Abstract
MicroRNA (miRNA) is an endogenous non-protein coding small RNA molecule that negatively regulates gene expression by the degradation of messenger RNA (mRNA) or the suppression of mRNA translation. miRNA plays important roles in physiologic processes such as cellular development, differentiation, proliferation, apoptosis, and stem cell self-renewal. Studies show that deregulation of miRNA expression is closely associated with tumorigenicity, invasion, and metastasis. The functionality of aberrant miRNAs in cancer could act either as oncogenes or tumor suppressors during tumor initiation and progression. Similar to protein-coding gene regulation, dysregulation of miRNAs may be related to changes in miRNA gene copy numbers, epigenetic modulation, polymorphisms, or biogenesis modifications. Elucidation of the miRNA expression profiles (miRNomes) of many types of cancers is starting to decode the regulatory network of miRNA-mRNA interactions from a systems biology perspective. Experimental evidence demonstrates that modulation of specific miRNA alterations in cancer cells using miRNA replacement or anti-miRNA technologies can restore miRNA activities and repair gene regulatory networks affecting apoptotic signaling pathways or drug sensitivity, and improve the outcome of treatment. Numerous animal studies for miRNA-based therapy offer the hope of targeting miRNAs as an alternative cancer treatment. Developing the small molecules to interfere with miRNAs could be of great pharmaceutical interest in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1. Introduction
The discovery of microRNAs (miRNAs or miRs) heralded a new and an exciting era in biology and began a new chapter in human gene regulation. The miRNAs, a class of endogenous, small, non-coding RNAs (∼22 nucleotides [nt] in length), fine tune gene expression at the post-transcriptional level, mainly through binding to the 3′ untranslated region (UTR) of messenger RNAs (mRNAs). They are involved in stem-cell self-renewal, cellular development, differentiation, proliferation, and apoptosis.[1]
Small miRNAs have big impacts in the cancer development. Among the many miRNAs, a subset has been identified as regulators of neoplastic transformation, tumor progression, invasion, and metastasis as well as of tumor-initiating cells (cancer stem cells). The widespread deregulation of miRNA expression profiles (miRNomes) in diverse types of cancers compared with normal tissues has been unveiled.[2] The oncogenic miRNAs (oncomirs), tumor suppressive miRNAs, and miRNAs associated to cancer metastasis comprise of an important part of cancer genome, and confer pivotal diagnostic and prognostic significance.[3] Moreover, cancer-associated miRNAs are proving worthwhile for developing effective cancer biomarkers for individualized medicine, and as potential therapeutic targets. Akin to small interfering RNA technology applications, numerous miRNAs have been assessed in research laboratories for treating various diseases, including cancers, some of which have exhibited promising results in cultured cells (reviewed by Wang and Wu[4] ). Approaches to effective delivery of miRNAs are still under investigation, and many challenges still remain.
2. MicroRNA (miRNA) Regulation
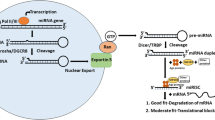
miRNAs are endogenous, small, non-coding RNA molecules that are often encoded within the intronic region of protein-coding genes, but can also be found as independent transcription units or in polycystronic clusters.[5] The miRNA genes are transcribed in the nucleus predominantly by RNA polymerase II into primary miRNAs (pri-miRNAs), which are hundreds to thousands of nucleotides in length and contain a 5′ 7-methylguanosine cap and a 3′ polyA tail. The microprocessor complex formed by the RNAse III enzyme Drosha and DGCR8 (also known as Pasha) cleaves the pri-miRNAs into ∼70 nt fragments of precursor miRNAs (pre-miRNAs) with a two nucleotide 3′ hydroxyl overhang in the nucleus. With the help of RAN-guanosine triphosphate (GTP) and exportin-5 in the nuclear membrane, pre-miRNAs are transported to the cytoplasm where a secondary cleavage is performed by another RNase III enzyme, Dicer, and the ∼21 nt duplex of mature miRNAs is formed. One strand of a mature miRNA is integrated into the RNA-induced silencing complex (RISC), containing Argonaute proteins to target the mRNAs. In the vast majority of cases, the mature miRNAs bind to the 3′ UTRs of mRNAs. Occasionally, the 5′ UTR[6] or even the amino acid coding sequence[7] can also be the binding sites. Gene silencing through degradation of mRNA or translational repression is a general consequence of miRNA binding, although in rare cases, a miRNA could increase gene expression.[8]
The molecular mechanisms of miRNA regulation are not clear to date. Recent studies of pathologic processes such as cancer development are aiding our understanding of miRNA function. Similar to the regulation of protein-coding genes, each step of miRNA regulation could be affected during the progression from genetic makeup to functional molecules. Emerging evidence shows that deregulation of miRNA molecules in cancer cells seems to involve the interplay of multiple mechanisms, as outlined in sections 2.1–2.4.
2.1 Genomic miRNA Copy Number Changes
Changes in miRNA copy number in the genome correlate with miRNA expression level. The gain of miRNA function through the gene amplification is documented by several miRNAs such as miR-21[9] and the miR-17-92 gene cluster.[10] The miR-17-92 gene cluster was mapped into a chromosome region that is frequently amplified in a subset of human B-cell lymphoma[11] and overexpressed in a variety of other human cancers. Loss of miRNA function is often due to the deletion of miRNA genes in chromosomal sites. For example, miRNA15a and miR-16 are deleted in the majority of chronic lymphocytic leukemias and in a subset of mantle cell lymphoma and prostate cancers.[12] A high frequency of genomic miRNA copy number changes was found in solid tumors as well. For example, changes in gene copy number for 41 miRNAs have been observed in breast cancer, ovarian cancer, and melanoma.[9] Interestingly, Dicer 1, Argonaute 2, and other miRNA-associated genes were also found to change in copy number in cancer cells.[9] This may partly contribute to the changes in miRNA expression observed.
2.2 Epigenetic Regulation of miRNAs
Epigenetic changes contribute to aberrant miRNA expression in several malignancies. Changes in DNA methylation of miRNA gene promoters or chromatin histone deacetylases (HDAC) of several miRNAs in various cancer cell lines or cancer samples have been reported.[13,14] Moreover, treatment with DNA methylation inhibitors (5-aza-2′-deoxycytidine) or HDAC inhibitors were shown to significantly change the miRNA expression profile.[15]
2.3 Transcriptional Regulation of miRNAs
Accumulating evidence demonstrates that a subset of miRNA genes is regulated by known transcription factors. It is clear that MYC regulates the expression of the miR-17-92 cluster through directly binding to E-boxes of the promoter of the miR-17-92 gene.[16] Tumor protein p53 (TP53) binds directly to the miR-34 gene promoter to activate its transcription.[17] However, the factors initiating transcription of other miRNAs remain largely unknown.
2.4 Polymorphisms and Mutations in miRNAs
The mutation of mature RNAs changes the miRNA-mRNA interaction and specificity, and can abrogate miRNA regulatory effects. In particular, drug-related miRNA polymorphisms have drawn much attention and are discussed below (section 3.2). We expect that a larger-scale cancer genome sequencing will provide more information on miRNA mutations in different types of cancers.
3. miRNA Modulation
Given the importance of miRNAs in regulating cellular differentiation and proliferation, it is not surprising that their dysregulation is linked to cancer. In cancer, miRNAs function as regulatory molecules, acting as oncogenes or tumor suppressors.[18] Either miRNA replacements or miRNA inhibitors could be introduced into the cells to restore the physiologic function of miRNAs.[4] Progress has been made to target every step of miRNA regulation, from endogenous induction of miRNA gene expression with small molecules, to enzymatic modification (i.e. Drosha or Dicer, Argonautes) involved in miRNA biogenesis.[19] Some new and creative approaches include miRNA mimics,[20] multiple-target anti-miRNAs,[21] and chemical inhibitors (e.g. miR-21 inhibitor diazobenzene),[22] to increase the specificity and decrease the off-target effects and other potential adverse effects. Modulation of miRNA by various strategies could lead to different biologic effects for certain disease treatments. The major applications in cancer are described in sections 3.1–3.4.
3.1 Modulation of miRNAs Mediates Apoptosis
miRNA is strongly related to the apoptosis signaling, and multiple miRNAs have been found to enhance or inhibit the apoptotic pathway following their changes during cancer development. miR-21 is overexpressed in various tumors, and is functionally considered as an oncomir; multiple targets of miR-21 have been identified and mapped to different signaling pathways including the anti-apoptotic signaling pathways (reviewed by Selcuklu et al.[23] ). Many studies have focused on inhibiting miR-21, and have demonstrated that such inhibition leads to the induction of programmed cell death, suggesting a promising miRNA treatment for cancer.[24–26] On the other hand, reduced expression of miR-15, miR-16, and let-7 has been observed in different types of cancers, and as one consequence, anti-apoptotic genes are activated in these cancer cells.[27,28] Restoration of these miRNAs triggers activation of apoptotic signaling pathways.[29,30] Other examples have been reported; for example, miR-491 induces apoptosis by targeting BCL-X(L) in colorectal cancer cells,[31] and transfection of anti-miR-24 oligonucleotides induces apoptosis in several cell lines.[7] Collectively, apoptotic and anti-apoptotic genes are the core hub of the gene network regulated by miRNAs. This well studied signaling pathway is one of the mechanisms of action for modulating miRNAs.
3.2 Modulation of miRNAs Sensitizes Chemo- or Radiotherapy
Multidrug resistance of cancer cell subpopulations to conventional chemo- or radiotherapy is the main cause of recurrence or relapse. Seeking agents or molecules to enhance cancer cell sensitivity to therapy is the long-term goal to improve the therapeutic efficacy.[32] By definition of a sensitivity enhancer, the given miRNAs themselves (at certain expression levels) may not have any or have only minor effects on cancer cell proliferation, apoptosis, or cell cycle; however, in combination with another treatment, additive or synergistic effects can be observed. This effect of miRNAs provides a novel platform for targeting resistant cancer cells. For example, knockdown of miR-221 and/or miR-222 sensitized MDA-MB-468 cells to tamoxifen-induced cell growth arrest and apoptosis.[33] miR15b and miR-16 maintain the sensitivity of gastric cancer cells to chemotherapy through inhibition of BCL1.[33] miR-326 is inversely related to multidrug resistance-associated protein (MRP1/ABCC1) expression in the VP-16-resistant, multidrug-resistant cell line MCF-7/VP.[34] It is possible that targeting miR-326 could be utilized for preventing and reversing multidrug resistance in tumor cells. miR-34 restoration in pancreatic cancer stem cells increases the sensitivity to docetaxel, cisplatin, and gemcitabine treatment, and irradiation exposure. The effect of miRNAs inducing apoptosis or increasing the sensitivity to treatment is likely to be determined by the expression level of miRNAs. When a given miRNA expression level goes over the critical threshold, the cell death process will take place. At this point, we are unable to quantify miRNA dose-dependent effects with the biological phenomenon, but a mathematical model could be proposed to test this idea. Taken together, increasing the cancer cell sensitivity by miRNAs provides an additional layer of miRNA modulation to treat cancer cells, at least in vitro.
3.3 Modulation of miRNAs Induces Cancer Cell Differentiation
Emerging evidence suggests that miRNA can regulate cell-fate decisions. A subgroup of miRNAs is markedly reduced in the stem-cell state and increased during differentiation. As the cancer stem-cell theory is beginning to be accepted, induction of cancer stem-cell differentiation is being recognized as a future direction for developing new anticancer agents.[35] miRNA expression data revealed that miRNAs are differentially expressed in breast cancer stem cells and in differentiated breast cancer cells.[36] By this comparison, let-7 was found to be a gatekeeper for cancer cell differentiation. Breast cancer stem cells derived from either cultured mammosphere cells or clinical cancer specimens expressed lower levels of let-7 and higher levels of RAS and HMGA2 proteins, which are negatively regulated by let-7. In contrast, when these cells were undergoing differentiation, the let-7 expression increased. Moreover, let-7 expression inhibits tumorigenesis and metastasis in nonobese diabetic (NOD)/severe combined immunodeficiency (SCID) mouse models.[37] More recently, miR-200c was linked to breast cancer stem cell differentiation through inhibition of the stem cell self-renewal factor BMI1 gene.[38] Taulli et al.[39] reported that the muscle-specific miR-206 blocks human rhabdomyosarcoma growth and promotes myogenic differentiation in xenotransplanted mice; furthermore, it appears that the met gene is the primary target of miR-206. Taken together, increasing evidence supports that the re-expression of specific miRNAs could induce cancer cell differentiation, which could be used for cancer therapy.
3.4 Individualized Therapy
Cancer cells are heterogeneous, even within individual tumors, and subcloncal cell populations give rise to different treatment responses, suggesting the need for personalized medicine. miRNA appears to be a reliable biomarker for diagnosis, prognosis, and treatment evaluation in different types of cancers.[40–42] miRNA expression profiles could be used to define the cell types,[2,3] thereby allowing detection of the miRNA biomarker or expression signatures from patients in order to develop individualized therapeutic plans, possibly even targeting specific miRNAs for a given patient. Furthermore, the understanding of how an individual’s genetic inheritance of miRNA polymorphisms affects the body’s response to certain drugs will be key to creating drugs with greater efficacy and safety. miRNA polymorphisms can occur in the miRNA itself or in its binding site in the mRNA 3′ UTR, resulting in loss of miRNA function. Some drug-target related miRNA polymorphisms have been reported. For example, the existence of a miR-24 binding site single-nucleotide polymorphism (SNP) 829C/T in the dihydrofolate reductase 3′ UTR contributes to dihydrofolate reductase overexpression and methotrexate resistance.[43] Thus, the monitoring or detection of miRNA polymorphisms could lead to the development of successful miRNA therapeutics and improved evaluation of the pharmacologic response.
4. The Complexity of miRNA Regulatory Network
The discovery of miRNAs has revolutionized the regulation of gene expression. These tiny RNA molecules likely act as molecular switches in the extensive regulatory web that involves thousands of transcripts. Most importantly, accumulating evidence suggests that numerous miRNAs are aberrantly expressed in human cancers. The roles of miRNAs as molecular switches in the integrated circuit of cancer cell regulation are emerging. The initial theoretical analysis indicated that as many as 30% of genes in the human genome may be the targets of miRNA.[44] More recently, this number has been estimated as >60%[45] or even 90% of the human genes that are the target of miRNAs.[46] Miranda et al.[46] found that most of the target genes encode proteins functioning as transcription factors, receptors, hydrolase and nucleotide binding proteins, and are involved in biologic processes such as apoptosis, the cell cycle, developmental processes, and signal transduction. Furthermore, miRNAs work as a cluster to co-target specific pathways. The miR-72-93 cluster includes eight members, which are co-localized on chromosome 13 within a 10 kb span, and act in a combinational fashion to exert a biologic effect.[47] In humans, 13 different let-7 family members[48] are associated with cellular development and differentiation. The redundant and compensatory role of the let-7 family exhibits the complexity of miRNA regulation. Thus, it is time to rethink miRNA function from a systems biology point of view, taking the miRNA network into account to design new therapeutic medicines.[49] Current miRNA-based therapy in vitro and in vivo only targets single miRNAs, which may not be sufficient to reach the maximum potential effects. Several methods have begun to address this bottleneck limitation. Innovative ideas such as ‘miRNA sponges’[50] and ‘multiple-target anti-miRNAs’[21] provide a starting point. The tools are available to manipulate a group of miRNAs if we know the exact miRNA cluster co-targeting similar pathways, which will open up an exciting avenue of miRNA-based treatment for several diseases, including cancer.
5. miRNA Modulation Towards Clinical Trials
miRNAs are central components of the gene regulatory network, and aberrant expression of miRNAs contribute to a wide variety of diseases, including cancer. Numerous experiments in cultured cells and xenograft animal models demonstrate that modulating miRNAs could serve a therapeutic purpose.[51–53] Delivery techniques developed for small interfering RNA therapeutics have been adapted to miRNA-based experimental therapy. The recent systemic delivery of miRNAs in animal models via various routes is listed in table I. No acute or sub-acute toxicity was observed in mice and non-human primates treated with locked nucleotide acid (LNA) anti-miRs.[58,59] Due to the advancement of the delivery technology and the relative safety of miRNAs-based treatment in the preclinical trial,[4] a clinical trial of LNA-antimiR™-122 (SPC3649) in human subjects was started in 2008.[60] This first miRNA-based therapy in clinical trials for hepatitis C virus will pave the way for treating other diseases, including cancer. As with other gene therapies, cautious optimism is recommended, because few gene-modifying drugs have been successfully launched in the market.
6. Conclusion and Outlook
Non-coding RNAs in the genome, particularly miRNAs, which were previously considered ‘junk’ DNA, have now been deciphered as important gene expression regulators. miRNAs play important roles in cellular development, the cell cycle, stem cell self-renewal, proliferation, and apoptosis, and are involved in certain diseases such as cardiac disorders, diabetes mellitus, and cancer. A predicted ∼1000 miRNAs in the human genome could regulate more than 60% of the ∼25 000 protein coding genes.[45] Elucidation of the broad miRNA-mRNA interaction has revolutionized our understanding of genetic regulatory networks and the development of diseases such as cancer. The aberrant expression of miRNAs in almost all types of cancers has been intensively scrutinized relative to normal tissues. Defining the driving force of abnormal changes of miRNAs in tumorigenicity, and modulation of these key miRNAs in the complex gene regulatory network could trigger apoptotic pathways, enhance the sensitivity of cancer cells to chemo- or radiotherapy, and induce cancer cell differentiation, possibly even providing individualized therapy for cancer patients. Systemic delivery of targeted miRNAs in vivo has provided the pharmacologic tool to develop miRNA drugs, clinical trials of which are eagerly awaited.
References
Ambros V. MicroRNA pathways in flies and worms: growth, death, fat, stress, and timing. Cell 2003; 113: 673–6
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature 2005; 435: 834–8
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer 2006; 6: 857–66
Wang V, Wu W. MicroRNA-based therapeutics for cancer. BioDrugs 2009; 23: 15–23
Kim VN, Nam JW. Genomics of microRNA. Trends Genet 2006 Mar; 22 (3): 165–73
Lytle JR, Yario TA, Steitz JA. Target mRNAs are repressed as efficiently bymicroRNA-binding sites in the 5’ UTR as in the 30 UTR. Proc Natl Acad Sci U S A 2007; 104: 9667–72
Qin W, Shi Y, Zhao B, et al. miR-24 Regulates Apoptosis by Targeting the Open Reading Frame (ORF) Region of FAF1 in Cancer Cells. PLoS One 2010; 5: e9429
Vasudevan S, Tong Y, Steitz JA. Switching from repression to activation: microRNAs can up-regulate translation. Science 2007; 318: 1931–4
Zhang L, Huang J, Yang N, et al. microRNAs exhibit high frequency genomic alterations in human cancer. Proc Natl Acad Sci U S A 2006; 103: 9136–41
Hayashita Y, Osada H, Tatematsu Y, et al. A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res 2005; 65 (21): 9628–32
Ota A, Tagawa H, Karnan S, et al. Identification and characterization of a novel gene, C13orf25, as a target for 13q31-q32 amplification in malignant lymphoma. Cancer Res 2004; 64: 3087–95
Calin GA, Dumitru CD, Shimizu M, et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A 2002; 99: 15524–9
Scott GK, Mattie MD, Berger CE, et al. Rapid alteration of microRNA levels by histone deacetylase inhibition. Cancer Res 2006; 66: 1277–81
Lujambio A, Ropero S, Ballestar E, et al. Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res 2007; 67: 1424–9
Saito Y, Liang G, Egger G, et al. Specific activation of micro-RNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 2006; 9: 435–43
O’Donnell KA, Wentzel EA, Zeller KI, et al. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005; 435: 839–43
Chang TC, Wentzel EA, Kent OA, et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell 2007; 26: 745–52
Wu W, Sun M, Zou GM, et al. MicroRNA and cancer: current status and prospective. Int J Cancer 2007; 120 (5): 953–60
Winter J, Jung S, Keller S, et al. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol 2009; 11: 228–34
Xiao J, Yang B, Lin H, et al. Novel approaches for genespecific interference via manipulating actions of microRNAs: examination on the pacemaker channel genes HCN2 and HCN4. J Cell Physiol 2007; 212: 285–92
Lu Y, Xiao J, Lin H, et al. A single anti-microRNA antisense oligodeoxyribonucleotide (AMO) targeting multiple microRNAs offers an improved approach for microRNA interference. Nucleic Acids Res 2009; 37: e24
Gumireddy K, Young DD, Xiong X, et al. Small-molecule inhibitors of microRNA miR-21 function. Angew Chem Int Ed Engl 2008; 47: 7482–4
Selcuklu SD, Donoghue MT, Spillane C. miR-21 as a key regulator of oncogenic processes. Biochem Soc Trans 2009; 37: 918–25
Meng F, Henson R, Wehbe-Janek H, et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007; 133 (2): 647–58
Zhu S, Si ML, Wu H, et al. MicroRNA-21 targets the tumor suppressor gene tropomyosin 1 (TPM1). J Biol Chem 2007; 282 (19): 14328–36
Frankel LB, Christoffersen NR, Jacobsen A, et al. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J Biol Chem 2008; 283 (2): 1026–33
Bottoni A, Piccin D, Tagliati F, et al. miR-15a and miR-16-1 down-regulation in pituitary adenomas. J Cell Physiol 2005; 204: 280–5
Takamizawa J, Konishi H, Yanagisawa K, et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 2004; 64: 3753–6
Cimmino A, Calin GA, Fabbri M, et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci U S A 2005; 102: 13944–9
Johnson CD, Esquela-Kerscher A, Stefani G, et al. The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res 2007; 67: 7713–22
Nakano H, Miyazawa T, Kinoshita K, et al. Functional screening identifies a microRNA, miR-491 that induces apoptosis by targeting Bcl-X(L) in colorectal cancer cells. Int J Cancer. Epub 2009 Dec 28
Frank NY, Schatton T, Frank MH. The therapeutic promise of the cancer stem cell concept. J Clin Invest 2010; 120 (1): 41–50
Zhao JJ, Lin J, Yang H, et al. MicroRNA-221/222 negatively regulates estrogen receptor alpha and is associated with tamoxifen resistance in breast cancer. J Biol Chem 2008; 283: 31079–86
Liang Z, Wu H, Xia J, et al. Involvement of miR-326 in chemotherapy resistance of breast cancer through modulating expression of multidrug resistance-associated protein 1. Biochem Pharmacol 2010; 79: 817–24
Gupta PB, Onder TT, Jiang G, et al. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 2009; 138 (4): 645–59
Wang V, Wu W. MicroRNA: a new player in breast cancer development. J Cancer Mol 2007; 3: 133–8
Yu F, Yao H, Zhu P, et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 2007; 131: 1109–23
Shimono Y, Zabala M, Cho RW, et al. Downregulation of miRNA-200c links breast cancer stem cells with normal stem cells. Cell 2009; 138: 592–603
Taulli R, Bersani F, Foglizzo V, et al. The muscle-specific microRNA miR-206 blocks human rhabdomyosarcoma growth in xenotransplanted mice by promoting myogenic differentiation. J Clin Invest 2009; 119: 2366–78
Heneghan HM, Miller N, Lowery AJ, et al. Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Ann Surg 2010; 251: 499–505
Hu Z, Chen X, Zhao Y, et al. Serum microRNA signatures identified in a genome-wide serum microRNA expression profiling predict survival of non-small-cell lung cancer. J Clin Oncol 2010; 28 (10): 1721–6
Gee HE, Camps C, Buffa FM, et al. hsa-miR-210 is amarker of tumor hypoxia and a prognostic factor in head and neck cancer. Cancer 2010; 116 (9): 2148–58
Mishra PJ, Humeniuk R, Longo-Sorbello GS, et al. A miR-24 microRNA binding-site polymorphism in dihydrofolate reductase gene leads to methotrexate resistance. Proc Natl Acad Sci U S A 2007; 104: 13513–8
Lewis BP, Shih IH, Jones-Rhoades MW, et al. Prediction of mammalian microRNA targets. Cell 2003; 115: 787–98
Friedman RC, Farh KK, Burge CB, et al. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 2009; 19 (1): 92–105
Miranda KC, Huynh T, Tay Y, et al. A pattern-based method for the identification of microRNA binding sites and their corresponding heteroduplexes. Cell 2006; 126: 1203–17
Ivanovska I, Cleary MA. Combinatorial microRNAs: working together to make a difference. Cell Cycle 2008; 7: 3137–42
Ruby JG, Jan C, Player C, et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 2006; 127: 1193–207
Sotiropoulou G, Pampalakis G, Lianidou E, et al. Emerging roles of microRNAs as molecular switches in the integrated circuit of the cancer cell. RNA 2009; 15: 1443–61
Ebert MS, Neilson JR, Sharp PA. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat Methods 2007; 4: 721–6
Si ML, Zhu S, Wu H, et al. miR-21-mediated tumor growth. Oncogene 2007; 26 (19): 2799–803
Park SM, Shell S, Radjabi AR, et al. Let-7 prevents early cancer progression by suppressing expression of the embryonic gene HMGA2. Cell Cycle 2007; 6: 2585–90
Takeshita F, Patrawala L, Osaki M, et al. Systemic delivery of synthetic microRNA-16 inhibits the growth of metastatic prostate tumors via downregulation of multiple cellcycle genes. Mol Ther 2010; 18: 181–7
Esquela-Kerscher A, Trang P, Wiggins JF, et al. The let-7 microRNA reduces tumor growth in mouse models of lung cancer. Cell Cycle 2008; 7: 759–64
Kumar MS, Erkeland SJ, Pester RE, et al. Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc Natl Acad Sci U S A 2008; 105: 3903–8
Trang P, Medina PP, Wiggins JF, et al. Regression of murine lung tumors by the let-7 microRNA. Oncogene 2010; 29 (11): 1580–7
Kota J, Chivukula RR, O’Donnell KA, et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 2009; 137: 1005–17
Stenvang J, Lindow M, Kauppinen S. Targeting of microRNAs for therapeutics. Biochem Soc Trans 2008; 36 (Pt 6): 1197–200
Elmén J, Lindow M, Schütz S, et al. LNA-mediated microRNA silencing in non-human primates. Nature 2008; 452 (7189): 896–9
Santaris Pharma begins human clinical testing of the world’s first medicine targeted at a human microRNA [media release]. 2008 May 28 [online] Available from URL: http://www.fier cebiotech.com/press-releases/santaris-pharma-begins-human-clinical-testing-worlds-first-medicine-targeted-human-mi
Acknowledgments
I thank all colleagues who have done work in this rapidly changing field, and apologize to the colleagues whose work in the field was not directly cited in this review due to the highly selected topic. The author has no conflicts of interest that are directly relevant to the content of this review.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Wu, W. MicroRNA: Potential Targets for the Development of Novel Drugs?. Drugs R D 10, 1–8 (2010). https://doi.org/10.2165/11537800-000000000-00000
Published:
Issue Date:
DOI: https://doi.org/10.2165/11537800-000000000-00000