摘要
目的
通过转录组学分析挖掘与新疆梨果中绿原酸生物合成相关的功能基因, 为未来新疆梨的功能性育种提供科学依据。
创新点
明确了在三种新疆梨果发育过程中, 与绿原酸含量变化趋势和绿原酸生物合成相关的差异基因及关键限制性酶。
方法
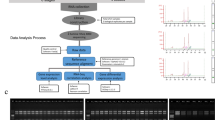
在鸭梨、库尔勒香梨和圆黄等三种梨果的不同发育阶段采集果实样品, 经液相色谱法分析不同发育期果品的绿原酸含量和熊果苷含量差异, 然后选择花后88、103和163天这三个绿原酸含量变化典型的时间点样品提取RNA, 进行转录组测序, 采取无参转录组学方法挖掘梨果中绿原酸生物合成途径的关键酶基因。
结论
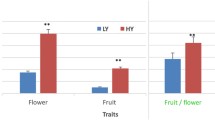
绿原酸含量总体随着梨果的发育呈下降趋势。筛选得到10个与绿原酸生物合成相关的差异基因, 分别为2个4CL基因、1个C4H基因、2个C3’H基因和5个HCT基因, 其中绿原酸生物合成下游途径的C3’H和HCT是关键的限速酶基因。
Similar content being viewed by others
References
Covington MF, Maloof JN, Straume M, et al., 2008. Global transcriptome analysis reveals circadian regulation of key pathways in plant growth and development. Genome Biol, 9:R130. https://doi.org/10.1186/gb-2008-9-8-r130
Dixon RA, Paiva NL, 1995. Stress-induced phenylpropanoid metabolism. Plant Cell, 7(7):1085–1097. https://doi.org/10.1105/tpc.7.7.1085
Dong XG, Wang Z, Tian LM, et al., 2020. De novo assembly of a wild pear (Pyrus betuleafolia) genome. Plant Biotechnol J, 18(2):581–595. https://doi.org/10.1111/pbi.13226
Durazzo A, Lucarini M, 2019. Editorial: the state of science and innovation of bioactive research and applications, health, and diseases. Front Nutr, 6:178. https://doi.org/10.3389/fnut.2019.00178
Han JY, Hwang HS, Choi SW, et al., 2012. Cytochrome P450 CYP716A53v2 catalyzes the formation of protopanaxatriol from protopanaxadiol during ginsenoside biosynthesis in Panax ginseng. Plant Cell Physiol, 53(9): 1535–1545. https://doi.org/10.1093/pcp/pcs106
Hoffmann L, Besseau S, Geoffroy P, et al., 2004. Silencing of hydroxycinnamoy-coenzyme A shikimate/quinate hydroxycinnamoyltransferase affects phenylpropanoid biosynthesis. Plant Cell, 16(6):1446–1465. https://doi.org/10.1105/tpc.020297
Legrand G, Delporte M, Khelifi C, et al., 2016. Identification and characterization of five BAHD acyltransferases involved in hydroxycinnamoyl ester metabolism in chicory. Front Plant Sci, 7:741. https://doi.org/10.3389/fpls.2016.00741
Li B, 2019. Diet-related NCDs in China: more needs to be done. Lancet Public Health, 4(12):E606. https://doi.org/10.1016/S2468-2667(19)30218-X
Li X, Wang TT, Zhou B, et al., 2014. Chemical composition and antioxidant and anti-inflammatory potential of peels and flesh from 10 different pear varieties (Pyrus spp.). Food Chem, 152:531–538. https://doi.org/10.1016/j.foodchem.2013.12.010
Liu F, Chen JR, Tang YH, et al., 2018. Isolation and characterization of cinnamate 4-hydroxylase gene from cultivated ramie (Boehmeria nivea). Biotechnol Biotechnol Equip, 32(2):324–331. https://doi.org/10.1080/13102818.2017.1418675
Liu GQ, Li WS, Zheng PH, et al., 2012. Transcriptomic analysis of ‘Suli’ pear (Pyrus pyrifolia white pear group) buds during the dormancy by RNA-Seq. BMC Genomics, 13:700. https://doi.org/10.1186/1471-2164-13-700
MacLean DD, Murr DP, Deell JR. et al., 2007. Inhibition of PAL, CHS, and ERS1 in ‘Red d’Anjou’ pear (Pyrus communis L.) by 1-MCP. Postharvest Biol Technol, 45(1):46–55. https://doi.org/10.1016/j.postharvbio.2007.01.007
McLean KJ, Hans M, Munro AW, 2012. Cholesterol, an essential molecule: diverse roles involving cytochrome P450 enzymes. Biochem Soc Trans, 40(3):587–593. https://doi.org/10.1042/BST20120077
Sullivan ML, Zarnowski R, 2010. Red clover coumarate 3’-hydroxylase (CYP98A44) is capable of hydroxylating p-coumaroyl-shikimate but not p-coumaroyl-malate: implications for the biosynthesis of phaselic acid. Planta, 231(2):319–328. https://doi.org/10.1007/s00425-009-1054-8
Tuominen LK, Johnson VE, Tsai CJ, 2011. Differential phylogenetic expansions in BAHD acyltransferases across five angiosperm taxa and evidence of divergent expression among Populus paralogues. BMC Genomics, 12:236. https://doi.org/10.1186/1471-2164-12-236
Villegas RJA, Kojima M, 1986. Purification and characterization of hydroxycinnamoyl D-glucose: quinate hydroxycinnamoyl transferase in the root of sweet potato, Ipomoea batatas Lam. J Biol Chem, 261(19):8729–8733. https://doi.org/10.1016/S0021-9258(19)84441-1
Wang D, Hou JX, Wan JD, et al., 2021. Dietary chlorogenic acid ameliorates oxidative stress and improves endothelial function in diabetic mice via Nrf2 activation. J Int Med Res, 49(1):300060520985363. https://doi.org/10.1177/0300060520985363
Wu J, Wang ZW, Shi ZB, et al., 2013. The genome of the pear (Pyrus bretschneideri Rehd.). Genome Res, 23(2):396–408. https://doi.org/10.1101/gr.144311.112
Zhou L, Li CJ, Niu JX, et al., 2018. Identification of miRNAs involved in calyx persistence in Korla fragrant pear (Pyrus sinkiangensis Yu) by high-throughput sequencing. Sci Hortic, 240:344–353. https://doi.org/10.1016/j.scienta.2018.06.026
Acknowledgments
This work was supported by the Major Scientific and Technological Projects of the Xinjiang Production and Construction Corps (Nos. 2017DB006 and 2020KWZ-012). We thank Prof. Liang LIU from Department of Statistics, the University of Georgia, the United States, for his helps on designing research and revising manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Author contributions
Hao WEN performed the experimental research and data analysis, wrote and edited the manuscript. Xi JIANG, Wenqiang WANG, Minyu WU, and Hongjin BAI collected and analyzed the data. Cuiyun WU and Lirong SHEN designed the experimental research, and wrote and revised the manuscript. All authors have read and approved the final manuscript, and therefore, have full access to all the data in the study and take responsibility for the integrity and security of the data.
Compliance with ethics guidelines
Hao WEN, Xi JIANG, Wenqiang WANG, Minyu WU, Hongjin BAI, Cuiyun WU, and Lirong SHEN declare that they have no conflict of interest.
This article does not contain any studies with human or animal subjects performed by any of the authors.
Supplementary information
Tables S1–S3; Figs. S1–S5
Supplementary Information
11585_2022_601_MOESM1_ESM.pdf
Comparative transcriptome analysis of candidate genes involved in chlorogenic acid biosynthesis during fruit development in three pear varieties of Xinjiang Uygur Autonomous Region
Rights and permissions
About this article
Cite this article
Wen, H., Jiang, X., Wang, W. et al. Comparative transcriptome analysis of candidate genes involved in chlorogenic acid biosynthesis during fruit development in three pear varieties of Xinjiang Uygur Autonomous Region. J. Zhejiang Univ. Sci. B 23, 345–351 (2022). https://doi.org/10.1631/jzus.B2100673
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1631/jzus.B2100673