Abstract
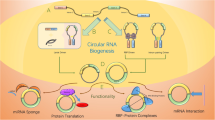
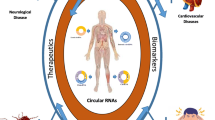
circRNAs have emerged as one of the key regulators in many cellular mechanisms and pathogenesis of diseases. However, with the limited knowledge and current technologies for circRNA investigations, there are several challenges that need to be addressed for. These include challenges in understanding the regulation of circRNA biogenesis, experimental designs, and sample preparations to characterize the circRNAs in diseases as well as the bioinformatics pipelines and algorithms. In this chapter, we discussed the above challenges and possible strategies to overcome those limitations. We also addressed the differences between the existing applications and technologies to study the circRNAs in diseases. By addressing these challenges, further understanding of circRNAs roles and regulations as well as the discovery of novel circRNAs could be achieved.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Haque S, Harries LW (2017) Circular RNAs (CircRNAs) in health and disease. Genes 8(12):1–17
Jeck WR, Sharpless NE (2014) Detecting and characterizing circular RNAs. Nat Biotechnol 32(5):453–461
Pamudurti NR, Bartok O, Jens M et al (2017) Translation of CircRNAs. Mol Cell 66(1):9–21 e27
Qu S, Liu Z, Yang X et al (2018) The emerging functions and roles of circular RNAs in cancer. Cancer Lett 414(2018):301–309
Tay Y, Rinn J, Pandolfi PP (2014) The multilayered complexity of ceRNA crosstalk and competition. Nature 505(7483):344–352
Lee SM, Kong HG, Ryu CM (2017) Are circular RNAs new kids on the block? Trends Plant Sci 22(5):357–360
Liu T, Zhang L, Chen G et al (2017) Identifying and characterizing the circular RNAs during the lifespan of Arabidopsis leaves. Front Plant Sci 8(July):1–9
Dou Y, Cha DJ, Franklin JL et al (2016) Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci Rep 6(October 2015):1–11
Meng S, Zhou H, Feng Z et al (2017) CircRNA: functions and properties of a novel potential biomarker for cancer. Mol Cancer 16(1):1–8
Wang Y, Wang Z (2015) Efficient backsplicing produces translatable circular mRNAs. RNA 21(2):172–179
Zhong Z, Lv M, Chen J (2016) Screening differential circular RNA expression profiles reveals the regulatory role of circTCF25-miR-103a-3p/miR-107-CDK6 pathway in bladder carcinoma. Sci Rep 6(November 2015):1–12
Zhang ZC, Guo XL, Li X (2018) The novel roles of circular RNAs in metabolic organs. Genes Dis 5(1):16–23
Yang P, Qiu Z, Jiang Y et al (2016) Silencing of cZNF292 circular RNA suppresses human glioma tube formation via the Wnt/β-catenin signaling pathway. Oncotarget 7(39):63449–63455
Wan L, Zhang L, Fan K et al (2016) Circular RNA-ITCH suppresses lung cancer proliferation via inhibiting the Wnt/β-Catenin pathway. Biomed Res Int 2016:1579490
Geng H-H, Li R, Su Y-M et al (2016) The circular RNA Cdr1as promotes myocardial infarction by mediating the regulation of miR-7a on its target genes expression. PLoS One 11(3):e0151753
Li F, Zhang L, Li W et al (2015) Circular RNA ITCH has inhibitory effect on ESCC by suppressing the Wnt/β-catenin pathway. Oncotarget 6(8):6001–6013
Kristensen LS, Hansen TB, Venø MT et al (2018) Circular RNAs in cancer: opportunities and challenges in the field. Oncogene 37(5):555–565
Nan A, Chen L, Zhang N et al (2017) A novel regulatory network among LncRpa, CircRar1, MiR-671 and apoptotic genes promotes lead-induced neuronal cell apoptosis. Arch Toxicol 91(4):1671–1684
Li Y, Zheng F, Xiao X, et al (2017) CircHIPK3 sponges miR-558 to suppress heparanase expression in bladder cancer cells. EMBO Rep:e201643581-e201643581.
Zheng Q, Bao C, Guo W et al (2016) Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun 7:1–13
Chen J (2016) Circular RNA profile identifies circPVT1 as a proliferative factor and prognostic marker in gastric cancer. (December):2016–2016
Hansen TB, Jensen TI, Clausen BH et al (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495:384
Wang Y, Liu J, Liu C et al (2013) MicroRNA-7 regulates the mTOR pathway and proliferation in adult pancreatic β-cells. Diabetes 62(3):887–895
Salzman J, Chen RE, Olsen MN et al (2013) Cell-type specific features of circular RNA expression. PLoS Genet 9(9):e1003777
Darbani B, Noeparvar S, Borg S (2016) Identification of circular RNAs from the parental genes involved in multiple aspects of cellular metabolism in barley. Front Plant Sci 7(June):1–8
Ye CY, Chen L, Liu C et al (2015) Widespread noncoding circular RNAs in plants. New Phytol 208(1):88–95
Sun X, Wang L, Ding J et al (2016) Integrative analysis of Arabidopsis thaliana transcriptomics reveals intuitive splicing mechanism for circular RNA. FEBS Lett 590(20):3510–3516
Han Y-N, Xia S-Q, Zhang Y-Y et al (2017) Circular RNAs: a novel type of biomarker and genetic tools in cancer. Oncotarget 8(38):64551–64563
Yang W, Du WW, Li X et al (2016) Foxo3 activity promoted by non-coding effects of circular RNA and Foxo3 pseudogene in the inhibition of tumor growth and angiogenesis. Oncogene 35(30):3919–3931
Ashwal-Fluss R, Meyer M, Pamudurti NR et al (2014) CircRNA biogenesis competes with pre-mRNA splicing. Mol Cell 56(1):55–66
Kramer MC, Liang D, Tatomer DC et al (2015) Combinatorial control of Drosophila circular RNA expression by intronic repeats, hnRNPs, and SR proteins. Genes Dev 29(20):2168–2182
Du WW, Yang W, Chen Y et al (2017) Foxo3 circular RNApromotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J 38(18):1402–1412
Du WW, Yang W, Liu E et al (2016) Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res 44(6):2846–2858
Braunschweig U, Barbosa-Morais NL, Pan Q et al (2014) Widespread intron retention in mammals functionally tunes transcriptomes. Genome Res 24(11):1774–1786
Yang Y, Fan X, Mao M et al (2017) Extensive translation of circular RNAs driven by N 6 -methyladenosine. Cell Res 27(5):626–641
Cui X, Niu W, Kong L et al (2016) hsa_circRNA_103636: potential novel diagnostic and therapeutic biomarker in major depressive disorder. Biomark Med 10(9):943–952
Guo X-Y, Chen J-N, Sun F et al (2017) circRNA_0046367 prevents Hepatoxicity of lipid peroxidation: an inhibitory role against hepatic steatosis. Oxid Med Cell Longev 2017:3960197
Hansen TB, Kjems J, Damgaard CK (2013) Circular RNA and miR-7 in Cancer. Cancer Res 73(18):5609–5612
Tang C-M, Zhang M, Huang L et al (2017) CircRNA_000203 enhances the expression of fibrosis-associated genes by derepressing targets of miR-26b-5p, Col1a2 and CTGF, in cardiac fibroblasts. Sci Rep 7:40342
Hansen TB, Venø MT, Damgaard CK et al (2016) Comparison of circular RNA prediction tools. Nucleic Acids Res 44(6):e58–e58
Szabo L, Salzman J (2016) Detecting circular RNAs: Bioinformatic and experimental challenges. Nat Rev Genet 17:679–692
Quail MA, Kozarewa I, Smith F et al (2008) A large genome centre’s improvements to the Illumina sequencing system. Nat Methods 5(12):1005–1010
Houseley J, Tollervey D (2010) Apparent non-canonical trans-splicing is generated by reverse transcriptase in vitro. PLoS One 5(8):e12271
Luo GX, Taylor J (1990) Template switching by reverse transcriptase during DNA synthesis. J Virol 64(9):4321–4328
Kelleher CD, Champoux JJ (1998) Characterization of RNA strand displacement synthesis by Moloney murine leukemia virus reverse transcriptase. J Biol Chem 273(16):9976–9986
Panda AC, De S, Grammatikakis I et al (2017) High-purity circular RNA isolation method (RPAD) reveals vast collection of intronic circRNAs. Nucleic Acids Res 45(12):e116–e116
Barrett SP, Salzman J (2016) Circular RNAs: analysis, expression and potential functions. Development 143(11):1838
Zheng Y, Zhao F (2018) Detection and reconstruction of circular RNAs from transcriptomic data. In: Dieterich C, Papantonis A (eds) Circular RNAs: methods and protocols. Springer, New York, pp 1–8. https://doi.org/10.1007/978-1-4939-7562-4_1
Jakobi T, Dieterich C (2018) Deep computational circular RNA analytics from RNA-seq data. In: Dieterich C, Papantonis A (eds) Circular RNAs: methods and protocols. Springer, New York, pp 9–25. https://doi.org/10.1007/978-1-4939-7562-4_2
Bachmayr-Heyda A, Reiner AT, Auer K et al (2015) Correlation of circular RNA abundance with proliferation – exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep 5:8057
Memczak S, Jens M, Elefsinioti A et al (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495:333
Zhang Y, Zhang X-O, Chen T et al (2013) Circular intronic long noncoding RNAs. Mol Cell 51(6):792–806
Zhang X-O, Wang H-B, Zhang Y et al (2014) Complementary sequence-mediated exon circularization. Cell 159(1):134–147
Guo JU, Agarwal V, Guo H et al (2014) Expanded identification and characterization of mammalian circular RNAs. Genome Biol 15(7):409
Jeck WR, Sorrentino JA, Wang K et al (2013) Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19(2):141–157
Cooper DA, Cortés-López M, Miura P (2018) Genome-wide circRNA profiling from RNA-seq data. In: Dieterich C, Papantonis A (eds) Circular RNAs: methods and protocols. Springer, New York, pp 27–41. https://doi.org/10.1007/978-1-4939-7562-4_3
Zhang Y, Liang W, Zhang P et al (2017) Circular RNAs: emerging cancer biomarkers and targets. J Exp Clin Cancer Res 36(1):152
Heumüller AW, Boeckel J-N (2018) Characterization and validation of circular RNA and their host gene mRNA expression using PCR. In: Dieterich C, Papantonis A (eds) Circular RNAs: methods and protocols. Springer, New York, pp 57–67. https://doi.org/10.1007/978-1-4939-7562-4_5
Panda AC, Abdelmohsen K, Gorospe M (2017) RT-qPCR detection of senescence-associated circular RNAs. Methods Mol Biol (Clifton, N.J.) 1534:79–87
Chen D-F, Zhang L-J, Tan K et al (2018) Application of droplet digital PCR in quantitative detection of the cell-free circulating circRNAs. Biotechnol Biotechnol Equip 32(1):116–123
Capel B, Swain A, Nicolis S et al (1993) Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 73(5):1019–1030
Schneider T, Schreiner S, Preußer C et al (2018) Northern blot analysis of circular RNAs. In: Dieterich C, Papantonis A (eds) Circular RNAs: methods and protocols. Springer New York, New York, pp 119–133. https://doi.org/10.1007/978-1-4939-7562-4_10
Kocks C, Boltengagen A, Piwecka M et al (2018) Single-molecule fluorescence in situ hybridization (FISH) of circular RNA CDR1as. In: Dieterich C, Papantonis A (eds) Circular RNAs: methods and protocols. Springer New York, New York, pp 77–96. https://doi.org/10.1007/978-1-4939-7562-4_7
Zirkel A, Papantonis A (2018) Detecting circular RNAs by RNA fluorescence in situ hybridization. In: Dieterich C, Papantonis A (eds) Circular RNAs: methods and protocols. Springer, New York, pp 69–75. https://doi.org/10.1007/978-1-4939-7562-4_6
Huang M-S, Zhu T, Li L et al (2018) LncRNAs and CircRNAs from the same gene: masterpieces of RNA splicing. Cancer Lett 415:49–57
Song X, Zhang N, Han P et al (2016) Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res 44(9):e87–e87
Izuogu OG, Alhasan AA, Alafghani HM et al (2016) PTESFinder: a computational method to identify post-transcriptional exon shuffling (PTES) events. BMC Bioinformatics 17(1):31
Hoffmann S, Otto C, Doose G et al (2014) A multi-split mapping algorithm for circular RNA, splicing, trans-splicing and fusion detection. Genome Biol 15(2):R34
Wang K, Singh D, Zeng Z et al (2010) MapSplice: accurate mapping of RNA-seq reads for splice junction discovery. Nucleic Acids Res 38(18):e178–e178
Chuang T-J, Wu C-S, Chen C-Y et al (2016) NCLscan: accurate identification of non-co-linear transcripts (fusion, trans-splicing and circular RNA) with a good balance between sensitivity and precision. Nucleic Acids Res 44(3):e29–e29
Gao Y, Wang J, Zhao F (2015) CIRI: an efficient and unbiased algorithm for de novo circular RNA identification. Genome Biol 16(1):4
Szabo L, Morey R, Palpant NJ et al (2015) Statistically based splicing detection reveals neural enrichment and tissue-specific induction of circular RNA during human fetal development. Genome Biol 16(1):126
Gao Y, Zhang J, Zhao F (2017) Circular RNA identification based on multiple seed matching. Brief Bioinform: bbx014–bbx014
Westholm JO, Miura P, Olson S et al (2014) Genomewide analysis of Drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell Rep 9(5):1966–1980
Cheng J, Metge F, Dieterich C (2016) Specific identification and quantification of circular RNAs from sequencing data. Bioinformatics 32(7):1094–1096
Gao Y, Zhao F (2018) Computational strategies for exploring circular RNAs. Trends Genet 34:389
Zeng X, Lin W, Guo M et al (2017) A comprehensive overview and evaluation of circular RNA detection tools. PLoS Comput Biol 13(6):e1005420
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with bowtie 2. Nat Methods 9(4):357–359
Langmead B, Trapnell C, Pop M et al (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10(3):R25–R25
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows–wheeler transform. Bioinformatics 25(14):1754–1760
Dobin A, Davis CA, Schlesinger F et al (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21
Hansen TB (2018) Improved circRNA identification by combining prediction algorithms. Front Cell Dev Biol 6:20
Baruzzo G, Hayer KE, Kim EJ et al (2017) Simulation-based comprehensive benchmarking of RNA-seq aligners. Nat Methods 14(2):135–139
Kim D, Pertea G, Trapnell C et al (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14(4):R36
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25(9):1105–1111
Glažar P, Papavasileiou P, Rajewsky N (2014) circBase: a database for circular RNAs. RNA 20(11):1666–1670
Ghosal S, Das S, Sen R et al (2013) Circ2Traits: a comprehensive database for circular RNA potentially associated with disease and traits. Front Genet 4:283
Liu Y-C, Li J-R, Sun C-H et al (2016) CircNet: a database of circular RNAs derived from transcriptome sequencing data. Nucleic Acids Res 44(Database issue):D209–D215
Gao Y, Wang J, Zheng Y et al (2016) Comprehensive identification of internal structure and alternative splicing events in circular RNAs. Nat Commun 7:12060
Metge F, Czaja-Hasse LF, Reinhardt R et al (2017) FUCHS—towards full circular RNA characterization using RNAseq. PeerJ 5:e2934
Meng X, Chen Q, Zhang P et al (2017) CircPro: an integrated tool for the identification of circRNAs with protein-coding potential. Bioinformatics 33(20):3314–3316
Feng J, Xiang Y, Xia S, et al (2017) CircView: a visualization and exploration tool for circular RNAs. Brief Bioinform:bbx070-bbx070
Dudekula DB, Panda AC, Grammatikakis I et al (2016) CircInteractome: a web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol 13(1):34–42
Grant GR, Farkas MH, Pizarro AD et al (2011) Comparative analysis of RNA-Seq alignment algorithms and the RNA-Seq unified mapper (RUM). Bioinformatics 27(18):2518–2528
Streit S, Michalski CW, Erkan M et al (2008) Northern blot analysis for detection and quantification of RNA in pancreatic cancer cells and tissues. Nat Protoc 4:37
PCvdC M, Roeland WD, PMvG R et al (1998) Sensitive mRNA detection by fluorescence in situ hybridization using horseradish peroxidase-labeled oligodeoxynucleotides and tyramide signal amplification. J Histochem Cytochem 46(11):1249–1259
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Sulaiman, S.A., Abdul Murad, N.A., Mohamad Hanif, E.A., Abu, N., Jamal, R. (2018). Prospective Advances in Circular RNA Investigation. In: Xiao, J. (eds) Circular RNAs. Advances in Experimental Medicine and Biology, vol 1087. Springer, Singapore. https://doi.org/10.1007/978-981-13-1426-1_28
Download citation
DOI: https://doi.org/10.1007/978-981-13-1426-1_28
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-1425-4
Online ISBN: 978-981-13-1426-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)