Abstract
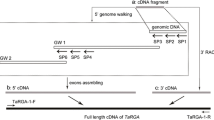
Powdery mildew (Blumeria graminis f. sp. tritici) is one of the most devastating wheat diseases. The wheat line N9134 contains PmAS846 that was transferred to N9134 from wild emmer wheat, and is still one of the most effective resistance genes in China. A full-length wheat RPM1 gene was obtained by rapid amplification of cDNA ends (RACE) based on the up-regulated probe sequence from differentially expressed transcripts during the N9134 and powdery mildew interaction. The gene was named TaRPM1, and the open reading frame (ORF) is 2721 nucleotides and encodes a polypeptide of 907 amino acids with a predicted isoelectric point of 4.86. Phylogenetic analysis revealed that TaRPM1 was highly homologous on both Aegilops tauschii and Triticum urartu at both the nucleotide and protein level. Using real-time quantitative PCR, the TaRPM1 gene expression level in wheat leaves was found to be sharply up-regulated, while the transcript level was lowly induced in the root and stem. Under the powdery mildew treatment, the transcription profile of TaRPM1 was very strongly expressed at 48 hour post inoculation (hpi), which increased again to 96 hpi and reaching a high level at 120 hpi. Based on sequence similarities and positions, we inferred that the TaRPM1 gene was on wheat chromosome 3D. These results suggested that TaRPM1 plays an important role in the mechanism of innate immunity to infection by the powdery mildew pathogen.
Similar content being viewed by others
References
Amelinetorregrosa, C., Wang, B.B., O’Bleness, M.S., Deshpande, S., Zhu, H., Roe, B., Young, N.D., Cannon, S.B. 2008. Identification and characterization of nucleotide-binding site-leucine-rich repeat genes in the model plant Medicago truncatula. Plant. Physiol. 146:5–21.
Appels, R., Eversole, K., Feuillet, C. 2018. Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:Eaar 7191.
Belkhadir, Y., Nimchuk, Z., Hubert, D.A., Mackey, D., Dangl, J.L. 2004. Arabidopsis RIN4 negatively regulates disease resistance mediated by RPS2 and RPM1 downstream or independent of the NDR1 signal modulator and is not required for the virulence functions of bacterial type III effectors AvrRpt2 or AvrRpm1. Plant.cell. 16:2822–2835.
Boyes, D.C., Nam, J., Dangl, J.L. 1998. The Arabidopsis thaliana RPM1 disease resistance gene product is a peripheral plasma membrane protein that is degraded coincident with the hypersensitive response. Proc. Natl. Acad. Sci. USA. 95:15849–15854.
Cao, A., Xing. L., Wang, X., Yang, X., Wang, W., Sun, Y., Qian, C., Ni, J., Chen, Y., Liu, D., Wang, X., Chen, P. 2011. Serine/threonine kinase gene Stpk-V, a key member of powdery mildew resistance gene Pm21, confers powdery mildew resistance in wheat. Proc. Natl. Acad. Sci. USA. 108:7727–7732.
Collins, N., Park, R.W., Ellis, J., Pryor, A.J. 2001. Resistance gene analogs in barley and their relationship to rustresistance genes. Genome 44:375–381.
Gong, C., Cao, S., Fan, R., Wei, B., Chen, G., Wang, X., Li, Y., Zhang, X. 2013. Identification and phylogenetic analysis of a CC-NBS-LRR encoding gene assigned on chromosome 7B of wheat. Int. J. Mol. Sci. 14:15330–15347.
Grant, M.R., Godiard, L., Straube, E., Ashfield, T., Lewald, J., Sattler, A., Innes, R.W., Dang, J.L. 1995. Structure of the Arabidopsis RPM1 gene enabling dual specificity disease resistance. Science 269:843–846.
Gu, L., Si, W., Zhao, L., Yang, S., Zhang, X. 2015. Dynamic evolution of NBS-LRR genes in bread wheat and its progenitors. Mol. Genet. Genomic. 290:727–738.
Hulbert, S.H., Webb, C.A., Smith, S.M., Sun, Q. 2001. Resistance gene complexes: evolution and utilization. Annu. Rev. Phytopathol. 39:285–312.
Ji, W.Q., Xue, X.Z., Wang, Q.Y., Wang, C.Y., Zhao, H.X., Zhang, H. 1999. Transfer of the powdery mildew resistant gene from T. dicoccoides into T. aestivum and its RAPD analysis. Acta Bot. Bor.Occid. Sin. 19:31–36.
Jiang, L.Y., Liu, J., Liu, X.Y., Wang, Z.Y. 2015. Cloning and functional verification of wheat resistant related gene TaRPM1 against powdery mildew. In: The 6th wheat genomics and molecular breeding conference, Yanglin, Shaanxi, China. pp. 86.
Li, F., Zhou, L., Xiao, F., Liao, K., Hu, J. 2011. Characterization of the psoRPM1 gene for resistance to root-knot nematodes in wild myrobalan plum (Prunus sogdiana). Afr. J. Biotechnol. 10:12859–12867.
Li, F., Zhu, X., Qiao, F., Chen, X.F., Li, H., Hu, J.F. 2013. psoRPM1 Gene from Prunus sogdiana Indicated Resistance to Root-knot Nematode in Tobacco. Acta. Horti. Sinica. 40:2497–2504.
McHale, L., Tan, X., Koehl, P., Michelmore, R.W. 2006. Plant NBS-LRR proteins: adaptable guards. Genome. Biol. 7:1–11.
Periyannan, S., Moore, J., Ayliffe, M., Bansal, U., Wang, X., Huang, L., Deal, K., Luo, M., Kong, X., Bariana, H., Mago, R., McIntosh, R., Dodds, P., Dvorak, J., Lagudah, E. 2013. The gene Sr33, an ortholog of barley Mla genes, encodes resistance to wheat stem rust race Ug99. Science 341:786–788.
Qiao, L.Y., Chang, J.Z., Guo, H.J., Gao, J.G., Zheng, J., Chang, Z.J. 2016. Genome-wide analysis of TaNBS resistance genes and development of chromosome 2AL-specific NBS-SSR markers in wheat. Acta Agron. Sinica. 42:795–802.
Saintenac, C., Zhang, W., Salcedo, A., Rouse, M.N., Trick, H.N., Akhunov, E., Dubcovsky. J. 2013. Identification of wheat gene Sr35 that confers resistance to Ug99 stem rust race group. Science 341:783– 786.
Srichumpa, P., Brunner, S., Keller, B., Yahiaoui, N. 2005. Allelic series of four powdery mildew resistance genes at the Pm3 locus in hexaploid bread wheat. Plant. Physiol. 139:885–895.
Swiderski, M.R., Birker, D., Jones, J.D.G. 2009. The TIR domain of TIR-NB-LRR resistance proteins is a signaling domain involved in cell death induction. Mol. Plant. Microbe. In. 22:157–165.
Thurau, T., Kifle, S., Jung, C., Cai, D. 2003. The promoter of the nematode resistance gene Hs1pro-1 activates a nematode-responsive and feeding site-specific gene expression in sugar beet (Beta vulgaris L.) and Arabidopsis thaliana. Plant. Mol. Biol. 52:643–660.
Tommasini, L., Yahiaoui, N., Srichumpa, P., Keller, B. 2006. Development of functional markers specific for seven Pm3 resistance alleles and their validation in the bread wheat gene pool. Theor. Appl. Genet. 114:165–175.
Wang, C.Y., Ji, W.Q., Zhang, G.S., Wang, Q.Y., Cai, D.M., Xue, X.Z. 2007. SSR markers and preliminary chromosomal location of a powdery mildew resistance gene in common wheat germplasm N9134. Acta Agron. Sin. 33:163–166.
Xue, F. 2012. Molecular mapping of the powdery mildew resistance gene and transcriptome analysis of the wheat-blumeria graminis f. sp. tritici interaction. PhD thesis. College of Agronomy, Northwest A&F University. Yangling, China.
Xue, F., Ji, W., Wang, C., Zhang, H., Yang, B. 2012. High-density mapping and marker development for the powdery mildew resistance gene PmAS846 derived from wild emmer wheat (Triticum turgidum var. dicoccoides). Theor. Appl. Genet. 124:1549–1560.
Yahiaoui, N., Srichumpa, P., Dudler, R., Keller, B. 2004. Genome analysis at different ploidy levels allows cloning of the powdery mildew resistance gene Pm3b from hexaploid wheat. Plant. J. 37:528–538.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Mohan
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Nie, Y.B., Ji, W.Q. Cloning and Characterization of Disease Resistance Protein RPM1 Genes against Powdery Mildew in Wheat Line N9134. CEREAL RESEARCH COMMUNICATIONS 47, 473–483 (2019). https://doi.org/10.1556/0806.47.2019.27
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/0806.47.2019.27