Abstract
MicroRNAs (miRNAs) from milk whey have been considered for their potential as noninvasive biomarkers for milk quality control and disease diagnosis. However, standard protocols for miRNA isolation and quantification from milk whey are not well established. The objective of this study was to compare two methods for the isolation of miRNAs from milk whey. These two methods were modified phenol-based technique (Trizol LS® followed by phenol precipitation, the TP method) and combined phenol and column-based approach (Trizol LS® followed by cleanup using the miRNeasy kit, the TM method). Yield and quality of RNA were rigorously measured using a NanoDrop ND-1000 spectrophotometer and then the distribution of RNA was precisely detected in a Bioanalyzer 2100 instrument by microchip gel electrophoresis. Several endogenous miRNAs (bta-miR-141, bta-miR-146a, bta-miR-148a, bta-miR-200c, bta-miR-362, and bta-miR-375) and an exogenous spike-in synthetic control miRNA (cel-miR-39) were quantified by real-time polymerase chain reaction (PCR) to examine the apparent recovery efficiency of milk whey miRNAs. Both methods could successfully isolate sufficient small RNA (<200 nt) from milk whey, and their yields were quite similar. However, the quantification results show that the total miRNA recovery efficiency by the TM method is superior to that by the TP method. The TM method performed better than the TP for recovery of milk whey miRNA due to its consistency and good repeatability in endogenous and spike-in miRNA recovery. Additionally, quantitative recovery analysis of a spike-in miRNA may be more accurate to reflect the milk whey miRNA recovery efficiency than using traditional RNA quality analysis instruments (NanoDrop or Bioanalyzer 2100).
概要
目的
系统比较两种从液态乳清中分离提取miRNA 的方法。
创新点
首次比较发现提取柱吸附法分离乳清miRNA 的效果优于醇沉淀法。
方法
采用柱吸附法和醇沉淀法分别对乳清中的miRNA进行提取, 采用NanoDrop ND-1000 分光光度计和基于Bioanalyzer 2100 的微芯片凝胶电泳检测RNA 提取质量。通过荧光定量聚合酶链式反应 (PCR) 检测外源加入的miRNA 及乳清内源的miRNA 丰度。确定最优方法后扩大样本容量, 采用20 份牛奶样本进行提取, 并通过荧光定量PCR检测了外源miRNA 提取的稳定性及牛奶内源miRNA 提取的重复性。
结论
乳清miRNA 提取柱吸附法优于醇沉淀法, 且重复性好。
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
MicroRNAs (miRNAs), a class of noncoding RNAs, play important roles in regulating gene expression at the post-transcriptional level (Bartel, 2004). Various types of miRNAs were detected in body fluids, including blood, urine, saliva, and milk (Weber et al., 2010). Milk, a primary nutrition source for infants and a widely used food, contains more than 400 miRNAs (Chen et al., 2010; Zhou et al., 2012). The constitution of miRNAs in milk varies among different species (Hata et al., 2010; Munch et al., 2013; Izumi et al., 2014), the lactation period (Chen et al., 2010), and maternal dietary manipulation (Hata et al., 2010). Moreover, some miRNAs that are abundant in milk have been evidenced for their important functional roles in various biological processes, including immune response (Kosaka et al., 2010; Sun et al., 2013), postnatal growth (Melnik et al., 2013), and lipid metabolism (Fernández-Hernando et al., 2011). Recently, miRNAs in milk have been studied as potential biomarkers for milk quality control (Chen et al., 2010). miRNAs originating from milk whey and the lipid fraction are stable after repeated thawing (Baier et al., 2014). This stability might be attributed to the protective effects of exosomes in the milk whey (Zhang et al., 2011). It is confirmed that milk miRNAs are tolerant to some severe conditions, such as the stomach acidic system and RNase treatment (Zhang et al., 2011), similar to the roles of plant miR168a in mammalians. All of these studies will surely shed light on developing diagnostic biomarkers for breast diseases and provide deep insight into understanding the functional importance of milk (Farazi et al., 2011). Despite the promise for developing milk miRNA as biomarkers, there are still some obstacles to be overcome, for instance, the reproducible recovery from milk samples and reliable measurement methods for the isolated miRNAs.
Several extraction methods have been used in recent studies describing milk miRNAs (Izumi et al., 2013; Sun et al., 2013). Current protocols for isolation of milk miRNAs can be divided into two categories that are mainly distinguished by whether there is an exosome enrichment step. However, the reported exosome isolation methods, for instance, ultracentrifugation, sucrose-density gradient ultracentrifugation, filtration and size-exclusion chromatography, need complicated equipment and are difficult to execute. Moreover, it has been shown that after ultracentrifugation, more than 20% of miRNAs still exist in the supernatant (Izumi et al., 2013; Sun et al., 2013). Therefore, isolating miRNAs from liquid milk whey may be applicable and reliable. In the studies describing RNA extraction from cell-free fluids, such as milk whey, two alternative RNA extraction methods have already been performed, i.e. the phenol-based technique (Chen et al., 2010) and column-based approach (Trizol followed by cleanup using the modified RNeasy, miRNeasy or mirVana kit) (Izumi et al., 2012). Despite some discrepancies between these two isolation methods in other body fluid samples (McAlexander et al., 2013; Moret et al., 2013; Duy et al., 2015), none of these studies compared and evaluated the quantity and quality of the isolated miRNAs from milk using these two methods.
In this study, two representative methods for the isolation of cell-free fluid miRNA, namely, the phenol-based technique (Trizol LS®, the TP method) and combined phenol and column-based approach (Trizol LS® followed by cleanup using miRNeasy kit, the TM method), were compared to obtain good-quality and high-yield miRNAs from milk whey. Based on these comparisons, the commercial protocols were further modified to improve the yield and quality of milk whey miRNAs.
2 Materials and methods
2.1 Milk sample collection and preparation
Fresh milk samples were collected from Hangzhou Hangjiang Dairy Farm, Zhejiang Province, China. The modern automated milking facilities and good quality of milk products from this farm meet the national milk quality standards of China. The nutrition of the herds in this experiment was based on a total mixed ration that was prepared at a central regional feeding center. The experimental procedures were approved by the Animal Care Committee of Zhejiang University (Hangzhou, China) and were in accordance with the guidelines of the university?s guidelines for animal research.
Raw milk samples were taken under sterile conditions using Waikato Milking System meters (Waikato Milking Systems NZ Ltd., Hamilton, New Zealand) while milking at noon. Milk was collected from 24 multiparous cows at approximately 150 d in milk with diethylpyrocarbonate (DEPC)-treated tubes, stored at −20 °C immediately after collection, transferred to the laboratory within 2 h, and stored at −80 °C until further use. Milk pre-preparation procedures were performed according to Izumi et al. (2012)’s protocol with some modifications. Briefly, samples were centrifuged twice (1200×g, 4 °C, 10 min), syringed into the middle defatted layer under RNase-free conditions, and dispensed into 1 ml aliquots per tube. Thereafter, the defatted milk fraction was centrifuged again (21500×g, 4 °C, 1 h), and the clear supernatant was collected and filtered through 0.22-μm syringe filters. Additionally, clear liquid supernatants (250 μl) were collected for RNA isolation.
2.2 Milk whey homogenization and RNA spike-in standard
A total of 250 μl of milk whey was homogenized with 750 μl of Trizol LS® Reagent (Qiagen, Valencia, CA, USA), mixed thoroughly by vortexing, and then incubated for 5 min at room temperature to denature the milk sample. Next, 2 μl of 20 fmol cel-miR-39 (Caenorhabditis elegans miScript miRNA Mimic; Qiagen, Valencia, CA, USA) was added into 1 ml homogenate.
2.3 Total RNA extraction
The homogenate was mixed with 200 μl of chloroform, vortexed thoroughly, incubated for 3 min at 25 °C, and centrifuged (12000×g) for 15 min at 4 °C. The aqueous phase was subjected to either the phenol-based technique (according to the Trizol LS® protocol) or column-based approach (according to the miRNeasy serum/plasma handbook) to extract total RNA.
2.3.1 Phenol-based technique
According to the manufacturers’ protocols for Trizol LS®, 0.5 μl of RNase-free glycogen (20 mg/ml; Aidlab, Beijing, China) was added to the aqueous phase before being mixed with 500 μl of isopropanol and incubated at −20 °C for 2 h. The RNA pellet was collected at the bottom of the tube after being centrifuged at 12000×g for 30 min at 4 °C and then was washed with 1 ml of 80% ethanol and re-enriched at the bottom of the tube using the same centrifugation procedure. The air-dried RNA pellet from 250 μl of milk whey was finally re-suspended in 20 μl of RNasefree water and stored at −80 °C until further use.
2.3.2 Column-based approach
Based on the manufacturers’ protocols in the miRNeasy serum/plasma handbook, the aqueous phase was added to 1.5 volumes of 100% ethanol, and the mixture was mixed thoroughly by pipetting. The supernatants were then transferred to the RNeasy Mini spin column by centrifugation. Thereafter, the column was washed with 700 μl of RWT, 500 μl of RPE, and 500 μl of 80% ethanol, and then dried by centrifugation at 12000×g for 5 min. Finally, the RNA on the membrane was eluted with 22 μl of RNase-free water and stored at −80 °C until further use.
2.4 Total RNA yield and quality
The RNA concentration and purity were measured using a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA). The quality of the RNA was further evaluated on a Bioanalyzer 2100 instrument (Agilent, Santa Clara, CA, USA) using an RNA 6000 Pico Kit (Agilent, Santa Clara, CA, USA).
2.5 Quantification of miRNA by real-time PCR
A facile mean for quantifying miRNA expression by real-time polymerase chain reaction (PCR) was used in the experiment (Shi and Chiang, 2005). The milk whey miRNA (5 μl) was polyadenylated with adenosine 5'-triphosphate (ATP) by poly(A) polymerase (PAP) at 37 °C for 1 h in a 10-μl reaction mixture following the manufacturers’ directions (Poly A Tailing Kit; Ambion Inc., Woodward St., Austin, TX, USA). The polyadenylated RNA was then directly reverse-transcribed using a PrimeScript RT Reagent Kit (TaKaRa, Dalian, China) in a 20-μl reaction mixture (10 μl of polyadenylated RNA reaction mix, 4 μl of 5× PrimeScript buffer, 5 μl of 10 μmol/L poly T adaptor, and 1 μl of PrimeScript RT Enzyme Mix I). The complementary DNA (cDNA) was diluted 25 times for real-time PCR analysis. The forward primers used for specific miRNAs were designed with Primer 5 software (Premier Biosoft, USA) and are listed in Table 1. The SYBR green quantitative PCR was carried out using a 7500c real-time PCR detection system (ABI, USA) with SYBR Premix EX Taq (TaKaRa, Dalian, China) using the following three-step reaction: one cycle of 95 °C for 30 s for initial denaturation, followed by 40 cycles of denaturation at 95 °C for 5 s, annealing at 55 °C for 30 s and extension at 60 °C for 30 s. After PCR amplification, melting curve analysis was performed to verify the specificity of the PCR products. All of the real-time PCR products were verified by agarose gel electrophoresis. The real-time PCR efficiency was calculated as the formula: efficiency=10(-1/slope)-1.
2.6 Statistical analysis
Data are expressed as mean±standard deviation (SD) for the indicated number of independently performed experiments. Statistical comparison of the data was performed by Student’s t-test. P-values less than 0.05 were considered statistically significant or relevant. All of the statistical tests were carried out using SPSS 17.0 (SPSS Inc., Chicago, IL, USA).
3 Results
3.1 Total RNA yield and quality were comparable by the two isolation methods
As shown in Table 2, the RNA concentrations from the two methods were comparable over a wide range. The average quantity of the recovered total RNA from 1 ml of milk whey was (1536±963.4) and (1525±990.8) ng by the TP and TM methods, respectively. However, low ratios of absorbance at 260 nm/280 nm (A260/A280) and absorbance at 260 nm/230 nm (A260/A230) were also found in all of the RNA samples isolated from milk whey.
As shown in Fig. 1, bovine milk whey contained a substantial amount of small RNA (<300 nt). Only a small amount of ribosomal RNA (18S and 28S rRNA) was found, with no significant differences between the two methods.
Photo and image for RNA isolated from milk whey using the phenol-based Trizol LS® technique (TP method) and column-based approach with Trizol LS® followed by cleanup using the miRNeasy kit (TM method)
(a) A representative photo taken after the step of 21500××g centrifugation showing the fraction of milk used for miRNAisolation. (b) Representative electropherogram and (c) fluorescence intensity distribution images from twelve repeat extractions of the RNA isolated by the two methods
3.2 cel-miR-39 is a proper spike-in control for comparing the efficiency of miRNA recovery
The real-time PCR results using the cel-miR-39 forward primer showed a good amplification efficiency (105.7%) and a high correlation coefficient value (R2=0.9967) (Fig. 2). When this forward primer was used to determine the amount of spike-in cel-miR-39, analysis of RNA (isolated from three non-spike-in milk whey mix samples) spiked with a 2-μl serial dilution of cel-miR-39 resulted in a linear curve with a low correlation coefficient value (R2=0.82) and amplification efficiency (16.5%), suggesting that cel-miR-39 cannot be used to perform the absolute quantification for recovery efficiency. Importantly, the cycle threshold value and amount of cel-miR-39 showed significant relevance (P<0.05), indicating that the CT value can be used to compare the miRNA recovery efficiency. In addition, 20 fmol of the spike-in cel-miR-39, equivalent to the CT value at 17.2±0.42, was used in subsequent experiments.
Determination of the synthetic miRNA spike-in control
(a) PCR efficiency (E) and correlation coefficient (R2) values were determined using the gradient dilution PCR product of cel-miR-39. The average CT values from quadruplicate wells were used to reproduce the linear equation. (b) The average CT value of different concentrations of spike-in cel-miR-39 in the background of milk whey RNA without spike-in miRNAs. Different concentrations of spike-in cel-miR-39 (2 μl) and milk whey RNA (3 μl) without spike-in miRNAs were reverse-transcribed and used in real-time PCR, as described in Section 2.5. The average CT values from quadruplicate wells were used to produce the linear equation
3.3 TM method shows better miRNA recovery efficiency
The CT value of cel-miR-39 (spike-in control) in the TP group (20.8±0.11) was significantly higher than that in the TM group (17.7±0.09) (Fig. 3a) and close to the value generated from 20 fmol of cel-miR-39 directly added to the milk whey RNA sample (17.2±0.42). The same difference (P<0.001) could be found in the CT values of bta-miR-141, bta-miR-148a, bta-miR-200c, and bta-miR-375 using these two methods, although specific enrichments of these miRNAs in milk whey were different (Fig. 3b). Additionally, the CT values of endogenous miRNAs were normalized to validate that these two methods have no bias concerning the miRNA recovery (Fig. 3c). Overall, the TM method was further evaluated in the following experiments for its good performance in milk whey miRNA recovery.
Better miRNA recovery efficiency in TM method
The mean CT values and SD of cel-miR-39 (a), bta-miR-141, bta-miR-148a, bta-miR-200c, and bta-miR-375 (b) from milk whey RNA isolated by the two methods (TM, Trizol LS® followed by cleanup using the miRNeasy; TP, Trizol LS® followed by phenol precipitate; n=12). (c) The \(\mathop 2\nolimits^{\mathop { - \Delta \Delta C}\nolimits_{\rm{T}} }\) method was used to normalize the data in (b). cel-miR-39 was used as a house-keeping reference and TP as a control group. The results are shown as the mean±SD (n=4). The Student’s t-test was performed to evaluate the statistical significance between the two methods. ns (not significant), P>0.05; * P<0.05, ** P<0.01, *** P<0.001, compared with the TM group
3.4 TM method shows good repeatability
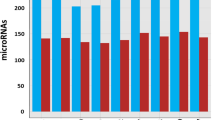
The CT value of cel-miR-39 fluctuated by approximately 17.7 in the 40 samples (Fig. 4a), close to the CT value of the spike-in control cel-miR-39 (17.2±0.42). The overall SD from 40 samples was only 0.26, indicating that the spike-in cel-miR-39 was hardly affected by sample variations. More importantly, the CT values of bta-miR-148a and bta-miR-375 in two milk samples from the same cow showed no differences between each replication (P>0.05; Figs. 4b and 4c).
Good consistency in the recovery of spike-in control miRNA and good repeatability in endogenous miRNAs in TM method
(a) The CT values of cel-miR-39 from 40 milk samples. Each spot shows the mean value from three real-time PCR reactions. (b, c) The CT values of bta-miR-148a (b) and bta-miR-375 (c) from 20 paired samples. Each pair represented replicate milk samples from the same dairy cow. Each spot represents paired CT values of bta-miR-148a (b) or bta-miR-375 (c) from two independent milk whey miRNA isolation procedures from the same milk sample. R1, replication 1; R2, replication 2
4 Discussion
Milk is an important biological fluid that can be easily obtained and used for biomarker identification studies (Saleh et al., 1998; Warensjö et al., 2010; Klein et al., 2012). Milk is also an abundant source of dietary miRNAs (Baier et al., 2014). Most of the previous studies have mainly focused on methods for miRNA isolation directly from body fluids, such as serum (Li and Kowdley, 2012), plasma (Moret et al., 2013), urinary sediment (Wang and Szeto, 2013), and cerebrospinal fluid (McAlexander et al., 2013). However, few attempts have been made to identify suitable methods for extracting miRNA from milk. In this study, it was demonstrated that a combined phenol and column-based approach (Trizol LS® followed by cleanup using the miRNeasy kit) is suitable for miRNA isolation from small-volume milk whey samples, and that exogenously added cel-miR-39 (20 fmol) can be a proper spike-in control to quantify miRNA recovery.
RNA has been found to exist in cow milk whey, but its concentration varies considerably (Weber et al., 2010; Izumi et al., 2012). The individual variations of RNA concentrations in milk whey can be attributed to the different milk compositions among individual cows. Low values of the A260/A280 and A260/A230 ratios found in the present study agreed with the work by others performing miRNA extraction from body fluids (Moret et al., 2013). This finding could also explain the low PCR efficiency of miRNA. Currently, NanoDrop and the Bioanalyzer are widely used to analyze the quality of RNA samples (Fleige and Pfaffl, 2006; Zhang et al., 2012). Among these, the Bioanalyzer 2100 instrument with an RNA 6000 Pico Kit (based on electropherograms) can also be used to analyze the distribution of RNA size in each RNA sample. No differences were found between the aforementioned two methods, contrasting with the results of the other studies using real spike-in miRNA to evaluate the recovery efficiency. The cause may be the instrument limitation of the RNA 6000 Pico Kit, but this limitation can be remedied by a small RNA kit (Agilent) that can distinguish RNA smaller than 25 nt. Combined with the results of the real recovery efficiency of the two methods, it is suggested that NanoDrop and Bioanalyzer 2100 (RNA 6000 Pico Kit) analysis, two commonly used methods to evaluate RNA quality, may not be suitable for evaluation of milk whey RNA quality. To analyze the miRNA recovery of these two methods, a synthetic miRNA spike-in control was used to monitor the isolation procedures of endogenous miRNAs (Kroh et al., 2010). However, this method could only be used to compare relatively the recovery efficiency because the standard curves could not be established, possibly because of the specific residues in milk whey extraction inhibiting enzymatic reactions such as reverse transcription or real-time PCR.
Both the TP and TM approaches have been used in isolation of body fluid RNA (Moret et al., 2013). The TM method showed better recovery efficiency, with lower CT values of specific endogenous miRNAs and spike-in miRNAs than the TP method, in line with the result in human plasma samples (Moret et al., 2013). It is reported that this situation may be due to the low efficiency of phenol in precipitating small RNAs because low doses of an RNA carrier before starting the extraction process lead to the improvement of the miRNA purification (Kroh et al., 2010; Moret et al., 2013). However, it seems that addition of a carrier molecule is beneficial for some but not all isolation methods (Guo and Lu, 2010; McAlexander et al., 2013), and different carriers result in a significant bias over miRNA profiles (Bartel, 2004). Additionally, it can be used to explain the failure of our modified phenol-based technique, including the glycogen supplement procedure. However, both methods show no bias in the types of recovered miRNAs, suggesting that the combined phenol and column-based approach is superior to the modified phenol-based technique in recovery of overall miRNAs. Using a larger set of milk samples, the combined phenol and column-based approach also showed excellent repeatability regarding the recovery of these endogenous miRNAs. Moreover, the recovery of spike-in miRNA was consistent among 40 different samples, strongly supporting the modified method for miRNA isolation from cow milk whey.
In conclusion, the combined phenol and column-based approach is superior to the modified phenol-based technique in recovering milk whey miRNAs. NanoDrop and Bioanalyzer 2100 (RNA 6000 Pico Kit) analysis may not reflect the real miRNA recovery efficiency. Exogenously added cel-miR-39 can be used as a proper spike-in control for isolation of miRNA from milk whey. Taken together, these results show that the use of the combined phenol and column-based approach is recommended to isolate miRNAs from milk whey.
Compliance with ethics guidelines
Xiao-lu JIN, Zi-hai WEI, Lan LIU, Hong-yun LIU, and Jian-xin LIU declare that they have no conflict of interest.
This article does not contain any studies with human or animal subjects performed by any of the authors.
References
Baier, S.R., Nguyen, C., Xie, F., et al., 2014. MicroRNAs areabsorbed in biologically meaningful amounts from nutritionally relevant doses of cow milk and affect gene expressionin peripheral blood mononuclear cells, HEK-293kidney cell cultures, and mouse livers. J. Nutr., 144(10):1495–1500. [doi:10.3945/jn.114.196436]
Bartel, D.P., 2004. Micro RNAs: genomics, biogenesis, mechanism,and function. Cell, 116(2):281–297. [doi:10.1016/S0092-8674(04)00045-5]
Chen, X., Gao, C., Li, H., et al., 2010. Identification andcharacterization of microRNAs in raw milk during different periods of lactation, commercial fluid, and powderedmilk products. Cell Res., 20(10):1128–1137.[doi:10.1038/cr.2010.80]
Duy, J., Koehler, J.W., Honko, A.N., et al., 2015. Optimized microRNA purification from TRIzol-treated plasma. BMC Genomics, 16(1):95. [doi:10.1186/s12864-015-1299-5]
Farazi, T.A., Horlings, H.M., Jelle, J., et al., 2011. MicroRNAsequence and expression analysis in breast tumors bydeep sequencing. Cancer Res., 71(13):4443–4453. [doi:10.1158/0008-5472.CAN-11-0608]
Fernández-Hernando, C., Suá rez, Y., Rayner, K.J., et al., 2011.MicroRNAs in lipid metabolism. Curr. Opin. Lipidol.,22(2):86. [doi:10.1097/MOL.0b013e3283428d9d]
Fleige, S., Pfaffl, M.W., 2006. RNA integrity and the effect onthe real-time qRT-PCR performance. Mol. Aspects Med.,27(2-3):126–139. [doi:10.1016/j.mam.2005.12.003]
Guo, L., Lu, Z., 2010. The fate of miRNA* strand throughevolutionary analysis: implication for degradation asmerely carrier strand or potential regulatory molecule? PLoS ONE, 5(6):e11387. [doi:10.1371/journal.pone.0011387]
Hata, T., Murakami, K., Nakatani, H., et al., 2010. Isolation ofbovine milk-derived microvesicles carrying mRNAs andmicroRNAs. Biochem. Biophys. Res. Commun., 396(2):528–533. [doi:10.1016/j.bbrc.2010.04.135]
Izumi, H., Kosaka, N., Shimizu, T., et al., 2012. Bovine milkcontains microRNA and messenger RNA that are stableunder degradative conditions. J. Dairy Sci., 95(9):4831–4841. [doi:10.3168/jds.2012-5489]
Izumi, H., Kosaka, N., Shimizu, T., et al., 2013. Puri fication ofRNA from milk whey. Methods Mol. Biol., 1024:191-201.[doi:10.1007/978-1-62703-453-1_15]
Izumi, H., Kosaka, N., Shimizu, T., et al., 2014. Time dependent expression profiles of microRNAs and mRNAsin rat milk whey. PLoS ONE, 9(2):e88843. [doi:10.1371/journal.pone.0088843]
Klein, M.S., Buttchereit, N., Miemczyk, S.P., et al., 2012.NMR metabolomic analysis of dairy cows reveals milkglycerophosphoch oline to phosphocholine ratio as prognosticbiomarker for risk of ketosis. J. Proteome Res.,11(2):1373–1381. [doi:10.1021/pr201017n]
Kosaka, N., Izumi, H., Sekine, K., et al., 2010. microRNA as anew immune-regulatory agent in breast milk. Silence,1(1):7. [doi:10.1186/1758-907X-1-7]
Kroh, E.M., Parkin, R.K., Mitchell, P.S., et al., 2010. Analysisof circulating microRNA biomarkers in plasma and serumusing quantitative reverse transcription-PCR (qRT-PCR). Methods, 50(4):298–301. [doi:10.1016/j.ymeth.2010.01.032]
Li, Y., Kowdley, K.V., 2012. Method for microRNA isolationfrom clinical serum samples. Anal. Biochem., 431(1):69–75. [doi:10.1016/j.ab.2012.09.007]
McAlexander, M.A., Phillips, M.J., Witwer, K.W., 2013.Comparison of methods for miRNA extraction fromplasma and quantitative recovery of RNA from cerebrospinal fluid. Front. Genet., 4:83[doi:10.3389/fgene.2013.00083]
Melnik, B.C., John, S.M., Schmitz, G., 2013. Milk is not justfood but most likely a genetic transfection system activatingmTORC1 signaling for postnatal growth. Nutr. J.,12(1):103. [doi:10.1186/1475-2891-12-103]
Moret, I., Sánchez-Izquierdo, D., Iborra, M., et al., 2013.Assessing an improved protocol for plasma microRNAextraction. PLoS ONE, 8(12):e82753. [doi:10.1371/journal.pone.0082753]
Munch, E.M., Harris, R.A., Mohammad, M., et al., 2013.Transcriptome profiling of microRNA by Next-Gen deepsequencing reveals known and novel miRNA species inthe lipid fraction of human breast milk. PLoS ONE,8(2):e50564. [doi:10.1371/journal.pone.0050564]
Saleh, M., Afify, A.M., Kamel, A., 1998. Mother’s milk protein profile, a possible biomarker for human exposure topersistent insecticides. J. Environ. Sci. Health B, 33(6):645–655. [doi:10.1080/03601239809373169]
Shi, R., Chiang, V.L., 2005. Facile means for quantifyingmicroRNA expression by real-time PCR. Biotechniques,39(4):519–525. [doi:10.2144/000112010]
Sun, Q., Chen, X., Yu, J., et al., 2013. Immune modulatoryfunction of abundant immune-related microRNAs in microvesiclesfrom bovine colostrum. Protein Cell, 4(3):197–210. [doi:10.1007/s13238-013-2119-9]
Wang, G., Szeto, C.C., 2013. Methods of microRNA quantificationin urinary sediment. Methods Mol. Biol., 1024:211–220. [doi:10.1007/978-1-62703-453-1_17]
Warensjö, E., Jansson, J.H., Cederholm, T., et al., 2010.Biomarkers of milk fat and the risk of myocardial infarctionin men and women: a prospective, matchedcase-control study. Am. J. Clin. Nutr., 92(1):194–202.[doi:10.3945/ajcn.2009.29054]
Weber, J.A., Baxter, D.H., Zhang, S., et al., 2010. ThemicroRNA spectrum in 12 body fluids. Clin. Chem.,56(11):1733–1741. [doi:10.1373/clinchem.2010.147405]
Zhang, L., Hou, D., Chen, X., et al., 2011. Exogenous plantMIR168a specifically targets mammalian LDLRAP1:evidence of cross-kingdom regulation by microRNA. Cell Res., 22(1):107–126. [doi:10.1038/cr.2011.158]
Zhang, W.Y., Liu, W., Zhou, Y.M., et al., 2012. AlteredmicroRNA expression profile with miR-27b downregulationcorrelated with disease activity of oral lichenplanus. Oral Dis., 18(3):265–270. [doi:10.1111/j.1601-0825.2011.01869.x]
Zhou, Q., Li, M.Z., Wang, X.Y., et al., 2012. Immune-relatedmicroRNAs are abundant in breast milk exosomes. Int. J.Biol. Sci., 8(1):118–123. [doi:10.7150/ijbs.8.118]
Acknowledgements
The authors sincerely acknowledge Bing WANG, Hui-zeng SUN, and Bin YANG from the Institute of Dairy Science, College of Animal Sciences, Zhejiang University, Hangzhou, China and all of the staff from Hangzhou Hangjiang Dairy Farm, Zhejiang Province, China, for their assistance during milk sample collections.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Project supported by the Zhejiang Provincial Key Science and Technology Innovation Team (No. 2011R50025) and the National Natural Science Foundation of China (No. 31328022)
ORCID: Hong-yun LIU, http://orcid.org/0000-0001-8806-6204
Rights and permissions
About this article
Cite this article
Jin, Xl., Wei, Zh., Liu, L. et al. Comparative studies of two methods for miRNA isolation from milk whey. J. Zhejiang Univ. Sci. B 16, 533–540 (2015). https://doi.org/10.1631/jzus.B1400355
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1631/jzus.B1400355