Abstract
Hypoxia in tumors is primarily a pathophysiologic consequence of structurally and functionally disturbed microcirculation with inadequate supply of oxygen. Tumor hypoxia is strongly associated with tumor propagation, malignant progression, and resistance to therapy. Aberrant epigenetic regulation plays a crucial role in the process of hypoxia-driven malignant progression. Convert of 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC) by ten-eleven translocation (TET) family enzymes plays important biological functions in embryonic stem cells, development, aging and disease. Recent reports showed that level of 5hmC and TET proteins was altered in various types of cancers. There is a strong correlation between loss of 5hmC and cancer development but research to date indicates that loss of TET activity is associated with the cancer phenotype but it is not clear whether TET proteins function as tumor suppressors or oncogenes. While loss of TET1 and TET2 expression is associated with solid cancers, implying a tumor suppressor role, TET1 exhibits a clear oncogenic role in the context of genomic rearrangements such as in MLL-fusion rearranged leukemia. Interestingly, hypoxia increases global 5hmC levels and upregulates TET1 expression in a HIF1α-dependent manner. Recently, hypoxia-induced TET1 has been demonstrated to play another important role for regulating hypoxia-responsive gene expression and epithelial-mesenchymal transition (EMT) by serving as a transcription co-activator. Furthermore, hypoxia-induced TET1 also regulates glucose metabolism and hypoxia-induced EMT through enhancing the expression of insulin induced gene 1 (INSIG1). The roles and mechanisms of action of 5hmC and TET proteins in ES cell biology and during embryonic development, as well as in cancer biology, will be the main focus in this review.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1. Introduction
Insufficient oxygen availability, so-called hypoxia, is a microenvironmental event that plays a critical role in various biological processes including development, metabolism, inflammation, tumor progression and cancer stemness [1, 2]. DNA methylation is one of the epigenetic modifications that plays important roles in numerous cellular processes, including genomic imprinting, X-chromosome inactivation, regulation of gene expression, and maintenance of epigenetic memory [3]. Compared with reversible modifications of histone proteins, DNA methylation was a relatively stable epigenetic modification. The recent finding that teneleven translocation (TET) proteins are 5-methylcytosine (5mC) oxidases has provided the mechanism for the reversal of DNA methylation. In the 1980’s, global hypomethylation was first observed in colorectal cancer cell lines [4]. In addition, genomewide DNA hypomethylation during tumor hypoxia also have been observed [5, 6]. Moreover, several reports demonstrated that hypoxia would enhance TET1 expression and lead to global DNA hypomethylation in different type of cancer [7-9]. Indeed, TET proteins might participate in hypoxia-mediated hypomethylation and lead to enhanced malignancy. Therefore, genome-wide maps of methylation must be systemically established to study the correlation between TET proteins and tumor progression. In this review, we discuss the current knowledge about the mechanism and functions of DNA methylation and TET protein. We also review our current understanding of epigenetic alterations that take place under hypoxia. We then discuss the role of TET proteins and DNA demethylation under hypoxia in epithelial-mesenchymal transition and tumorigenesis.
2. Epigenetics and DNA methylation
The important role of epigenetic processes in cancer progression and treatment has been emphasized for the past decades. Epigenetic alterations are leading candidates for cancer detection, diagnosis and prognosis [10]. Cancer research in epigenetics in the 1990s was led by a focus on the discoveries and understanding of DNA methylation abnormalities in the 1980s [11]. Abnormal gains of DNA methylation in normally unmethylated gene promoters, leading to transcriptional repression or loss of gene function, are the most widely studied epigenetic alterations in cancer [12]. Moreover, in certain cancers, there is an increasing list of candidate tumor suppressor genes that are silenced by hypermethylation in their promoters [13]. In addition, global hypomethylation has also been implicated in the development and progression of cancer [14]. DNA methylation is established by DNA methyltransferases (DNMTs). The classical model of methylation, which was presented more than 30 years ago, involves two types of DNMTs: maintenance and de novo DNMTs [15, 16]. DNA demethylation involves the removal of the methyl group from 5mC in DNA. This process occurs through two pathways, the passive or active demethylation pathways [17]. Passive mechanisms involve a failure of the repair system to maintain DNA methylation patterns during replication or DNA synthesis and are associated with the dilution of hemimethylated CpG in subsequent replication cycles. Active DNA demethylation involves the replacement of 5mC by cytosine [18, 19]. It has been well recognized that the methyl group of 5mC can also be removed regardless of replication, particularly in the zygotic paternal genome or primordial germ cells. However, the mechanism that causes the active demethylation has remained enigmatic until the function of TET enzymes was identified recently (Figure 1) [20]. Subsequent studies have revealed new visions in the field of DNA methylation and raise many questions about the mechanism and contribution of TET proteins to development and cancer.
Mechanism of passive and active DNA demethylation. Passive DNA demethylation was thought to occur by a reduction in activity or absence of DNMTs. Active DNA demethylation involves 2 different enzyme families: (1) 5-methylcytosine (5mC) is hydroxylated by TET to form 5 hydroxymethylcytosine (5hmC) and further oxidize 5hmC to 5-formylcytosine (5fC) and 5-carboxycytosine (5caC). (2) Thymine DNA glycosylase (TDG) converted and repaired 5fC/5caC to unmodified cytosines.
3. Domain structures of TET family proteins
The mammalian TET family contains three members, TET1, TET2 and TET3, all of which share a high degree of homology within their C-terminal catalytic domain - CD domain (Cys-rich and DSBH regions) that belongs to the Cupin-like dioxygenase superfamily and exhibits 2-oxoglutarate (2-OG) - and iron (II)- dependent dioxygenase activity [20, 21]. TET proteins oxidize 5mC into 5-hydroxymethylcytosin (5hmC) through these CD domains and require α-ketoglutarate as a co-substrate for enzymatic activity. Subsequent studies revealed the ability of TET family proteins to further oxidize 5hmC to 5-formylcytosine (5fC) and 5-carboxycytosine (5caC) (Figure 1) [22, 23]. Another mark of TET family proteins is the CXXC zinc-finger domain, which was first identified and defined in DNMT1 [24]. The CXXC domain would be found only in the N-terminus of TET1 and TET3 but not in TET2. Although CXXC domain is known to distinguish between methylated and unmethylated DNA [25], the function of this domain in TET1 and TET3 is largely unknown. It is known that the CXXC domain of TET1 recognizes not only unmodified cytosine but also 5mC and 5hmC, and it favors binding to regions in the genome of high CpG content [26]. In addition to the described functional domains, there is a spacer region that links the two parts of the disconnected DSBH enzymatic domain. This unique spacer region is common to all TET family members, although its length varies. The functional importance of this spacer region is currently unknown (Figure 2). In summary, TET family proteins, through their CD domains, have an enzymatic capability to convert 5mC to 5hmC, 5fC or 5caC depending on the company of co-factors, such as ATP. Furthermore, except CD domain and CXXC domain, TET proteins also contain various additional domains with as yet undefined functions. Its potential regulatory function for modulating TET enzymatic activities deserves future investigation.
Structure and function of TET family proteins. The TET protein family contains three members: TET1, TET2 and TET3. All three members have a C-terminal CD domain (containing the Cys-rich and DSBH regions) that obtains 2-oxoglutarate (2-OG)- and iron (II)-dependent dioxygenase activity. The CD domain also includes a spacer region, the length of which varies between TET family proteins and its function remains unknown. The N-termini of TET1 and TET3, but not TET2, contain a CXXC domain, which mediates their direct DNA-binding ability.
4. The biological function of TET proteins
Although all TET family members possess 5mC oxidation activity, their expression levels are very different in various cell types and tissues. For example, TET1 and TET2 are highly expressed in mouse ES cells, but TET3 is more enriched in oocytes and onecell zygotes [21]. Evidence suggested that oxidation of 5mC in the paternal genome in fertilized eggs by TET3 initiates DNA demethylation and facilitates the activation of the paternal copy of early embryonic genes [27]. Concomitant with the rapid reduction of TET3 expression at the two-cell stage, the expression of TET1 is rapidly up-regulated at later stage [28, 29]. Result of genome-wide location analyses in mouse ES cells indicated that TET1 and 5hmC are enriched at promoter regions of several pluripotency factors, including Nanog, Tcl1, and Esrrb [30]. Several genes related to pluripotency are also down-regulated in the double knockdown of TET1 and TET2 of mouse ES cells [31]. Recently, Jaenisch and colleagues generated the TET1-null mouse ES cells and mice to further study the function of TET1 in ES cell maintenance and in vivo development, and discovered that TET1 null ES cells maintain their self-renewal ability under mouse ES cell culture conditions in vitro and develop normally in vivo [32]. Furthermore, TET1 knockout mice and TET2 knockout mice appear to develop normally, and appear healthy through adulthood and are fertile [32, 33]. It is still unclear that the purpose of TET1/TET2 and 5hmC are maintained at high levels in mouse ES cells. As such, the biological role of TET1/TET2 or 5hmC in embryonic development remains unclear. Furthermore, evidence suggests a role for TET enzymes in the generation of pluripotent stem cells (iPSCs) that are phenotypically similar to embryonic stem cells (ESCs) [34]. As shown recently, formation of 5hmC is very important for brain development. In DNA from human brain cortex, the level of 5hmC is about 1% of all cytosines or 20 to 25% of all 5mC bases [35]. TET3 is most highly expressed in the developing mouse brain cortex followed by TET2, while the levels of TET1 are very low in this tissue. An increase in the levels of TET2, TET3 and 5hmC in differentiating neurons corresponds to a decrease in the Polycomb histone H3 lysine 27 (H3K27)-specific methyltransferase EZH2 and loss of H3K27me3 marker at gene promoter. Besides, decreasing the levels of TET2 and TET3 or increasing EZH2 expression results in imperfect neuronal differentiation [36]. Thus, formation of 5hmC promotes neuronal differentiation by modulating the expression of genes most critical in this important developmental transition.
5. TET proteins in cancer
In 2011, a chemical labeling technique for determining the genome-wide distribution of 5hmC in human cell lines, as well as in the mouse cerebellum, was developed. A genome-wide study observed that an enrichment of 5hmC in genes linked to hypoxia and angiogenesis [37]. Aberrant DNA methylation is a hallmark of cancer; growing evidence has suggested that an imbalance in TET-mediated DNA demethylation may participate in carcinogenesis. The first reports implicating a role for TET proteins in cancer showed that TET1 is fused to the mixed lineage leukemia (MLL) gene in a case of pediatric AML containing the t(10;11) (q22;q23) [38]. Recent report indicated that TET1 is significantly up-regulated in MLL-rearranged leukemia and is a direct target gene of MLL-fusion proteins. MLL fusions would bind to the promoter region of TET1 to promote its expression directly and result in a global increase of 5hmC. Moreover, together with MLL, TET1 activates the homeobox A9 (Hoxa9)/myeloid ecotropic viral integration 1 (Meis1)/pre-B-cell leukemia homeobox 3 (Pbx3) signaling pathway, which subsequently promotes cell proliferation and inhibits apoptosis/cell differentiation, thereby leading to cell transformation and leukemogenesis [39]. In contrast, many mutations, including nonsense/missense, deletions and frameshift somatic in TET2 were identified in myelodysplastic syndrome (MDS) and other types of leukemia [40, 41]. Some of TET2 mutations could damage the catalytic activity, but many mutations appear unrelated to enzymatic activity [42]. Therefore, the molecular mechanism underlying loss-of-function mutation in TET2 is still unknown and requires further investigation. In addition to AML, TET2 mutations and/or deletions have been observed in other types of tumor, including bladder, breast, kidney, liver, lung and uterine cancers [43]. Since the role of mutated TET proteins in solid cancers has not yet been firmly established, it should be possible to resolve this point by crossing the many available TET-deficient mouse models with the various mouse cancer models available. More importantly, the up-regulation or downregulation of TET gene expression, which is often associated with 5hmC levels, has been observed in numerous solid cancers (Table 1). So far, a direct correlation between TET3 and cancer has not been reported. A possible explanation for this is that TET3 is the most important regulator and critical TET3 mutations or dysregulation might lead to lethality.
6. The role of hypoxia and hypoxia-induced factors in tumor progression
The definition of hypoxia is that a reduction of tissue oxygen tension compares to the normal level. Hypoxia usually occurs during acute and chronic vascular disease, pulmonary disease and cancer [44]. Although hypoxia is toxic to both cancer cells and normal cells, cancer cells undergo genetic and adaptive changes to promote cell survival and even proliferate in a hypoxic environment [45]. Hypoxic tumors are associated with a poor prognosis and resistance to treatments. Well-oxygenated tumor cells have a threefold greater sensitivity to radiation than hypoxic cells [46]. Recently, hypoxia has been identified as an important factor that is correlated with tumor progression including an increasing probability of recurrence, locoregional spread, and distant metastasis [47]. Furthermore, recent studies suggest that tumor hypoxia is associated with malignant biological phenotype such as angiogenesis, migration, invasion and metastasis [48]. The key factor that is involved in adaptive responses to cellular hypoxia is hypoxiainducible factor-1 (HIF-1) and its activity is tightly regulated by the cellular oxygen tension [49, 50]. HIF-1 is a heterodimeric transcription factor that constitutes of a hypoxia-inducible HIF- 1α subunit and a ubiquitously expressed HIF-1β subunit. HIF-1 binds to a 5’-ACGTG-3’ hypoxia-response element (HRE) in the promoter or enhancer of various hypoxia-inducible genes specifically under hypoxia [51]. HIF-1α has been identified as a major regulator of adaptation to hypoxia and implicated in the malignant progression of cancers [52]. Many studies have associated hypoxia and HIF-1α expression with cancer progression. The increased HIF-1α protein level is associated with patient mortality, poor prognosis and treatment resistance in head-and-neck, ovary, colon, breast and lung cancers [53].
7. Hypoxia-induced epigenetics
Epigenetic regulation plays an important role in regulating transcriptional changes in hypoxia. Hypoxia in tumor cells display a decrease in the levels of histone acetylation that is associated global transcriptional repression [54]. HDAC expression and activity have been shown to be up-regulated in hypoxia [55]. Hypoxia treatment induces HDAC1 activity and expression. Treatment with TSA, a specific HDAC inhibitor, inhibits hypoxia-induced angiogenesis in the Lewis lung carcinoma model [55]. Some evidences indicate that hypoxia enhances HDACs function and HDACs are implicated in hypoxia-induced metastasis through suppression of hypoxia-responsive tumor suppressor genes [56]. HDAC1 cooperates with HIF-1 downstream target and contribute to suppression of STAT1 [57]. HDAC7 has been found to interact with HIF-1 under hypoxia and increases transcriptional activity of HIF-1 [58]. HDAC4 and 6 also have been found to complex with HIF-1 and associate with HIF-1 stability and transcriptional activity [59]. Under hypoxia, HDAC3, directly activated by HIF-1, would interact with hypoxia-induced WDR5 and recruits the histone methyltransferase (HMT) complex to increase histone H3 lysine 4 (H3K4)-specific HMT activity [60]. Furthermore, treatment with HDAC inhibitor represses HIF-1 induction in response to hypoxia. Hypoxia stimulated proliferation, invasion, migration, and neovascularization are suppressed by treatment with HDAC inhibitors [61]. The O2, Fe(II), and α-ketoglutaratedependent jumonji domain (JMJD) histone demethylases are transcriptionally upregulated in hypoxia, and global changes in many histone modifications, especially at the sites of H3 lysine 4, 9, and 36, have also been reported [62-64]. Site-specific changes in histone modifications have been observed at hypoxia-induced genes including CA9, LDHA, and PDK1 [65, 66]. These results support the regulation of hypoxia-responsive gene expression through various histone modifications mediated by histone modifiers.
8. DNA methylation status under hypoxia
Recent studies have demonstrated that oxygen levels significantly influence a change in another epigenetic mark, DNA methylation. Hypermethylation of CpG islands in the promoter region can block the expression of HIF-mediated gene expression in hypoxic cells. For example, Bcl-2/adenovirus E1B 19 kDa interacting protein 3 (BNIP3), a regulator of hypoxia-induced cell death, was found to be repressed by DNA methylation in pancreatic, colorectal and gastric cancer [67, 68]. Prolyl hydroxylase (PHD1, PHD2 and PHD3) and factor inhibiting HIF-1 (FIH) had been known as negative regulators of HIF-1 [69]. In the invasive breast carcinoma and colorectal cancer, only PHD3 expression was associated with increased DNA methylation levels in the CpG islands of its promoter [70, 71]. Different from the hypoxia-induced DNA hypermethylation observed at certain loci, hypoxia has been linked to a global reduction in DNA methylation. DNA hypomethylation during hypoxia by examining the amount of 5mC by HPLC was reported in human colorectal and melanoma cell lines [6]. Specifically, CAIX overexpression has been associated with promoter DNA hypomethylation in gastric cancer, and CAIX expression correlates with tumor advancement and metastasis [72]. Moreover, in human hepatoma cell lines, hypoxia induces genomic DNA demethylation through the direct activation of Methionine adenosyltransferase 2A (MAT2A) that maintains the S-adenosylmethionine (SAM)/S-adenosylhomocysteine (SAH) ratio, a critical marker of genomic methylation status [73]. Tumor-associated CpG demethylation leads to increased HIF-1 binding to the HRE and enhanced HIF-1-mediated effects on tumor progression in the HCT116 colon cancer cell line [74]. Human cardiac fibroblast cells exposed to prolonged 1% hypoxia caused a pro-fibrotic state, which was associated with global DNA hypermethylation and increased expression of the DNMTs, such as DNMT1 and DNMT3B [75]. In contrast, down-regulation of DNMT1, DNMT3A and DNMT3B was reported, which contributed to DNA hypomethylation at the proximal promoter region of p16INK4a under hypoxia in human colorectal cancer (HCT116, 379.2) cell lines [76]. These findings support that epigenetic modification, whether global or sitespecific DNA methylation, plays a role in regulation gene expression under hypoxia-induced tumor progression.
9. Hypoxia-induced TET proteins upregulation
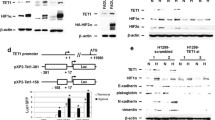
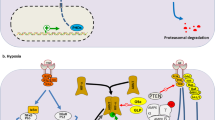
Hypoxia has also been linked to the HIF-dependent up-regulation of TET1, which catalyzes the hydroxylation of 5mC to 5hmC in tumorigenic N-type neuroblastoma cells exposed to 1% oxygen. TET1 activity essentially leads to DNA demethylation and production of 5hmC, a modification that is associated with active transcription [7]. Recently, Wu et al. found that hypoxia would regulate TET1 and TET3 expression through HIF-1α, leading to increased level of global DNA hydroxymethylation that was associated with tumor malignancy in the breast cancer. Furthermore, the results also demonstrated that hypoxia-induced TET1/TET3 proteins made a great contribution to the activation of the TNFα- p38-MAPK pathway in regulating cancer stemness. Histological analysis demonstrated that the levels of 5hmC, TET1 and TET3 were significantly associated with tumor hypoxia, tumor aggressiveness and poor prognosis [9]. TET1 up-regulation leading to global DNA hypomethylation in a HIF-independent manner was also demonstrated in scleroderma fibroblasts [77]. However, the unconventional role of TET proteins in transcription regulation independent of its enzymatic activity has been described in other studies. Many of the genomic regions are inactivated by TET1 through recruitment of the polycomb repressive complex 2 (PRC2), which catalyzes the formation of H3K27me3 and represses gene transcription [78], or it can directly bind to Sin3a histone deacetylase repressive complex to inhibit transcription [79]. Moreover, several groups reported that TET2 and TET3 play an essential part in recruiting O-linked b-N-acetylglucosamine transferase (OGT) to H3K4me3-positive CpG-rich promoters, thus enabling O-GlcNAc modification of histones [80-82]. These findings demonstrate the potential for crosstalk between TET proteins and pathways involved in glucose metabolism. Aberrant glucose metabolism in cancer cells may alter O-GlcNAcylation of TET proteins and therefore affect their stability. Recently, Tsai et al. explores the role of TET1 under hypoxia and also proves that TET1 plays another role in serving as a transcriptional coactivator and interacts with HIF-1α and HIF-2α to enhance their transactivation activity independent of its enzymatic activity [8]. It was also reported that TET1 may form a complex with HIF-1/CBP, or with OGT, to facilitate hypoxia-mediated gene expressions [8]. Combined results of RNA sequencing and 5hmC sequencing comparing TET1 knockdown cells under normoxia with those under hypoxia showed that 98 genes were regulated by TET1 and also had increased levels of 5hmC in their promoters. INSIG1 (insulin induced gene 1), a major regulator of cholesterol biosynthesis [83, 84], was inside this list. Furthermore, knockdown of TET1 or INSIG1 diminished a set of hypoxia-induced genes involved in epithelial-mesenchymal transition (EMT), providing a link between lipid metabolism and EMT [8]. Because of the capability of INSIG1 to inhibit cholesterol biosynthesis, it may inhibit lipid synthesis and turn to glucose utilization similar to the role of AMPK that also inhibits cholesterol synthesis [85]. These results demonstrated that the activation of INSIG1 expression through hypoxia-induced TET1 also contributes to Warburg effect, linking hypoxia-regulated TET1/5hmC to metabolism and EMT [8].
10. Conclusion and perspectives
The discovery of TET enzymes and the oxidative derivatives of 5mC are a great step for epigenetic research. Studies in the past few years focused on the role of active DNA demethylation in development and disease. Therefore, not surprisingly, changes in TET expression and 5hmC levels have been observed for numerous cancers. However, a deficiency in understanding the mechanism of decreased 5hmC levels and its role in transcriptional control during tumor malignancy still exists. Moreover, TET proteins are sources of 5fC and 5caC. TET protein expression levels and activities as well as the possible readers of all oxidized 5mC derivatives in cancer should be investigated. A potential link between TET proteins and cancer will open a new road for identifying potential therapeutic tools to treat cancer. Moreover, it is now well established that TET proteins are mutated or their expression levels or activities are dysregulated in numerous cancers. Furthermore, investigations are needed to distinguish the catalytic activity-dependent and -independent functions of TET proteins in order to offer new strategies for anti-cancer drug development and cancer therapy.
References
Bertout JA, Patel SA, Simon MC. The impact of O2 availability on human cancer. Nat Rev Cancer 2008; 8: 967–75.
Semenza GL. Hypoxia-inducible factors: mediators of cancer progression and targets for cancer therapy. Trends Pharmacol Sci. 2012; 33: 207–14.
Bird A. DNA methylation patterns and epigenetic memory. Genes Dev 2002; 16: 6–21.
Feinberg AP, Vogelstein B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature 1983; 301: 89–92.
Pal A, Srivastava T, Sharma MK, Mehndiratta M, Das P, Sinha S, et al. Aberrant methylation and associated transcriptional mobilization of Alu elements contributes to genomic instability in hypoxia. J Cell Mol Med 2010; 14: 2646–54.
Shahrzad S, Bertrand K, Minhas K, Coomber BL. Induction of DNA hypomethylation by tumor hypoxia. Epigenetics 2007; 2: 119–25.
Mariani CJ, Vasanthakumar A, Madzo J, Yesilkanal A, Bhagat T, Yu Y, et al. TET1-mediated hydroxymethylation facilitates hypoxic gene induction in neuroblastoma. Cell Rep 2014; 7: 1343–52.
Tsai YP, Chen HF, Chen SY, Cheng WC, Wang HW, Shen ZJ, et al. TET1 regulates hypoxia-induced epithelial-mesenchymal transition by acting as a co-activator. Genome Biol 2014; 15: 513.
Wu MZ, Chen SF, Nieh S, Benner C, Ger LP, Jan CI, et al. Hypoxia Drives Breast Tumor Malignancy through a TET-TNFalpha-p38- MAPK Signaling Axis. Cancer Res 2015; 75: 3912–24.
Baylin SB, Jones PA. A decade of exploring the cancer epigenome - biological and translational implications. Nat Rev Cancer 2011; 11: 726–34.
Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev Genet 2002; 3: 415–28.
Jones PA, Baylin SB. The epigenomics of cancer. Cell 2007; 128: 683–92.
Esteller M. CpG island hypermethylation and tumor suppressor genes: a booming present, a brighter future. Oncogene 2002; 21: 5427–40.
Ehrlich M. DNA hypomethylation in cancer cells. Epigenomics 2009; 1: 239–59.
Holliday R, Pugh JE. DNA modification mechanisms and gene activity during development. Science 1975; 187: 226–32.
Riggs AD. X inactivation, differentiation, and DNA methylation. Cytogenet Cell Genet 1975; 14: 9–25.
Ehrlich M, Lacey M. DNA hypomethylation and hemimethylation in cancer. Adv Exp Med Biol. 2013; 754: 31–56.
Shen L, Song CX, He C, Zhang Y. Mechanism and function of oxidative reversal of DNA and RNA methylation. Annu Rev Biochem 2014; 83: 585–614.
Wu H, Zhang Y. Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell 2014; 156: 45–68.
Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y, et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science. 2009; 324(5929): 930–5.
Ito S, D’Alessio AC, Taranova OV, Hong K, Sowers LC, Zhang Y. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell selfrenewal and inner cell mass specification. Nature 2010; 466: 1129–33.
Ito S, Shen L, Dai Q, Wu SC, Collins LB, Swenberg JA, et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 2011; 333: 1300–3.
He YF, Li BZ, Li Z, Liu P, Wang Y, Tang Q, et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 2011; 333: 1303–7.
Bestor TH. Activation of mammalian DNA methyltransferase by cleavage of a Zn binding regulatory domain. EMBO J 1992; 11: 2611–7.
Allen MD, Grummitt CG, Hilcenko C, Min SY, Tonkin LM, Johnson CM, et al. Solution structure of the nonmethyl-CpG-binding CXXC domain of the leukaemia-associated MLL histone methyltransferase. EMBO J 2006; 25: 4503–12.
Zhang H, Zhang X, Clark E, Mulcahey M, Huang S, Shi YG. TET1 is a DNA-binding protein that modulates DNA methylation and gene transcription via hydroxylation of 5-methylcytosine. Cell Res 2010; 20: 1390–3.
Gu TP, Guo F, Yang H, Wu HP, Xu GF, Liu W, et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 2011; 477: 606–10.
Iqbal K, Jin SG, Pfeifer GP, Szabo PE. Reprogramming of the paternal genome upon fertilization involves genome-wide oxidation of 5-methylcytosine. Proc Natl Acad Sci USA 2011; 108: 3642–7.
Wossidlo M, Nakamura T, Lepikhov K, Marques CJ, Zakhartchenko V, Boiani M, et al. 5-Hydroxymethylcytosine in the mammalian zygote is linked with epigenetic reprogramming. Nat Commun 2011; 2: 241.
Wu H, Zhang Y. Tet1 and 5-hydroxymethylation: a genome-wide view in mouse embryonic stem cells. Cell Cycle 2011; 10: 2428–36.
Ficz G, Branco MR, Seisenberger S, Santos F, Krueger F, Hore TA, et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 2011; 473: 398–402.
Dawlaty MM, Ganz K, Powell BE, Hu YC, Markoulaki S, Cheng AW, et al. Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 2011; 9: 166–75.
Ko M, Bandukwala HS, An J, Lamperti ED, Thompson EC, Hastie R, et al. Ten-Eleven-Translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc Natl Acad Sci USA 2011; 108: 14566–71.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006; 126: 663–76.
Jin SG, Jiang Y, Qiu R, Rauch TA, Wang Y, Schackert G, et al. 5-Hydroxymethylcytosine is strongly depleted in human cancers but its levels do not correlate with IDH1 mutations. Cancer Res 2011; 71: 7360–5.
Hahn MA, Qiu R, Wu X, Li AX, Zhang H, Wang J, et al. Dynamics of 5-hydroxymethylcytosine and chromatin marks in Mammalian neurogenesis. Cell Rep 2013; 3: 291–300.
Song C-X, Szulwach KE, Fu Y, Dai Q, Yi C, Li X, et al. Selective chemical labeling reveals the genome-wide distribution of 5-hydroxymethylcytosine. Nat Biotech 2011; 29: 68–72.
Lorsbach RB, Moore J, Mathew S, Raimondi SC, Mukatira ST, Downing JR. TET1, a member of a novel protein family, is fused to MLL in acute myeloid leukemia containing the t(10; 11)(q22; q23). Leukemia 2003; 17: 637–41.
Huang H, Jiang X, Li Z, Li Y, Song CX, He C, et al. TET1 plays an essential oncogenic role in MLL-rearranged leukemia. Proc Natl Acad Sci USA 2013; 110: 11994–9.
Langemeijer SM, Kuiper RP, Berends M, Knops R, Aslanyan MG, Massop M, et al. Acquired mutations in TET2 are common in myelodysplastic syndromes. Nat Genet 2009; 41: 838–42.
Delhommeau F, Dupont S, Della Valle V, James C, Trannoy S, Masse A, et al. Mutation in TET2 in myeloid cancers. N Engl J Med 2009; 360: 2289–301.
Ko M, Huang Y, Jankowska AM, Pape UJ, Tahiliani M, Bandukwala HS, et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 2010; 468: 839–43.
Wang Y, Xiao M, Chen X, Chen L, Xu Y, Lv L, et al. WT1 recruits TET2 to regulate its target gene expression and suppress leukemia cell proliferation. Mol Cell 2015; 57: 662–73.
Harris AL. Hypoxia—a key regulatory factor in tumour growth. Nat Rev Cancer 2002; 2: 38–47.
Vaupel P. The role of hypoxia-induced factors in tumor progression. Oncologist. 2004; 9 Suppl 5: 10–7.
Gray LH, Conger AD, Ebert M, Hornsey S, Scott OC. The concentration of oxygen dissolved in tissues at the time of irradiation as a factor in radiotherapy. Br J Radiol 1953; 26: 638–48.
Brown JM. Exploiting the hypoxic cancer cell: mechanisms and therapeutic strategies. Mol Med Today 2000; 6: 157–62.
Brown JM, Wilson WR. Exploiting tumour hypoxia in cancer treatment. Nat Rev Cancer 2004; 4: 437–47.
Wang GL, Semenza GL. Characterization of hypoxia-inducible factor 1 and regulation of DNA binding activity by hypoxia. J Biol Chem 1993; 268: 21513–8.
Semenza GL. Regulation of mammalian O2 homeostasis by hypoxiainducible factor 1. Annu Rev Cell Dev Biol 1999; 15: 551–78.
Semenza GL. HIF-1, O(2), and the 3 PHDs: how animal cells signal hypoxia to the nucleus. Cell 2001; 107: 1–3.
Semenza GL. Targeting HIF-1 for cancer therapy. Nat Rev Cancer 2003; 3: 721–32.
Zhong H, De Marzo AM, Laughner E, Lim M, Hilton DA, Zagzag D, et al. Overexpression of hypoxia-inducible factor 1alpha in common human cancers and their metastases. Cancer Res. 1999; 59: 5830–5.
Johnson AB, Denko N, Barton MC. Hypoxia induces a novel signature of chromatin modifications and global repression of transcription. Mutat Res 2008; 640: 174–9.
Kim MS, Kwon HJ, Lee YM, Baek JH, Jang JE, Lee SW, et al. Histone deacetylases induce angiogenesis by negative regulation of tumor suppressor genes. Nat Med 2001; 7: 437–43.
Liu T, Kuljaca S, Tee A, Marshall GM. Histone deacetylase inhibitors: multifunctional anticancer agents. Cancer Treat Rev 2006; 32: 157–65.
Ivanov SV, Salnikow K, Ivanova AV, Bai L, Lerman MI. Hypoxic repression of STAT1 and its downstream genes by a pVHL/HIF-1 target DEC1/STRA13. Oncogene 2007; 26: 802–12.
Kato H, Tamamizu-Kato S, Shibasaki F. Histone deacetylase 7 associates with hypoxia-inducible factor 1alpha and increases transcriptional activity. J Biol Chem 2004; 279: 41966–74.
Qian DZ, Kato Y, Shabbeer S, Wei Y, Verheul HM, Salumbides B, et al. Targeting tumor angiogenesis with histone deacetylase inhibitors: the hydroxamic acid derivative LBH589. Clin Cancer Res 2006; 12: 634–42.
Wu MZ, Tsai YP, Yang MH, Huang CH, Chang SY, Chang CC, et al. Interplay between HDAC3 and WDR5 is essential for hypoxiainduced epithelial-mesenchymal transition. Mol Cell 2011; 43: 811–22.
Kwon HJ, Kim MS, Kim MJ, Nakajima H, Kim KW. Histone deacetylase inhibitor FK228 inhibits tumor angiogenesis. Int J Cancer. 2002; 97: 290–6.
Beyer S, Kristensen MM, Jensen KS, Johansen JV, Staller P. The histone demethylases JMJD1A and JMJD2B are transcriptional targets of hypoxia-inducible factor HIF. J Biol Chem 2008; 283: 36542–52.
Pollard PJ, Loenarz C, Mole DR, McDonough MA, Gleadle JM, Schofield CJ, et al. Regulation of Jumonji-domain-containing histone demethylases by hypoxia-inducible factor (HIF)-1alpha. Biochem J 2008; 416: 387–94.
Xia X, Lemieux ME, Li W, Carroll JS, Brown M, Liu XS, et al. Integrative analysis of HIF binding and transactivation reveals its role in maintaining histone methylation homeostasis. Proc Natl Acad Sci USA 2009; 106: 4260–5.
Luo W, Chang R, Zhong J, Pandey A, Semenza GL. Histone demethylase JMJD2C is a coactivator for hypoxia-inducible factor 1 that is required for breast cancer progression. Proc Natl Acad Sci USA 2012; 109: E3367–76.
van den Beucken T, Koritzinsky M, Niessen H, Dubois L, Savelkouls K, Mujcic H, et al. Hypoxia-induced expression of carbonic anhydrase 9 is dependent on the unfolded protein response. J Biol Chem 2009; 284: 24204–12.
Okami J, Simeone DM, Logsdon CD. Silencing of the hypoxiainducible cell death protein BNIP3 in pancreatic cancer. Cancer Res 2004; 64: 5338–46.
Murai M, Toyota M, Suzuki H, Satoh A, Sasaki Y, Akino K, et al. Aberrant methylation and silencing of the BNIP3 gene in colorectal and gastric cancer. Clin Cancer Res 2005; 11: 1021–7.
Jokilehto T, Jaakkola PM. The role of HIF prolyl hydroxylases in tumour growth. J Cell Mol Med 2010; 14: 758–70.
Huang KT, Mikeska T, Dobrovic A, Fox SB. DNA methylation analysis of the HIF-1alpha prolyl hydroxylase domain genes PHD1, PHD2, PHD3 and the factor inhibiting HIF gene FIH in invasive breast carcinomas. Histopathology 2010; 57: 451–60.
Rawluszko AA, Bujnicka KE, Horbacka K, Krokowicz P, Jagodzinski PP. Expression and DNA methylation levels of prolyl hydroxylases PHD1, PHD2, PHD3 and asparaginyl hydroxylase FIH in colorectal cancer. BMC Cancer 2013; 13: 52–6.
Nakamura J, Kitajima Y, Kai K, Hashiguchi K, Hiraki M, Noshiro H, et al. Expression of hypoxic marker CA IX is regulated by site-specific DNA methylation and is associated with the histology of gastric cancer. Am J Pathol 2011; 178: 515–24.
Liu Q, Liu L, Zhao Y, Zhang J, Wang D, Chen J, et al. Hypoxia induces genomic DNA demethylation through the activation of HIF-1- alpha and transcriptional upregulation of MAT2A in hepatoma cells. Mol Cancer Ther 2011; 10: 1113–23.
Koslowski M, Luxemburger U, Tureci O, Sahin U. Tumor-associated CpG demethylation augments hypoxia-induced effects by positive autoregulation of HIF-1alpha. Oncogene 2011; 30: 876–82.
Watson CJ, Collier P, Tea I, Neary R, Watson JA, Robinson C, et al. Hypoxia-induced epigenetic modifications are associated with cardiac tissue fibrosis and the development of a myofibroblast-like phenotype. Hum Mol Genet 2014; 23: 2176–88.
Skowronski K, Dubey S, Rodenhiser D, Coomber B. Ischemia dysregulates DNA methyltransferases and p16INK4a methylation in human colorectal cancer cells. Epigenetics 2010; 5: 547–56.
Hattori M, Yokoyama Y, Hattori T, Motegi SI, Amano H, Hatada I, et al. Global DNA hypomethylation and hypoxia-induced expression of the ten eleven translocation (TET) family, TET1, in scleroderma fibroblasts. Exp Dermatol. 2015.
Wu H, D’Alessio AC, Ito S, Xia K, Wang Z, Cui K, et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 2011; 473: 389–93.
Williams K, Christensen J, Pedersen MT, Johansen JV, Cloos PA, Rappsilber J, et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 2011; 473: 343–8.
Chen Q, Chen Y, Bian C, Fujiki R, Yu X. TET2 promotes histone OGlcNAcylation during gene transcription. Nature 2013; 493: 561–4.
Deplus R, Delatte B, Schwinn MK, Defrance M, Mendez J, Murphy N, et al. TET2 and TET3 regulate GlcNAcylation and H3K4 methylation through OGT and SET1/COMPASS. EMBO J 2013; 32: 645–55.
Vella P, Scelfo A, Jammula S, Chiacchiera F, Williams K, Cuomo A, et al. Tet proteins connect the O-linked N-acetylglucosamine transferase Ogt to chromatin in embryonic stem cells. Mol Cell 2013; 49: 645–56.
Yang T, Espenshade PJ, Wright ME, Yabe D, Gong Y, Aebersold R, et al. Crucial step in cholesterol homeostasis: sterols promote binding of SCAP to INSIG-1, a membrane protein that facilitates retention of SREBPs in ER. Cell 2002; 110: 489–500.
Sever N, Yang T, Brown MS, Goldstein JL, DeBose-Boyd RA. Accelerated degradation of HMG CoA reductase mediated by binding of insig-1 to its sterol-sensing domain. Mol Cell 2003; 11: 25–33.
Shaw RJ. Glucose metabolism and cancer. Curr Opin Cell Biol 2006; 18: 598–608.
Yang H, Liu Y, Bai F, Zhang JY, Ma SH, Liu J, et al. Tumor development is associated with decrease of TET gene expression and 5-methylcytosine hydroxylation. Oncogene 2013; 32: 663–9.
Hsu CH, Peng KL, Kang ML, Chen YR, Yang YC, Tsai CH, et al. TET1 suppresses cancer invasion by activating the tissue inhibitors of metalloproteinases. Cell Rep 2012; 2: 568–79.
Liu C, Liu L, Chen X, Shen J, Shan J, Xu Y, et al. Decrease of 5-hydroxymethylcytosine is associated with progression of hepatocellular carcinoma through downregulation of TET1. PLoS One 2013; 8: e62828.
Muller T, Gessi M, Waha A, Isselstein LJ, Luxen D, Freihoff D, et al. Nuclear exclusion of TET1 is associated with loss of 5-hydroxymethylcytosine in IDH1 wild-type gliomas. Am J Pathol 2012; 181: 675–83.
Orr BA, Haffner MC, Nelson WG, Yegnasubramanian S, Eberhart CG. Decreased 5-hydroxymethylcytosine is associated with neural progenitor phenotype in normal brain and shorter survival in malignant glioma. PLoS One 2012; 7: e41036.
Lian CG, Xu Y, Ceol C, Wu F, Larson A, Dresser K, et al. Loss of 5-hydroxymethylcytosine is an epigenetic hallmark of melanoma. Cell 2012; 150: 1135–46.
Kudo Y, Tateishi K, Yamamoto K, Yamamoto S, Asaoka Y, Ijichi H, et al. Loss of 5-hydroxymethylcytosine is accompanied with malignant cellular transformation. Cancer Sci 2012; 103: 670–6.
Rawluszko-Wieczorek AA, Siera A, Horbacka K, Horst N, Krokowicz P, Jagodzinski PP. Clinical significance of DNA methylation mRNA levels of TET family members in colorectal cancer. J Cancer Res Clin Oncol 2015; 141: 1379–92.
Frycz BA, Murawa D, Borejsza-Wysocki M, Marciniak R, Murawa P, Drews M, et al. Decreased expression of ten-eleven translocation 1 protein is associated with some clinicopathological features in gastric cancer. Biomed Pharmacother 2014; 68: 209–12.
Acknowledgments
We apologize to the authors whose work could not be cited due to the space constraint of reference citation. We declare no competing financial interests. This work was supported in part to K.J.W. by Ministry of Science and Technology Summit grant [MOST 104-2745-B-039-001-ASP]; center of excellence for cancer research at Taipei Veterans General Hospital [MOHW104-TDU-B- 211-124-001]; and National Health Research Institutes [NHRIEX104- 10230SI].
Author information
Authors and Affiliations
Corresponding author
Additional information
Research Center for Tumor Medical Science and Graduate Inst. of Cancer Biology, China Medical University, Taichung 404, Taiwan
*Corresponding author. Research Center for Tumor Medical Science, Graduate Institute of Cancer Biology, China Medical University, Taichung 404, Taiwan.
E-mail address: wukj@mail.cmu.edu.tw (K.-J. Wu).
Open Access This article is distributed under terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided original author(s) and source are credited.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, HF., Wu, KJ. Epigenetics, TET proteins, and hypoxia in epithelial-mesenchymal transition and tumorigenesis. BioMed 6, 1 (2016). https://doi.org/10.7603/s40681-016-0001-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.7603/s40681-016-0001-9