Abstract
Anaerobaculum mobile Menes and Muxí 2002 is one of three described species of the genus Anaerobaculum, family Synergistaceae, phylum Synergistetes. This anaerobic and motile bacterium ferments a range of carbohydrates and mono- and dicarboxylic acids with acetate, hydrogen and CO2 as end products. A. mobile NGAT is the first member of the genus Anaerobaculum and the sixth member of the phylum Synergistetes with a completely sequenced genome. Here we describe the features of this bacterium, together with the complete genome sequence, and annotation. The 2,160,700 bp long single replicon genome with its 2,053 protein-coding and 56 RNA genes is part of the Genomic Encyclopedia of Bacteria and Archaea project.
Similar content being viewed by others
Introduction
Strain NGAT (= DSM 13181 = ATCC BAA-54 = JCM 12221) is the type strain of Anaerobaculum mobile [1]. The genus, described for a thermophilic, citrate-fermenting anaerobe with A. thermoterrenum as the type species [2], was emended twice [1,3]. The third species is the recently described A. hydrogeniformans, forming H2 and acetate from glucose [3] while the other two species form CO2. A. mobile was isolated from a 10−7 dilution of an oleic acid-degrading consortium, originating from the sludge of a wool-scouring wastewater. Enrichment was performed in BCYT medium (basal BC medium [4] with tryptone and yeast extract), supplemented with crotonate [1]. Here we present a summary classification and a set of features for A. mobile NGAT together with the description of the complete genomic sequencing and annotation.
Classification and features
16S rRNA gene sequence analysis
A representative genomic 16S rRNA gene sequence of A. mobile NGAT was compared using NCBI BLAST [5,6] under default settings (e.g., considering only the high-scoring segment pairs (HSPs) from the best 250 hits) with the most recent release of the Greengenes database [7] and the relative frequencies of taxa and keywords (reduced to their stem [8]) were determined, weighted by BLAST scores. The most frequently occurring genera were Anaerobaculum (81.9%), Acetomicrobium (9.9%), Thermovirga (4.5%) and Dethiosulfovibrio (3.6%) (12 hits in total). Regarding the two hits to sequences from members of the species, the average identity within HSPs was 99.9%, whereas the average coverage by HSPs was 100.0%. Regarding the seven hits to sequences from other members of the genus, the average identity within HSPs was 97.1%, whereas the average coverage by HSPs was 99.1%. Among all other species, the one yielding the highest score was Acetomicrobium flavidum (FR733692), which corresponded to an identity of 99.8% and an HSP coverage of 100.0%. (Note that the Greengenes database uses the INSDC (= EMBL/NCBI/DDBJ) annotation, which is not an authoritative source for nomenclature or classification.) The highest-scoring environmental sequence was FN436106 (Greengenes name ‘Molekularbiologische Charakterisierung der bakteriellen Populationsdynamik eines thermophil betriebenen Biogasfermenters zur Vergaerung nachwachsender Rohstoffe thermophilic biogas reactor fed renewable biomass clone HAW-R60-B-745d-BC’), which showed an identity of 99.9% and an HSP coverage of 99.9%. The most frequently occurring keywords within the labels of all environmental samples which yielded hits were ‘digest’ (7.0%), ‘anaerob’ (4.6%), ‘thermophil’ (4.3%), ‘microbi’ (3.4%) and ‘mesophil’ (3.3%) (238 hits in total). The most frequently occurring keywords within the labels of those environmental samples which yielded hits of a higher score than the highest scoring species were ‘thermophil’ (12.3%), ‘reactor’ (8.8%), ‘bioga, fed’ (7.0%), ‘bakteriellen, betriebenen, biogasferment, biomass, charakterisierung, molekularbiologisch, nachwachsend, populationsdynamik, renew, rohstoff, vergaerung’ (5.3%) and ‘corn, microbi, silag, structur’ (1.8%) (4 hits in total).
A. flavidum is a species that was originally described without 16S rRNA gene sequencing [9]. The sequence FR733692 was only recently been generated by DSMZ staff in the course of the Living-Tree Project [10], yielding an unexpected placement of the species within the Synergistetes. An assessment of whether a contamination or confusion of strains has occurred, or whether the currently available culture deposits are biologically identical to the original description but the classification of A. flavidum needs to be revised. This work is currently in progress. (R. Pukall (DSMZ), personal communication).
Figure 1 shows the phylogenetic neighborhood of A. mobile in a 16S rRNA gene based tree. The sequences of the two identical 16S rRNA gene copies in the genome differ by one nucleotide from the previously published 16S rRNA gene sequence (AJ243189).
Phylogenetic tree highlighting the position of A. mobile relative to the type strains of the other species within the phylum ‘Synergistetes’. The tree was inferred from 1,360 aligned characters [11,12] of the 16S rRNA gene sequence under the maximum likelihood (ML) criterion [13]. Rooting was done initially using the midpoint method [14] and then checked for its agreement with the current classification (Table 1). The branches are scaled in terms of the expected number of substitutions per site. Numbers adjacent to the branches are support values from 1,000 ML bootstrap replicates [15] (left) and from 1,000 maximum-parsimony bootstrap replicates [16] (right) if larger than 60%. Lineages with type strain genome sequencing projects registered in GOLD [17] are labeled with one asterisk, those also listed as ‘Complete and Published’ with two asterisks [18–20]; the non-contiguous finished draft genomes of Aminomonas paucivorans [21] and Dethiosulfovibrio peptidovorans [22] lack the second asterisk.
Morphology and physiology
The Gram-negative rod-shaped cells of A. mobile are straight (0.5–1.0 × 2.0–4.0 µm; Figure 2), occurring singly or, predominantly in the exponential growth phase, in pairs. Longer cells (4.0-8.0 µm) are observed in older cultures. Spores or sheath formation were never observed. Cells are moderately thermophilic; motile by means of a single flagellum inserted in the lateral region of the cell. On BCTC agar, colonies (1.0–1.5 mm in diameter) are white, lens shaped and with entire margins. Temperature and pH range for growth is between 35–65°C (optimum between 55 and 60°C) and between 5.4 and 8.7 (optimum between 6.6 and 7.3), respectively. Growth occurs at NaCl concentrations of up to 1.5%, with an optimum of 0.8%.
Cells are strictly anaerobic, fermenting glucose, organic acids (pyruvate, tartrate and malate) and glycerol to acetate, H2 and, presumably, CO2 (the authors of [1] assumed that 2 mol CO2 was produced per mol glucose, tartrate and malate degraded, and 1 mol CO2 was produced per mol pyruvate and glycerol degraded). Gluconate, L-malate, glycerol, tryptone, L-arginine, L-leucin, L-phenylalanine and starch are utilized [3]. No growth occurs on citrate, 2-oxoglutarate, glutamate, mannose, pectin, lactose, xylose, galactose, maltose, sucrose, rhamnose, raffinose, malonate, lactate, succinate, xylan, dextrin, inulin, melibiose, adonitol, cellobiose, arabinose, polygalacturonate, cellulose, gelatin, butyrate or oleate [1], nor lactate, maltose, malonate, mannose, inositol and inulin [3].
In co-culture with Methanothermobacter thermautotrophicus glucose and malate degradation were enhanced. Crotonate is not fermented but used as an electron acceptor, reducing it to butyrate in the presence of yeast extract and tryptone, glucose, Casamino acids and leucine. Fumarate, acetate, sulfate, sulfite and nitrate are not used as electron acceptors in PY broth. The addition of crotonate, sulfur, cystine and thiosulfate enhances growth from tryptone and yeast extract, Casamino acids and glucose, as indicated by an increase in OD and by the production of reduction compounds (butyrate and sulfide, respectively) [1].
Chemotaxonomy
The cell wall is of the Gram-negative type as judged by transmission electron micrography [1]. Polar lipid composition includes diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine, three unknown phospholipids and four unknown aminophospholipids [3]. Major fatty acids (>20%) are iso-C15:0 and iso-C11:0; iso-C13:0 3-OH is found in smaller amounts (<10%) [3]. The DNA G+C content was previously reported with 51.5 mol% [1].
Genome sequencing and annotation
Genome project history
This organism was selected for sequencing on the basis of its phylogenetic position [29], and is part of the Genomic Encyclopedia of Bacteria and Archaea project [30]. The genome project is deposited in the Genomes On Line Database [17] and the complete genome sequence is deposited in GenBank. Sequencing, finishing and annotation were performed by the DOE Joint Genome Institute (JGI) using state of the art sequencing technology [31]. A summary of the project information is shown in Table 2.
Growth conditions and DNA isolation
A mobile strain NGAT, DSM 13181, was grown anaerobically in DSMZ medium 104b (PYX, medium) at 55°C. DNA was isolated from 1–1.5 g of cell paste using a Jetflex Genomic DNA Purification Kit (GENOMED 600100) and following the standard protocol as recommended by the manufacturer with modification (an additional 10 µl proteinase K digestion for cell lysis was added; 40 min incubation at 58°C). DNA is available through the DNA Bank Network [32].
Genome sequencing and assembly
The genome was sequenced using a combination of Illumina and 454 sequencing platforms. All general aspects of library construction and sequencing can be found at the JGI website [33]. Pyrosequencing reads were assembled using the Newbler assembler (Roche). The initial Newbler assembly, consisting of 31 contigs in one scaffold, was converted into a phrap [34] assembly by making fake reads from the consensus, to collect the read pairs in the 454 paired end library. Illumina GAii sequencing data (2,649.5 Mb) was assembled with Velvet [35] and the consensus sequences were shredded into 1.5 kb overlapped fake reads and assembled together with the 454 data. The 454 draft assembly was based on 113.8 Mb 454 draft data and all of the 454 paired end data. Newbler parameters are -consed -a 50 -l 350 -g -m -ml 20. The Phred/Phrap/Consed software package [34] was used for sequence assembly and quality assessment in the subsequent finishing process. After the shotgun stage, reads were assembled with parallel phrap (High Performance Software, LLC). Possible mis-assemblies were corrected with gapResolution [33], Dupfinisher [36], or sequencing cloned bridging PCR fragments with subcloning. Gaps between contigs were closed by editing in Consed, by PCR and by Bubble PCR primer walks (J.-F. Chang, unpublished). A total of 68 additional reactions and 3 shatter libraries were necessary to close gaps and to raise the quality of the finished sequence. Illumina reads were also used to correct potential base errors and increase consensus quality using a software Polisher developed at JGI [37]. The error rate of the completed genome sequence is less than 1 in 100,000. Together, the combination of the Illumina and 454 sequencing platforms provided 1147.3 × coverage of the genome. The final assembly contained 237,629 pyrosequence and 32,103,764 Illumina reads.
Genome annotation
Genes were identified using Prodigal [38] as part of the DOE-JGI genome annotation pipeline [39], followed by a round of manual curation using the JGI GenePRIMP pipeline [40]. The predicted CDSs were translated and used to search the National Center for Biotechnology Information (NCBI) non-redundant database, UniProt, TIGR-Fam, Pfam, PRIAM, KEGG, COG, and InterPro databases. Additional gene prediction analysis and functional annotation was performed within the Integrated Microbial Genomes - Expert Review (IMG-ER) platform [41].
Genome properties
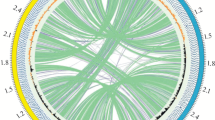
The genome statistics are provided in Table 3 and Figure 3. The genome consists of one chromosome with a total length of 2,160,700 bp and a G+C content of 48.0%. Of the 2,109 genes predicted, 2,053 were protein-coding genes, and 56 RNAs; 34 pseudogenes were also identified. The majority of the protein-coding genes (85.7%) were assigned a putative function while the remaining ones were annotated as hypothetical proteins. The distribution of genes into COGs functional categories is presented in Table 4.
References
Menes RJ, Muxi L. Anaerobaculum mobile sp. nov., a novel anaerobic, moderately thermophilic, peptide-fermenting bacterium that uses crotonate as an electron acceptor, and emended description of the genus Anaerobaculum. Int J Syst Evol Microbiol 2002; 52:157–164. PubMed
Rees GN, Pater BKC, Grassia GS, Sheehy AJ. Anaerobaculum thermoterrenum gen. nov., sp. nov., a novel, thermophilic bacterium which ferments citrate. Int J Syst Bacteriol 1997; 47:150–154. PubMed http://dx.doi.org/10.1099/00207713-47-1-150
MW Maune RS Tanner (2012) ArticleTitleDescription of Anaerobaculum hydrogeniformans sp. nov., an anaerobe that produces hydrogen from glucose, and emended description of the genus Anaerobaculum Int J Syst Evol Microbiol 62 832–838 Occurrence Handle10.1099/ijs.0.024349-0 Occurrence Handle1:CAS:528:DC%2BC38XotlCitbY%3D Occurrence Handle21602364
Touzel JP, Albagnac G. Isolation and characterization of Methanococcus mazeii strain MC3. FEMS Microbiol Lett 1983; 16:241–245. http://dx.doi.org/10.1111/j.15746968.1983.tb00295.x
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol 1990; 215:403–410. PubMed
Korf I, Yandell M, Bedell J. BLAST, O’Reilly, Sebastopol, 2003.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 2006; 72:5069–5072. PubMed http://dx.doi.org/10.1128/AEM.03006-05
Porter MF. An algorithm for suffix stripping. Program: electronic library and information systems 1980; 14:130–137.
Soutschek E, Winter J, Schindler F, Kandler O. Acetomicrobium flavidum, gen. nov., sp. nov., a thermophilic, anaerobic bacterium from sewage sludge, forming acetate, CO2 and H2 from glucose. Syst Appl Microbiol 1984; 5:377–390. http://dx.doi.org/10.1016/S0723-2020(84)80039-9
Munoz R, Yarza P, Ludwig W. Euzéby, Amann R, Schleifer K-H, Glöckner FO, Rossellö-Möra R. Release LTPs104 of the All-Species Living Tree. Syst Appl Microbiol 2011; 34:169–170. PubMed http://dx.doi.org/10.1016/j.syapm.2011.03.001
Lee C, Grasso C, Sharlow MF. Multiple sequence alignment using partial order graphs. Bioinformatics 2002; 18:452–464. PubMed http://dx.doi.org/10.1093/bioinformatics/18.3.452
Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 2000; 17:540–552. PubMed http://dx.doi.org/10.1093/oxfordjournals.molbev.a 026334
Stamatakis A, Hoover P, Rougemont J. A rapid bootstrap algorithm for the RAxML web-servers. Syst Biol 2008; 57:758–771. PubMed http://dx.doi.org/10.1080/10635150802429642
Hess PN, De Moraes Russo CA. An empirical test of the midpoint rooting method. Biol J Linn Soc Lond 2007; 92:669–674. http://dx.doi.org/10.1111/j.10958312.2007.00864.x
Pattengale ND, Alipour M, Bininda-Emonds ORP, Moret BME, Stamatakis A. How many bootstrap replicates are necessary? Lect Notes Comput Sci 2009; 5541:184–200. http://dx.doi.org/10.1007/978-3-642-02008-7_13
Swofford DL. PAUP*: Phylogenetic Analysis Using Parsimony (*and Other Methods), Version 4.0 b10. Sinauer Associates, Sunderland, 2002.
Pagani I, Liolios K, Jansson J, Chen IM, Smirnova T, Nosrat B, Markowitz VM, Kyrpides NC. The Genomes OnLine Database (GOLD) v.4: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res 2012; 40:D571–D579. PubMed http://dx.doi.org/10.1093/nar/gkr1100
Chovatia M, Sikorski J, Schröder M, Lapidus A, Nolan M, Tice H, Del Rio TG, Copeland A, Cheng JF, Lucas S, et al. Complete genome sequence of Thermanaerovibrio acidaminovorans type strain (Su883T). Stand Genomic Sci 2009; 1:254–261. PubMed http://dx.doi.org/10.4056/sigs.40645
Göker M, Saunders E, Lapidus A, Nolan M, Lucas S, Hammon N, Deshpande S, Cheng JF, Han C, Tapia R, et al. Genome sequence of the moderately thermophilic, amino-acid-degrading and sulfur-reducing bacterium Thermovirga lienii type strain (Cas60314T). Stand Genomic Sci 2012; 6:230–239. PubMed http://dx.doi.org/10.4056/sigs.2726028
Chertkov O, Sikorski J, Brambilla E, Lapidus A, Copeland A, Del Rio TG, Nolan M, Lucas S, Chen F, Tice H, et al. Complete genome sequence of Aminobacterium colombiense type strain (ALA-1T). Stand Genomic Sci 2010; 2:280–289. PubMed http://dx.doi.org/10.4056/sigs.902116
Pitluck S, Yasawong M, Held B, Lapidus A, Nolan M, Copeland A, Lucas S, Del Rio TG, Tice H, Cheng JF, et al. Non-contiguous finished genome sequence of Aminomonas paucivorans type strain (GLU-3T). Stand Genomic Sci 2009; 3:285–293. http://dx.doi.org/10.4056/sigs.1253298
LaButti K, Mayilraj S, Clum A, Lucas S, Del Rio TG, Nolan M, Tice H, Cheng JF, Pitluck S, Liolios K, et al. Permanent draft genome sequence of Dethiosulfovibrio peptidovorans type strain (SEBR 4207T). Stand Genomic Sci 2010; 3:85–92. PubMed http://dx.doi.org/10.4056/sigs.1092865
Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541–547. PubMed http://dx.doi.org/10.1038/nbt1360
Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 1990; 87:4576–4579. PubMed http://dx.doi.org/10.1073/pnas.87.12.4576
Jumas-Bilak E, Roudière L, Marchandin H. Description of ‘Synergistetes’ phyl. nov. and emended description of the phylum ‘Deferribacteres’ and of the family Syntrophomonadaceae, phylum ‘Firmicutes’. Int J Syst Evol Microbiol 2009; 59:1028–1035. PubMed http://dx.doi.org/10.1099/ijs.0.006718-0
Rees GN, Patel BKC, Grassia GS, Sheehy AJ. Anaerobaculum thermoterrenum gen. nov., sp. nov., a novel, thermophilic bacterium which ferments citrate. Int J Syst Bacteriol 1997; 47:150–154. PubMed http://dx.doi.org/10.1099/00207713-47-1-150
BAuA. 2010, Classification of bacteria and archaea in risk groups. http://www.baua.de TRBA 466, p. 24.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. Gene ontology: tool for the unification of biology. Nat Genet 2000; 25:25–29. PubMed http://dx.doi.org/10.1038/75556
Klenk HP, Göker M. En route to a genome-based classification of Archaea and Bacteria? Syst Appl Microbiol 2010; 33:175–182. PubMed http://dx.doi.org/10.1016/j.syapm.2010.03.003
Wu D, Hugenholtz P, Mavromatis K, Pukall R, Dalin E, Ivanova NN, Kunin V, Goodwin L, Wu M, Tindall BJ, et al. A phylogeny-driven genomic encyclopaedia of Bacteria and Archaea. Nature 2009; 462:1056–1060. PubMed http://dx.doi.org/10.1038/nature08656
Mavromatis K, Land ML, Brettin TS, Quest DJ, Copeland A, Clum A, Goodwin L, Woyke T, Lapidus A, Klenk HP, et al. The fast changing landscape of sequencing technologies and their impact on microbial genome assemblies and annotation. PLoS ONE 2012; 7:e48837;. PubMed http://dx.doi.org/10.1371/journal.pone.0048837
Gemeinholzer B, Dröge G, Zetzsche H, Haszprunar G, Klenk HP, Güntsch A, Berendsohn WG, Wägele JW. The DNA Bank Network: the start from a German initiative. Biopreserv Biobank 2011; 9:51–55. http://dx.doi.org/10.1089/bio.2010.0029
JGI website. http://www.jgi.doe.gov
The Phred/Phrap/Consed software package. http://www.phrap.com
Zerbino DR, Birney E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 2008; 18:821–829. PubMed http://dx.doi.org/10.1101/gr.074492.107
Han C, Chain P. Finishing repeat regions automatically with Dupfinisher. In: Proceeding of the 2006 international conference on bioinformatics & computational biology. Arabnia HR, Valafar H (eds), CSREA Press. June 26–29, 2006: 141–146.
Lapidus A, LaButti K, Foster B, Lowry S, Trong S, Goltsman E. POLISHER: An effective tool for using ultra short reads in microbial genome assembly and finishing. AGBT, Marco Island, FL, 2008.
Hyatt D, Chen GL, LoCascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 2010; 11:119. PubMed http://dx.doi.org/10.1186/1471-2105-11-119
Mavromatis K, Ivanova NN, Chen IM, Szeto E, Markowitz VM, Kyrpides NC. The DOE-JGI Standard operating procedure for the annotations of microbial genomes. Stand Genomic Sci 2009; 1:63–67. PubMed http://dx.doi.org/10.4056/sigs.632
Pati A, Ivanova NN, Mikhailova N, Ovchinnikova G, Hooper SD, Lykidis A, Kyrpides NC. GenePRIMP: a gene prediction improvement pipeline for prokaryotic genomes. Nat Methods 2010; 7:455–457. PubMed http://dx.doi.org/10.1038/nmeth.1457
Markowitz VM, Ivanova NN, Chen IMA, Chu K, Kyrpides NC. IMG ER: a system for microbial genome annotation expert review and curation. Bioinformatics 2009; 25:2271–2278. PubMed http://dx.doi.org/10.1093/bioinformatics/btp393
Acknowledgements
We would like to gratefully acknowledge the help of Maren Schröder for growing A. mobile cultures, Evelyne-Marie Brambilla for DNA extraction and quality control, and Cathrin Spröer and Rüdiger Pukall for helpful comments (all at DSMZ). This work was performed under the auspices of the US Department of Energy Office of Science, Biological and Environmental Research Program, and by the University of California, Lawrence Berkeley National Laboratory under contract No. DE-AC02-05CH11231, Lawrence Livermore National Laboratory under Contract No. DE-AC52-07NA27344, and Los Alamos National Laboratory under contract No. DE-AC02-06NA25396, UT-Battelle and Oak Ridge National Laboratory under contract DE-AC05-00OR22725.
Author information
Authors and Affiliations
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Mavromatis, K., Stackebrandt, E., Held, B. et al. Complete genome sequence of the moderate thermophile Anaerobaculum mobile type strain (NGAT). Stand in Genomic Sci 8, 47–57 (2013). https://doi.org/10.4056/sigs.3547050
Published:
Issue Date:
DOI: https://doi.org/10.4056/sigs.3547050