidance on observation and indicator systems (UNESCO 2012).
The principal framework for aggregating coral reef data to global levels has been the Global Coral Reef Monitoring Network (GCRMN) of the International Coral Reef Initiative (ICRI), which was initiated in 1995. The establishment of the GCRMN coincided with the largest global impact to reefs ever recorded, the 1997–98 El Niño event, giving strong impetus for global reporting for a decade. However funding for this level of reporting has been difficult to sustain, forcing the GCRMN to focus on regional level reporting, such as in the Caribbean (Jackson et al. 2014) and currently underway in the Western Indian Ocean. The GCRMN regions closely match those of the UNEP Regional Seas programmes, and inform countries regarding fisheries and food security. The GCRMN provides guidance for three levels of monitoring effort: citizen volunteer-focused, ‘intermediate’ and ‘expert’ (Wilkinson and Hill 2004). The challenges across these levels include data reliability and quality, replication and representation, and taxonomy, the latter exacerbated by the high diversity of coral reef taxa. The intermediate level of monitoring is most frequently applied and is implemented through technical staff (e.g., marine rangers), students and experienced volunteers, and focused on functional group or genus-level identifications for principal benthic taxa (e.g., hard corals, algae) and family or genus level identification for fish. The basic sampling unit recommended by the GCRMN has been line transects or photoquadrats for benthic cover, 50 m belt transects (2 or 5 m width) for fish and narrower belt transects or quadrats for mobile invertebrates. The configuration of these samples varies greatly among programmes. Expert-level monitoring has been the domain of professional researchers, often with genus-level identification for corals and species-level identification for fish. Due to the popularity of coral reefs for SCUBA diving, sampling by volunteers has been feasible, with the most widespread methods being those of Reef Check (Hodgson 1999), REEF (Francisco-Ramos and Arias-González 2013), and the Reef Life Survey (see below). In volunteer programs, assessments are generally restricted to indicator species and more rapid estimates of variables such as benthic cover, and lower levels of replication are accepted than in intermediate and expert monitoring. Though variable in quality and coverage, the resulting data can be invaluable in broad scale scientific assessments of reef status (Bruno and Selig 2007).
The urgency for accurate and reliable monitoring of coral reefs, that can serve both national (local) and international (global) needs is high, due to the poor performance of coral reef targets in the mid-term assessment of Aichi Target performance (GBO 2014). The GCRMN is developing with involvement from GEO BON and GOOS to become a mature observation network (UNESCO 2012), to better report on global targets (Aichi Target 10 on climate-sensitive ecosystems, and 14 on Oceans), and to feed into management, such as through the IUCN Red Lists of species and ecosystems. At the same time, extending citizen science contributions, and establishing a more open-data philosophy for monitoring data to maximise its accessibility, for example, through OBIS (Table 6.2), are emerging priorities.
6.3.3 The Reef Life Survey (RLS)
The Reef Life Survey (RLS) was established in 2007 to test the concept that a rigorous scientific approach to marine biodiversity monitoring could be developed within a citizen science framework (Edgar and Stuart-Smith 2014). The primary aim was to engage recreational divers to obtain scientific data from biodiversity observations that spanned geographic, temporal and taxonomic scales too costly for scientists to collect. It also aimed to extend other citizen science programs such as Reef Check and the REEF (see Sect. 6.3.2) that collected less detailed data (Edgar et al. 2016). Following establishment of the charitable Reef Life Survey Foundation (www.reeflifesurvey.com) to oversee field activities, appropriate data collection methodology, training, data entry and management procedures were developed, and different mechanisms for data collection were tested. Field survey methods were based on those applied over two decades by University of Tasmania researchers in Marine Protected Area (MPA) monitoring studies (Edgar and Barrett 1999; Barrett et al. 2009).
Three coincident elements of biodiversity are documented along 50 m long underwater transect lines. Divers record abundances and sizes of all fish, and abundances of all large (>2.5 cm length) mobile invertebrates (echinoderms, crustaceans and gastropods) and cryptic fishes. The area covered by sessile invertebrates, macrophytes and abiotic habitat is quantified through digitisation of photoquadrats (e.g., using Coral Point Count; Kohler and Gill 2006). Divers are trained on a one-on-one basis, each novice diver following behind a trained diver and duplicating transect blocks until the required level of expertise is reached. A comparison of data collected by trained volunteers and experienced scientists at the same sites showed that the variation attributable to diver experience was not significant, and negligible (<1 %) relative to differences between sites and regions (Edgar and Stuart-Smith 2009). The RLS program possesses a degree of self-regulation, where the keenest volunteers tend to also collect the best data, participate most frequently and persist longest (Edgar and Stuart-Smith 2009). A network of over 100 active RLS divers has now been established worldwide.
Application of RLS methods has allowed the first global analyses using standardised site-based procedures that are quantitative, species-level and cover multiple higher taxa. Data have been obtained for over 4500 species, 2800 sites, 600,000 species abundance records, 43 countries, and 83 marine ecoregions including Antarctica (e.g., Stuart-Smith et al. 2013, 2015). Many sites have been surveyed on multiple occasions, in some cases annually since 2007. These data add enormous contextual value to local surveys, and provide sufficient replication to disentangle many interactive and non-linear threats to marine biodiversity, including impacts of climate change, fishing and invasive species. For example, Edgar et al. (2014) included an order of magnitude more MPAs than any previously attempted using standardised field data. They found no detectable differences between fish communities present in most of the 87 MPAs investigated when compared with comparable fished communities (i.e., most MPAs were ‘paper parks’). However, some MPAs were extremely effective, with many large fishes and high conservation success. The RLS data are expected to be increasingly useful for (i) assessing ecosystem impacts of global threats to species at all levels of the food web from primary producers to higher predators, (ii) quantifying population trends for threatened species, and (iii) tracking international commitments associated with marine biodiversity in shallow reef ecosystems.
6.3.4 Harmful Algal Blooms (HAB)
Proliferation of microalgae in marine or brackish waters can cause massive fish kills, contaminate seafood with toxins, and alter ecosystems in ways that humans perceive as harmful. These phenomena are referred to as harmful algal blooms (HAB). Data on the distribution of toxic and harmful microalgae are collected through national surveillance programmes aimed at protecting public health, wild and cultured fish and shellfish, and bathing water quality. Sampling methods include plankton net hauls, water samples and molecular tools to detect species or genus-specific algal toxins in fish and shellfish. Benthic HAB species are collected from sediment, corals, seaweed or standardised screens. The detection of HAB species is challenging as many are difficult or impossible to identify even by using a light microscope. The challenge of maintaining a consistent microalgal taxonomy is addressed in the IOC Taxonomic Reference List of Toxic Plankton Algae within the World Register of Marine Species (Moestrup et al. 2009). The Intergovernmental Oceanographic Commission (IOC) of UNESCO has for two decades facilitated research to improve observations of harmful algae, provided training opportunities for their improved monitoring, as well as supported regional and global networks for knowledge and data sharing. The provision of method manuals and guides is central to observations of HAB species. The manual on HAB (Hallegraeff et al. 2003) is a base reference for methods and has been complemented by Babin et al.’s (2008) monograph on real-time observation systems, Karlson et al.’s (2010) intercomparison of quantative methods, and Reguera et al.’s (2011) sampling and analysis manual.
Global data on HAB species occurrences and their impacts are stored in the Harmful Algae Event Data Base (HAEDAT) in OBIS (Table 6.2). This international compiling and sharing of HAB data was initiated in the 1980s and is now accelerating and will provide the basis for a ‘Global HAB Status Report’ with the aims of compiling an overview of HAB events and their societal impacts; providing a worldwide appraisal of the occurrence of toxin-producing microalgae; and assessing the status and probability of change in HAB frequencies, intensities, and distribution resulting from environmental changes at the local and global scale. Linkages will be established with the International Panel on Climate Change (IPCC) reporting on the biological impacts of climate change. The Status report will provide the scientific community as well as decision makers with a reference on HAB occurrence and impacts on ecosystem services. IOC UNESCO project partners include the International Atomic Energy Agency (IAEA), the International Council for Exploration of the Sea (ICES), the North Pacific Marine Science Organization (PICES) and the International Society for the Study of Harmful Algae (ISSHA).
6.4 Data Management
Field data may be mapped to geographic areas, seascapes, habitats and against environmental parameters. Similar, globally applicable systems for the classification of marine habitats have been developed in Europe (Connor et al. 2004; Costello and Emblow 2005; Anon. 2014) and USA (Anon. 2012). The former leads to species-level biotopes (i.e., habitat + community), while the latter does not go to biotope level but does include seascape features (reviewed by Costello 2009). These can be presented as hierarchical lists and two-dimensional matrices (Fig. 6.1). The term habitat is highly context dependent and loosely used. Strictly speaking habitats are the immediate physical environment repeatedly associated with a species or distinct assemblage (or community) of species. The lowest level of habitat classifications are thus characterised by particular species. In contrast, related concepts of seascapes (landscapes, topographic features) and ecosystems will contain a variety of habitats (Costello 2009). These can be mapped over larger areas using remote sensing methods, whereas habitats usually need in situ sampling to identify their characteristic species, although exceptions exist in locations with biogenic habitat structure (e.g., seagrass beds, mangrove forests) (e.g., Andréfouët et al. 2001).
A biotope matrix with the most important physical habitat features on the axes. This illustrates the relationship of shore height (littoral) and sea depth (sublittoral, offshore), with substratum (rock and grades of sediment), and the exposure of rocky habitats to wave action. These factors distinguish biotopes at the upper levels of the habitat classification. Within the matrix the characteristic species of the communities occurring in the habitats are indicated. Adapted from Costello and Emblow (2005)
Knowing which species are present at a place and time is fundamental to biodiversity studies. Usually species are classified taxonomically because this is convenient and closely related species tend to have similar functional roles in ecosystems. However, ecologists may also classify species by their ecological traits (e.g., Wahl et al. 2013). Thus WoRMS (see Sect. 6.4.1) is developing a standardised approach to apply biological and ecological traits to marine species (Costello et al. 2015a).
A necessary step in organizing marine biodiversity data in integrated information systems is the development of appropriate thesauri and classification systems, as well as implementing quality control and feedback mechanisms. When integrating quantitative and qualitative natural history and distributional data, the use of both authoritative taxonomic and geographical hierarchical schema is essential. Here we introduce the leading taxonomic and geographic standards databases for the marine environment (Table 6.2).
6.4.1 World Register of Marine Species (WoRMS)
WoRMS is an open-access online database that provides an authoritative and comprehensive list of names of all marine organisms, including information on higher classification, synonymy, images and links to other information (Costello et al. 2013c). It currently contains over 240,000 accepted species names (Boxshall et al. 2015). While highest priority goes to valid names, other names in use are included so that this register is a guide to interpret taxonomic literature. Automated tools allow users to upload their species lists and match and classify their names against WoRMS. WoRMS makes use of the Aphia infrastructure which is designed to capture taxonomic and related data and information (Vandepitte et al. 2015a). WoRMS was a development from the European Register of Marine Species (ERMS) (Costello 2000; Costello et al. 2001), and thus its content is controlled by an Editorial Board of taxonomic and thematic experts who elect a governing and steering committee, and invite colleagues to assist them. A permanent host institution provides professional computational support for the database, including monthly archiving. As of January 2016, there were 393 editors from 273 institutions in 50 countries actively involved in the management and quality control of the WoRMS content. Through this editorial community, communication and collaboration within and beyond this community is facilitated (e.g., Appeltans et al. 2012), which can lead to increased rates of species discoveries and synonym names, which in turn can lead to a reduced rate of creating new synonyms and homonyms. WoRMS uses Life Science Identifiers (LSIDs) as persistent, location-independent, resource identifiers for each species name (Costello et al. 2013a). WoRMS forms the taxonomic backbone for OBIS, meaning that each taxon name in OBIS is matched against WoRMS to verify its validity and spelling (Vandepitte et al. 2011, 2015a, b). WoRMS is also a major contributor to the Catalogue of Life, Encyclopedia of Life, and LifeWatch Marine Virtual Research Environment (http://marine.lifewatch.eu). Species can be grouped with WoRMS to form Global, Regional and Thematic Databases. For example, the World Register of Introduced Marine Species (WRIMS) provides an entry point and experts to manage information on alien species (Pagad et al. 2015).
6.4.2 Marine Regions
Marine Regions (www.marineregions.org) hierarchically organises over 30,000 geographic areas from national and global marine gazetteers and databases (Claus et al. 2014). It contains spatial information of 264 different physical (e.g., sandbank, seamount, island, bay) and administrative (e.g., Exclusive Economic Zones, Marine Protected Area, Fisheries Zones or Biogeographic Regions) kinds of places. Both marine (e.g., seamounts, canyons, guyots, fracture zones, banks, ridges, basins) and coastal features (e.g., bays, fjords, cliffs, lagoons, beaches) are included. In order to preserve the identity of the marine geographic objects from the database, and to name and locate the geographic resources on the web, each geographic object is allocated a Marine Region Identifier, or MRGID. This unique persistent resource identifier is comparable to a LSID, being a unique identifier to locate the item on the World Wide Web.
6.4.3 Ocean Biogeographic Information System (OBIS)
OBIS is the world’s largest database on the distribution and abundance of marine life. In 2009, IOC Member States recognised the importance of knowledge of the ocean’s biodiversity to national and global environmental policies when they adopted it from the Census of Marine Life (Costello and Vanden Berghe 2006; Costello et al. 2007; O’Dor et al. 2012). OBIS operates through a network of national, regional and thematic nodes, and a secretariat based at the IOC’s International Oceanographic Data and Information Exchange (IODE) programme office in Oostende, Belgium. This office provides training and technical assistance, guides new data standards and technical developments, and encourages international cooperation to foster the group benefits of the network.
OBIS is a global science alliance that facilitates free and open access to data and information on marine biodiversity. It provides a single access point to over 45 million observations of 114,000 marine species, collected on 4.6 million sampling events from 3.2 million sampling stations, integrated from over 1900 datasets provided by nearly 500 institutions in 56 countries, It grows by about 3 million records per year. Data are subject to a series of quality control steps, including for taxonomic nomenclature and geography (Vandepitte et al. 2011, 2015a, b; IODE Steering Group for OBIS 2013).
Communities associated with OBIS include OBIS-SEAMAP (Spatial Ecological Analysis of Megavertebrate Populations) focusing on megafauna, and MICROBIS (http://icomm.mbl.edu/microbis) on microbes. The latter collects molecular observations of marine microbial organisms at taxonomic ranks from phyla to genus, together with their contextual physical and biochemical data measured in situ or from remote sensing. It has developed tools for extracting diversity measures, as well as other ecologically relevant statistics, from molecular datasets (Giongo et al. 2010; Buttigieg and Ramette, 2014). More comprehensive taxon based databases include the pioneering FishBase (Froese and Pauly 2015).
So far, 1000 publications have cited OBIS and on average 10 more each month (e.g., Basher et al. 2014a, b; Saeedi and Costello 2012; Costello et al. 2015a). OBIS directly contributes to several international activities, such as the UN Convention on Biological Diversity (for the identification of Ecologically or Biologically Significant Areas), the UN Food and Agriculture Organization (for the identification of Vulnerable Marine Ecosystems), the UN World Ocean Assessment, and the Global Environment Fund Transboundary Water Assessment. The Global Biodiversity Information Facility (GBIF) and OBIS use the same data standards and data sharing protocol (i.e., GBIF’s Integrated Publishing Toolkit). GBIF contains all OBIS and additional marine data (e.g., Costello et al. 2013d). Most data in OBIS are available from the north-west and north-east Atlantic, South Africa and New Zealand, and some other locations (Fig. 6.2). The potential of data published through OBIS for time-series analysis was highlighted in a recent global scale analysis (Dornelas et al. 2014).
An important development that will aid time series analysis, ecological niche modelling and climate change studies is currently underway as part of a two-year IODE project called ‘Expanding OBIS with environmental data’ (OBIS-ENV-DATA), which started in March 2015. The project is working on a solution to retain data in biological datasets that hold more than just species occurrence data, such as providing environmental and ecological context and data. The new approach will be based on the new Darwin Event Core and a modified ‘MeasurementorFact’ extension. The major change is that it will bring OBIS from a purely species occurrence database to one that can handle hierarchical sampling event structure with additional environmental and biometric measurements as well as details on the nature of the observations, measurements, and data collection methods, including equipment, data processing and sampling efforts.
6.4.4 Time-Series Data Availability
At present there are 20 monitoring programmes that have targeted species for more than five years that have been entered in OBIS. The focus of these efforts is on economically valuable and charismatic species (e.g., Antarctic krill, American lobster, marine mammals and seabirds). By contrast there are many more monitoring programmes targeting marine communities that have data for at least five years; 216 community monitoring programmes have uploaded their data to OBIS. When these programmes are combined, 16,616 stations have been monitored, encompassing most coastlines of the world, with less data available in developing countries or remote regions (Fig. 6.3a). The accumulation of time-series data has been exponential (Fig. 6.3b), reflecting both increasing monitoring efforts and global coordination. There may be an increasing willingness of scientists and institutions to share their data, with programmes such as the European Groundfish Survey showing up as being an important source of biodiversity data in the mid-1990s on a global scale (Fig. 6.3b). Even so, there are relatively fewer new stations that are being added to OBIS in comparison to the number of stations where monitoring surveys have ceased (Fig. 6.3b), leading to a net loss of time-series from OBIS in this decade. Explanations for this trend may be delays in data deposition, and/or perhaps the scope of specific monitoring efforts is increasing in extent and coordination.
a Map of station locations where monitoring surveys have been conducted for at least five years presently held in OBIS. b The left axis illustrates the number of stations where time-series data has been collected versus the year of the first survey (black histogram). Overall there has been an increase in monitoring. However, since the start of this century there has been a relative decrease in the number of stations being added to OBIS, evidenced by (right axis) the proportional difference in the number of new stations being added to OBIS versus those reaching completion. The blue line indicates where more monitoring stations were gained than lost from OBIS in a given year, while the red line indicates a loss
6.4.5 Global Marine Environment Datasets (GMED)
GMED is a compilation of more than 60 publicly available climatic, biological and geophysical environmental layers featuring present, past and future environmental conditions (Basher et al. 2015). Marine biologists increasingly utilise geo-spatial techniques with modelling algorithms to visualise and predict species biodiversity at a global scale. Marine environmental datasets available for species distribution modelling (SDM) have different spatial resolutions and are frequently provided in assorted file formats. This makes data assembly one of the most time-consuming parts of any study using multiple environmental layers for biogeography visualisation or SDM applications. GMED covers the widest available range of environmental layers from in situ measured, remote-sensed, and modelled datasets for a broad range of quantitative environmental variables from the surface to the deepest part of the ocean. It has a uniform spatial extent, high-resolution land mask (to eliminate land areas in the marine regions), and high spatial resolution (5 arc-minute, ca. 9.2 km near equator). The free online availability of GMED enables rapid map overlay of species of interest (e.g., endangered or invasive) against different environmental conditions of the past, present and the future, and expedites mapping distribution ranges of species using popular SDM algorithms (e.g., Basher et al. 2014a, 2015; Basher and Costello 2016).
6.5 Data Analysis
Although marine biodiversity data analysis requires its own taxonomic, geographic and environmental information context, such as provided by WoRMS, Marine-Regions, OBIS, and GMED, the methods of data analysis are similar to biodiversity in other environments. The data are categorical (i.e., species, habitats, biotopes), numerical (e.g., species abundance, cover, biomass), and cartographic. Thus metrics of ‘biodiversity’ include species richness and abundance, phylogenetic structure (e.g., taxonomic distinctness; Warwick and Clarke 1998), indicator species, habitat and/or biotope richness in an area (Costello 2001). Data may be presented on maps, graphs, tables and as matrices (e.g., Figs. 6.1 and 6.2). Numerous software tools are available for this analysis, including PRIMER-E (www.primer-e.com), PAST (http://folk.uio.no/ohammer/past), MODESTR (www.ipez.es/ModestR), SAGA (System for Automated Geoscientific Analyses; www.saga-gis.org) and DIVA-GIS (www.diva-gis.org). The open-source software R has the benefit that the analytical process is documented and can be published to aid reproducibility of the analyses.
The massive size of modern datasets, such as in OBIS and GMED, can lead to a new set of difficulties in analysis and interpretation. These difficulties include processing times that can exceed the capabilities of extant computers, propagation of undetected errors, unfamiliarity with analytical assumptions (e.g., spatial autocorrelation), and difficulties in visualisation (Edgar et al. 2016). Fortunately, big-data techniques applied in other fields, such as high-performance and parallel computing, are helping to solve many of these problems. In addition packages to overcome significant challenges in compiling large datasets and maintaining these data through time are being improved. For example, the R package ‘taxize’ (Chamberlain and Szocs 2013), which relies on accessing freely available and accurate information on species taxonomy, including from WoRMS. This emphasises the benefits of scientists and institutes publishing monitoring data in order to advance our understanding of biodiversity change.
6.6 Discussion
Global marine biological databases are well-established for quality assurance of species nomenclature and associated information (WoRMS) and distribution data (OBIS) (Costello and Wieczorek 2014; Costello et al. 2015b). The coverage and quality of global marine environmental layers improves each year through a combination of remotely sensed, in situ, and modelling data. These layers and maps of marine regions are also freely available online at GMED and marineregions.org. Species trait information is being added to WoRMS, and more sample information can be added to OBIS so users can select datasets suitable for their purposes. The mapping of available data in OBIS shows how more sampling has been conducted in northern hemisphere and coastal environments compared to open-ocean, deep-sea and developing countries (Fig. 6.3a). However, because neither biodiversity nor human impacts are homogenously distributed, neither should it be expected that global sampling programmes will be. Sampling of particular guilds of biodiversity should thus be stratified to represent its spatial variation.
A major obstacle to engaging more scientists and citizens in recording marine biodiversity is the availability of guides to the identification of species. Generally, these are only widely available for vertebrates (Costello et al. 2006, 2015b). To identify invertebrates often requires numerous papers to be obtained, sometimes in different languages. The most useful publications are reviews of the taxonomy of particular taxa in a region that include images, drawings and keys that synthesise information on many species (Costello et al. 2013a, b, 2014a, b). The best long-term solution would be an online, pictorial, guide to all marine species accessible to people in several languages and scripts (Costello et al. 2015b).
All methods have their biases and this needs to be recognised in data analysis rather than assume a conventional method is representative of all biodiversity. In fact, it may be that pooling different sampling processes to gain insights into different aspects of biodiversity will create the most comprehensive understanding of how biodiversity is changing in the ocean. Methods must be selected that are ‘best fit for the purpose’ and limitations imposed by costs and environmental conditions should be considered in the interpretation of the samples obtained. Standardised methods have the advantage of apparent comparability between study locations and over time. However, this assumes the behaviour of animals is the same between species, and even within a species between locations and over time. This is not necessarily the case. Being ectothermal, fish appetite and activity is strongly temperature dependent (e.g., Darwall et al. 1993; Costello et al. 1995). Thus seasonal changes in the catch of fish and other mobile species may not reflect fish abundance or changing distribution, but rather their activity. Animal behaviour also needs to be considered. For example, fish are wary of people in places they are fished, especially spear-fished. However, in marine reserves they lose this fear and can be approached closely (Costello 2014). Where mammals, birds, fish and other animals may be fed, they become attracted to people. This mirrors the behaviour of animals on land. Thus not only do the physical features of sampling methods need to be considered in terms of bias, so do the behavioural responses of animals.
More recently developed methods, such as using photography, hydrophones, and tagging, avoid killing the species of interest. Artificial substrata, light-traps, hook and line, and traps can avoid killing unwanted by-catch species. However, most netting and trawling methods result in by-catch, and seabed dredging and trawling also damage habitat. It seems likely that scientific sampling will come under increasing ethical pressure to minimise habitat damage, by-catch and stress to species, especially in nature conservation areas and where species are threatened. Thus new in situ observation methods such as still and video image capture, seafloor observatories, and sensors, are likely to become more important because they cause less disturbance of biodiversity.
In addition to the CPR, RLS and GEOHAB programmes reviewed here, new networking initiatives, marine biodiversity observation networks (mBON) in the USA (Muller-Karger et al. 2014), marine station networks and related organisations (Costello et al. 2015c), and groups of scientists interested in the biological and ecological effects of climate change, may establish globally coordinated marine biodiversity monitoring programmes. In addition, several international ocean observing systems, initially focused on the collection of physical and chemical ocean data, are now including biological data as well. These are comprised of the Australian led Integrated Marine Observing Systems (IMOS; www.imos.org.au) and the Southern Ocean Observing System (SOOS; www.soos.au), and NOAA’s Integrated Ocean Observing System (IOOS; www.ioos.noaa.gov). Some international efforts have a regional focus. For example, the Circumpolar Biodiversity Monitoring Program (CBMP), under the auspices of the Arctic Council, has an Arctic Marine Biodiversity Monitoring Plan (www.caff.is/marine).
Dornelas et al. (2014) compiled the first global time-series data base for analysis of trends in marine biodiversity. As described earlier, biodiversity data are available for many taxa and regions of the world and the challenge remains to access, compile and curate these data. A major obstacle is therefore not only the difficulty in maintaining funding for monitoring or data synthesis efforts, but fostering motivation for institutes and scientists to publish their data and overcoming communication and cultural differences. Building collaborative networks may be one means to begin to surmount these challenges to collate data across scientists, institutions, and data repositories. While efforts to collate the data that has been collected by the global monitoring community is certainly the best hope for generating historical knowledge, purpose-built global biodiversity platforms are fundamental for ensuring the capacity to track biodiversity change into the future. For example, MarineGEO (Duffy 2014) is establishing observatories where multiple components of biodiversity, including benthic and pelagic communities and food webs, will be monitored using globally standardised methods and experimentation, including artificial substrata (e.g., ARMS). Associated initiatives focus on global studies on seagrass (http://zenscience.org/about-zen; Reynolds et al. 2014) and kelp (www.kelpecosystems.org) habitats. Standard methods for these habitats have been published (e.g., Edgar et al. 2001; Davies et al. 2001). Such projects may utilise the Zooniverse platform for citizen science crowd sourcing (www.zooniverse.org).
Global sampling of surface water marine microbes is also underway utilising genomic methods, including synchronised sampling of hundreds of stations on ‘Ocean Sampling Day’ (www.microb3.eu/osd) (e.g., Davies et al. 2012a, b; Kopf et al. 2015). These and related research into molecular indicators may fill gaps that complement more conventional metrics of biodiversity (Leray and Knowlton 2015). Although there are issues to be resolved in the interpretation of DNA found in the environment (eDNA), including contamination, accuracy of matching results to species, and uncertainty about live versus dead material, it may prove invaluable in detecting rare and/or microscopic species that are otherwise hard to sample (Thomsen and Willerslev 2015).
There are two established and several emerging globally coordinated marine biodiversity monitoring programmes, covering surface plankton (CPR), mobile rocky and coral reef fauna (RLS), seagrass and kelp habitats, and pelagic microbes. There are similar sampling methods used internationally for other guilds of species; including mammals, whale sharks and birds; small fish and crustaceans in fishery trawls; macro-invertebrate infauna of coastal sediments; and sessile and sedentary biota on rocky seashores. For example, programmes such as the ICES North Sea Benthos Survey (e.g., Duineveld et al. 1991; Basford et al. 1993) and NaGISA (Benedetti-Cecchi et al. 2010; Cruz-Motta et al. 2010; Konar et al. 2010; Pohle et al. 2011; Miloslavich et al. 2013) could be continued and expanded internationally. NaGISA was one of several projects within the decade-long Census of Marine Life, the largest global collaboration in marine biology covering coastal to deep-sea, and polar to tropical environments, and which established OBIS (O’Dor et al. 2012). Thus opportunities exist to design globally standardised programmes for these ecological guilds that would be comparable with historic data. For example, the IOC–UNESCO endorsed IndiSeas (www.indiseas.org) has begun to provide indicators of biodiversity (including ecosystem health) related to fisheries and environment.
Gaps in time-series may be partly filled by using microfossils from sediment cores and specimen collections in museums, and also by revisiting places sampled in the past without continuous time-series. In addition, video cameras (baited and unbaited) are widely used for recording scavenging megafauna from coastal to deep-sea habitats (e.g., Costello et al. 2005). Gaps in these programmes include the species rich epi-benthic crustaceans and molluscs which together comprise one quarter of all marine species (Appeltans et al. 2012). However, the use of artificial substrata such as ARMS and light-traps may be able to fill this gap. Additional guilds that could be considered for monitoring include sediment meiofauna and parasites.
A common concern in launching global initiatives is both the start-up and long-term funding (Costello et al. 2014c). It is notable that the CPR, RLS, WoRMS and FishBase established their own legal organisations to ‘own’ their initiatives, even though they are largely funded by government and hosted by particular institutes. This community ownership may address issues of financial liability of individuals and their institutions, ownership of intellectual property, and perceptions of who benefits from the research. The establishment of global programmes must consider these and other issues so as to maximise the likelihood of support from individual scientists, host institutions and governments in the long term (Costello et al. 2014c).
References
Andréfouët, S., Costello, M. J., Faith, D. P., Ferrier, S., Geller, G. N., Höft, R., et al. (2008a). The GEO biodiversity observation network concept document. Geneva, Switzerland: GEO—Group on Earth Observations.
Andréfouët, S., Costello, M. J., Rast, M., & Sathyendranath, S. (2008b). Earth observations for marine and coastal biodiversity. Remote Sensing of Environment, 112, 3297–3299.
Andréfouët, S., Muller-Karger, F. E., Hochberg, E. J., Hu, C., & Carder, K. L. (2001). Change detection in shallow coral reef environments using Landsat 7 ETM+ data. Remote Sensing of Environment, 78, 150–162.
Anonymous. (2012). Coastal and marine ecological classification standard. Chalreston SC: Federal Geographic Data Committee, FGDC-STD-018-2012. www.csc.noaa.gov/digitalcoast/_/pdf/CMECS_Version%20_4_Final_for_FGDC.pdf Cited 8 February 2016.
Anonymous. (2014). EUNIS habitat classification. Copenhagen: European Environment Agency. http://www.eea.europa.eu/themes/biodiversity/eunis/eunis-habitat-classification Cited 8 February 2016.
American Public Health Association (APHA), American Water Works Association, and Water Environment Federation. (2007) Standard methods for the examination of water and wastewater. American Public Health Association, American Water Works Association, and Water Environment Federation, Washington, DC., USA.
Appeltans, W., Ahyong, S.T., Anderson, G., Angel, M. V., Artois, T., & Bailly, N. et al. (2012). The magnitude of global marine species diversity. Current Biology 22, 2189–2202.
Ash, N., Jürgens, N., Leadley, P., Alkemade, R., Araújo, M. B., & Asner, G. P. (2009). bioDISCOVERY: Assessing, monitoring and predicting biodiversity. DIVERSITAS Report No. 7.
Atalah, J., Costello, M. J., & Anderson, M. (2007a). Temporal variability and intensity of grazing: A mesocosm experiment. Marine Ecology Progress Series, 341, 15–24.
Atalah, J., Otto, S., Anderson, M., Costello, M. J., Lenz, M., & Wahl, M. (2007b). Temporal variance of disturbance did not affect diversity and structure of a marine fouling community in north-eastern New Zealand. Marine Biology, 153, 199–211.
Atkinson, L. J., Leslie, R. W., Field, J. G., & Jarre, A. (2011). Changes in demersal fish assemblages on the west coast of South Africa, 1986–2009. African Journal of Marine Science, 33, 157–170.
Babin, M., Cullen, J., & Roesler, C. (Eds.). (2008). Real time observations systems for marine ecosystem dynamics and harmful algal blooms: Theory, instrumentation and modelling. Paris, France: UNESCO Publishing.
Barrett, N. S., Buxton, C. D., & Edgar, G. J. (2009). Changes in invertebrate and macroalgal populations in Tasmanian marine reserves in the decade following protection. Journal of Experimental Marine Biology and Ecology, 370, 104–119.
Basford, D. J., Eleftheriou, A., Davies, I. M., Irion, G., & Soltwedel, T. (1993). The ICES North Sea benthos survey: The sedimentary environment. ICES Journal of Marine Science: Journal du Conseil, 50, 71–80.
Basher, Z. & Costello, M. J. (2016). The past, present and future distribution of a deep-sea shrimp in the Southern Ocean. PeerJ, 4, e1713 https://doi.org/10.7717/peerj.1713
Basher, Z., Bowden, D. A., & Costello, M. J. (2014a). Diversity and distribution of deep-sea shrimps in the Ross Sea region of Antarctica. PLoS ONE, 9, e103195.
Basher, Z. & Costello, M. J. (2014b). Chapter 5.22: Shrimps (Crustacea: Decapoda). In C. De Broyer, P. Koubbi, H. J. Griffiths, B. Raymond, C. d’Udekem d’Acoz, A. Van de Putte, B. Danis, B. David, S. Grant, J. Gutt, C. Held, G. Hosie, F. Huettemann, A. Post, Y. Ropert-Coudert (Eds.), Biogeographic Atlas of the Southern Ocean (pp. 190–194). Cambridge, UK: Scientific Committee on Antarctic Research.
Basher, Z., Bowden, D. A. & Costello, M. J. (2015). Global marine environment dataset (GMED). World Wide Web electronic publication. Version 1.0 (Rev.01.2014). http://gmed.auckland.ac.nz Cited 4 February 2015.
Benedetti-Cecchi, L., Iken, K., Konar, B., Cruz-Motta, J., Knowlton, A., et al. (2010). Spatial relationships between polychaete assemblages and environmental variables over broad geographical scales. PLoS ONE, 5, e12946.
Benoít, H. P., & Swain, D. P. (2003). Accounting for length-and depth-dependent diel variation in catchability of fish and invertebrates in an annual bottom-trawl survey. ICES Journal of Marine Science, 60, 1298–1317.
Bianchi, G., Gislason, H., Graham, K., Hill, L., Jin, X., Koranteng, K., et al. (2000). Impact of fishing on size composition and diversity of demersal fish communities. ICES Journal of Marine Science: Journal du Conseil, 57, 558–571.
Boero, F., Dupont, S., & Thorndyke, M. (2015). Make new friends, but keep the old: Towards a transdisciplinary and balanced strategy to evaluate good environmental status. Journal of the Marine Biological Association of the United Kingdom, 95, 1069–1070.
Bork, P., Bowler, C., de Vargas, C., Gorsky, G., Karsenti, E., & Wincker, P. (2015). Tara Oceans studies plankton at planetary scale. Science, 348, 873.
Boxshall, G., Mees, J., Costello, M. J., Hernandez, F., Gofas, S., & Hoeksema, B. W. (2015). World register of marine species. Available via VLIZ. http://www.marinespecies.org Cited 23 December 2015.
Bruno, J. F., & Selig, E. R. (2007). Regional decline of coral cover in the indo-pacific: Timing, extent, and subregional comparisons. PLoS ONE, 2, e711.
Buttigieg, P. L., & Ramette, A. (2014). A guide to statistical analysis in microbial ecology: A community-focused, living review of multivariate data analyses. FEMS Microbiology Ecology, 90, 543–550.
Chamberlain, S. & Szocs, E. (2013). Taxize—taxonomic search and retrieval in R. F1000Research 2, 191.
Claus, S., De Hauwere, N., Vanhoorne, B., Deckers, P., Souza Dias, F., Hernandez, F., et al. (2014). Marine regions: Towards a global standard for georeferenced marine names and boundaries. Marine Geodesy, 37, 99–125.
Connor, D. W., Allen, J. H., Golding, N., Howell, K. L., Lieberknecht, L. M., & Northen, K. O. et al. (2004). The marine habitat classification for Britain and Ireland version 04.05. Peterborough, UK: Joint Nature Conservation Committee (JNCC). ISBN 1 861 07561 8 (internet version).
Connor, D. W., Gilliland, P. M., Golding, N., Robinson, P., Todd, D., & Verling, E. (2006). UKSeaMap: The mapping of seabed and water column features of UK seas. Peterborough, UK: Joint Nature Conservation Committee.
Costello, M. J. (1992). Abundance and spatial overlap of gobies in Lough Hyne, Ireland. Environmental Biology of Fishes, 33, 239–248.
Costello, M. J. (2000). Developing species information systems: the European register of marine species. Oceanography, 13(3), 48–55.
Costello, M. J. (2001). To know, research, manage, and conserve marine biodiversity. Océanis, 24(4), 25–49.
Costello, M. J. (2009). Distinguishing marine habitat classification concepts for ecological data management. Marine Ecology Progress Series, 397, 253–268.
Costello, M. J. (2014). Long live marine reserves: A review of experiences and benefits. Biological Conservation, 176, 289–296.
Costello, M. J. (2015). Biodiversity: The known, unknown and rates of extinction. Current Biology, 25(9), 368–371.
Costello, M. J., & Thrush, S. F. (1991). Colonization of artificial substrata as a multi-species bioassay of marine environmental quality. In D. W. Jeffrey & B. Madden (Eds.), Bioindicators and environmental management (pp. 401–418). London, UK: Academic Press.
Costello, M. J., & Emblow, C. (2005). A classification of inshore marine biotopes. In J. G. Wilson (Ed.), The intertidal ecosystem: The value of Ireland’s shores (pp. 25–35). Dublin, Ireland: Royal Irish Academy.
Costello, M. J., & Vanden Berghe, E. (2006). “Ocean biodiversity informatics” enabling a new era in marine biology research and management. Marine Ecology Progress Series, 316, 203–214.
Costello, M. J., & Baker, C. S. (2011). Who eats sea meat? Expanding human consumption of marine mammals. Biological Conservation, 144, 2745–2746.
Costello, M. J., & Myers, A. A. (1996). Turnover of transient species as a contributor to the richness of a stable amphipod (Crustacea) fauna in a sea inlet. Journal of Experimental Marine Biology and Ecology, 202, 49–62.
Costello, M. J., & Wieczorek, J. (2014). Best practice for biodiversity data management and publication. Biological Conservation, 173, 68–73.
Costello, M. J., Darwall, W. R. & Lysaght S. (1995). Activity patterns of north European wrasse (Pisces, Labridae) and precision of diver survey techniques. In A. Eleftheriou, A. D. Ansell, C. J. Smith (Eds.), Biology and ecology of shallow coastal waters. Proceedings of the 28th European Marine Biology Symposium, IMBC, Hersonissos, Crete 1993 (pp. 343–350). Fredensborg, Denmark: Olsen and Olsen Publ..
Costello, M. J., Emblow, C. & White, R. (Eds.). (2001). European register of marine species. A check-list of marine species in Europe and a bibliography of guides to their identification. Patrimoines Naturels 50, 1–463.
Costello, M. J., Pohle G., & Martin A. (2004). Evaluating biodiversity in marine environmental assessments. Research and Development Monograph Series 2001, Canadian Environmental Assessment Agency, Ottawa, Canada. http://www.collectionscanada.gc.ca/webarchives/20071213100057/www.ceaa.gc.ca/015/001/019/print-version_e.htm
Costello, M. J., McCrea, M., Freiwald, A., Lundälv, T., Jonsson, L., Bett, B. J., et al. (2005). Role of cold-water Lophelia pertusa coral reefs as fish habitat in the NE Atlantic. In A. Freiwald & J. M. Roberts (Eds.), Cold-water corals and ecosystems (pp. 771–805). Berlin Heidelberg, Germany: Springer.
Costello, M. J., Emblow, C. S., Bouchet, P., & Legakis, A. (2006). European marine biodiversity inventory and taxonomic resources: State of the art and gaps in knowledge. Marine Ecology Progress Series, 316, 257–268.
Costello M. J., Stocks K., Zhang Y., Grassle J. F. & Fautin D. G. (2007). About the Ocean biogeographic information system. Retrieved from http://hdl.handle.net/2292/236
Costello, M. J., Cheung, A., & De Hauwere, N. (2010a). Topography statistics for the surface and seabed area, volume, depth and slope, of the world’s seas, oceans and countries. Environmental Science and Technology, 44, 8821–8828.
Costello, M. J., Coll, M., Danovaro, R., Halpin, P., Ojaveer, H., & Miloslavich, P. (2010b). A census of marine biodiversity knowledge, resources and future challenges. PLoS ONE, 8, e12110.
Costello, M. J., May, R. M., & Stork, N. E. (2013a). Can we name Earth’s species before they go extinct? Science, 339, 413–416.
Costello, M. J., May, R. M., & Stork, N. E. (2013b). Response to comments on “can we name Earth’s species before they go extinct?”. Science, 341, 237.
Costello, M. J., Bouchet, P., Boxshall, G., Fauchald, K., Gordon, D. P., Hoeksema, B. W., et al. (2013c). Global coordination and standardisation in marine biodiversity through the World Register of Marine Species (WoRMS) and related databases. PLoS ONE, 8(1), e51629.
Costello, M. J., Michener, W. K., Gahegan, M., Zhang, Z.-Q., & Bourne, P. (2013d). Data should be published, cited and peer-reviewed. Trends in Ecology & Evolution, 28(8), 454–461.
Costello, M. J., Appeltans, W., Bailly, N., Berendsohn, W. G., de Jong, Y., Edwards, M., et al. (2014a). Strategies for the sustainability of online open-access biodiversity databases. Biological Conservation, 173, 155–165.
Costello, M. J., Houlding, B., & Wilson, S. (2014b). As in other taxa, relatively fewer beetles are being described by an increasing number of authors: Response to Löbl & Leschen. Systematic Entomology, 39, 395–399.
Costello, M. J., Houlding, B. & Joppa, L. (2014c). Further evidence of more taxonomists discovering new species, and that most species have been named: response to Bebber et al. (2014). New Phytologist 202, 739–740.
Costello, M. J., Claus, S., Dekeyzer, S., Vandepitte, L., Tuama, É. Ó., & Lear, D. et al. (2015a). Biological and ecological traits of marine species. PeerJ 3, e1201.
Costello, M. J., Vanhoorne, B., & Appeltans, W. (2015b). Conservation of biodiversity through taxonomy, data publication and collaborative infrastructures. Conservation Biology, 29(4), 1094–1099.
Costello, M. J., Archambault, P., Chavanich, S., Miloslavich, P., Paterson, D. M., Phang, S.-W., et al. (2015c). Organizing, supporting and linking the world marine biodiversity research community. Journal of the Marine Biological Association of the United Kingdom, 95(3), 431–433. doi:10.1017/S0025315414001969
Cruz-Motta, J. J., Miloslavich, P., Palomo, G., Iken, K., Konar, B., Pohle, G., et al. (2010). Patterns of spatial variation of assemblages associated with intertidal rocky shores: A global perspective. PLoS ONE, 12, e14354.
Daan, N., Gislason, H., Pope, J. G., & Rice, J. C. (2005). Changes in the North Sea fish community: Evidence of indirect effects of fishing? ICES Journal of Marine Science, 62(2), 177–188.
Darwall, W. R. T., Costello, M. J., Donnelly, R., & Lysaght, S. (1993). Implications of life history strategies for a new wrasse fishery. Journal of Fish Biology, 41B, 111–123.
Davies, J., Baxter, J., Bradley, M., Connor, D., Khan, J., & Murray, E. et al. (2001), Marine monitoring handbook (405 p.). Peterborough: Joint Nature Conservation Committee.
Davies, N., Field, D., & Genomic Observatories Network. (2012a). Sequencing data: A genomic network to monitor Earth. Nature, 481(7380), 145.
Davies, N., Meyer, C., Gilbert, J. A., Amaral-Zettler, L., Deck, J., Bicak, M., et al. (2012b). A call for an international network of genomic observatories (GOs). GigaScience, 1(1), 5.
De Monte, S., Soccodato, A., Alvain, S., & d’Ovidio, F. (2013). Can we detect oceanic biodiversity hotspots from space? ISME Journal, 7, 2054–2056.
Doherty, P. J. (1987). Light-traps: Selective but useful devices for quantifying the distributions and abundances of larval fishes. Bulletin of Marine Science, 41(2), 423–431.
Dornelas, M., Gotelli, N. J., McGill, B., Shimadzu, H., Moyes, F., Sievers, C., et al. (2014). Assemblage time series reveal biodiversity change but not systematic loss. Science, 344(6181), 296–299.
Duffy, J. E. (2014). Sustaining coastal resilience. Pan-European Networks: Science and Technology, 12, 1–4.
Duineveld, G. C. A., Künitzer, A., Niermann, U., De Wilde, P. A. W. J., & Gray, J. S. (1991). The macrobenthos of the North Sea. Netherlands Journal of Sea Research, 28(1–2), 53–65.
Edgar, G. J., & Barrett, N. S. (1999). Effects of the declaration of marine reserves on Tasmanian reef fishes, invertebrates and plants. Journal of Experimental Marine Biology and Ecology, 242, 107–144.
Edgar, G. J., & Stuart-Smith, R. D. (2009). Ecological effects of marine protected areas on rocky reef communities: a continental-scale analysis. Marine Ecology Progress Series, 388, 51–62.
Edgar, G. J., & Stuart-Smith, R. D. (2014). Systematic global assessment of reef fish communities by the Reef Life Survey program. Scientific Data, 1(140007), 1–8.
Edgar, G. J., Orth, R. J., & Mukai, H. (2001). Fish, crabs, shrimps and other large mobile epibenthos: Measurement methods for their biomass and abundance in seagrass. In F. T. Short & R. G. Coles (Eds.), Global seagrass research methods (pp. 255–270). Amsterdam, The Netherlands: Elsevier.
Edgar, G. J., Stuart-Smith, R. D., Willis, T. J., Kininmonth, S., Banks, S., Barrettk, N. S., et al. (2014). Global conservation outcomes depend on marine protected areas with five key features. Nature, 506, 216–220.
Edgar, G. J., Bates, A. E., Bird, T. J., Jones, A. H., Kininmonth, S., Stuart-Smith, R. D., et al. (2016). New approaches to marine conservation through the scaling up of ecological data. Annual Review of Marine Science, 8, 435–461.
Edwards, M., Beaugrand, G., Hays, G. C., Koslow, J. A., & Richardson, A. J. (2010). Multi-decadal oceanic ecological datasets and their application in marine policy and management. Trends in Ecology & Evolution, 25, 602–610.
Edwards, M., Helaouet, P., Johns, D. G., Batten, S., Beaugrand, G., & Chiba, S. et al. (2012). Global marine ecological status report: Results from the global CPR survey 2010/2011. SAHFOS Technical Report 9, 1–40. Plymouth, U.K.
Eleftheriou, A. (Ed.). (2013). Methods for the study of marine benthos. Chichester, UK: Wiley-Blackwell.
Elliott, M., & Hemingway, K. (Eds.). (2002). Fishes in estuaries. Oxford, UK: Blackwell Science.
English, S. S., Wilkinson, C. C. & Baker, V. V. (1994). Survey manual for tropical marine resources. Townsville, Australia: Australian Institute of Marine Science (AIMS).
Facon, M., Pinault, M., Obura, D., Pioch, S., Pothin, K., Bigot, L., et al. (2016). A comparative study of the accuracy and effectiveness of line and point intercept transect methods for coral reef monitoring in the southwestern Indian Ocean islands. Ecological Indicators, 60, 1045–1055.
Francisco-Ramos, V., & Arias-González, J. E. (2013). Additive partitioning of coral reef fish diversity across hierarchical spatial scales throughout the Caribbean. PLoS ONE, 8(10), e78761.
Fraser, H. M., Greenstreet, S. P. R., & Piet, G. J. (2007). Taking account of catchability in groundfish survey trawls: Implications for estimating demersal fish biomass. ICES Journal of Marine Science, 64, 1800–1819.
Fraser, H. M., Greenstreet, S. P. R., Fryer, R. J., & Piet, G. J. (2008). Mapping spatial variation in demersal fish species diversity and composition in the North Sea: accounting for species- and size-related catchability in survey trawls. ICES Journal of Marine Science, 65, 531–538.
Froese, R. & Pauly, D. (2015). Fishbase. World Wide Web database. Accessed at http://www.fishbase.org on 19th February 2015.
GBO. (2014). Global biodiversity outlook 4. Secretariat of the convention on biological diversity. Montréal. 155 p. https://www.cbd.int/GBO4/
Giongo, A., Crabb, D. B., Davis-Richardson, A. G., Chauliac, D., Mobberley, J. M., Gano, K. A., et al. (2010). PANGEA: Pipeline for analysis of next generation amplicons. The ISME journal, 4(7), 852–861.
Gotelli, N. J., & Colwell, R. K. (2001). Quantifying biodiversity: Procedures and pitfalls in the measurement and comparison of species richness. Ecology Letters, 4(4), 379–391.
Haegeman, B., Hamelin, J., Moriarty, J., Neal, P., Dushoff, J., & Weitz, J. S. (2013). Robust estimation of microbial diversity in theory and in practice. The ISME Journal, 7, 1092–1101.
Hallegraeff, G. M., Anderson, D. M. & Cembella, A. D. (Eds.). (2003). Manual on harmful marine microalgae (794 p.). Paris: UNESCO Publishing.
Harris, S. A., Shears, N. T., & Radford, C. A. (2015). Ecoacoustic indices as proxies for biodiversity on temperate reefs. Methods in Ecology and Evolution.
Hiscock, K. (2014). Marine biodiversity conservation: A practical approach. Abingdon: Routledge. 289 pp.
Hodgson, G. (1999). A global assessment of human effects on coral reefs. Marine Pollution Bulletin, 38(5), 345–355.
Holmes, J. M. C., & O’Connor, J. P. (1988). A portable light-trap for collecting marine crustaceans. Journal of the Marine Biological Association of the United Kingdom, 68(2), 235–238.
Hungerford, H. B., Spangler, P. J. & Walker, N. A. (1955). Subaquatic light traps for insects and other animal organisms. Transactions of the Kansas Academy of Science (1903–) 58(3), 387–407.
Hunt, G., Cronin, T. M., & Roy, K. (2005). Species–energy relationship in the deep sea: A test using the quaternary fossil record. Ecology Letters, 8, 739–747.
Inniss, L., Simcock, A., Ajawin, A. Y., Alcala, A. C., Bernal, P., & Calumpong, H. P. et al. (2016). The first global integrated marine assessment. United Nations. Accessed at on 5th February 2016 http://www.un.org/Depts/los/global_reporting/WOA_RegProcess.htm
IODE Steering Group for OBIS (SG-OBIS). (2013). Reports of meetings of experts and equivalent bodies. Third session, IOC Project Office for IODE, 4–6 December 2013, IOC/IODE-SG-OBIS-III/3. UNESCO, Oostende. 40 p.
Jackson, J., Donovan, M., Cramer, K., & Lam, V. (2014). Status and trends of Caribbean coral reefs: 1970–2012 (304 p.). Washington, D.C: Global Coral Reef Monitoring Network.
Karlson B., Cusack C. & Bresnan, E. (Eds.). (2010). Microscopic and molecular methods for quantitative phytoplankton analysis. Paris: Intergovernmental Oceanographic Commission of UNESCO. 110 p.
Kingsford, M., & Battershill, C. (Eds.). (1998). Studying temperate marine environments: a handbook for ecologists ( 355 p.). Canterbury University Press.
Kohler, K. E., & Gill, S. M. (2006). Coral point count with excel extensions (CPCe): A Visual Basic program for the determination of coral and substrate coverage using random point count methodology. Computers & Geosciences, 32, 1259–1269.
Konar, B., Iken, K., Cruz-Motta, J. J., Benedetti-Cecchi, L., Knowlton, A., Pohle, G., et al. (2010). Current patterns of macroalgal diversity and biomass in northern hemisphere rocky shores. PLoS ONE, 5(10), e13195.
Kopf, A., Bicak, M., Kottmann, R., Schnetzer, J., Kostadinov, I., Lehmann, K., et al. (2015). The ocean sampling day consortium. GigaScience, 4(1), 1–5. doi:10.1186/s13742-015-0066-5
Kotwicki, S., De Robertis, A., von Szalay, P., & Towler, R. (2009). The effect of light intensity on the availability of walleye pollock (Theragra chalcogramma) to bottom trawl and acoustic surveys. Canadian Journal of Fisheries and Aquatic Sciences, 66(6), 983–994.
Leleu, K., Remy-Zephir, B., Grace, R., & Costello, M. J. (2012). Mapping habitat change after 30 years in a marine reserve shows how fishing can alter ecosystem structure. Biological Conservation, 155, 193–201.
Leray, M., & Knowlton, N. (2015). DNA barcoding and metabarcoding of standardized samples reveal patterns of marine benthic diversity. Proceedings of the National Academy of Sciences, 201424997.
Leujak, W., & Ormond, R. F. G. (2007). Comparative accuracy and efficiency of six coral community survey methods. Journal of Experimental Marine Biology and Ecology, 351(1), 168–187.
McGeoch, A. M., Genovesi, P., Bellingham, P. J., Costello, M. J., McGrannachan, C., & Sheppard, A. (2016). Prioritizing species, pathways, and sites to achieve conservation targets for biological invasion. Biological Invasions, 18(2), 299–314.
Meekan, M. G., Wilson, S. G., Halford, A., & Retzel, A. (2001). A comparison of catches of fishes and invertebrates by two light trap designs, in tropical NW Australia. Marine Biology, 139(2), 373–381.
Miller, T. J. (2013). A comparison of hierarchical models for relative catch efficiency based on paired-gear data for US Northwest Atlantic fish stocks. Canadian Journal of Fisheries and Aquatic Sciences, 70(9), 1306–1316.
Miloslavich, P., Cruz-Motta, J. J., Klein, E., Iken, K., Weinberger, V., Konar, B., et al. (2013). Large-scale spatial distribution patterns of gastropod assemblages in rocky shores. PLoS ONE, 8(8), e71396. doi:10.1371/journal.pone.0071396
Moestrup, Ø., Akselmann, R., Fraga, S., Hansen, G., Hoppenrath, M., & Iwataki, M., et al. (Eds.). (2009 onwards). IOC-UNESCO taxonomic reference list of harmful micro algae. Accessed at on 18 January 2015 http://www.marinespecies.org/hab/
Muller-Karger, F. E., Kavanaugh, M. T., Montes, E., Balch, W. M., Breitbart, M., Chavez, F. P., et al. (2014). A framework for a marine biodiversity observing network within changing continental shelf seascapes. Oceanography, 27(2), 18–23.
Neilson, B., & Costello, M. J. (1999). The relative lengths of seashore substrata around the coastline of Ireland as determined by digital methods in a geographical information system. Estuarine, and Coastal Shelf Sciences, 49, 501–508.
O’Dor, R., Boustany, A. M., Chittenden, C. M., Costello, M. J., Moustahfid, H., Payne, J., et al. (2012). A census of fishes and everything they eat: How the census of marine life advanced fisheries science. Fisheries, 37(9), 398–409.
Øresland, V. (2007). Description of the IMR standard light trap and the vertical distribution of some decapod larvae (Homarus and Nephrops). Western Indian Ocean Journal of Marine Science, 6(2), 225–231.
Pagad S., Hayes K, Katsanevakis S., & Costello M. J. (2015). World Register of Introduced Marine Species (WRIMS). Accessed at on 12 May 2015 http://www.marinespecies.org/introduced
Pereira, H. M., Ferrier, S., Walters, M., Geller, G. N., Jongman, R. H. G., Scholes, R. J., et al. (2013). Essential biodiversity variables. Science, 339, 277–278.
Perry, A. L., Low, P. J., Ellis, J. R., & Reynolds, J. D. (2005). Climate change and distribution shifts in marine fishes. Science, 308(5730), 1912–1915.
Pohle, G., Iken, K., Clarke, K. R., Trott, T., Konar, B., Cruz-Motta, J. J., et al. (2011). Aspects of benthic decapod diversity and distribution from rocky nearshore habitat at geographically widely dispersed sites. PLoS ONE, 6(4), e18606.
Queirolo, D., Gaete, E., Montenegro, I., Soriguer, M. C., & Erzini, K. (2012). Behaviour of fish by-catch in the mouth of a crustacean trawl. Journal of Fish Biology, 80(7), 2517–2527.
Reguera, B., Alonso, R., Moreira, A. & Méndez, S. 2011. Guía para el diseño y puesta en marcha de un plan de seguimiento de microalgas productoras de toxinas. IOC Manuals and Guides, no. 59 – (IOC/2011/MG/59). COI de UNESCO y OIEA, Paris. 65 p.
Remy-Zephir, B., Leleu, K., Grace, R. & Costello, M.J. (2012). Geographical information system (GIS) files of seabed habitats and biotope maps from Leigh marine reserve in 1977 and 2006. Accessed at Dryad data archive http://dx.doi.org/10.5061/dryad.6vr28
Reynolds, P. L., Richardson, J. P., & Duffy, J. E. (2014). Field experimental evidence that grazers mediate transition between microalgal and seagrass dominance. Limnology and Oceanography, 59(3), 1053–1064.
Saeedi, H., & Costello, M. J. (2012). Aspects of global distribution of six marine bivalve mollusc families. In F. da Costa (Ed.), Clam fisheries and aquaculture (pp. 27–44). New York: Nova Science Publishers Inc.
Santhanam, R. & Srinivasan, A. (1994). A manual of marine zooplankton. Oxford and IBH Pub. Co.
Shackell, N. L., & Frank, K. T. (2003). Marine fish diversity on the Scotian Shelf, Canada. Aquatic Conservation: Marine and Freshwater Ecosystems, 13(4), 305–321.
Sistiaga, M., Herrmann, B., Grimaldo, E., Larsen, R. B., & Tatone, I. (2015). Effect of lifting the sweeps on bottom trawling catch efficiency: A study based on the Northeast arctic cod (Gadus morhua) trawl fishery. Fisheries Research, 167, 164–173.
Stuart-Smith, R. D., Bates, A. E., Lefcheck, J. S., Duffy, J. E., Baker, S. C., Thomson, R. J., et al. (2013). Integrating abundance and functional traits reveals new global hotspots of fish diversity. Nature, 501(7468), 539–542.
Stuart-Smith, R. D., Edgar, G. J., Barrett, N. S., Kininmonth, S. J., & Bates, A. E. (2015). Thermal biases and vulnerability to warming in the world’s marine fauna. Nature, 528, 88–92.
Sudia, W. D., & Chamberlain, R. W. (1988). Battery-operated light trap, an improved model. Journal of the American Mosquito Control Association, 4(4), 536–538.
Szentkiralyi, F. (2002). Fifty year long insect survey in Hungary: T. Jermy’s contributions to light-trapping. Acta Zoologica Academiae Scientiarum Hungaricae, 1, 85–105.
Tait, R. V. & Dipper, F. (1998). Elements of marine ecology (462 p.). Oxford: Butterworth-Heinemann.
Thomsen, P. F., & Willerslev, E. (2015). Environmental DNA—an emerging tool in conservation for monitoring past and present biodiversity. Biological Conservation, 183, 4–18.
UNESCO. (2012). A framework for ocean observing. By the task team for an integrated framework for sustained ocean observing. UNESCO, IOC/INF-1284. doi:10.5270/OceanObs09-FOO
Vandepitte, L., Hernandez, F., Claus, S., Vanhoorne, B., De Hauwere, N., Deneudt, K., et al. (2011). Analysing the content of the European ocean biogeographic information system (EurOBIS): Available data, limitations, prospects and a look at the future. Hydrobiologia, 667, 1–14.
Vandepitte, L., Bosch, S., Tyberghein, L., Waumans, F., Vanhoorne, B., & Hernandez, F. et al. (2015a). Fishing for data and sorting the catch: Assessing the data quality, completeness and fitness for use of data in marine biogeographic databases. Database 2015.
Vandepitte, L., Vanhoorne, B., Decock, W., Dekeyzer, S., Trias Verbeeck, A., Bovit, L., et al. (2015b). How Aphia—The platform behind several online and taxonomically oriented databases—can serve both the taxonomic community and the field of biodiversity informatics. Journal of Marine Science and Engineering, 3(4), 1448–1473.
Venter, J. C., Remington, K., Heidelberg, J. F., Halpern, A. L., Rusch, D., Eisen, J. A., et al. (2004). Environmental genome shotgun sequencing of the Sargasso Sea. Science, 304(5667), 66–74.
Wahl, M., Link, H., Alexandridis, N., Thomason, J. C., Cifuentes, M., Costello, M. J., et al. (2011). Re-structuring of marine communities exposed to environmental change: A global study on the interactive effects of species and functional richness. PLoS ONE, 6(5), e19514.
Warwick, R. M., & Clarke, K. R. (1998). Taxonomic distinctness and environmental assessment. Journal of Applied Ecology, 35(4), 532–543.
Webb, T. J., & Mindel, B. L. (2015). Global patterns of extinction risk in marine and non-marine systems. Current Biology, 25(4), 506–511.
Weinberg, K. L., & Kotwicki, S. (2008). Factors influencing net width and sea floor contact of a survey bottom trawl. Fisheries Research, 93(3), 265–279.
Wilkinson, C., & Hill, J. (2004). Methods for ecological monitoring of coral reefs. Townsville, Australia: Australian Institute of Marine Science.
Winger, P. D., Eayrs, S., & Glass, C. W. (2010). Fish behaviour near bottom trawls. In P. He (Ed.), Behaviour of marine fishes: Capture processes and conservation challenges (pp. 67–103). Oxford, UK: Wiley-Blackwell.
Yasuhara, M., & Cronin, T. M. (2008). Climatic influences on deep-sea ostracode (Crustacea) diversity for the last three million years. Ecology, 89, S52–S65.
Yasuhara, M., Cronin, T. M., deMenocal, P. B., Okahashi, H., & Linsley, B. K. (2008). Abrupt climate change and collapse of deep-sea ecosystems. Proceedings of the National Academy of Sciences of the United States of America, 105, 1556–1560.
Yasuhara, M., Hunt, G., Cronin, T. M., & Okahashi, H. (2009). Temporal latitudinal-gradient dynamics and tropical instability of deep-sea species diversity. Proceedings of the National Academy of Sciences of the United States of America, 106, 21717–21720.
Yasuhara, M., Okahashi, H., Cronin, T. M., Rasmussen, T. L., & Hunt, G. (2014). Deep-sea biodiversity response to deglacial and Holocene abrupt climate changes in the North Atlantic Ocean. Global Ecology and Biogeography, 23, 957–967.
Yasuhara, M., Tittensor, D. P., Hillebrand, H., & Worm, B. (2015). Combining marine macroecology and palaeoecology in understanding biodiversity: Microfossils as a model. Biological Reviews. doi:10.1111/brv.12223.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is distributed under the terms of the Creative Commons Attribution-Noncommercial 2.5 License (http://creativecommons.org/licenses/by-nc/2.5/) which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
The images or other third party material in this chapter are included in the work’s Creative Commons license, unless indicated otherwise in the credit line; if such material is not included in the work’s Creative Commons license and the respective action is not permitted by statutory regulation, users will need to obtain permission from the license holder to duplicate, adapt or reproduce the material.
Copyright information
© 2017 The Author(s)
About this chapter
Cite this chapter
Costello, M.J. et al. (2017). Methods for the Study of Marine Biodiversity. In: Walters, M., Scholes, R. (eds) The GEO Handbook on Biodiversity Observation Networks. Springer, Cham. https://doi.org/10.1007/978-3-319-27288-7_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-27288-7_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-27286-3
Online ISBN: 978-3-319-27288-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)