Abstract
The dwarf shrub Cressa cretica is a thermocosmopolitan halophilic species. Different mechanisms confer salt stress tolerance such as tissue and osmotic tolerance and ion exclusion, as well as the associated microbiota. The aims were (i) to investigate the best conditions for C. cretica seed germination and to examine the tolerance of germinated seeds and seedlings to different salt concentrations and (ii) to characterize the rhizosphere and bulk soil microbiota. Germination and growth experiments were conducted to address plant salt tolerance, and with Illumina sequencing the microbiota of rhizosphere and bulk soil was investigated. While high salt concentrations (600, 800, and 1000 mM NaCl) inhibited C.cretica seed germination, recovery of ungerminated seeds under non-saline conditions was high, indicating osmotic rather than toxic effects of high salt concentrations. The microbiota found in rhizosphere and bulk soil showed high similarity with that found in previous studies on halophyte-associated microbiota, among the Planctomyces, Halomonas and Jeotgalibacillus. Concluding, salt stress responses on the plant level were shown, as has the involvement of associated halotolerant bacteria. Still, the potential role for mitigating saline stress by the bacterial associates of C. cretica, most of them microbial dark matter, will have to be investigated, as will the contributions of archaea and fungi.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Nowadays, soil salinity has become an increasing problem for agriculture, and salt tolerance during germination stage is critical for the establishment of plant growth on saline soil (Ungar 1995; Coleman-Derr and Tringe 2014). Upward movement of soil solution followed by soil surface evaporation causes a high salt concentration in the seed-planting zone and often results in the failure of germination. Understanding salt tolerance of halophytic plants has recently become augmented by improved knowledge on the involved plant associated microbiota (Yuan et al. 2016).
Halophytes are plants which naturally grow and survive in environments with salt concentrations as high as 1 M NaCl (Flowers and Colmer 2008). These plants have evolved various strategies including the accumulation of Na+ in the vacuole, the exclusion of Na+ from cells and the increase of the osmotic pressure in the cytoplasm (Flowers and Colmer 2008). Understanding salt tolerance of halophytic plants has recently become augmented by improved knowledge on the involved plant associated microbiota (Yuan et al. 2016). Researchers suggested that the halophytic plant-associated microbiota can be the key factor for the adaptation of plants to salinity (Etesami and Beattie 2018). The phenomenon of habitat-adapted symbiosis is an example of an adaptation of plants to adverse environments through the symbiosis with non-mycorrhizal fungal endophytes (Rodriguez et al. 2008). Furthermore, it has been reported that plant growth-promoting rhizobacteria (PGPR) are able to promote the systematic tolerance of plants by inducing the physical and chemical changes of their hosts that results in tolerance improvement to abiotic stress (Kohler et al. 2009). Some of these microorganisms have the potential to mitigate the salinity stress (Akhtar et al. 2015; Upadhyay and Singh 2015; Wang et al. 2016). Halotolerant PGPR are able to survive and grow in saline environments with a wide range from 1 to 33% NaCl (Etesami and Beattie 2018). Therefore, they are well-suited to grow in the rhizosphere of halophytic plants. For instance, Kushneria sp. YCWA18, as a high phosphorus-solubilizing halotolerant PGPR, was isolated from the sediment of Daqiao saltern on the eastern coast of China and able to grow in media with salinity concentrations of 20 % (w/v) NaCl (Zhu et al. 2011). Arthrobacter sp., Bacillus pumilus, Halomonas sp., Nitrinicola lacisaponensis and Pseudomonas mendocinawere also isolated as PGPRs with ability to tolerate 2 – 25% NaCl (Tiwari et al. 2011). Moreover, Bacillus polymyxa BcP26, Mycobacterium phlei MbP18 and Pseudomonas alcaligens P5A15 were also shown to be capable of tolerating high salt concentrations (Egamberdieva2009). Moreover, previous studies also introduced some PGPR to improve the growth of bean, canola, lettuce, pepper and tomato under saline conditions (Yildirim et al. 2006; Barassi et al. 2006). Since the survival of single species of beneficial microorganisms is often hampered due to various reasons, it was hypothesized that it is a combination of various members of the microbiota that together would exert beneficial effects (Qin et al. 2016).
The diversity of plants species in the Gav Khooni region in Isfahan, Iran is high. Many plants belong to Chenopodiaceae (Halostachys, Salsola,) and/or Tamaricaceae (Tamarix) as well as the Convolvulaceae family like the dwarf shrub Cressa cretica, a thermocosmopolitan halophilic species reaching up to 38 cm height is (Priyashree et al. 2010). C. cretica is a shrub to be utilized in landscaping as a ground cover, and it is well known as a medicinal plant (Weber et al. 2007). Salinity reduces the growth of the plant, however the plant has different mechanisms to tolerate this stress such as tissue tolerance, osmotic tolerance and ion exclusion (Roy et al. 2014). Moreover, many fundamental studies have been done to characterize salt stress-related genes in the plant using genetic modification methods, but only limited success from these approaches has been reported (Coleman-Derr and Tringe 2014; Yuan et al. 2016).
The purpose of the present study was, firstly, to investigate the optimum condition for C. cretica seed germination and to examine the tolerance of germinated seeds and seedlings to different salt concentrations. Secondly, by studying the C. cretica habitat-specific rhizosphere and soil microbiota we tried to understand possible interactions with microorganisms concerning the complex adaptation of C. cretica to its highly saline habitat.
Material and methods
Seed collection and storage
Fruits of C. cretica were collected during September 2015 from around Kamal Abad (Lat. 32° 31’ N, 51°51′ E) near Gav Khooni swamp, Isfahan, Iran. The soil of this region belongs to the group of Aquisalids with an electrical conductivity exceeding 200 dS m−1 (Roozitalab et al. 2018). The seed collection site was a dry saline plain between sandy hills and swamp. This semiarid region receives 110 mm annual rainfall, and the average annual evaporation is 1900 mm. Seeds were separated from infructescences and stored in paper bags at 4 °C.
Seed scarification
C. cretica seeds were scarified with sulfuric acid (96 %) for different durations (0, 20, 40, 60, 80, 100, 120 min). Each treatment comprised four replicates and each replicate contained 25 seeds. Seeds were washed three times with running tap water for 2 min to remove any trace of acid. Seed germination was carried out in 9.5 cm Petri dishes with two Whatman filter papers in each one. Twenty five seeds were placed on each dish and 5 mL of distilled water were added. Then, after placing the lid, the dishes were sealed with parafilm to avoid evaporation and incubated at room temperature (varied between 20 and 30 °C). The seed germination rate was counted after 20 days.
Effects of temperature on seed germination
The seeds were scarified with H2SO4 (96 %) for 60 min and washed with distilled water three times, then the seeds were surface sterilized with 5 % commercial Clorox solution (contains 5.25 % sodium hypochlorite) for 10 min and rinsed three times with distilled water. Seed germination was carried out as above; this time, however, the experiment was conducted in incubators set at 5, 10, 15, 20, 25 and 30 °C in 12 light/12 dark cycles with 4 replicates each. The counting of germinated seeds was done every day (seeds were considered to be germinated upon emergence (2 mm) of radicle) for 20 days, and in case of low or no germination, the recovery test for ten days (see below) was done. The statistical analysis based on percentage of seed germination and T50 (days to germination of 50 % of all germinated seeds) was done by GERMINATOR software (Joosen et al. 2010).
Effects of salinity on germination and plantlet
Seeds were provided for germination as detailed above. The seeds were subjected to five NaCl concentrations (200, 400, 600, 800 and 1000 mM) and distilled water as a control (four replicates each). Germination was carried out in an incubator (PERCIVAL SE – 41AR2; CLF Plant Climatics, Wertingen, Germany) set at 25 °C in 12 light/12 dark cycles. The germination was recorded daily for up to 20 days. After that, the un-germinated seeds were transferred from the salt solution to distilled water for 10 days for a recovery test. Seedlings were sown in 13x13x13 cm polyethylene pots (Göttinger, Göttingen, Germany) filled with substrate (sand, peat, quartz sand). Pots were sub-irrigated with 100 mL of tap water as a control and five different salt concentrations (200, 400, 600, 800 and 1000 mM NaCl) every week. A half-strength Hoagland nutrient solution was used every two weeks. The pots were placed in a climate chamber (Heraeus HQI – BT 400 W/D E40 FLH1) at 25 °C, with 50% humidity, 16/8 (d/n) light and 130 – 170 umolm2 s−1 light intensity. The height of each plantlet (three replicates per treatment) was measured after 5, 10 and 15 weeks. The statistical analysis for percentage of seed germination in different salt concentrations was done by GERMINATOR software (Joosen et al. 2010). Plantlet data were tested for statistical significance using an analysis of variance (ANOVA). Means were compared by Tukey test at p < 0.05 with the statistical program R (package multcomView) (R Core Team 2015).
Soil sampling and DNA extraction
Rhizosphere and bulk soil samples were taken in three replicates at least 10 m apart and stored at −20 °C until analysis. Soil samples (0.5 g fresh weight, fw) were mixed with 1.5 mL of 100 mM PBS (phosphate-buffered saline) used as a prewashing agent, followed by vortexing for 3 min. The mixture was then incubated for 5 min at 5 °C, centrifuged at 10.000 ɡ for 5 min, and the resulting supernatant was discarded. This pre-washing step with PBS was repeated three times for each sample. Afterwards, total DNA was extracted from each pre-washed sample (0.4 g, fw) using the NucleoSpin Soil kit (Macherey-Nagel, Germany) according to the manufacturer’s protocol. DNA quality was checked on 2 % agarose gel and DNA extracts were stored in low-DNA binding tubes (Genuine Axygen Quality 1.7 mL Maximum recovery, Axygen, USA) at −20 °C until use.
Illumina MiSeq sequencing workflow and analysis
An amplicon sequencing using Illumina’s MiSeq platform was performed to study the taxonomic structure of bacteria and archaea using three replicates per treatment. An identified primer set (515F and 806R) for amplifying a fragment of 16S rRNA gene capturing the V4 – V5 region was used (Caporaso et al. 2011). The PCR cycling initiated with denaturation at 95 °C for 3 min, followed by 20 cycles (first step) and 15 cycles (second step) of denaturation at 98 °C for 20 s, primer annealing at 56 °C for 30 s, and extension at 72 °C for 30 s with a final elongation step at 72 °C for 5 min. PCR products from all replicates were then purified, quantified and pooled in equimolar concentrations for sequencing on an Illumina MiSeq instrument using the 2 × 250 bp paired-end approach (Microsynth AG, Switzerland). Raw Illumina MiSeq paired-end reads were processed through the following workflow. Paired-End reads were assembled with default settings of PandaSeq software, version 2.88 (Masella et al. 2012). Low-quality reads, defined as reads with an average quality score below 25, with more than one ambiguous base and a length < 250 and > 260, were removed using the Prinseq program, version 0.20.4 (Schmieder and Edwards 2011). In addition, barcodes and primers were trimmed by Prinseq software. The processing of filtered reads to operational taxonomic units (OTUs) was done with the LotuS program using USEARCH at 97% similarity (Hildebrand et al. 2014). The processing of taxonomic affiliation of each OTU was obtained by using SILVA. The output of data was double-checked with the workflow for microbiome data analysis proposed by Callahan et al. (2016).
After data preprocessing, the final OTU table was proportionally normalized to the median of total sample reads. Further microbiome analysis and visualization were done by Phyloseq package (McMurdie and Holmes 2013, 2014; R Core Team 2015). To account for heterogeneity in reading counts among samples, we employed the DESeq2 statistical package according to Love et al. (2014) to assess differential OTU abundances.
Results
Optimization of seed germination of Cressa cretica
The optimum duration of seed scarification was 60 min in sulfuric acid which led to significantly higher final seed germination compared to the control and the other incubation periods (Fig. 1). Figure 2 shows the effect of temperature on the kinetics of germination of scarified seeds. Within 20 days, we did not record any germinated seeds at 5 °C and 10 °C. Germination at 15 °C (56 %) was significantly lower than at 20, 25 and 30 °C with 79, 89 and 88 % germination, respectively (Fig. 3a). Germination at 20, 25 and 30 °C was not significantly different, however, T50 revealed differences (Fig. 3b). The lowest T50 was recorded at 30 °C, and the highest was found at 15 °C. Although at 30 °C seed germination was high and T50 was low, the root tips of germinated seeds were necrotic (data not shown). Therefore, we chose 25 °C as the optimum temperature for germination.
Seeds that had not germinated at 5 and 10 °C showed a complete recovery when transferred to 25 °C (Fig. 4). This result confirmed that germination was 94 % and 84 % in the recovery test and was not significantly different to the 25 °C and 30 °C treatments (Fig. 4a). The lowest T50 in the recovery test was obtained from the seeds from the 10 °C treatment, which were transferred to 25 °C (Fig. 4b).
The effect of salinity on seed germination and plantlet
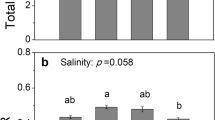
Salinity treatment significantly (p < 0.05) affected the final percent germination of seeds, and the highest value was recorded in distilled water (Fig. 5). There were no significant differences between the final germination of seeds treated with 200 mM NaCl and the control treatment. At the concentration of 400 mM NaCl, the germination percentage significantly decreased (Fig. 5a), while the T50 of germination increased at this NaCl concentration (Fig. 5b).
Salt concentrations higher than 400 mM (600, 800 and 1000 mM) totally inhibited seed germination. Thus, after the germination test, seeds were transferred to distilled water to study the recovery of germination. After ten days the seeds showed a complete recovery (Fig. 6a). Although the germination percentage in the recovery test for 600, 800 and 1000 mM NaCl was statistically similar to the control, T50 of 600 and 800 mM was significantly less than the control (Fig. 6b).
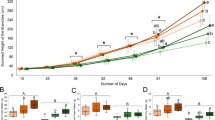
In the experiment on the effect of salinity on shoot growth no differences were found after five weeks. In the following weeks, however, shoot growth declined with increasing salinity (Fig. 7). Out of a total of 6 plantlet replications per concentration, one plantlet in 800 mM NaCl and two plantlets in 1000 mM NaCl died and growth in 400 and 600 mM NaCl ceased (Supp. 1).
Microbiome of Cressa cretica rhizosphere and bulk soil
The microbiomes of C. cretica rhizosphere and bulk soil were analyzed by 16S Illumina sequencing. Overall, after removing single- and doubleton operational taxonomic units (OTUs), 3420 OTUs (16S rDNA at 97 % similarity cut-off) were identified in the final dataset with a median of 48,274 reads per sample. More than 70 % of the OTUs were not classified (unknown) at the genus level (Fig. 8).
Focusing on the OTUs allocated to the known, a total of 364 OTUs was found in the six samples. The total reads for each sample were normalized proportionally to the median of 364 OTUs equaling 12,467 reads. The composition of bacterial communities at the phylum level revealed Proteobacteria (30–35 %), Firmicutes (29–30 %) and Planctomycetes (16–20 %), which dominated with more than 80% abundance in all samples (Fig. 9).
The four most abundant genera in the rhizosphere were Bacillus, Planctomyces, Halomonas and Jeotgalibacillus (Fig. 10a). However, Paenisporosarcina, Bacillus, Pir4_lineage and Planctomyces were the four most abundant genera in the bulk soil (Fig. 10b).
To assess differential OTU abundances between rhizosphere and bulk soil, we applied the DESeq2. The result shows that nine genera such as Saccharospirillum, Marinobacterium, and Halomonas were significantly more abundant in the rhizosphere compared to the bulk soil. On the other hand, we found ten genera, which were significantly more abundant in the bulk soil (Fig. 11, Supp. 2).
Discussion
C. cretica grows in the most saline regions around Gav Khooni swamp. It produces seeds during September and October and germinates during spring while the salt marsh area drains after rainfall and upon increasing temperature. The seed germination rate of C. cretica in its natural habitat is very low, which is attributed to several environmental factors such as temperature and salinity (Badger and Ungar 1989; Ungar 1995; Khan et al. 2002; Qu et al. 2008). In our study, percent seed germination of C. cretica increased with time of exposure to sulfuric acid up to 60 min but declined thereafter. Scarification by sulfuric acid has been commonly used to promote germination of several hard seeded species (Tigabu and Oden 2001; Vilela and Ravetta 2001; Aliero 2004; Fang et al. 2006). Therefore, the positive responses of seeds to the scarification treatment indicate that the hard seed coat by preventing imbibition of water is one of the factors for low germination rates of the untreated seeds as has been found for other members of the Convolvulaceae (Ajmal Khan and Ungar 1998; Tetetay 1998).
Low temperatures ceased (5 °C and 10 °C) or delayed (15 °C) germination, but with increasing temperature there was a visible improvement in germination. It has been reported that cold temperature inhibits mitosis and maintains the dormancy of seeds (Simon et al. 1976; Khan and Ungar 1984). Several species from arid zones have shown higher germination percentages from 20 °C to 30 °C (Rojas-Aréchiga et al. 1998; Khan and Gulzar 2003; Rojas-Aréchiga and Vázquez-Yanes 2000; Manzoor et al. 2017) which is in agreement with our study where the optimum germination occurred at 25 °C. Seeds held for 20 days at 5 °C and 10 °C have been shown to have a higher germination and germination percentage while transferred to 25 °C. Following such a transfer, seeds immediately entered the second phase of growth and their germination rate increased compared to an incubation temperature of 25 °C from the beginning (Simon et al. 1976).
C. cretica germinated well under non-saline and 200 mM NaCl treatment conditions. Accordingly, previous studies have reported that the majority of halophyte seeds had better germination under non-saline or slightly saline conditions (Bewley and Black 1994; Manzoor et al. 2017). Recovery of germination of un-germinated seeds (600, 800, and 1000 mM NaCl) after alleviation of salinity stress is a well-known indicator of osmotic effects, while seed fatality refers to the toxic effects of salinity (Pujol et al. 2000; Song et al. 2005; Rasool et al. 2016). In this study, un-germinated seeds of C. cretica from the different salinity treatments showed high recovery of germination when transferred to distilled water. A high percentage of seed germination in the recovery test indicates that the dormancy is caused by osmotic constraints rather than by salt toxicity (Qu et al. 2008; Lin et al. 2016; Rasool et al. 2016). The dormancy of seeds while keeping the viability is an essential adaptation of halophytes to stress conditions (Gul et al. 2013; Pujol et al. 2000; Qu et al. 2008). Similar results were obtained by Mahmoud et al. (1983) for Limonium axillare, which showed 95 % recovery for 60 %–100 % seawater treatments. Ajmal Khan and Ungar (1998) also observed a quick recovery in Suaeda fruticosa seeds and Manzoor et al. (2017) reported similar behavior for Zygophyllum propinquum. However, seeds of species such as Sporobolus ioclados showed poor recovery response when transferred to distilled water after 20 days salinity (Khan and Gulzar 2003).
Therefore, seed survival could be a more proper measurement to evaluate the success of plants under hypersaline conditions rather than germination, since the ability of seeds to germinate after the alleviation of salinity would be a selective advantage (Ungar 1995). This finding is consistent with the high seedling emergence of C. cretica in its natural habitat after rainfall in spring when soil salinity, temperature, and moisture become more favorable for germination. Khan (1999) organized/classified the halophytes into three categories as marginally tolerant (able to germinate at 125 mM NaCl), moderately tolerant (able to germinate at 500 mM NaCl) and highly tolerant (able to germinate at 800 mM NaCl or higher). We have shown that the seeds and plantlets of C. cretica exhibited moderate tolerance to NaCl. On the other hand, Ajmal Khan and Ungar (1998) reported the growth promotion of C. cretica at 425 mM salinity; moreover, they stated that a salinity level of 850 mM NaCl did not have any significant effect on C. cretica growth. Altogether, we hypothesize that not only C. cretica as a halophyte plant, but also its natural habitat (i.e., soil microbiota) around Gav Khooni may have a vital role to increase the salinity tolerance of the plant.
Several species of the most abundant genera found in the rhizosphere of C. cretica have earlier been found to be halotolerant or halophytic (see review by Ruppel et al. 2013), among them Bacillus spp., Halomonas (Mapelli et al. 2013) and Planctomyces (e.g. P. brasiliensis; Schlesner 1989). For several plants it has been found that microorganisms in the rhizosphere (PGPRs, plant growth promoting rhizobacteria) are able to confer tolerance to salts, among them Bacillus with mungbean (Islam et al. 2016), microbial consortia with Achromobacter, Enterobacter, Microbacterium, Pseudomonas and Serratia, in avocado (Barra et al. 2016) or Bacillus thuringiensis with chickpea (Egamberdieva et al. 2017). Also, Jeotgalibacillus is a genus well known for its salt tolerance (Yaakop et al. 2016). Pelagibius has been found in Jerusalem artichoke, a highly salt tolerant plant (Yang et al. 2016). Except for Halomonas, the above genera have also been found in the rhizosphere, along with Paenisporosarcina that has previously been described as highly salt tolerant (Van Horn et al. 2014). Further, Truepera is also known to comprise halophilic species, like T. radiovictrix (Albuquerque et al. 2005).
However, to which extent the abovementioned halophilic microorganisms may confer tolerance to salinity is still uncertain. In this sense, several mechanisms have been proposed.
Osmotic and ionic stress in plants lead to nutritional imbalance, morphological damages and reduced photosynthesis (Ahmad et al. 2013). It is attributed to PGPRs to increase the ability of the crop to reduce salt stress for the plants (Shrivastava and Kumar 2015). Some of the PGPR effects involve increased germination rate, root and shoot growth, leaf area index, chlorophyll content, protein content, nutrient availability, and delayed leaf senescence stresses (Habib and Kausar, 2016). PGPRs have been found, amongst many other crops, for wheat, chickpea, onion and groundnut (Dasgupta et al. 2015) and it has been suggested that they promote plant growth by altering the selectivity of Na+, K+, and Ca2+ and help to sustain a higher K+/Na+ ratio in plants under salt stress (Hamdia et al. 2004).
In the present study, various halotolerant bacteria were recorded in the soil and rhizosphere samples such as Inmirania sp., a Gamma-Proteobacterium known for halophilic representatives (e.g. the thermophilic, facultative autotrophic, sulfur-oxidizing I. thermothiophila; Slobodkina et al. 2016) as well as Zhihengiuella, a halophilic genus found in tidal flats (Baik et al. 2011). In addition, most of the other genera contributing to the differentiation of the rhizosphere and bulk soils are also known from other saline habitats. Little information, however, is available concerning their respective roles for plant growth and vigor. Tepidanerobacter species are known for their syntrophic interactions with methanogens (Westerholm et al. 2011) suggesting that in this habitat anaerobiosis may occur that eventually leads to liberation of methane, and a similar role may be attributed to the hydrogen-releasing Hydrogenispora (Liu et al. 2014).
Conclusion
Our results provide evidence of salt stress responses of C. cretica at the plant level, and characteristics of the microbiota in the rhizosphere and bulk soil of this habitat, which is rich in various halotolerant bacteria. Still, the potential role for mitigating saline stress by the bacterial associates of C. cretica, most of them microbial dark matter that is yet unculturable, requires further investigation. Also, the contributions of archaea and eukaryotic microorganisms like fungi will have to be investigated.
References
Ahmad M, Zahir ZA, Khalid M, Nazli F, Arshad M (2013) Efficacy of rhizobium and Pseudomonas strains to improve physiology, ionic balance and quality of mung bean under salt-affected conditions on farmer's fields. Plant Physiol Biochem 63:170–176. https://doi.org/10.1016/j.plaphy.2012.11.024
Ajmal Khan M, Ungar I (1998) Germination of the salt tolerant shrub Suaeda fruticosa from Pakistan: salinity and temperature responses. Seed Sci Technol 26:657–667
Akhtar SS, Andersen MN, Naveed M, Zahir ZA, Liu F (2015) Interactive effect of biochar and plant growth-promoting bacterial endophytes on ameliorating salinity stress in maize. Funct Plant Biol 42:770–781. https://doi.org/10.1071/FP15054
Albuquerque L, Simoes C, Nobre MF, Pino NM, Battista JR, Silva MT, Rainey FA, de Costa MS (2005) Truepera radiovictrix gen. Nov., sp. nov., a new radiation resistant species and the proposal of Trueperaceae fam. Nov. FEMS Microbiol Lett 247:161–169. https://doi.org/10.1016/j.femsle.2005.05.002
Aliero B (2004) Effects of sulphuric acid, mechanical scarification and wet heat treatments on germination of seeds of African locust bean tree, Parkia biglobosa. Afr J Biotechnol 3:179–181. https://doi.org/10.5897/AJB2004.000-2031
Badger KS, Ungar IA (1989) The effects of salinity and temperature on the germination of the inland halophyte Hordeum jubatum. Can J Bot 67:1420–1425. https://doi.org/10.1139/b89-188
Baik KS, Lim CH, Park SC, Choe HN, Kim HJ, Kim D, Lee KH, Seong CN (2011) Zhihengliuella aestuarii sp. nov., isolated from tidal flat sediment. Int J Syst Evol Microbiol 61:1671–1676. https://doi.org/10.1099/ijs.0.026245-0
Barassi C, Ayrault G, Creus C, Sueldo R, Sobrero M (2006) Seed inoculation with Azospirillum mitigates NaCl effects on lettuce. Sci Hortic 109:8–14. https://doi.org/10.1016/j.scienta.2006.02.025
Barra PJ, Inostroza NG, Acuña JJ, Mora ML, Crowley DE, Jorquera MA (2016) Formulation of bacterial consortia from avocado (Persea americana mill.) and their effect on growth, biomass and superoxide dismutase activity of wheat seedlings under salt stress. Appl Soil Ecol 102:80–91. https://doi.org/10.1016/j.apsoil.2016.02.014
Bewley J, Black M (1994) Seeds: physiology of development and germination. Plenum Press, New York. https://doi.org/10.1007/978-1-4899-1002-8_9
Callahan BJ, Sankaran K, Fukuyama JA, McMurdie PJ, Holmes SP (2016) Bioconductor workflow for microbiome data analysis: from raw reads to community analyses. F1000Research 5. https://doi.org/10.12688/f1000research.8986.1
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci 108:4516–4522. https://doi.org/10.1073/pnas.1000080107
Coleman-Derr D, Tringe SG (2014) Building the crops of tomorrow: advantages of symbiont-based approaches to improving abiotic stress tolerance. Front Microbiol 5:283. https://doi.org/10.3389/fmicb.2014.00283
Dasgupta D, Ghati A, Sarkar A, Sengupta C, Paul G (2015) Application of plant growth promoting rhizobacteria (PGPR) isolated from the rhizosphere of Sesbania bispinosa on the growth of chickpea (Cicer arietinum L.). Int J Curr Microbiol App Sci 4:1033–1042
Egamberdieva D (2009) Alleviation of salt stress by plant growth regulators and IAA producing bacteria in wheat. Acta Physiol Plant 31:861–864. https://doi.org/10.1007/s11738-009-0297-0
Egamberdieva D, Wirth SJ, Shurigin VV, Hashem A, Abd_Allah EF (2017) Endophytic Bacteria Improve Plant Growth, Symbiotic Performance of Chickpea (Cicer arietinum L.) and Induce Suppression of Root Rot Caused by Fusarium solani under Salt Stress. Frontiers in Microbiology 8:1887. https://doi.org/10.3389/fmicb.2017.01887
Etesami H, Beattie GA (2018) Mining halophytes for plant growth-promoting halotolerant bacteria to enhance the salinity tolerance of non-halophytic crops. Frontiers in microbiology 9(148). https://doi.org/10.3389/fmicb.2018.00148
Fang S, Wang J, Wei Z, Zhu Z (2006) Methods to break seed dormancy in Cyclocarya paliurus (Batal) Iljinskaja. Sci Hortic 110:305–309. https://doi.org/10.1016/j.scienta.2006.06.031
Flowers TJ, Colmer TD (2008) Salinity tolerance in halophytes. New Phytol 179:945–963. https://doi.org/10.1111/j.1469-8137.2008.02531.x
Gul B, Ansari R, Flowers TJ, Khan MA (2013) Germination strategies of halophyte seeds under salinity. Environ Exp Bot 92:4–18. https://doi.org/10.1016/j.envexpbot.2012.11.006
Habib SH, Kausar H (2016) Saud HM (2016) plant growth-promoting rhizobacteria enhance salinity stress tolerance in okra through ROS-scavenging enzymes. Biomed Res Int. https://doi.org/10.1155/2016/6284547
Hamdia MAE-S, Shaddad M, Doaa MM (2004) Mechanisms of salt tolerance and interactive effects of Azospirillum brasilense inoculation on maize cultivars grown under salt stress conditions. Plant Growth Regul 44:165–174. https://doi.org/10.1023/B:GROW.0000049414.03099.9b
Hildebrand F, Tadeo R, Voigt AY, Bork P, Raes J (2014) LotuS: an efficient and user-friendly OTU processing pipeline. Microbiome 2:30. https://doi.org/10.1186/2049-2618-2-30
Islam F, Yasmeen T, Arif MS, Ali S, Ali B, Hameed S, Zhou W (2016) Plant growth promoting bacteria confer salt tolerance in Vigna radiata by up-regulating antioxidant defense and biological soil fertility. Plant Growth Regul 80:23–36. https://doi.org/10.1007/s10725-015-0142-y
Joosen RV, Kodde J, Willems LA, Ligterink W, van der Plas LH, Hilhorst HW (2010) Germinator: a software package for high-throughput scoring and curve fitting of Arabidopsis seed germination. Plant J 62:148–159. https://doi.org/10.1111/j.1365-313X.2009.04116.x
Khan MA (1999) Comparative influence of salinity and temperature on the germination of subtropical halophytes. Halophyte uses in different climates I: ecological and ecophysiological studies 13:77–88
Khan MA, Gulzar S (2003) Light, salinity, and temperature effects on the seed germination of perennial grasses. Am J Bot 90:131–134. https://doi.org/10.3732/ajb.90.1.131
Khan MA, Ungar IA (1984) Seed polymorphism and germination responses to salinity stress in Atriplex triangularis Willd. Bot Gaz 145:487–494. https://doi.org/10.1086/337483
Khan M, Gul B, Weber D (2002) Seed germination in relation to salinity and temperature in Sarcobatus vermiculatus. Biol Plant 45:133–135. https://doi.org/10.1023/A:1015133515568
Kohler J, Hernández JA, Caravaca F, Roldán A (2009) Induction of antioxidant enzymes is involved in the greater effectiveness of a PGPR versus AM fungi with respect to increasing the tolerance of lettuce to severe salt stress. Environ Exp Bot 65:245–252. https://doi.org/10.1016/j.envexpbot.2008.09.008
Lin J, Shao S, Wang Y, Qi M, Lin L, Yan X (2016) Germination responses of the halophyte Chloris virgata to temperature and reduced water potential caused by salinity, alkalinity and drought stress. Grass Forage Sci 71:507–514. https://doi.org/10.1111/gfs.12218
Liu Y, Qiao J-T, Yuan X-Z, Guo R-B, Qiu Y-L (2014) Hydrogenispora ethanolica gen. Nov., sp. nov., an anaerobic carbohydrate-fermenting bacterium from anaerobic sludge. Int J Syst Evol Microbiol 64:1756–1762. https://doi.org/10.1099/ijs.0.060186-0
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Mahmoud A, El-Sheikh A, Abd El Basit S (1983) Germination of Artemisia abyssinica sch. King Saud University, Bip. Journal of the College of Science
Manzoor S, Hameed A, Khan MA, Gul B (2017) Seed germination ecology of a medicinal halophyte Zygophyllum propinquum: responses to abiotic factors. Flora 233:163–117. https://doi.org/10.1016/j.flora.2017.06.004
Mapelli F, Marasco R, Rolli E, Barbato M, Cherif H, Guesmi A, Ouzari I, Daffonchio D (2013) Borin S (2013) potential for plant growth promotion of rhizobacteria associated with Salicornia growing in Tunisian hypersaline soils. Biomed Res Int. https://doi.org/10.1155/2013/248078
Masella AP, Bartram AK, Truszkowski JM, Brown DG, Neufeld JD (2012) PANDAseq: paired-end assembler for illumina sequences. BMC Bioinformatics 13:31. https://doi.org/10.1186/1471-2105-13-3
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PloS one 8:e61217. https://doi.org/10.1371/journal.pone.0061217
McMurdie PJ, Holmes S (2014) Shiny-phyloseq: web application for interactive microbiome analysis with provenance tracking. Bioinformatics 31:282–283. https://doi.org/10.1093/bioinformatics/btu616
Priyashree S, Jha S, Pattanayak S (2010) A review on Cressa cretica Linn.: a halophytic plant. Pharmacogn Rev 4: 161. https://doi.org/10.4103/0973-7847.70910
Pujol JA, CALVO JF, Ramirez-Diaz L (2000) Recovery of germination from different osmotic conditions by four halophytes from southeastern Spain. Ann Bot 85:279–286. https://doi.org/10.1006/anbo.1999.1028
Qin Y, Druzhinina IS, Pan X, Yuan Z-L (2016) Microbially mediated plant salt tolerance and microbiome-based solutions for saline agriculture. Biotechnol Adv 34(7):1245–1259. https://doi.org/10.1016/j.biotechadv.2016.08.005
Qu X, Baskin JM, Wang L, Huang Z (2008) Effects of cold stratification, temperature, light and salinity on seed germination and radicle growth of the desert halophyte shrub, Kalidium caspicum (Chenopodiaceae). Plant Growth Regul 54:241–248. https://doi.org/10.1007/s10725-007-9246-3
R Core Team (2015) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Rasool SG, Hameed A, Ahmed MZ, Khan MA, Gul B (2016) Comparison of seed germination and recovery responses of a salt marsh halophyte Halopeplis perfoliata to osmotic and ionic treatments. Pak J Bot 48:1335–1343
Redman RS, Sheehan KB, Stout RG, Rodriguez RJ, Henson JM (2002) Thermotolerance generated by plant/fungal symbiosis. Science 298:1581–1581. https://doi.org/10.1126/science.1072191
Rodriguez RJ, Henson J, Van Volkenburgh E, Hoy M, Wright L, Beckwith F, Kim Y-O, Redman RS (2008) Stress tolerance in plants via habitat-adapted symbiosis. The ISME Journal 2:404. https://doi.org/10.1038/ismej.2007.106
Rojas-Aréchiga M, Vázquez-Yanes C (2000) Cactus seed germination: a review. J Arid Environ 44:85–104. https://doi.org/10.1006/jare.1999.0582
Rojas-Aréchiga M, Vázquez-Yanes C, Orozco-Segovia A (1998) Seed response to temperature of Mexican cacti species from two life forms: an ecophysiological interpretation. Plant Ecol 135:207–214. https://doi.org/10.1023/A:1009757227493
Roozitalab MH, Siadat H, Farshad A (2018) The soils of Iran. Springer. https://doi.org/10.1007/978-3-319-69048-3
Roy SJ, Negrão S, Tester M (2014) Salt resistant crop plants. Curr Opin Biotechnol 26:115–124. https://doi.org/10.1016/j.copbio.2013.12.004
Ruppel S, Franken P, Witzel K (2013) Properties of the halophyte microbiome and their implications for plant salt tolerance. Funct Plant Biol 40:940–951. https://doi.org/10.1071/FP12355
Schlesner H (1989) Planctomyces brasiliensis sp. nov., a halotolerant bacterium from a salt pit. Syst Appl Microbiol 12:159–161. https://doi.org/10.1016/S0723-2020(89)80008-6
Schmieder R, Edwards R (2011) Quality control and preprocessing of metagenomic datasets. Bioinformatics 27:863–864. https://doi.org/10.1093/bioinformatics/btr026
Shrivastava P, Kumar R (2015) Soil salinity: a serious environmental issue and plant growth promoting bacteria as one of the tools for its alleviation. Saudi Journal of Biological Sciences 22:123–131. https://doi.org/10.1016/j.sjbs.2014.12.001
Simon E, Minchin A, McMenamin MM, Smith J (1976) The low temperature limit for seed germination. New Phytol 77:301–311. https://doi.org/10.1111/j.1469-8137.1976.tb01519.x
Slobodkina GB, Baslerov RV, Novikov AA, Viryasov MB, Bonch-Osmolovskaya EA, Slobodkin AI (2016) Inmirania thermothiophila gen. Nov., sp. nov., a thermophilic, facultatively autotrophic, sulfur-oxidizing gammaproteobacterium isolated from a shallow-sea hydrothermal vent. Int J Syst Evol Microbiol 66:701–706. https://doi.org/10.1099/ijsem.0.000773
Song J, Feng G, Tian C, Zhang F (2005) Strategies for adaptation of Suaeda physophora, Haloxylon ammodendron and Haloxylon persicum to a saline environment during seed-germination stage. Ann Bot 96:399–405. https://doi.org/10.1093/aob/mci196
Tetetay D (1998) Germination of Acacia origena, Acacia pilispina and Pterolobium stellatum in response to different pre-sowing seed treatments, temperature and light https://doi.org/10.1006/jare.1997.0332
Tigabu M, Oden P (2001) Effect of scarification, gibberellic acid and temperature on seed germination of two multipurpose Albizia species from Ethiopia. Seed Sci Technol 29:11–20
Tiwari S, Singh P, Tiwari R, Meena K. K, Yandigeri M, Singh D. P, et al. (2011). Salt-tolerant rhizobacteria-mediated induced tolerance in wheat (Triticum aestivum) and chemical diversity in rhizosphere enhance plant growth. Biol. Fertil. Soils 47:907 https://doi.org/10.1007/s00374-011-0598-5
Ungar I (1995) Seed germination and seed-Bank ecology in halophytes. In: Kigel J, Galili G (eds) Seed development and germination. Marcel Dekker, New York, pp 599–628. https://doi.org/10.1201/9780203740071-23
Upadhyay S, Singh D (2015) Effect of salt-tolerant plant growth-promoting rhizobacteria on wheat plants and soil health in a saline environment. Plant Biol 17:288–293. https://doi.org/10.1111/plb.12173
Van Horn DJ, Okie JG, Buelow HN, Gooseff MN, Barrett JE, Takacs-Vesbach CD (2014) Soil microbial responses to increased moisture and organic resources along a salinity gradient in a polar desert. Appl Environ Microbiol 80:3034–3043. https://doi.org/10.1128/AEM.03414-13
Vilela A, Ravetta D (2001) The effect of seed scarification and soil-media on germination, growth, storage, and survival of seedlings of five species of Prosopis L.(Mimosaceae). J Arid Environ 48:171–184. https://doi.org/10.1006/jare.2000.0735
Wang Q, Dodd IC, Belimov AA, Jiang F (2016) Rhizosphere bacteria containing 1-aminocyclopropane-1-carboxylate deaminase increase growth and photosynthesis of pea plants under salt stress by limiting Na+ accumulation. Funct Plant Biol 43:161–172. https://doi.org/10.1071/FP15200
Weber D, Ansari R, Gul B, Khan MA (2007) Potential of halophytes as source of edible oil. J Arid Environ 68:315–321. https://doi.org/10.1016/j.jaridenv.2006.05.010
Westerholm M, Roos S, Schnürer A (2011) Tepidanaerobacter acetatoxydans sp. nov., an anaerobic, syntrophic acetate-oxidizing bacterium isolated from two ammonium-enriched mesophilic methanogenic processes. Syst Appl Microbiol 34:260–266. https://doi.org/10.1016/j.syapm.2010.11.018
Yaakop AS, Chan K-G, Ee R, Lim YL, Lee S-K, Manan FA, Goh KM (2016) Characterization of the mechanism of prolonged adaptation to osmotic stress of Jeotgalibacillus malaysiensis via genome and transcriptome sequencing analyses. Sci Rep 6:33660. https://doi.org/10.1038/srep33660
Yang H, Hu J, Long X, Liu Z, Rengel Z (2016) Salinity altered root distribution and increased diversity of bacterial communities in the rhizosphere soil of Jerusalem artichoke Scientific Reports 6. https://doi.org/10.1038/srep20687
Yildirim E, Taylor A; Spittler TD (2006) Ameliorative effects of biological treatments on growth of squash plants under salt stress. Sciencia Horticulturae 111, 1–6. https://doi.org/10.1016/j.scienta.2006.08.003
Yuan Z, Druzhinina IS, Labbé J, Redman R, Qin Y, Rodriguez R, Zhang C, Tuskan GA, Lin F (2016) Specialized microbiome of a halophyte and its role in helping non-host plants to withstand salinity. Sci Rep 6:32467. https://doi.org/10.1038/srep32467
Zhu F., Qu L, Hong X, Sun X. (2011). Isolation and characterization of a phosphate-solubilizing halophilic bacterium Kushneria sp. YCWA18 from Daqiao Saltern on the coast of Yellow Sea of China. Evid. Based Complement. Altern. Med.. https://doi.org/10.1155/2011/615032
Acknowledgements
Open access funding provided by University of Innsbruck and Medical University of Innsbruck.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Etemadi, N., Müller, M., Etemadi, M. et al. Salt tolerance of Cressa cretica and its rhizosphere microbiota. Biologia 75, 355–366 (2020). https://doi.org/10.2478/s11756-020-00416-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-020-00416-y