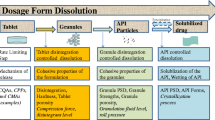
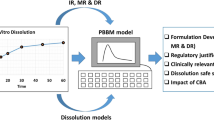
Abstract
Assessing in vivo performance to inform formulation selection and development decisions is an important aspect of drug development. Biopredictive dissolution methodologies for oral dosage forms have been developed to understand in vivo performance, assist in formulation development/optimization, and forecast the outcome of bioequivalence studies by combining them with simulation tools to predict plasma profiles in humans. However, unlike compendial dissolution methodologies, the various biopredictive methodologies have not yet been harmonized or standardized. This manuscript presents the initial phases of an effort to develop best practices and move toward standardization of the biopredictive methodologies through the Product Quality Research Institute (PQRI, https://pqri.org) entitled “The standardization of in vitro predictive dissolution methodologies and in silico bioequivalence study Working Group.” This Working Group (WG) is comprised of participants from 10 pharmaceutical companies and academic institutes. The project will be accomplished in a total of five phases including assessing the performance of dissolution protocols designed by the individual WG members, and then building “best practice” protocols based on the initial dissolution profiles. After refining the “best practice” protocols to produce equivalent dissolution profiles, those will be combined with physiologically based biopharmaceutics models (PBBM) to predict plasma profiles. In this manuscript, the first two of the five phases are reported, namely generating biopredictive dissolution profiles for ibuprofen and dipyridamole and using those dissolution profiles with PBBM to match the clinical plasma profiles. Key experimental parameters are identified, and this knowledge will be applied to build the “best practice” protocol in the next phase.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, many investigations to better understand the in vivo behavior of immediate release (IR) orally dosage forms have been carried out in both the pharmaceutical industry and academia. The prediction of in vivo dissolution has been substantially improved by incorporating aspects of the human gastrointestinal (GI) physiology into increasing advanced biopredictive dissolution methodologies, noting that such predictions are not possible using the traditional compendial dissolution methodologies for low solubility drugs. Biopredictive dissolution methodologies have been widely adopted to evaluate the in vivo performance of test oral dosage forms, under consideration of quality by design (QbD) concepts, as a part of overall efforts to optimize oral dosage form performance.
Compendial dissolution experiments are generally conducted under highly regulated standard conditions of temperature, volume, hydrodynamics, and medium composition for specific compounds and may be useful to predict in vivo dissolution for highly soluble oral drugs (1). The United States Pharmacopoeia (USP) standard dissolution methods, which are largely harmonized with those of other developed countries, are mainly performed for quality control purposes to assure the quality of oral drug products and to evaluate formulation variables such as excipients, batch-to-batch variation, and manufacturing procedure/changes. Although the compendial methodologies are valuable for these purposes, they typically do not aim to reflect in vivo dissolution behavior, mostly because they are constructed to determine the quality of the dosage form and how well the dose can be released from it, rather than specifically taking into account the physiological parameters (e.g., the pH profile, bile salts concentrations, hydrodynamics, GI transit, digestive enzymes) of the GI tract which largely affect drug dissolution after oral administration.
For dissolution testing to be a more useful predictor of in vivo drug dissolution and, hence, oral drug absorption, it should reflect the physiological conditions of the GI tract more closely than the compendial methodologies. Academia and pharmaceutical industries have advanced dissolution testing to adopt the use of biorelevant media, which were first proposed by Dressman et al. in 1998 (2), incorporating the pH, ionic strength, and buffer and bile component concentration changes that occur along the GI tract to produce more meaningful dissolution profiles in terms of predicting the in vivo behavior of oral dosage forms (3,4,5,6,7,8).
A key addition to the range of biopredictive dissolution methodologies is the transfer methodology, which was first introduced by Kostewicz et al. in 2004 (4). The transfer model enables the researcher to model the behaviour of a drug/dosage form as it travels from the stomach into the small intestine and thus assess whether a poorly soluble weakly basic drug (e.g., Biopharmaceutical Classification System (BCS) class IIb according to the sub-classification), might precipitate when it reaches the higher pH environment in the small intestine, or how long it will take for a poorly soluble acidic drug (e.g., BCS class IIa) to dissolve and become available for absorption (9).
Over the last two decades, individual research groups have typically created/adapted their dissolution methodologies to suit their research objectives. Therefore, even with the same drug and oral dosage form, published biopredictive dissolution profiles may differ due to variations in the reported experimental conditions, e.g., buffer species, buffer volumes, and buffer pH. Thus, it is hard to identify key experimental conditions and to establish the best biorelevant dissolution practices. Scientists from industry and academia came together with the support of the PQRI, which is a non-profit consortium of organization, to create a collaborative opportunity for scientists from industry, academia, and regulatory agencies, to address the biopharmaceutical issues. In this study, each research group was invited to perform its own variation of biopredictive dissolution methodology on the same batches of two oral drug products to address the following questions: (1) can the scientific community, as exemplified by PQRI WG members, identify elements that are key to dissolution performance by comparing their dissolution profiles, (2) do the results enable us to move toward harmonizing experimental methodologies, and (3) are the dissolution profiles projected to predict bioequivalence (BE) with clinical data using a physiologically based biopharmaceutics modeling (PBBM) approach. The rationale behind these studies was that, by comparing results and identifying any differences in the detailed dissolution methodologies, it would be possible to tighten and harmonize the requirements for the biopredictive dissolution methodologies, resulting in clearer and more definitive interpretation of results for both the pharmaceutical industry and regulatory agencies. The dissolution profiles from experiments using the “best practice” methodologies will then inputted into PBBMs to determine whether they produce BE or non-BE plasma profiles when compared against clinical results. Potentially the identification of “best practice” methodology will require a “learn and confirm” approach, whereby several iterations are needed to arrive at globally applicable “best practice” methodologies.
The overarching goals of the PQRI WG studies are to advance and harmonize biopredictive tools. The studies are divided into five phases to achieve these goals:
-
(1)
Using ibuprofen (400 mg), a widely used weakly acid drug, biopredictive dissolution profiles are generated and compared among the WG members to determine whether the results among the participating research groups are equivalent or not in terms of generating plasma profiles with PBBM which are similar to those observed in clinical pharmacokinetic studies.
-
(2)
Using dipyridamole (50 mg), a widely used weakly base drug, biopredictive dissolution profiles are generated with the same methodologies used in (1) and be compared among the WG members to determine whether the results among the participating research groups are equivalent or not in terms of generating plasma profiles with PBBM which are similar to those observed in clinical pharmacokinetic studies.
-
(3)
Using a higher dose of dipyridamole (200 mg), biopredictive dissolution profiles will be generated with the same methods used in (1) and be compared among the WG members to determine whether the results among the participating research groups are equivalent or not in terms of generating plasma profiles with PBBM which are similar to those observed in clinical pharmacokinetic studies.
-
(4)
If the results from (3) are not equivalent among participants, dissolution profiles would be generated with “best practice” biopredictive protocols at the 200 mg dipyridamole dose. If not all dissolution profiles obtained using this methodology are able to simulate the plasma profiles observed in vivo, it will be further revised in a “learn and confirm” cycle.
-
(5)
Throughout these four phases, WG members will discuss how to incorporate the biopredictive dissolution profiles into PBBM. The goal is to recognize the important parameters in biopredictive dissolution testing, and then harmonize the protocols to achieve uniform dissolution profiles regardless of site and researcher/operator. In turn, this will enable optimization of PBBM settings.
Since the purpose of this PQRI WG is to harmonize the biopredictive dissolution methodologies, the same batch/lots of ibuprofen and dipyridamole were supplied to every participant in order to evaluate different experimental conditions and dissolution methodologies. This manuscript presents the results from the first two phases described in the overall study plan: comparison of ibuprofen and dipyridamole dissolution results from tests performed by WG members according to their individual protocols. The results will assist in identifying key experimental parameters to developing a “best practice” protocol throughout those two phases.
Materials
A total of 400 mg ibuprofen tablets (Lot# 7708221A, Stride Pharma Science Ltd, Bangalore, India) and 50 mg dipyridamole tablets (Lot# 200203A, Rising Pharmaceuticals, East Brunswick, NJ, USA) were purchased and distributed to all members of the PQRI working group. For the preparation of biopredictive media, FaSSGF/FeSSIF/FaSSIF was purchased in powder form from Biorelevant.com (Biorelevant.com, London, UK) and prepared by each WG before the biopredictive dissolution study. All other chemicals were of analytical grade or HPLC grade.
Methods
Participants in this study were AstraZeneca, Boehringer Ingelheim Pharmaceuticals, Bristol-Myers Squibb Company, Fraunhofer Institute for Translational Medicine Pharmacology, Goethe Universität, MSD, Schrodinger Inc., Sawai Pharmaceutical Co. Ltd, University of Minnesota, and Miguel Hernandez University.
In each case, the WG was instructed to use their usual in-house biopredictive methodologies for two stage dissolution methods and/or the transfer method, incorporating their usual choices of experimental condition and buffers (Scheme 1). Those dissolution methods were loosely regulated to maintain parameters within the wide range of experimental conditions shown below:
-
Fasted gastric conditions:
-
The buffer: 0.01–0.1 M HCl, SGF, or FaSSGF
-
The pH range: pH 1–2.5
-
The volume 50–500 mL (buffer only or buffer + water)
-
The mixing: not specified
-
-
Fasted intestinal conditions:
-
The buffer: 5–10 mM maleate, 25–100 mM phosphate buffer, or FaSSIF
-
The pH range: pH 6.0–7.5
-
The volume: 40–500 mL
-
The mixing: not specified
-
-
Transfer rate for transfer method:
-
Transfer rate: 1–62 mL/min (zero-order or first-order rate)
-
Dissolution Methodologies
The WG’s dissolution methodologies for ibuprofen and dipyridamole are summarized in Tables I and II, respectively. The theories behind some of the methodologies and more detailed descriptions can be found in the literature (5, 10, 11).
Modeling Approach
The dissolution profiles obtained using the participant’s biopredictive methods were coupled with an in silico method, GastroPlus™ version 9.8 (SimulationPlus, Inc., Lancaster, CA), to simulate human plasma profiles, as follows. In order to minimize any version or variation in modeling, all simulations were performed centrally by the same operator.
Oral drug absorption was computed from the physicochemical, pharmacokinetic, and dissolution properties of ibuprofen, a BCS class IIa drug, and dipyridamole, a BCS class IIb drug, using previously reported simulation methods (8, 12). Briefly, single simulations were performed with the biopredictive dissolution profiles to compare with the pharmacokinetic profile of 400 mg ibuprofen IR tablet and 50 mg dipyridamole IR tablet ingested under fasted conditions. Those biopredictive dissolution profiles were incorporated into the in silico software as the controlled release profiles to specify the drug release profiles. No drug absorption from the stomach was assumed in this set of predictions. The duration of the simulations was 24 h. Input parameters for in silico simulations to predict the plasma profiles of ibuprofen and dipyridamole were obtained from the literature and the values are summarized in Table III (1, 13,14,15,16,17,18,19,20,21). The predictions were performed using the GastroPlus™ standard physiological conditions: Human Physiological-Fasted and Opt LogD Model SA/V 6.1.
The predicted plasma profiles were compared with clinical trial results for ibuprofen and dipyridamole pharmacokinetics to evaluate whether incorporating the results from the various dissolution studies into GastroPlus™ was able to predict the in vivo performance (22,23,24,25).
Results
Ibuprofen Dissolution
The dissolution profiles of ibuprofen are summarized in Fig. 1 for the two-stage methodology and Fig. 2 for the transfer methodology. In two-stage dissolution methods, the dissolution curves obtained using experimental conditions within the above-mentioned constraints exhibited similar profiles regardless of buffer volume, buffer species, buffer pH, rotational speed, and duration of the first (gastric) stage (Fig. 1). In the transfer method dissolution profiles, only the dissolution profiles in the small intestinal vessel(s) are plotted (Fig. 2). The dissolution profiles from Institutes D and G both achieved 100% dissolution, in line with the solubility of ibuprofen at intestinal pH. By contrast, the profile from Institute F reached only 50% dissolution of ibuprofen, which was explained post hoc by the fact that 50% of the drug was retained in the gastric stage rather than being transferred. The profile from Institute H exhibited even lower dissolution of ibuprofen because the second (“duodenal”) chamber was held at a constant volume of 30 mL, by discarding fluid transferred out of that chamber. Since Institute H only measured the drug concentration in the second chamber, ibuprofen dissolution was essentially measured in just 30 mL of the 0.01N HCl/SIF mixture.
Since ibuprofen is a weakly acidic drug and the small intestinal environment is favorable for its dissolution, for modeling purposes, it is necessary to account for all transferred material from the gastric to the small intestinal stage. For this reason, the observed concentration and the volume out of duodenal chamber at each time point were used to generate the full dissolution profiles in the small intestinal region for the Institute H results (Fig. 2).
Dipyridamole Dissolution
The dissolution profiles of dipyridamole are summarized in Fig. 3 for the two-stage methodologies and in Fi, 4 for the transfer methodologies. Using two-stage dissolution methodologies, three institutes (Institute A, B, and D) displayed complete dissolution of dipyridamole and no or close to no precipitation regardless of buffer species and capacity, while Institute C observed precipitation following pH adjustment to 6.8 with NaOH (Fig. 3).
Dipyridamole dissolution profiles generated using the two-stage method. The first stage dissolution was conducted for either the first 20 or 30 min before the second, small intestinal condition was initiated. *The buffer at the second stage: SIF – simulated intestinal fluid, 1/10 SIF – 1/10th concentration of SIF, Maleate – 7 mM Maleate buffer
In transfer methodologies, only the dissolution profiles in the small intestinal vessel(s) are plotted (Fig. 4). The results from Institutes D, E, and I exhibited more than 80% dissolution of dipyridamole, while Institutes G and F reported only 50–60% of dipyridamole dissolution at the end of the experiment. As for ibuprofen, Institute H reported very low dissolution of dipyridamole because, once again, the dissolution profile was calculated based on the volume in the second (“duodenal”) chamber multiplied by the drug concentration at the time points of sample withdrawal. Dipyridamole is a weak base drug and the gastric environment, pH, stirring speed, volume, and the residence time might all be important to describe its dissolution from the tablet. Since the pKa of dipyridamole (pKa 6.4) is close to the pH of the dissolution buffers representing the small intestine (pH 6.5–7.5, depending on the institute), the buffer capacity, species and volume are all expected to affect its dissolution. This was reflected by the large range of results recorded.
Ibuprofen Modeling
The plasma profiles of ibuprofen were simulated using the dissolution profiles produced by the WG as the in vivo dissolution input. The purpose of this simulation was not to provide a fully accurate prediction of the observed clinical plasma profile (e.g., no attempt was made to further optimize, e.g., disposition parameters in the model) but rather directed at assessing the criticality of different in vitro conditions by different members of the WG (including pH, buffer species, buffer capacity, volumes of media, media change conditions and stirring rate). The prediction of ibuprofen absorption after oral dosing of 400 mg with the biopredictive dissolution profiles from the two-stage test was all BE with the clinical data (Fig. 5). The simulation results presented in Table IV suggest that all of the two-stage dissolution conditions studied were indeed biopredictive for ibuprofen, even though there were some differences in buffers, transfer rate/timing and other experimental conditions. Part of the reason for this is that for weakly acidic drugs like ibuprofen, the environment in the small intestine is far more favorable than the one in the stomach for dissolution, so as long as the buffer pH is higher than the drug’s pKa and the pH is adequately maintained over the experiment, modest differences in dissolution conditions are unlikely to affect the outcome of the simulation with respect to BE.
Predicted plasma profiles of ibuprofen based on biopredictive dissolution profiles. Dark blue dotted lines represent 80% and 125% of the average clinical plasma profile (23). a represents simulated results based on two-stage dissolution methodologies and b represents simulated results based on transfer methodologies
With respect to the transfer test, only the results from Institutes D, G, and H fell within the BE limits (Fig. 5). The failure of results from Institute F to meet the BE limits can be traced back to the experimental design: in the Institute F experiments, only 50% of ibuprofen was transferred to the small intestinal chamber from the gastric chamber. Even though the dissolution profile was re-generated to assume the same concentration was achieved in the rest of the small intestine as in the duodenal chamber, the concentration of ibuprofen in the duodenal chamber used to generate the simulated in vivo dissolution profile might have been too low to evaluate the biorelevant dissolution. Since a large volume of acid comes from the gastric chamber into the duodenal chamber (which holds only 30 mL of the small intestinal buffer), the pH in the duodenal chamber is lowered, and thus limits the extent of ibuprofen dissolution, leading to an underestimation of the predicted plasma profile of ibuprofen. Since ibuprofen dissolves well at intestinal pH (as shown in Institute D, and G setups) and is completely absorbed in vivo (23, 26), it seems that the short duration of dissolution, lowered pH and small volume used to represent the small intestinal environment in the Institute H setup, combine to produce a less extensive dissolution than would occur in the small intestine in vivo.
Dipyridamole Modeling
The plasma profiles of dipyridamole were simulated using the dissolution profiles produced by the WG as the in vivo dissolution input, analogous to the approach used for ibuprofen. Again, the purpose of this simulation was not to provide a fully accurate prediction of the observed clinical plasma profile but rather directed toward assessing the criticality of differences in the in vitro conditions among the different institutes. The results are shown in Fig. 6a and b.
Predicted plasma profiles of dipyridamole based on biopredictive dissolution profiles. Dark blue dot lines represent 80% and 125% of the calculated mean clinical plasma profile (43). a represents simulated results based on two-stage dissolution methodologies and b represents simulated results based on transfer methodologies
While nine of the eleven simulated plasma profiles satisfied equivalency with the clinical data with respect to Cmax but overestimated AUC, the dissolution profile by Institute F led to a simulation which satisfied the BE requirements for AUC0-24 but missed on Cmax (Table V). An issue with all of the simulations is that the clinical data chosen as the reference in vivo data exhibited a Tmax of 0.5 h, while the in silico simulations based on the biorelevant dissolution profiles consistently predicted a longer Tmax of ~1.5 h (27). BCS class IIb drugs like dipyridamole dissolve well in the gastric environment and the earlier Tmax suggests a faster gastric emptying time. Indeed, the stomach has a rapid emptying mechanism described as “Magenstrasse,” which might potentially explain the relatively faster Tmax observed in the reference clinical data (28, 29). It also should be mentioned that other clinical data reported by Gregov et al. exhibited a Tmax of 1.06–1.58 h over the 25–200 mg dose range, in agreement with the simulation results, and, as seen, high inter-study and subject variabilities in the PK study of dipyridamole have been reported (10, 24, 30).
The dose of 50 mg of dipyridamole in this set of experiments is relatively low and its pKa is high (pKa 6.4). Since the total aqueous volume available for dissolution at the end of the experiment was ~300 to 500 mL in all cases (except for Institute H), most of the dipyridamole dose dissolved, with little precipitation, and hence more absorption was predicted by most simulations than was observed clinically.
Although it is beyond the scope of the current evaluation, more investigation of the precipitation kinetics and of the intestinal pH, along with buffer capacity, would be required to fully simulate the entire plasma concentration profile for this poorly soluble but highly permeable weak base. Since dipyridamole is a weakly basic drug, the gastric environment is the more favorable for its dissolution, and it is consequently necessary to model the pH and dissolution time in the stomach (via the gastric emptying kinetics) in a way that reflects the physiology closely in order to attain a more predictive dissolution and simulation. Additionally, media volumes and hydrodynamics of the test method may need to be adjusted to adequately reflect the supersaturation and precipitation kinetics.
Discussion
As dissolution technology has advanced, academia and pharmaceutical companies have tried to predict how oral formulations would perform with regard to dissolution in the human GI tract, such that oral absorption can be predicted with in silico technology and the oral formulation can be optimized. A meaningful prediction of bioperformance for oral dosage forms would bring huge benefits, and, yet, unlike compendial dissolution methodologies, there is currently no harmonization of biopredictive dissolution methodology and its use in regulatory applications. This is likely because each pharmaceutical company and academic institute has developed its own version of biopredictive methodology to test their oral compounds of interest and reasonably predict their performance.
Many BCS class II and IV drugs have pH-dependent solubilities in the physiological pH range. Those oral drugs, especially BCS class IIb drugs like dipyridamole, should be sequentially studied in different pH buffers, gastric and intestinal pH conditions, to adequately predict bioperformance of those oral drug formulations. There are multiple different methodologies to predict the bioperformance of oral drug products (3, 7, 31,32,33,34). However, largely, biorelevant methodologies, or in vivo predictive dissolution (iPD), can be divided into two major types: two-stage dissolution and transfer methodologies. Even these two approaches are subject to a wide variety of experimental conditions employed with respect to buffer species and volumes, buffer pH, rotational speeds, dissolution time and, for transfer methodologies, the transfer rate (Tables I and II). Various institutes have successfully utilized biopredictive dissolution results to predict the in vivo performance of the dosage forms they tested and have also optimized the input of their results into in silico models (6, 31, 35, 36).
The most popular biopredictive dissolution testing procedure appears to be the two-stage dissolution methodology, which involves two different biorelevant media conditions (the gastric conditions, i.e., acidic, as the first stage, and the intestinal conditions, i.e. close to neutral, as the second stage) in sequence, to evaluate the bioperformance of drug substances and/or oral formulations when those are introduced to two different pH environments (3, 10, 34). Another biopredictive dissolution testing procedure is the transfer model methodology, which has the same principle as the two-stage dissolution methodology but may involve more than two chambers, with mechanical transfer of the dissolution buffers from one to the next chamber (5, 8, 11, 36). Different dissolution media have been used in those dissolution methodologies to mimic the gastric conditions, namely 0.01–0.1N hydrochloric acid (HCl), simulated gastric fluid (SGF), fasted state simulated gastric fluid (FaSSGF), and less acidic buffers such as maleate buffer to represent achlorhydric conditions (pH 4.0 to 6.0). To mimic the intestinal conditions, various concentrations of phosphate buffer like simulated intestinal fluid (SIF) and fasted state simulated intestinal fluid (FaSSIF), including its different versions, within the pH range of 6.5 to 7.5 (37,38,39,40). Those biorelevant dissolution methodologies have been mainly used to investigate the supersaturation and the precipitation behavior of poorly soluble drugs or drugs which have pH-dependent solubility.
As seen in Tables I and II, the conditions of biorelevant / iPD methodologies vary among the PQRI WG, even though this WG understands the human GI physiology well and the methods used are similar in their approach. Rather than adapting a “one method fits all” approach, scientists in each institute tend to customize the methods and conditions for dissolution testing to the specific drug and formulation properties they are tasked to work with, in order to predict the in vivo performance of the formulation. In the past few years, it has also been broadly recognized that the in vitro experimental conditions used to represent in vivo conditions should be selected in such a way that the dissolution profiles can be easily connected to PBPK models in a PBBM to realize successful in vivo predictions (27, 41,42,43,44,45). In the present studies, all data were entered into GastroPlus™ using the same approach and by the same operator, so that any differences in dissolution data would be reflected the same way in the simulations of the PK profile. This was important to understand which differences in methodology would lead to a difference in the simulated plasma profile and which differences would not.
The Oral Biopharmaceutics Tools (OrBiTo) project was carried out as a consortium in European countries. This collaborative effort has gained much knowledge and filled many gaps in our understanding of GI function pertaining to oral drug absorption and oral drug delivery, creating a framework for the development of the next predictive biopharmaceutical tools, and their validation (46,47,48,49,50,51). In the OrBiTo project, a ring study among both academic and industrial partners verified the reproducibility and reliability of biorelevant dissolution among sites, based on a fixed dissolution protocol.
By contrast, in this PQRI project, biorelevant dissolution was carried out using different biopredictive dissolution methods to see how comparable those results are in terms of being able to simulate in vivo performance using PBBM. This enabled us to identify which experimental parameters might have a big impact on the simulated plasma profiles. The ibuprofen results demonstrated BE with the clinical data for the majority of the PQRI WG (Table IV and Fig. 5). Given that ibuprofen is a weakly acidic drug, the conditions over the intestinal transit time of ~3 h are favorable for its dissolution. On the other hand, weakly basic drugs like dipyridamole must dissolve within a comparably short time in the stomach since the gastric emptying time is ~0.5–1 h (52). Therefore, the range of dissolution results that will satisfy BE criteria might be expected to be wider for a poorly soluble weak acid than for a poorly soluble weak base. This was borne out by the simulation results for ibuprofen and dipyridamole. Still, the results of these first two phases of the project have given the WG the confidence to proceed to the next phase of the project. As a next step, the WG will discuss and determine the most important experimental parameters for biopredictive dissolution and narrow the range of experimental conditions with a view to developing “best practice” biopredictive dissolution methodologies. Then, the WG will perform further dissolution experiments with the same batches used to generate the results in the current studies, with the aim of harmonizing the biopredictive tools and conditions and identifying best practices in PBBM for successful prediction of in vivo performance.
Conclusion
Biopredictive dissolution is, by its nature, a surrogate methodology and since surrogate methodologies are designed to prevent clinical failures, the availability of well-designed and validated biopredictive dissolution methodologies could even lead to viewing clinical PK studies as confirmatory methodologies. In the next steps of these PQRI studies, key experimental parameters will be identified, and the WG will develop “best practice” biopredictive dissolution methodologies, which in turn will be verified by testing their ability to predict plasma profiles in the framework of PBBM. The availability and application of “best practice” biopredictive dissolution methodologies will lead to efficiencies in oral product development, a reduction of the number of animal studies in preclinical development, as well as potentially increasing the success rate of clinical studies. The PQRI studies constitute the first big step toward harmonizing biorelevant dissolution methodologies.
Change history
08 June 2023
A Correction to this paper has been published: https://doi.org/10.1208/s12248-023-00827-0
References
Siepmann J, Siepmann F. Mathematical modeling of drug dissolution. Int J Pharm. 2013;453(1):12–24.
Dressman JB, et al. Dissolution testing as a prognostic tool for oral drug absorption: immediate release dosage forms. Pharm Res. 1998;15(1):11–22.
Fiolka T, et al. Biorelevant two-stage in vitro testing for rDCS classification and in PBPK Modeling-case example ritonavir. J Pharm Sci. 2020;109(8):2512–26.
Kostewicz ES, et al. Predicting the precipitation of poorly soluble weak bases upon entry in the small intestine. J Pharm Pharmacol. 2004;56(1):43–51.
Kourentas A, et al. An in vitro biorelevant gastrointestinal transfer (BioGIT) system for forecasting concentrations in the fasted upper small intestine: Design, implementation, and evaluation. Eur J Pharm Sci. 2016;82:106–14.
Matsui K, et al. Utilization of gastrointestinal simulator, an in vivo predictive dissolution methodology, coupled with computational approach to forecast oral Absorption of Dipyridamole. Mol Pharm. 2017;14(4):1181–9.
Minekus, M. The TNO Gastro-Intestinal Model (TIM). In: The impact of food bioactives on health: in vitro and ex vivo models, K. Verhoeckx, et al., Editors. 2015: Cham (CH). p. 37–46.
Patel S, et al. Integration of precipitation kinetics from an in vitro, multicompartment transfer system and mechanistic oral absorption modeling for pharmacokinetic prediction of weakly basic drugs. J Pharm Sci. 2019;108(1):574–83.
Tsume Y, et al. The biopharmaceutics classification system: subclasses for in vivo predictive dissolution (IPD) methodology and IVIVC. Eur J Pharm Sci. 2014;57:152–63.
Segregur D, et al. Evaluating the impact of acid-reducing agents on drug absorption using biorelevant in vitro tools and PBPK modeling - case example dipyridamole. Eur J Pharm Sci. 2021;160: 105750.
Takeuchi S, et al. Evaluation of a three compartment in vitro gastrointestinal simulator dissolution apparatus to predict in vivo dissolution. J Pharm Sci. 2014;103(11):3416–22.
Hens B, et al. Dissolution challenges associated with the surface ph of drug particles: integration into mechanistic oral absorption modeling. AAPS J. 2022;24(1):17.
Abernethy DR, Greenblatt DJ. Ibuprofen disposition in obese individuals. Arthritis Rheum. 1985;28(10):1117–21.
Alvarez C, et al. Investigation on the possibility of biowaivers for ibuprofen. J Pharm Sci. 2011;100(6):2343–9.
Avdeef A, et al. PH-metric log P 11. pKa determination of water-insoluble drugs in organic solvent-water mixtures. J Pharm Biomed Anal. 1999;20(4):631–41.
Avdeef A, et al. pH-metric logP 10. Determination of liposomal membrane-water partition coefficients of ionizable drugs. Pharm Res. 1998;15(2):209–15.
Yu A, et al. Mechanistic deconvolution of oral absorption model with dynamic gastrointestinal fluid to predict regional rate and extent of GI drug dissolution. AAPS J. 2019;22(1):3.
Kalantzi L, et al. Canine intestinal contents vs. simulated media for the assessment of solubility of two weak bases in the human small intestinal contents. Pharm Res. 2006;23(6):1373–81.
Mahony C, et al. Dipyridamol kinetics. Clin Pharmacol Ther. 1982;31(3):330–8.
Mitra A, Fadda HM. Effect of surfactants, gastric emptying, and dosage form on supersaturation of dipyridamole in an in vitro model simulating the stomach and duodenum. Mol Pharm. 2014;11(8):2835–44.
Sugano K. Computational oral absorption simulation of free base drugs. Int J Pharm. 2010;398(1–2):73–82.
Albert KS, et al. Effects of age on the clinical pharmacokinetics of ibuprofen. Am J Med. 1984;77(1A):47–50.
Lockwood GF, et al. Pharmacokinetics of ibuprofen in man. I. Free and total area/dose relationships. Clin Pharmacol Ther. 1983;34(1):97–103.
Bjornsson TD, Mahony C. Clinical pharmacokinetics of dipyridamole. Thromb Res Suppl. 1983;4:93–104.
Russell TL, et al. Upper gastrointestinal pH in seventy-nine healthy, elderly, North American men and women. Pharm Res. 1993;10(2):187–96.
Atkinson HC, et al. Pharmacokinetics and bioavailability of a fixed-dose combination of ibuprofen and paracetamol after intravenous and oral administration. Clin Drug Investig. 2015;35(10):625–32.
Wagner C, et al. Predicting the oral absorption of a poorly soluble, poorly permeable weak base using biorelevant dissolution and transfer model tests coupled with a physiologically based pharmacokinetic model. Eur J Pharm Biopharm. 2012;82(1):127–38.
Grimm M, et al. Gastric Water emptying under fed state clinical trial conditions is as fast as under fasted conditions. Mol Pharm. 2017;14(12):4262–71.
Jefferson G. The human stomach and the Canalis Gastricus (Lewis). J Anat Physiol. 1915;49(Pt 2):165–81.
Gregov D, et al. Dipyridamole: pharmacokinetics and effects on aspects of platelet function in man. Br J Clin Pharmacol. 1987;24(4):425–34.
Hens B, et al. Gastrointestinal transfer: in vivo evaluation and implementation in in vitro and in silico predictive tools. Eur J Pharm Sci. 2014;63:233–42.
Lyng E, et al. Increased bioavailability of celecoxib under fed versus fasted conditions is determined by postprandial bile secretion as demonstrated in a dynamic gastrointestinal model. Drug Dev Ind Pharm. 2016;42(8):1334–9.
Shi Y, et al. Application of a biphasic test for characterization of in vitro drug release of immediate release formulations of celecoxib and its relevance to in vivo absorption. Mol Pharm. 2010;7(5):1458–65.
Takagi T, et al. In vitro sensitivity analysis of the gastrointestinal dissolution profile of weakly basic drugs in the stomach-to-intestine fluid changing system: explanation for variable plasma exposure after oral administration. Mol Pharm. 2021;18(4):1711–9.
Fotaki N, et al. A comparative study of different release apparatus in generating in vitro-in vivo correlations for extended release formulations. Eur J Pharm Biopharm. 2009;73(1):115–20.
Tsume Y, et al. The introduction of a new flexible in vivo predictive dissolution apparatus, GIS-Alpha (GIS-alpha), to study dissolution profiles of BCS class IIb drugs Dipyridamole and Ketoconazole. J Pharm Sci. 2020;109(11):3471–9.
Fuchs A, Dressman JB. Composition and physicochemical properties of fasted-state human duodenal and jejunal fluid: a critical evaluation of the available data. J Pharm Sci. 2014;103(11):3398–411.
Fuchs A, et al. Advances in the design of fasted state simulating intestinal fluids: FaSSIF-V3. Eur J Pharm Biopharm. 2015;94:229–40.
Jantratid E, et al. Dissolution media simulating conditions in the proximal human gastrointestinal tract: an update. Pharm Res. 2008;25(7):1663–76.
Perez de la Cruz Moreno M, et al. Characterization of fasted-state human intestinal fluids collected from duodenum and jejunum. J Pharm Pharmacol. 2006;58(8):1079–89.
Mitra A, Kesisoglou F, Dogterom P. Application of absorption modeling to predict bioequivalence outcome of two batches of etoricoxib tablets. AAPS PharmSciTech. 2015;16(1):76–84.
Otsuka K, Shono Y, Dressman J. Coupling biorelevant dissolution methods with physiologically based pharmacokinetic modelling to forecast in-vivo performance of solid oral dosage forms. J Pharm Pharmacol. 2013;65(7):937–52.
Tsume Y, et al. In silico prediction of drug dissolution and absorption with variation in intestinal pH for BCS class II weak acid drugs: ibuprofen and ketoprofen. Biopharm Drug Dispos. 2012;33(7):366–77.
Vuletic L, et al. Development of a Clinically Relevant Dissolution Method for Metaxalone Immediate Release Formulations Based on an IVIVC Model. Pharm Res. 2018;35(8):163.
Zhang S, et al. Evaluating the bioequivalence of metronidazole tablets and analyzing the effect of in vitro dissolution on in vivo absorption based on PBPK modeling. Drug Dev Ind Pharm. 2019;45(10):1646–53.
Abrahamsson B, et al. Six years of progress in the oral biopharmaceutics area - A summary from the IMI OrBiTo project. Eur J Pharm Biopharm. 2020;152:236–47.
Andreas CJ, et al. Introduction to the OrBiTo decision tree to select the most appropriate in vitro methodology for release testing of solid oral dosage forms during development. Eur J Pharm Biopharm. 2018;130:207–13.
Kostewicz ES, et al. In vitro models for the prediction of in vivo performance of oral dosage forms. Eur J Pharm Sci. 2014;57:342–66.
Lennernas H, et al. Oral biopharmaceutics tools - time for a new initiative - an introduction to the IMI project OrBiTo. Eur J Pharm Sci. 2014;57:292–9.
Mann J, et al. Validation of dissolution testing with biorelevant media: an OrBiTo Study. Mol Pharm. 2017;14(12):4192–201.
Berben P, et al. Biorelevant dissolution testing of a weak base: interlaboratory reproducibility and investigation of parameters controlling in vitro precipitation. Eur J Pharm Biopharm. 2019;140:141–8.
Krieg BJ, et al. In vivo predictive dissolution: transport analysis of the CO2, bicarbonate in vivo buffer system. J Pharm Sci. 2014;103(11):3473–90.
Acknowledgements
The authors acknowledge the Product Quality Research Institute (PQRI) and its member organizations for providing this forum to make this collaboration possible. Authors also thank Jillian Brady and Dede Godstrey for their help and support.
Author information
Authors and Affiliations
Contributions
All authors contributed to the content, review, and editing of the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
All authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original article has been corrected to update the missing beaker picture in Scheme 1 (a) "Gastric Compartment".
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tsume, Y., Ashworth, L., Bermejo, M. et al. Harmonizing Biopredictive Methodologies Through the Product Quality Research Institute (PQRI) Part I: Biopredictive Dissolution of Ibuprofen and Dipyridamole Tablets. AAPS J 25, 45 (2023). https://doi.org/10.1208/s12248-023-00793-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-023-00793-7