Abstract
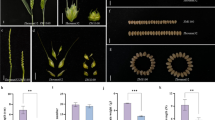
A wheat (Triticum aestivum L.) immature spike culture system was used to expeditiously generate mutations for use in wheat improvement programs. Wheat immature spikes in culture were treated with three concentrations of ethylmethane sulphonate (EMS) to generate a spike culture derived variant (SCDV) population. EMS in a concentration dependent manner affected seed development in wheat immature spike cultures. Based on the number of seeds produced, inclusion of EMS (25 mM) for three hours in immature spike culture medium generated variants in wheat cv. AC Nanda. The wheat AC Nanda SCDV population showed variation in several phenotypic characters. Flag leaf (length, angle and sheath length), length of first and second internode, spike length, number of spikes, number of seeds per spike and seed weight, showed variation below and above the non-treated controls. A molecular screening technique combining simple sequence repeat (SSR) oligonucleotide primers with high resolution melt (HRM) PCR with EvaGreen was used to identify the variants. Screening for starch branching enzyme IIb (SbeIIb) revealed 75 lines with point mutations. Combining SSR and SbeIIb, a total of 100 Kbp portion of wheat DNA was screened. The estimated mutation frequency in SbeIIb was one per 20.8 Kbp. The spike culture system utilizes very small amounts of EMS for a brief period, thus needs minimal handling of EMS and saves one generation of plant growth in a greenhouse. The morphological variants observed are similar to those reported for seed-derived variants using EMS.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Botticella, E., Sestili, F., Hernandez-Lopez, A., Phillips, A., Lafiandra, D. 2011. High resolution melting analysis for the detection of EMS induced mutations in wheat SbeIIa genes. BMC Plant Biol. 11:156.
Campbell, J., Zhang, H., Giroux, M.J., Feiz, L., Jin, Y., Wang, M., Chen, X., Huang, L. 2012. A mutagenesis-derived broad-spectrum disease resistance locus in wheat. Theor. Appl. Genet. 125:391–404.
Chen, L., Huang, L., Min, D., Phillips, A., Wang, S., Madgwick, P.J., Parry, M.A.J., Hu,Y.G. 2012. Development and characterization of a new TILLING population of common bread wheat (Triticum aestivum L.). Plos One 7:e41570.
Chibbar, R.N., Ganeshan, S., Baga, M. 2007. In planta novel starch synthesis. In: Ranalli, P. (ed.), Improvement of Crop Plants for Industrial Uses. Springer Verlag. Berlin, Germany. pp. 181–208.
Colbert, T., Till, B.J., Tompa, R., Reynolds, S., Steine, M.N., Yeung, A.T., McCallum, C.M., Comai, L., Henikoff, S. 2001. High-throughput screening for induced point mutations. Plant Physiol. 126:480–484.
Dhaliwal, A.K., Mohan, A., Sidhu, G., Maqbool, R., Gill, K.S. 2015. An ethylmethane sulfonate mutant resource in pre-green revolution hexaploid wheat. Plos One 10:e0145227.
Dong, C., Dalton-Morgan, J., Vincent, K., Sharp, P. 2009. A modified TILLING method for wheat breeding. The Plant Genome 2:39–47.
Ganeshan, S., Baga, M., Limin, A.E., Ratnayaka, I., Chodaparambil, S.V., Fowler, D.B., Chibbar, R.N. 2007. Biotechnological approaches for improving the tolerance of wheat to abiotic stress. In: Singh, D.P., Tomar, V.S., Behl, R.K., Upadhyaya, S.D., Bhale, M.S., Khare, D. (eds), Int. Conf. on Sustainable Crop Production in Stress Environments: Management and Genetic Options. Agrobios International. Jodhpur, India. pp. 297–309.
Ganeshan, S., Drinkwater, J.M., Repellin, A., Chibbar, R.N. 2010. Selected carbohydrate metabolism genes show coincident expression peaks in grains of in vitro-cultured immature spikes of wheat (Triticum aestivum L.). J. Agric. Food Chem. 58:4193–4201.
Ganeshan, S., Leis, M., Drinkwater, J.M., Madsen, L.T., Jain, J.C., Chibbar, R.N. 2011. In vitro-cultured wheat spikes provide a simplified alternative for studies of cadmium uptake in developing grains. J. Sci. Food Agric. 92:1740–1747.
Henry, I.M., Nagalakshmi, U., Lieberman, M.C., Ngo, K.J., Krasileva, K.V., Vasquez-Gross, H., Akhunova, A., Akhunov, E., Dubcovsky, J., Tai, T.H., Comai, L. 2014. Efficient genome-wide detection and cataloging of EMS-induced mutations using exome capture and next-generation sequencing. Plant Cell 26:1382–1397.
Hogg, A.C., Gause, K., Hofer, P., Martin, J.M., Graybosch, R.A., Hansen, L.E., Giroux, M.J. 2013. Creation of a high-amylose durum wheat through mutagenesis of starch synthase II (SSIIa). J. Cereal Sci. 57:377–383.
Jansson, C., Sun, C., Ganeshan, S., Chibbar, R.N. 2007. Plant breeding: antisense ODN inhibition in in vitro spike cultures as a powerful diagnostic tool in studies on cereal grain development. In: Esser, K., Lüttge, U.E., Beyschlag, W., Murata, J. (eds.), Progress in Botany. Springer Verlag. Berlin, Germany. pp. 179–190.
King, R., Bird, N., Ramirez-Gonzalez, R., Coghill, J.A., Patil, A., Hassani-Pak, K., Uauy, C., Phillips, A.L. 2015. Mutation scanning in wheat by exon capture and next-generation sequencing. Plos One 10:e0137549.
McCallum, C.M., Comai, L., Greene, E.A., Henikoff, S. 2000a. Targeting induced local lesions in genomes (TILLING) for plant functional genomics. Plant Physiol. 123:439–442.
McCallum, C.M., Comai, L., Greene, E.A., Henikoff, S. 2000b. Targeted screening for induced mutations. Nat. Biotechnol. 18:455–457.
Parry, M.A.J., Madgwick, P.J., Bayon, C., Tearall, K., Hernandez-Lopez, A., Baudo, M., Rakszegi, M., Hamada, W., Al-Yassin, A., Ouabbou, H., Labhilili, M., Phillips, A.L. 2009. Mutation discovery for crop improvement. J. Exp. Bot. 60:2817–2825.
Regina, A., Bird, A., Topping, D., Bowden, S., Freeman, J., Barsby, T., Kosar-Hashemi, B., Li, Z.Y., Rahman, S., Morell, M. 2006. High-amylose wheat generated by RNA interference improves indices of large-bowel health in rats. Proc. Natl Acad. Sci. U. S. A. 103:3546–3551.
Ririe, K.M., Rasmussen, R.P., Wittwer, C.T. 1997. Product differentiation by analysis of DNA melting curves during the polymerase chain reaction. Anal. Biochem. 245:154–160.
Sadasivaiah, R.S., Perkovic, S.M., Pearson, D.C., Postman, B. 2000. Registration of AC Nanda wheat. Crop Sci. 40:579–580.
Sestili, F., Botticella, E., Bedo, Z., Phillips, A., Lafiandra, D. 2010 Production of novel allelic variation for genes involved in starch biosynthesis through mutagenesis. Mol. Breed. 25:145–154.
Simon, M.R. 1999. Inheritance of flag-leaf angle, flag-leaf area and flag-leaf area duration in four wheat crosses. Theor. Appl. Genet. 98:310–314.
Slade, A.J., Fuerstenberg, S.I., Loeffler, D., Steine, M.N., Facciotti, D. 2005. A reverse genetic, nontransgenic approach to wheat crop improvement by TILLING. Nat. Biotechnol. 23:75–81.
Till, B.J., Reynolds, S.H., Greene, E.A., Codomo, C.A., Enns, L.C., Johnson, J.E., Burtner, C., Odden, A.R., Young, K., Taylor, N.E., Henikoff, J.G., Comai, L., Henikoff, S. 2003. Large-scale discovery of induced point mutations with high-throughput TILLING. Genome Res. 13:524–530.
Uauy, C., Paraiso, F., Colasuonno, P., Tran, R.K., Tsai, H., Berardi, S., Comai, L., Dubcovsky, J. 2009. A modified TILLING approach to detect induced mutations in tetraploid and hexaploid wheat. BMC Plant Biol. 9:115.
Acknowledgements
Canada Research Chairs and the Saskatchewan Agriculture Development Fund is acknowledged for financial support.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material (ESM)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Ganeshan, S., Chibbar, R.N. A Simple Novel Expedited Spike Culture-derived Variation Creation Strategy in Wheat. CEREAL RESEARCH COMMUNICATIONS 45, 539–548 (2017). https://doi.org/10.1556/0806.45.2017.047
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/0806.45.2017.047