Abstract
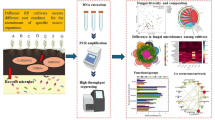
Activity and functional diversity of rhizosphere bacterial communities and fungal composition were studied in order to assess the effects of different genotypes (N8035, N224 and N8637) of Arabidopsis thaliana on these communities growing in different soils. Genotype effect and soil effect were studied independently. Also, the interactions between both factors (genotypes and soils) were considered. The activity was determined by thymidine and leucine incorporation analysis, and Biolog ECO plates were used to study bacterial functional diversity. Additionally, fungi groups (genera and/or species) were studied in the different rhizospheres. Statistical differences on thymidine incorporation between plant genotypes were only found in two of the soils. In addition, functional diversity (measured by Shanonn-Weaver index), showed statistical differences only in soil 1 for line N8035 (line B) vs. the other lines. Redundancy analysis (RDA) performed with Biolog data indicated and important effect of soil type, but also an effect of genotype since line N8035 (line B) was separated from the other lines within each soil in the RDA ordination, in spite of genotypic differences between them were minimum. Furthermore, carboxylic acids and amino acids were found to be the Biolog plate substrates with more influence in samples ordination in the Redundancy Analysis (RDA). However, fungi seem to be less labile to plant selection than bacteria probably due to a lower turn-over time of fungi than bacteria coupled with the short phenology of Arabidopsis. In this paper, plant-soil-microorganism relationships in the rhizosphere were studied, and the complex interactions between them were highlighted. More studies are necessary to go deep in these interactions and to be able to asses the impact of genetically modified plants.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- AWCD:
-
Average Well Colour Development
- CLPP:
-
Community Level Physiological Profiles
- DCA:
-
Detrended Correspondence Analysis
- GMP:
-
Genetically Modified Plants
- RDA:
-
Redundancy Analysis
References
Alisi, C., Lasinio, G.F., Dalmastri, C., Sprocati, A., Tabacchioni, S., Bevivino, A. and Chiarini, L. 2005. Metabolic profiling of Burk-holderia cenocepacia, Burkholderia ambifaria and Burk-holderia pyrrocinia isolates from maize rhizosphere. Microbial Ecol. 50: 385–395.
Arondel, V., Lemieux, B., Hwang, I., Gibson, S., Goodman, H.M. and Somerville, C.R. 1992. Map-based cloning of a gene controlling omega-3-fatty-acid desaturation in Arabidopsis. Science 258: 1353–1355.
Bååth, E. 1992. Thymidine incorporation into macro-molecules of bacteria extracted from soil by homogenisation-centrifugation. Soil Biol. Biochem. 24: 1157–1165.
Bååth, E. 1996. Thymidine incorporation of soil bacteria sequentially extracted from soil using repeated homogenization-centrifugation. Microbial Ecol. 31: 153–166.
Bååth, E., Pettersson, M. and Söderberg, K.H. 2001. Adaptation of a rapid and economical microcentrifugation method to measure thymidine and leucine incorporation of soil bacteria. Soil Biol. Biochem. 33: 1571–1574.
Baudoin, E., Benizri, E. and Guckert, A. 2001. Metabolic fingerprint of microbial communities from distinct rhizosphere compartments. Eur. J. Soil Biol. 37: 85–93.
Bolton, H.J., Fredrickson, J.K. and Elliott, L.F. 1992. Microbial ecology of the rhizosphere. In: Metting, F.B.J. (Ed.) Soil Microbial Ecology, Marcel Dekker, New York, pp 27–63.
Bowen, G.D. and Rovira , A.D., 1991. The rhizosphere, the hidden half of the hidden half. In: Waisel, Y., Eshel, A. and Kafkafi, U. (eds.), Plant Roots-The Hidden Half, Marcel-Dekker, New York, pp 641–649.
Bowen, G.D. and Rovira, A.D. 1999. The rhizosphere and its management to improve plant growth. Adv. Agron. 66: 1–102.
Bridge, P. and Spooner, B. 2001. Soil fungi: diversity and detection. Plant Soil 232: 147–154.
Brusetti, L., Francia, P., Bertolini, C., Pagliuca, A., Borin, C. Sorlini, C., Abruzzese, A., Sacchi, G., Viti, C., Giovannetti, E. Giuntini, E., Bazzicalupo, M. and Daffonchio, D. 2004. Bacterial communities associated with the rhizosphere oftransgenic Bt 176 maize (Zea mays) and its non-transgenic counterpart. Plant Soil 266: 11–21.
Castaldini, M., Turrini, A., Sbrana, C., Benedetti, A., Marchionni, M., Mocali, S., Fabiani, A., Landi, S., Santomassimo, F., Pietrangeli, B., Nuti, M.P., Miclaus, N. and Giovannetti, M. 2005. Impact of Bt corn on rhizospheric and soil eubacterial communities and on beneficial mycorrhizal symbiosis in experimental microcosms. Appl. Environ. Microbiol. 71: 6719–6729.
Cook, R.J. 1999. Science-based risk assessment for the approval and use of plants in agricultural and other environments. In: Persley, G.J. and Lantin, M.M. (eds) Agricultural Biotechnology and the Poor. Consultative Group on International Agricultural Research, Washington, DC. pp 123–130.
Crecchio, C., Gelsomino, A., Ambrosoli, R., Minati, J.L. and Ruggiero, P. 2004. Functional and molecular responses of soil microbial communities under differing soil management practices. Soil Biol. Biochem., 36: 1873–1883.
Davet, P. and Rouxel, F. 1997. Detection and Isolation of Soil Fungi. Science Publishers Inc, Enfield, USA.
de Boer, W., Folman, L.B., Summerbell, R.C. and Boddy, L. 2005. Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol. Rev. 29: 795–811.
Di Giovanni, G.D., Watrud, L.S., Seidler, R.J. and Widmer, F. 1999. Comparison of parental and transgenic alfalfa rhizosphere bacterial communities using biolog GN metabolic fingerprinting and enterobacterial repetitive intergenic consensus sequence-PCR (ERIC-PCR). Microbial Ecol. 37: 129–139.
Dunfield, K.E. and Germida, J.J. 2001. Diversity of bacterial communities in the rhizosphere and root interior of field-grown genetically modified Brassica napus. FEMS Microbiol. Ecol. 82: 1–9.
Eiland, F. 1983. A simple method for quantitative determination of ATP in soil. Soil Biol. Biochem. 15: 665–670.
Garbeva, P., van Elsas, J.D. and van Veen, J.A. 2008. Rhizosphere microbial community and its response to plant species and soil history. Plant Soil 302: 19–32.
Garland, J.L. 1997. Analysis and interpretation of community-level physiological profiles in microbial ecology. FEMS Microbiol. Ecol. 24:289–300.
Garland, J.L. and Mills, A.L. 1991. Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Appl. Environ. Microbiol. 57: 2351–2359.
Girvan, M.S., Bullimore, J., Pretty, J.N., Osborn, A.M. and Ball, A.S. 2003. Soil type is the primary determinant of the composition of the total and active bacterial communities in arable soils. Appl. Environ. Microbiol. 69 (3): 1800–1809.
Gomes, N.C.M., Fagbola, O., Costa, R.,. Rumjanek, N.G., Buchner, A., Mendona-Hagler, L. and Smalla, K. 2003. Dynamics of fungal communities in bulk and maize rhizosphere soil in the tropics. Appl. Environ Microbiol. 69: 3758–3766.
Grayston, S.J. and Campbell, C.D. 1996. Functional biodiversity of microbial communities in the rhizosphere of hybrid larch (Larix eurolepis) and Sitka spruce (Picea sitchensis). Tree Physiol. 16: 11–12.
Grayston, S.J., Griffith, G.S., Mawdsley, J.L., Campbell, C.D. and Bardgett, R.D. 2001. Accounting for variability in soil microbial communities of temperate upland grassland ecosystems. Soil Biol. Biochem. 33: 533–551.
Grayston, S.J., Vaughan, D. and Jones, D. 1996. Rhizosphere carbon flow in trees, in comparison with annual plants: the importance of root exudation and its impact on microbial activity and nutrient availability. Appl. Soil Ecol. 5: 29–56.
Grayston, S.J., Wang, S., Campbell, C.D. and Edwards, A.C. 1998. Selective influence of plant species on microbial diversity in the rhizosphere. Soil Biol. Biochem. 30: 369–378.
Griffiths, B.S., Ritz, K., Ebblewhite, N. and Dobson, G. 1999. Soil microbial community structure: effects of substrate loading rates. Soil Biol. Biochem. 31: 145–153.
Hertenberger, G., Zampach, P. and Bachmann, G. 2002. Plant species affect the concentration of free sugars and free amino acids in different types of soil. J. Plant Nutr. Soil Sci. 165:557–565.
Heuer, H. and Smalla, K. 1997. Evaluation of community-level catabolic profiling using BIOLOG GN microplates to study microbial community changes in potato phyllosphere. J. Microbiol. Methods 32: 49–61.
Heuer, H. and Smalla, K. 1999. Bacterial phyllosphere communities of Solanum tuberosum L. and T4-Lysozyme-producing transgenic variants. FEMS Microbiol. Ecol. 28: 357–371.
Information Systems for Biotechnology. 2006. Status of all field test permits 1987-present. Virginia Tech., Blacksburg, VA. https://doi.org/www.isb.vt.edu/cfdocs/ISBtables.cfm.
Huang, Z., Xu, Z. and Chen, C. 2008. Effect of mulching on labile soil organic matter pools, microbial community functional diversity and nitrogen transformations in two hardwood plantations of subtropical Australia. Appl. Soil Ecol. 40: 229–239.
James, C., 2004. Preview: Global Review of Commercialized Biotech/GM Crops: 2004. ISAAA Briefs No. 32. ISAAA, Ithaca, NY.
Jones, D.L. and Darrah, P.R. 1994. Amino-acid influx at the soil-root interface of Zea mays L. and its implications in the rhizosphere. Plant Soil 163: 1–12.
Jones, D.L, Hodge, A. and Kuzyakov, Y. 2004. Plant and mycorrhizal regulation of rhizodeposition. New Phytol. 163: 459–480.
Kemmitta, S.J., Lanyonb, C.V., Waiteb, I.S., Wenc, Q., Addiscotta, T.M., Birda, N.R.A., O’Donnellb, A.G. and Brookesa, P.C. 2007. Mineralization of native soil organic matter is not regulated by the size, activity or composition of the soil microbial biomass—a new perspective. Soil Biol. Biochem. 40: 61–73.
Liao, H., Wan, H.Y., Shaff, J., Wang, X.R., Yan, X.L. and Kochian, L.V. 2006. Phosphorus and aluminum interactions in soybean in relation to aluminum tolerance, exudation of specific organic acids from different regions of the intact root system. Plant Physiol. 141: 674–684.
Lippi, D., Paolis, M.R., Mattia, E.D., Pietrosanti, T. and Cacciari, I. 2004. Influence of substrate composition and flow rate on growth of Azospirillum brasilensis Cd in a co-culture with 3 sorghum rhizobacteria. Can. J. Microbiol. 50: 861–867.
Liu, B., Zeng, Q., Yan, F., Xu, H. and Xu, C. 2005. Effects of transgenic plants on soil microorganisms. Plant Soil 271: 1–13.
Lottmann, J. and Berg, G. 2001. Phenotypic and genotypic characterisation of antagonistic bacteria associated with roots of transgenic and non-transgenic potato plants. Microbiol. Res. 156: 75–82.
Lucas García, J.A., Barbas, C., Probanza, A., Barrientos, M.L. and Gutiérrez Mañero, F.J. 2001a. Low molecular weight organic acids (LOAs) and fatty acids in root exudates of two Lupinus cultivars at flowering and fruiting stages. Phytochem. Anal. 12: 305–311.
Lucas García, J.A., Probanza, A., Ramos, B. and Gutiérrez Mañero, F.J. 2001b. Genetic variability of rhizobacteria from wild populations of four Lupinus species based on PCR-RAPDs. J. Plant Nutr. Soil Sci. 164: 1–7.
Lynch, J.M. and Whipps, J.M. 1990. Substrate flow in the rhizosphere. Plant Soil 129: 1–10.
Marschner, H., 1996. Mineral Nutrition of Higher Plants, Second Edition. London: Academic.
McGregor, A.N. and Turner, M.A. 2000. Soil effects of transgenic agriculture: biological processes and ecological consequences. NZ Soil News 48(6): 166–169
Miethling, R., Ahrends, K. and Tebbe, C.C. 2003. Structural differences in the rhizosphere communities of legumes are not equally reflected in community-level physiological profiles. Soil Biol. Biochem. 35: 1405–1410.
Morse, C.C., Yevdokimov, I.V. and DeLuca, T.H. 2000. In situ extraction of rhizosphere organic compounds from contrasting plant communities. Comm. Soil Sci. Plant Anal. 31: 725–742.
National Research Council. 2000. Genetically modified pest-protected plants: science and regulation committee on genetically modified pest-protected plants. National Academy Press, Washington, DC.
Prosser, J.I. 2002. Molecular and functional diversity in soil microorganisms. Plant Soil 244: 9–17.
Rengel, Z. 2002. Genetic control of root exudation. Plant Soil 245: 59–70.
Rogers, B.F. and Tate, R.L. 2001. Temporal analysis of the soil microbial community along a toposequence in Pineland soils. Soil Biol. Biochem. 33, 1389–1401.
Ruiz Palomino, M., Lucas García, J.A., Ramos, B., Gutiérrez Mañero, F.J. and Probanza, A. 2005. Sucesional diversity changes in alder (Alnus glutinosa) culturable rhizobacterial communities throght a phenological cycle. Appl. Soil Ecol. 29: 215–224.
Saxena, D., Flores, S. and Stotzky, G. 2002. Vertical movement in soil of insecticidal Cry 1a protein from Bacillus thuringiensis. Soil Biol. Biochem. 34: 111–120.
Saxena, D. and Stotzky, G. 2000. Insecticidal toxin from Bacillus thuringiensis is released from roots of transgenic Bt corn in vitro and in situ. FEMS Microbiol. Ecol. 33: 35–39.
Saxena, D. and Stotzky, G. 2001. Bacillus thuringiensis (Bt) toxin released from root exudates and biomass of Bt corn has no apparent effect on earthworms, nematodes, protozoa, bacteria, and fungi in soil. Soil Biol. Biochem. 33: 1225–1230.
Schilling, G., Gransee, A. Deubel, A., Lezovic, G. and Ruppel, S. 1998. Phosphorus availability, root exudates, and microbial activity in the rhizosphere. Z. Pflanzenernähr Bodenk 161: 465–478.
Schutter, M. and Dick, R., 2001. Shifts in substrate utilization potential and structure of soil microbial communities in response to carbon substrates. Soil Biol. Biochem. 33: 1481–1491.
Sengeløv, G., Kristensen, K.J., Sørensen, A.H., Kroer, N. and Sørensen, S.J. 2001. Effect of genomic location on horizontal transfer of a recombinant gene cassette between Pseudomonas strains in the rhizosphere and spermosphere of barley seedlings. Cur. Microbiol. 42: 160–167.
Shen, R.F., Cai, H. and Gong, W.H. 2006. Transgenic Bt cotton has no apparent effect on enzymatic activities or functional diversity of microbial communities in rhizosphere soil. Plant Soil 285: 149–159.
Siciliano, S.D., Theoret, C.M., de Freitas, J.R., Hucl, P.J. and Germida, J.J. 1998. Differences in the microbial communities associated with the roots of different cultivars of canola and wheat. Can. J. Microbiol. 44: 844–851.
Smalla, K., Wachtendorf, U., Heuer, H., Liu, W.T. and Forney, L. 1998. Analysis of BIOLOG GN substrate utilization patterns by microbial communities. Appl. Environ. Microb. 64: 1220–1225.
Soderberg, K.H., Probanza, A., Jumpponen, A. and Bååth, E. 2004. The microbial community in the rhizosphere determined by community-level physiological profiles (CLPP) an direct soil and cfu-PLFA techniques. Appl. Soil Ecol. 25: 135–145.
Sokal, R.R. and Rohlf, F.J. 1980. Introducción a la bioestadística. Editorial Reverté S.A., Barcelona.
Swinnen, J., Van Veen, J.A. and Merckx, R. 1994. 14C pulse labelling of field grown spring wheat: an evaluation of its use in rhizosphere carbon budget estimation. Soil Biol. Biochem. 26: 161–170.
Taok, M., Cochet, N., Pauss, A. and Schoefs, O. 2007. Monitoring of microbial activity in soil using biological oxygen demand measurement and indirect impedancemetry. Eur. J. Soil Biol. 43 (5–6): 335–340.
ter Braak, C.J.F. and Šmilauer, P. 2002. CANOCO Reference Manual and CanoDraw for Windows User´s Guide. Software for Canonical Community Ordination (version 4.5). Biometris, Wageningen and Èeské Budìjovice,
Tesfaye, M., Denton, M.D., Samac, D.A. and Vance, C.P. 2005. Transgenic alfalfa secretes a fungal endochitinase protein to the rhizosphere. Plant Soil 269: 233–243.
Tesfaye, M., Dufault, N.S., Dornbusch, M.R., Allan, D.L., Vance, C.P. and Samac, D.A. 2003. Influence of enhanced malate dehydrogenase expression by alfalfa on diversity of rhizobacteria and soil nutriente availability. Soil Biol. Biochem. 35: 1103–1113.
Uhlirova, E. and Santruckova, H. 2003. Growth rate of bacteria is affected by soil texture and extraction procedure. Soil Biol. Biochem. 35: 217–224.
Van Veen, J.A., Liljeroth, E. and Lekkerkerk, L.J.A. 1991. Carbon fluxes in plant-soil systems at elevated atmospheric CO2 levels. Ecol.Appl. 1: 175–185.
Vance, E.D., Brookes, P.C. and Jenkinson, D.S. 1987. An extraction method for measuring soil microbial biomass C. Soil Biol. Biochem. 19: 703–707.
Watanabe, T. 1994. Pictorial Atlas of Soil and Seed Fungi. Lewis Publishers, Boca Raton, Florida, 411 pp.
Whipps, J.M. 1987. Effect of media on growth and interaction between a range of soil-borne glasshouse pathogens and antagonistic fungi. New Phytol. 107: 127–142.
Winding, A. and Hendriksen, N.B. 1997. Biolog substrate utilisation assay for metabolic fingerprints of soil bacteria: incubation effects. In: Insam, H. and Rangger, A. (eds.), Microbial Communities: Functional versus Structural Approaches, Springer, Heidelberg. pp. 195–205.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
García-Villaraco Velasco, A., Probanza, A., Gutierrez Mañero, F.J. et al. Functional diversity of rhizosphere microorganisms from different genotypes of Arabidopsis thaliana. COMMUNITY ECOLOGY 10, 111–119 (2009). https://doi.org/10.1556/ComEc.10.2009.1.13
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/ComEc.10.2009.1.13