Abstract
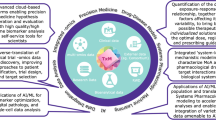
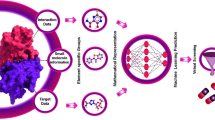
In recent decades, the advancement of computational algorithms and the availability of big data have enabled artificial intelligence (AI) to dramatically improve predictive performance in nearly all research areas. Specifically, machine learning (ML) techniques, a major branch of AI, have been widely used in many tasks of drug discovery and development, including predicting treatment effects, identifying target genes and functional pathways, as well as selecting potential biomarkers. However, in practice, blindly applying ML methods may lead to common pitfalls, including overfitting and lack of generalizability. Therefore, how to improve the robustness and prediction accuracy of ML methods has become a crucial problem for researchers. In this review, we summarize the application of ML models to drug discovery by introducing the top-performing methods developed from large-scale drug-related data challenges in recent years.


Similar content being viewed by others
References
Artificial intelligence abstracts. Artif Intell. 1987;32:414–5.
Ahmet C. Artificial intelligence: how advance machine learning will shape the future of our world. Shockwave Publishing via PublishDrive; 2018.
Michalski RS, Carbonell JG, Mitchell TM. Machine learning: an artificial intelligence approach: Springer Science & Business Media; 2013.
Alpaydin E. Machine learning: The New AI: MIT Press; 2016.
Bishop CM. Pattern recognition and machine learning: Springer; 2016.
Supervised Learning. Neural Smithing. 1999. https://doi.org/10.7551/mitpress/4937.003.0003.
Unsupervised Learning. Unsupervised learning. 1999. https://doi.org/10.7551/mitpress/7011.003.0002.
Deo RC. Machine learning in medicine. Circulation. 2015;132:1920–30.
Burbidge R, Trotter M, Buxton B, Holden S. Drug design by machine learning: support vector machines for pharmaceutical data analysis. Comput Chem. 2001;26:5–14.
Ekins S. The next era: deep learning in pharmaceutical research. Pharm Res. 2016;33:2594–603.
Flower DR. Drug discovery: today and tomorrow. Bioinformation. 2020;16:1–3.
Hughes JP, Rees S, Kalindjian SB, Philpott KL. Principles of early drug discovery. Br J Pharmacol. 2011;162:1239–49.
Mohs RC, Greig NH. Drug discovery and development: role of basic biological research. Alzheimers Dement. 2017;3:651–7.
Vamathevan J, et al. Applications of machine learning in drug discovery and development. Nat Rev Drug Discov. 2019;18:463–77.
Chen H, Engkvist O, Wang Y, Olivecrona M, Blaschke T. The rise of deep learning in drug discovery. Drug Discov Today. 2018;23:1241–50.
Hamet P, Tremblay J. Artificial intelligence in medicine. Metabolism. 2017;69S:S36–40.
Costello JC, et al. A community effort to assess and improve drug sensitivity prediction algorithms. Nat Biotechnol. 2014;32:1202–12.
Harrell FE Jr, Lee KL, Mark DB. Multivariable prognostic models: issues in developing models, evaluating assumptions and adequacy, and measuring and reducing errors. Stat Med. 1996;15:361–87.
Wilson CM, Li K, Yu X, Kuan P-F, Wang X. Multiple-kernel learning for genomic data mining and prediction. BMC Bioinformatics. 2019;20:426.
Bansal M, et al. A community computational challenge to predict the activity of pairs of compounds. Nat Biotechnol. 2014;32:1213–22.
Menden MP, et al. Community assessment to advance computational prediction of cancer drug combinations in a pharmacogenomic screen. Nat Commun. 2019;10.
Li H, Li T, Quang D, Guan Y. Network propagation predicts drug synergy in cancers. Cancer Res Canres. 2018;0740.2018. https://doi.org/10.1158/0008-5472.can-18-0740.
Li H, Hu S, Neamati N, Guan Y. TAIJI: approaching experimental replicates-level accuracy for drug synergy prediction. Bioinformatics. 2019;35:2338–9.
Cristianini N. Cross-Validation (K-Fold Cross-Validation, Leave-One-Out, Jackknife, Bootstrap). Dictionary of bioinformatics and computational biology. 2004. https://doi.org/10.1002/9780471650126.dob0148.pub2.
Elkins JM, et al. Comprehensive characterization of the published kinase inhibitor set. Nat Biotechnol. 2016;34:95–103.
Santos R, et al. A comprehensive map of molecular drug targets. Nat Rev Drug Discov. 2017;16:19–34.
Öztürk H, Özgür A, Ozkirimli E. DeepDTA: deep drug-target binding affinity prediction. Bioinformatics. 2018;34:i821–9.
LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521:436–44.
Pahikkala T, et al. Toward more realistic drug-target interaction predictions. Brief Bioinform. 2015;16:325–37.
He T, Heidemeyer M, Ban F, Cherkasov A, Ester M. SimBoost: a read-across approach for predicting drug-target binding affinities using gradient boosting machines. Aust J Chem. 2017;9:24.
Wang Z, Li H, Guan Y. Machine learning for cancer drug combination. Clin Pharmacol Ther. 2020;107:749–52.
Funding
This work is supported by NSF#1452656 and NIH R35-GM133346.
Author information
Authors and Affiliations
Corresponding author
Additional information
Guest Editors: Lawrence Yu, Hao Zhu and Qi Liu
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, Z., Li, H., Carpenter, C. et al. Challenge-Enabled Machine Learning to Drug-Response Prediction. AAPS J 22, 106 (2020). https://doi.org/10.1208/s12248-020-00494-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-020-00494-5