Abstract
Drug-transporter interactions could impact renal drug clearance and should ideally be detected in early stages of drug development to avoid toxicity-related withdrawals in later stages. This requires reliable and robust assays for which current high-throughput screenings have, however, poor predictability. Kidney-on-a-chip platforms have the potential to improve predictability, but often lack compatibility with high-content detection platforms. Here, we combined conditionally immortalized proximal tubule epithelial cells overexpressing organic anion transporter 1 (ciPTEC-OAT1) with the microfluidic titer plate OrganoPlate to develop a screenings assay for renal drug-transporter interactions. In this platform, apical localization of F-actin and intracellular tight-junction protein zonula occludens-1 (ZO-1) indicated appropriate cell polarization. Gene expression levels of the drug transporters organic anion transporter 1 (OAT1; SLC22A6), organic cation transporter 2 (OCT2; SLC22A2), P-glycoprotein (P-gp; ABCB1), and multidrug resistance-associated protein 2 and 4 (MRP2/4; ABCC2/4) were similar levels to 2D static cultures. Functionality of the efflux transporters P-gp and MRP2/4 was studied as proof-of-concept for 3D assays using calcein-AM and 5-chloromethylfluorescein-diacetate (CMFDA), respectively. Confocal imaging demonstrated a 4.4 ± 0.2-fold increase in calcein accumulation upon P-gp inhibition using PSC833. For MRP2/4, a 3.0 ± 0.2-fold increased accumulation of glutathione-methylfluorescein (GS-MF) was observed upon inhibition with a combination of PSC833, MK571, and KO143. Semi-quantitative image processing methods for P-gp and MRP2/4 was demonstrated with corresponding Z′-factors of 0.1 ± 0.3 and 0.4 ± 0.1, respectively. In conclusion, we demonstrate a 3D microfluidic PTEC model valuable for screening of drug-transporter interactions that further allows multiplexing of endpoint read-outs for drug-transporter interactions and toxicity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Excretion of metabolic waste products and drugs from blood into the urine takes place in the kidney and is a result of glomerular filtration, active tubular secretion, and reabsorption. It has been estimated that about one third of the commonly used drugs and compounds tested in clinical trials are being (partially) removed via the urine without further metabolism (1,2). Renal clearance thus is a pivotal parameter to take into account during drug disposition studies in drug development.
Active tubular secretion of drugs in the kidney takes place mainly by the proximal tubule epithelial cells (PTEC) of the nephron and involves transporters (3). Uptake from blood to PTEC is mediated by members of the solute carrier (SLC) transporter family, such as organic cation transporter 2 (OCT2/SLC22A2) and organic anion transporter 1 and 3 (OAT1/SLC22A6; OAT3/SLC22A8). To complete elimination via urine from compounds internalized via SLC-mediated transport over the basolateral membrane, a range of drug transporters is expressed at the apical membrane of PTEC. This energy-demanding process is facilitated by transporters classified in the ATP-binding cassette (ABC)-transporter family, such as P-glycoprotein (P-gp/ABCB1), breast cancer resistance protein (BCRP/ABCG2), and multidrug resistance-associated proteins 2 and 4 (MRP2/ABCC2; MRP4/ABCC4). Other drug transporters present at the apical membrane are the multidrug and toxin extrusion proteins 1 and 2-k (MATE1/SLC47A1; MATE2-k/SLC47A2). In drug development, investigating interactions with a sub-set of drug transporters is recommended by the regulatory agencies, such as the US Food and Drug Administration (FDA) and the European Medicines Agency (EMA) (4,5). Clinical studies have shown drug-transporter interactions with, e.g., digoxin, ritonavir, or valspodar (PSC833), and with tenofovir via P-gp and MRP2/4, respectively (6,7,8).
In the past decade, multiple renal immortalized cell lines have been established, allowing in vitro screening of nephrotoxicity and renal drug transporters. Conditionally immortalized PTEC (ciPTEC) are derived from a healthy volunteer and manifest a stable expression and functionality of OCT2, P-gp, and MRP4 (9). Further transduction resulted in stable expression and functionality of OAT1 or OAT3 (10). Multiple studies have shown the potential of ciPTEC for studying drug transport and drug-induced toxicity cultured under conventional static conditions, often referred to as two-dimensional (2D) cell culture (10,11,12,13). In addition, transepithelial barrier function, polarization, and transport of cations and anions have been demonstrated in ciPTEC cultured in a three-dimensional (3D) model using a tailor-made hollow fiber membrane system exposed to flow (13,14). Benefits of 3D microfluidic models incorporating fluid shear stress (FSS) and cell-extracellular matrix (ECM) interactions are often referred to as organ-on-a-chip or microphysiological systems. Using PTEC, kidney-on-a-chip platforms have been demonstrated to improve proximal tubule characteristics, such as increased expression of tight-junction protein zonula occludens-1 (ZO-1) and increased number of ciliated cells (15). Kidney-on-a-chip platforms could therefore advance predictability of large in vitro screenings, also referred to as high-throughput screening (HTS), which are typically employed during early phases in drug development.
The current study focuses on implementing a kidney-on-a-chip platform for the determination of drug-transporter interactions during drug development. To this end, we culture human ciPTEC-OAT1 in a microfluidic titer plate, the OrganoPlate, that enables 3D culture under FSS. The OrganoPlate consists of 40 or 96 chips and is based upon a 384-well microtiter plate format (16,17). Flow is induced by passive leveling by gravity of medium in medium-channel inlets and outlets of chips, creating a bi-directional flow. Using this platform, we assessed stability in drug transporter expression and functionality via fluorescence-based 3D drug efflux assays to evaluate P-gp and MRP2/4 transporter activities. In addition, we set up a user-friendly data analysis method using open-source software compatible with HTS. This approach resulted in a 3D microfluidic PTEC model suitable for HTS of drug-transporter interactions.
MATERIALS AND METHODS
Cell Culture
CiPTEC-OAT1 were obtained as described previously (9,10). Briefly, cells were retrieved from urine from a healthy volunteer in compliance with the guidelines of the Radboud Institutional Review Board and conditionally immortalized via transduction with the temperature-sensitive mutant of SV large T antigen (SV40T) and human telomerase reverse transcriptase (hTERT) (9). Transduction of OAT1 in ciPTEC was performed using lentiviral particles containing genes encoding for human OAT1 (10).
Cells were sub-cultured at a dilution of 1:2 to 1:6 at 33°C and 5% (v/v) CO2 to allow proliferation and were used from passage numbers 48 to 60 for experiments. Cells were cultured in a 1:1 mixture of Dulbecco’s modified Eagle’s medium (DMEM) and Ham’s F-12 nutrient mixture without phenol red (DMEM HAM’s F12, Life Technologies, Paisley, UK) and supplemented with insulin (5 μg/ml, Sigma-Aldrich, Zwijndrecht, the Netherlands); transferrin (5 μg/ml, Sigma-Aldrich); selenium (5 ng/ml, Sigma-Aldrich); hydrocortisone (36 ng/ml, Sigma-Aldrich); epidermal growth factor (10 ng/ml, Sigma-Aldrich); tri-iodothyrine (40 pg/ml, Sigma-Aldrich); 1% (v/v) penicillin/streptomycin (Life Technologies); and 10% (v/v) fetal calf serum (FCS, Greiner Bio One, Kremsmuenster, Austria), further referred to as PTEC complete medium.
For 2D static cell culture, cells were seeded at 55,000 cells/cm2 in 6-well plates and cultured for 1 day at 33°C 5% (v/v) CO2 without penicillin/streptomycin to allow proliferation. Then, cells were cultured for 7 days at 37°C 5% (v/v) CO2 to induce maturation into confluent cell monolayers. Cells were harvested on day 8 after seeding.
For 3D FSS cell culture, a 400 μm two-lane OrganoPlate (Mimetas, Leiden, the Netherlands) was used for culturing ciPTEC-OAT1 in a microfluidic platform. Prior to seeding, 50 μl of Hank’s balanced salt solution (HBSS, Life Technologies) was added to the observation window (Fig. 1a). An ECM mix containing collagen I (4 mg/ml, AMSbio Cultrex 3D Collagen I Rat Tail); sodium bicarbonate (3.7 mg/ml, Sigma-Aldrich); and HEPES (100 mM, Life Technologies) was prepared and 2 μl was added to the gel channel (17). Then, the OrganoPlate was placed at 37°C 5% (v/v) CO2 for 30 min to allow polymerization of the ECM. Proliferating cells were harvested from a culture flask using accutase (Sigma-Aldrich), centrifuged (300×g, 5 min) and then resuspended in PTEC complete medium. Per chip, 40,000 cells in a suspension of 2 μl were seeded in the medium-channel inlet (Fig. 1a, b). In chips used for background correction of fluorescent signal, 2 μl of medium was used. The OrganoPlate was then placed vertically at 33°C 5% (v/v) CO2 for 30 min with the gel channel facing down to allow cells to attach to the ECM (Fig. 1b). Next, 30 μl of PTEC complete medium was added to the medium channel inlet. The OrganoPlate was oriented horizontally again at 33°C 5% (v/v) CO2 for 4 to 6 h. Upon addition of another 30-μl PTEC complete medium to the medium channel inlet, 60-μl PTEC complete medium was added to the medium channel outlet. CiPTEC-OAT1 in the OrganoPlate was cultured at 33°C 5% (v/v) CO2 to allow proliferation without any FSS for 3 days. Medium was supplemented with 1% (v/v) penicillin/streptomycin when culturing at 33°C.
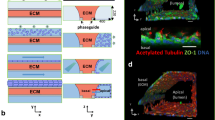
Overview of two-lane OrganoPlate and cell culture, assay and image analysis protocol. a Schematic overview of a two-lane OrganoPlate consisting of a gel channel (red) and medium channel (blue) and covering four wells of a 384-wells plate. Four wells are named 1 = gel channel inlet, 2 = medium channel inlet, 3 = observation window. and 4 = medium channel outlet. Dotted line indicates position of cross-sectional overview (b). b Cross-sectional overview of culture protocol of ciPTEC-OAT1 in the OrganoPlate: 1 = addition of ECM to the gel channel creating two membrane-free compartments because of the phaseguide; 2 = seeding of ciPTEC-OAT1; 3 = by placing the plate vertically, cells are allowed to attach to ECM by gravity; 4 = ciPTEC-OAT1 is allowed to mature under FSS; 5 = two compartments are created per chip consisting of an extracellular matrix (ECM) and an apical channel (medium channel). c Culture conditions and assay details of 3D P-gp, MRP2/4 drug efflux assay specified. Image analysis in Fiji contained background correction, image QC (visual observation) and normalization. Z stacks per chip were acquired using 10-μm intervals between each Z stack resulting in a total number of five Z stacks per chip. Cells on ECM and medium interface were selected for further analysis and calculation of the average intensity of Z stacks per chip
At day 3 after seeding, medium channels were refreshed with PTEC complete medium without penicillin/streptomycin and the OrganoPlate was placed on an interval rocker platform (angled at − 7° or + 7° from base, 8-min interval) at 37°C 5% (v/v) CO2 to allow the cells to maturate under FSS induced by a bi-directional flow (Fig. 1b, c). Experiments in the OrganoPlate were performed on day 8 after seeding, unless stated otherwise. Medium was refreshed every 2 to 3 days.
For long-term cell culture of ciPTEC-OAT1 in the OrganoPlate, chips were pre-coated (30 min, 37°C) prior to seeding with L-3,4-dihydroxydiphenylalanine (L-DOPA, 2 mg/ml in 10 mM Tris pH 8.5, Sigma-Aldrich) as earlier described (18). Cells were then cultured at 33°C on an interval rocker, as above described, for 20 days.
Immunofluorescence staining of ciPTEC-OAT1 in the OrganoPlate
At days 8 and 20 after seeding, PTEC complete medium was aspirated from the medium channel and cells were fixated with 3.7% (w/v) formaldehyde (Sigma-Aldrich) in HBSS. The plate was incubated for 15 min at room temperature. For all described incubation and wash steps, 50 μl was added to the inlet and outlet of the medium channel, and plate was placed at angle to allow perfusion, unless stated otherwise. Fixative was aspirated and medium channels were washed twice with washing solution (4% (v/v) FCS in HBSS). Then, cells were permeabilized by adding 0.3% (v/v) Triton-X100 (Sigma-Aldrich) in HBSS and incubated for 10 min at room temperature. Cells were subsequently washed with washing solution and then incubated with blocking solution (2% (v/v) FCS, 2% (w/v) bovine serum albumin (BSA, Sigma-Aldrich), 0.1% (v/v) Tween-20 (Sigma-Aldrich) in HBSS) for 45 min at room temperature. Primary and secondary antibodies were diluted in blocking solution and perfusion during incubation was allowed by adding 30 μl to the medium channel inlet and 10 μl to the outlet. Primary antibodies against zonula occludens-1 (ZO-1, 1:125, rabbit, Thermo Fisher), a tight-junction protein; acetylated-tubulin (clone 6-11B-1, 1:1000, mouse, Sigma-Aldrich), a constituent of the primary cilium; and pericentrin (1:500, rabbit, Abcam), a centrosome marker, were used and incubated for 60 min at room temperature. Next, cells were washed three times with washing solution and then incubated with secondary antibodies Alexa Fluor® 488 goat anti-rabbit (1:250, Life Technologies or Abcam), Alexa Fluor® 488 goat anti-mouse (1:250, Life Technologies), and Alexa Fluor® 647 goat anti-rabbit (1:250, Life Technologies) for 30 min at room temperature. Cells were then washed three times, followed by addition of probes in the last wash step to stain F-actin and nuclei with ActinRed 555 (two drops per milliliter in HBSS, Life Technologies) and DRAQ5 (1:1000 in HBSS, Abcam) or DAPI (300 nM in HBSS, Life Technologies), respectively. Fluorescent images of the ZO-1 staining were acquired with a Leica TCS SP5 laser point confocal microscope (Leica, Wetzlar, Germany) or a Zeiss LSM880 confocal scanning microscope (Carl Zeiss, Jena, Germany) using a ×20 objective. Z stacks were acquired using a 2 μm interval over a 200–220 μm range. To investigate polarization of the PTEC monolayer of ciPTEC-OAT1 in the OrganoPlate, fluorescent images were acquired of acetylated-tubulin and pericentrin staining with a Zeiss LSM880 confocal scanning microscope using a ×40 water immersion objective. A range of 45 μm was imaged using a 0.88-μm interval between Z stacks. Images were adjusted for windows and level, and then 3D images were reconstructed in Fiji (version 1.51n) (19).
Gene Expression of ciPTEC-OAT1
Cells were harvested from 6-well plates using Accutase, and then RNA was extracted using the RNeasy Mini kit (QIAGEN, Venlo, the Netherlands). RLT lysis buffer was added to the cell pellet and RNA was then extracted according to the protocol provided by the manufacturer.
RNA from cells in the OrganoPlate was isolated using the RNeasy Micro kit (QIAGEN). From each chip, RNA was isolated by adding 50 and 25 μl RLT lysis buffer to the medium channel inlet and outlet, respectively. Cell lysates of two chips were then pooled and RLT lysis buffer was added up to a volume of 350 μl. The extracted RNA was purified following the manufacturer’s protocol.
M-MLV reverse transcriptase (Promega, Madison, USA) was used to synthesize complementary DNA (cDNA) according the manufacturer’s protocol. Human kidney cortex material was used as reference. Tissues unsuitable for transplantation were obtained for research purposes from three donors, who gave permission and which was registered in the Dutch Donor registry. RNA isolation and cDNA synthesis were performed as previously described (20). The expression levels of mRNA were quantified using gene-specific primer-probe sets from Applied Biosystems (Bleiswijk, the Netherlands): GAPDH (hs99999905_m1), HPRT1 (hs02800695_m1), SLC22A6 (hs00537914_m1), ABCB (hs01067802_m1), ABCC2 (hs00166123_m1), ABCC4 (hs00195260_m1), SLC22A2 (hs01010723_m1), and TaqMan Universal polymerase chain reaction (PCR) Master Mix. Quantitative PCR reactions were carried out using CFX96-Touch Real-Time PCR System (BioRad, Veenendaal, the Netherlands) and analyzed with BioRad CFX Manager (version 3.1). Expression levels of GAPDH were used as reference gene.
3D Drug Efflux Assays in the OrganoPlate
Functionality of P-gp and MRP2/4 was assessed based upon accumulation of substrates, as previously described for ciPTEC in 2D (Fig. 1c) (12). Prior to measuring efflux activity of P-gp and MRP2/4 in the OrganoPlate, medium-channel inlet and outlets were washed with 50 μl KHH at 37°C. Medium channels were then perfused two times. Then, cells were exposed to substrates with or without inhibitors or corresponding vehicle for 1 h at 37°C.
For P-gp activity assessments, calcein-AM (2 μM, Life Technologies) was used as substrate and PSC833 (10 μM, Tocris, Bristol, UK) was used as inhibitor. Calcein-AM and PSC833 were dissolved in DMSO (Sigma-Aldrich). All dilutions were prepared in Krebs-Henseleit buffer (Sigma-Aldrich) supplemented with HEPES (10 mM, Sigma-Aldrich) and adjusted to pH 7.4 (KHH).
Chloromethylfluorescein-diacetate (CMFDA, 1.25 μM, Molecular Probes) was used to study MRP2/4 activity. A combination of PSC833 (10 μM); KO143 (10 μM, Sigma-Aldrich); and MK571 (10 μM, Sigma-Aldrich) was used to inhibit efflux of CMFDA and glutathione-methylfluorescein (GS-MF) via P-gp, BCRP, and MRP2/4, respectively. CMFDA and KO143 were dissolved in DMSO, while MK571 was dissolved in Milli-Q water, requiring appropriate vehicle controls. All dilutions were prepared in KHH.
After substrate incubations, cell nuclei were stained with Hoechst33342 by first washing with KHH, followed by staining with Hoechst33342 (10 μg/ml, Molecular Probes) for 10 min at room temperature. Cells were co-incubated with PSC833 (10 μM), KO143 (10 μM), and MK571 (10 μM) during staining with Hoechst33342 to inhibit all efflux transport processes. Subsequently, cells were washed and efflux of calcein-AM and GS-MF was arrested by adding ice-cold KHH to medium channel. FSS in medium channel during experiment and washing was assured by adding 80 μl to the medium channel inlet and 20 μl to the medium channel outlet.
Fluorescent Imaging in the OrganoPlate
Fluorescence of intracellular accumulation of calcein or GS-MF and Hoechst33342 in ciPTEC-OAT1 cultured in the OrganoPlate were imaged in situ using a Becton Dickinson (BD) Pathway 855 high-throughput microscope (BD Bioscience, Breda, The Netherlands) using a ×10 objective (Fig. 1c). The BD Pathway 855 microscope was compatible with the two-lane OrganoPlate and allowed to acquire images of the observation window of each chip using two different fluorescent dyes with wild-field microscopy and spinning disk confocal microscopy at room temperature. A macro was developed to acquire fluorescent and bright-field images using BD AttoVision (version 1.6, BD Bioscience). Z height was determined per experiment by determining the top and bottom of multiple chips in the OrganoPlate visually. The average height was used as starting point for acquiring five Z planes with a 10 μm interval, covering 50 μm in Z direction, generating focused images of cells cultured against ECM. First, calcein and GS-MF fluorescence images were acquired (488-nm excitation and 520-nm emission filters) with exposure time and gain set at 0.5 s and 100, respectively. Hoechst33342 images were acquired subsequently (360-nm excitation filter and 435-nm emission filter), with exposure time and gain set at 0.01 s and 100, respectively. Afterwards, bright-field images of each chip were acquired using an exposure time of 0.02 s.
Image Processing
An image analysis method (Fig. 1c) was set up to analyze the full data set using custom made macros in Fiji. Initially, the region of interest (ROI) was selected from bright-field images in which the gel channel, medium channel and the phase guide were clearly visible, by focusing at the interface of ECM and medium. The same area size was used for all images. All raw fluorescence images (uncompressed 12-bit TIFF) were cropped based upon the interface selection and converted from a pixel to micrometer scale. Then, the raw fluorescence images were corrected for background noise by subtracting images of chips with ECM without any cells exposed to either calcein-AM or CMFDA. Background-corrected images were saved to an uncompressed 16-bit TIFF file and the average intensity was calculated (see macro in Supplementary Material). In parallel, a mask was created for each image using 17 threshold algorithms available in Fiji (Table S1 in Supplementary Material) to automatically determine localization of intracellular accumulated fluorescent calcein or GS-MF. Upon running a median image filter (radius size five pixels), a cell selection mask was created via the selected algorithm to create a binary image. Pixels from the edges of the cell selection mask in the binary image were removed using “erode” in Fiji. The binary image represented a cell selection and was used to analyze fluorescence intensity of each Z stack per chip.
As part of the image quality control (QC), only in-focus images of cells cultured at the ECM and medium interface were selected and reviewed by three independent observers (Fig. 1c). An image was defined as out-of-focus if no in-focus cell in the ROI were scored by at least two of the three observers.
Manual interventions, such as determination of out-of-focus images are not compatible with HTS. Therefore, selected image QC metrics were established to automatically exclude out-of-focus images: focus score, image correlation, and power log-log slope (PLLS), which all have been described in literature for image analysis (21,22,23,24). An image QC method was set up using these image QC metrics in CellProfiler (version 2.2.0) (25,26). For the focus score, the deviation of pixel intensity was assessed upon normalization per image. The correlation of the grayscale value of neighboring pixel was evaluated for image correlation. Pixel size for image correlation and focus score was empirically based upon the area of the observed intracellular accumulation of calcein or GS-MF and set at 20, 50, 100, 200, and 500 pixels. For the PLLS, the slope of power spectral density versus the spatial frequency, both on a log scale, was plotted. All images were automatically binned based upon the image QC metrics in GraphPad Prism (version 5.03). Multiple cutoff values were compared per image QC metric. The efficiency of the automated image QC metrics was compared to visually reviewing of out-of-focus scoring as described earlier, which was defined as ground truth. For all image QC metrics, the precision and recall rates were determined, as well as the harmonic mean of the precision and recall rates, also known as F score. Precision, recall, and F score were calculated as follows:
where precision is:
and recall is:
Average intensities of visual observation in-focus images were calculated per chip (Fig. 1c). Two to eight chips were analyzed per condition and repeated in at least three independent experiments. All groups were checked for outliers using Grubbs’ test for outliers (using a significance level of α = 0.05) and finally normalized to average fluorescence intensity in vehicle condition per experiment.
The Z′ factor was used to index the thresholding algorithms and overall intensity per image and condition to assess quality and optimize the data analysis methods. The Z′-factor has been described as statistical parameter in HTS assays taking in account the variation between a signal in maximum response, the positive control, and minimum response, the negative control (27). Hence, the Z′ factor can be used as guideline to optimize semi-quantitative methods in order to improve discrimination between the positive and negative controls. Mean and standard deviation (SD) were determined for the positive control, inhibitor condition, and negative control, vehicle condition, to calculate the Z′ factor per experiment as follows:
Statistical Data Analysis
All statistics were performed using GraphPad Prism. Data is presented as mean ± SEM of at least three independent experiments (n = 3) and considered to be significantly different if p < 0.05 using an unpaired two-tailed Student’s t test, unless stated otherwise.
RESULTS
PTEC’s Formed a 3D Tubular Structure in the OrganoPlate
Upon seeding ciPTEC-OAT1 against the ECM in the two-lane OrganoPlate, cells formed a monolayer that migrated towards the top and bottom of the medium channel, creating a tubule-like structure at day 8 after seeding (Fig. 2a, Fig. S1). PTEC tubules could be cultured up to 20 days in the OrganoPlate (Fig. S2). ZO-1, a tight-junction protein, was localized between the cells in the monolayer of the PTEC tubules (Fig. 2b), indicative for polarization. Staining of the centrosome, or pericentrin, and acetylated-tubulin, showed apical expression of single cilia per cell (Fig. 2c; Fig. S3). Some migration of ciPTEC-OAT1 from the medium channel into the ECM was observed, likely influencing the monolayer integrity. Although the majority of cilia were mainly expressed at the apical side of the tubules, some cilia were observed on single cells that migrated into the ECM (Fig. S3).
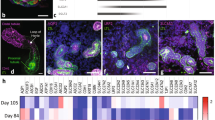
Immunofluorescence staining of ciPTEC-OAT1 in OrganoPlate. a Three-dimensional (3D) view of ciPTEC-OAT1 in OrganoPlate with F-actin (red) and nuclei (DAPI, blue) showing the formation of tubule structures after 8 days of seeding. The phase guide is visible as blue bar at the left side of the image. The lumen at the apical side and the extracellular matrix (ECM) are indicated. Scale bar represents 100 μm. b Staining of nuclei (DAPI, blue) and zonula occludens (ZO-1, green). Expression of ZO-1 visible between cell monolayer on top of PTEC tubules. Scale bar represents 100 μm. c Representative images of single cells attached to the extracellular matrix (ECM) selected from the tubular ciPTECOAT1 structures in the OrganoPlate, showing staining of nuclei (DAPI, blue) and primary cilia with pericentrin (magenta) and acetylated tubulin (green). Most cilia face towards the lumen (apical side), demonstrating polarization of the tubules. Scale bar represents 20 μm
Rotation of 360° of tubules at day 8 after seeding. Immunofluorescence staining with F-actin (red) and nuclei (DAPI, blue) in ciPTEC-OAT1 in the OrganoPlate. (MP4 18077 kb)
Cilia staining in ciPTEC-OAT1 demonstrates polarized tubular structures. Immunofluorescence staining of nuclei (DAPI, blue) and cilia by means of staining pericentrin (magenta) and acetylated-tubulin (green) in ciPTEC-OAT1 in the OrganoPlate at day 8 after seeding. Z stacks displayed as volume and 360° rotation of cell monolayer attached to the extracellular matrix (ECM). Rotation starts viewing from lumen (apical side) and rotates to viewing point from ECM (basolateral side) and back with the viewing point from lumen. Clearly visible is that cilia were mainly demonstrated on apical side, facing the lumen of the tubules. Cilia are also observed on few cells that migrated into the ECM. (MP4 10989 kb)
Drug Transporter Gene Expression in ciPTEC-OAT1 Cultured in the OrganoPlate
Drug transporter gene expression levels in ciPEC-OAT1 cultured in 2D static (6-well plate) and under FSS (OrganoPlate) were compared (Fig. 3). Expression levels of SLC transporters OAT1 and OCT2 and ABC transporters P-gp and MRP2/4 were found to be similar in ciPTEC-OAT1 cultured in the OrganoPlate as compared to 2D cultures. This indicates a maintained phenotype of ciPTEC-OAT1 when cultured in the OrganoPlate. Only expression levels of HPRT1, here assessed to be used as a reference gene, were lower in ciPTEC-OAT1 cultured in the OrganoPlate and, consequently, not valid as reference gene. On the other hand, reference gene GAPDH was stably expressed in ciPTEC-OAT1 cultured both in 6-well plate and in the OrganoPlate® with corresponding Ct values of 19.7 ± 0.1 and 19.6 ± 0.1, respectively. It is important to note that drug transporter gene expression levels presented here were derived from cells both located in the perfused channels, as well as the medium channel inlet and outlets. As the latter cells are not exposed to laminar flow, nor were cultured in 3D, gene expression profiling was based on a hybrid population and should be regarded as qualitative measurement.
The HPRT1, OAT1, OCT2, P-gp, MRP2, and MRP4 expression levels in matured ciPTEC-OAT1 cultured under FSS (OrganoPlate). Gene expression levels were normalized to GAPDH, used as reference gene, and here calculated a fold difference compared to gene expression levels in 2D static cell culture (6-well plate). **p < 0.01 compared to mRNA expression in ciPTEC-OAT1 cultured in a 6-well plate
In addition, expression levels of ciPTEC-OAT1 cultured in 6-well plate or in the OrganoPlate® were compared to expression levels in human kidney cortex samples (Table I). In human kidney cortex, OAT1 is most abundantly expressed relative to expression levels of other drug transporters assessed here (rank order OAT1 > MRP2 > MRP4 > OCT2 > P-gp). Expression of OAT1 was also abundantly expressed in ciPTEC-OAT1 when cultured either in a 6-well plate or in the OrganoPlate, although a different relative expression was observed (rank order OAT1 > MRP4 > P-gp > MRP2 > OCT2).
Fluorescence-based Drug Efflux Assays in 3D Tubule Structures
We set up 3D efflux assays for P-gp and MRP2/4 as proof-of-concept to demonstrate transport activity in 3D microfluidic PTEC model. The first step in our image analysis included the selection of the ROI per chip, which included cells at the ECM and medium interface. As shown in Fig. 4a, the gel channel, medium channel, and the phase guide were clearly visible. The bright-field images were used to objectively choose the ROI per chip. Cell layers formed at the ECM were demonstrated by the Hoechst staining (Fig. 4a).
Representative bright-field and fluorescent images of Hoechst and calcein or GS-MF. A cell monolayer is formed at the phase guide is clearly visible in the center of the bright-field images. The ECM and medium channel are positioned above and below the phase guide, respectively. Hoechst staining was used to improve cell monolayer visibility. a Accumulation of calcein was visible in control, vehicle, and PSC833-treated tubules. Accumulation of calcein was increased upon PSC833-mediated inhibition of P-gp. b Intracellular accumulation of GS-MF in control, vehicle, treated, and in presence of inhibitor cocktail (consisting of PSC833, KO143 and MK571). Accumulation of GS-MF was increased upon incubation with the cocktail. For visualization, gray values for bright-field, Hoechst, calcein, and GS-MF images were set from 250 to 1000, 0 to 500, 0 to 2000, and 0 to 500, respectively. Gray scale of calcein and GS-MF fluorescent images were converted to look-up table (LUT) color fire to enhance contrast. Calibration bar of LUT shows level of intensity of calcein or GS-MF. Scale bar in bright-field images represents 100 μm. See Fig. 5 for quantification of the signals
P-gp shows the highest affinity for uncharged and positively charged compounds (28,29). Calcein-AM was used as substrate to study P-gp efflux activity in ciPTEC-OAT1 cultured in the OrganoPlate. Calcein-AM can freely diffuse through the cell membrane and is a substrate with high affinity for P-gp. Intracellular, calcein-AM is metabolized through esterase activity into the fluorescent calcein. Slight accumulation of calcein was observed in the control and vehicle incubated tubules upon incubation with calcein-AM (Fig. 4a), which was strongly enhanced upon co-incubation with PSC833, suggesting inhibition of P-gp (Figs. 4a and 5a).
Average intensity measured per condition and chip is shown for control, solvent-treated and inhibitor-treated conditions. a Fold difference in calcein or in b GS-MF fluorescence. Values were normalized to vehicle per experiment. No threshold algorithm was used. Significant differences of the means were compared to vehicle using one-way ANOVA followed by Dunnett’s multi comparison post hoc test. ***p < 0.001 compared to vehicle
MRP2/4 show highest affinity for negatively charged compounds in the kidney (30,31,32,33). To study MRP2/4 activity, 5-chloromethylfluorescein-diacetate (CMFDA), which enters the cell via diffusion, was used. CMFDA is then metabolized to the fluorescent 5-chloromethylfluorescein (CMF) via esterase activity and then further metabolized into carboxylfluorescein-glutathione (GS-MF) through direct interaction with glutathione (GSH) or via transferase activity. CMF is completely transferred into GS-MF during incubation time used here (12). GS-MF is predominantly a substrate for MRP2/4, although some selectivity was demonstrated for BCRP and P-gp (12). We therefore used a combination of inhibitors, as described in the “Materials and Methods” section. In the control and vehicle tubules, minor accumulation of GS-MF was visible. After co-incubation with an inhibitor cocktail of PSC833, KO143, and MK571, a higher fluorescence intensity was demonstrated as compared to vehicle control, indicating active MRP2/4 in ciPTEC-OAT1 cultured in the OrganoPlate. Signal intensity was stable for at least 25 min for both calcein and GS-MF (data not shown).
Image QC and Image Analysis
Setting up a proper image QC could reduce the variation caused by factors such as out-of-focus images, signal-to-noise ratio, and cell density. In-focus images of the cell layer between ECM and medium interfaces were acquired in the majority of chips. Available image QC metrics in CellProfiler were applied to obtain an automated and fast image QC of our data set.
Each image was visually evaluated to be in-focus or out-of-focus. Of all images acquired for the 3D P-gp drug efflux assay (n = 245, from four different experiments), 14.3% (n = 35) were assigned to be out-of-focus. For the 3D MRP2/4 drug efflux assay, 21.7% (n = 52) of all images (n = 240, from four different experiments) were assigned to be out-of-focus. The visually evaluated in-focus and out-of-focus images were defined as ground truth.
Means of image QC metrics of in-focus and out-of-focus images were compared for the two transporter assays as defined for P-gp (Table S2 in Supplementary Material) and MRP2/4 (Table S3 in Supplementary Material). By calculating the F score per image QC metric, the false negatives, true and false positives were incorporated and compared with the ground truth. Focus score showed an F score of 0.46, the highest F score of all image QC metrics determined for the 3D P-gp drug efflux assay, the recall and precision score obtained were 0.33 and 0.77, respectively (Table S2 in Supplementary Material). For the 3D MRP2/4 drug efflux assay, using focus score with a pixel size 200 resulted in an F score of 0.53, the highest F score for this assay, with corresponding precision and recall scores of 0.36 and 0.94, respectively (Table S3 in Supplementary Material). For both assays, low precision scores were obtained, because of a high number of false positives compared to true positives, leading to low F scores.
To demonstrate proof-of-concept, further analysis was based on the visually determined out-of-focus images by three observers, since the image QC metrics determined here were less optimal to distinguish in-focus and out-of-focus images for our data set. Visually reviewed out-of-focus images were excluded from further calculations from this point.
Fluorescence Intensity in Image and Average per Chip
Fluorescence intensities per image were determined in Fiji to perform semi-quantitative analysis of P-gp and MRP2/4 drug efflux activities (Tables II and III). Masking using thresholding algorithms was applied to create a selection area incorporating the intracellular accumulation of calcein or GS-MF and compared to the intensity of the entire image. A higher intensity after co-incubation with inhibitor(s) was observed for all semi-quantitative methods used in both drug efflux assays (Tables II and III). No differences between vehicle-treated and control were observed in the drug efflux assays.
Z′ factor was calculated per experiment to compare and optimize the semi-quantitative analysis methods. Thresholding algorithm and the intensity of the entire image were compared in Tables II and III. Z′ factors are only meaningful when they fall within a range of − 1 < Z′ ≤ 1, which is the case for the majority of the thresholding algorithms (27). Remarkably, when using algorithm threshold moments for the 3D P-gp drug efflux assay, the mean of the negative control (vehicle treatment) was higher than the mean of positive control (PSC833) treatment the in two experiments, resulting in Z′ > 1. Without using a thresholding algorithm, this resulted in Z′ factor values of 0.1 ± 0.3 and 0.4 ± 0.1 for the 3D P-gp and MRP2/4 drug efflux assays, respectively. For all analysis methods, a significant increase in fluorescence intensity was found upon transport inhibition for both assays; however, thresholding algorithms resulted in large standard deviations. In general, Z′ factor values were < 0 when using a thresholding algorithm, because of an overlap in fluorescence intensity between the groups, indicating that image analysis using threshold algorithms as presented here is not suitable for HTS (27). Thresholding algorithms included more steps in the semi-quantitative analysis macro in Fiji (see Supplementary Material), which increased the processing time per image, and Z′ factors were lower compared to the intensity of the entire image. Therefore, measuring intensity in the entire image showed the most reliable results.
Upon establishing a valid semi-quantitative image analysis method for both assays, the average fluorescence intensities per chip were calculated, as shown in Fig. 5. PSC833-mediated inhibition of P-gp resulted in a 4.4 ± 0.2-fold increase fluorescence intensity compared to vehicle treated tubules (Fig. 5a). For the 3D MRP2/4 drug efflux assay, in tubules co-incubated with PSC833, KO143, and MK571 led to a 3.0 ± 0.2-fold increased compared to vehicle (Fig. 5b). Both assays demonstrate function of P-gp and MRP2/4 and the possibility to semi-quantify renal drug interaction via increased fluorescence intensity.
DISCUSSION
In this study, we developed a 3D microfluidic PTEC model demonstrated to be suitable for the evaluation of renal drug interactions with efflux transporters. The system has high potential for early drug screening due to its medium to high-throughput features and compatibility with high-content imaging platforms, as shown in the current study on 3D efflux assays as well as demonstrated by Vormann et al.(under revision at AAPSJ) and Suter-Dick et al. (accepted by AAPSJ). We cultured the human conditionally immortalized cell line ciPTEC-OAT1 in the OrganoPlate exposed to FSS induced by leveling, creating a 3D microfluidic PTEC model. The OrganoPlate enabled the formation of polarized tubule-like structures in a 96-well format.
A polarized epithelium was obtained upon culturing of ciPTEC-OAT1 in the OrganoPlate and showed apical expression of cilia, in line with an earlier publication of a perfusable printed 3D PTEC (34). Furthermore, gene expression levels of OAT1 (SLC22A6), OCT2 (SLC22A2), P-gp (ABCB1), MRP2 (ABCC2), and MRP4 (ABCC4) were confirmed. Representative monolayers were formed and functional activity and drug interactions for P-gp and MRP2/4 was demonstrated by means of fluorescence-based transport assays. Open-source imaging software was used to analyze fluorescence to make this platform user-friendly and suitable for screening of larger data sets.
Basal PTEC characteristics were shown to be improved in kidney-on-a-chip platforms, likely because of inclusion of a physiological FSS and cell- ECM interaction (15,34,35). Interestingly, enhanced P-gp efflux activity and transepithelial transport of para-aminohippurate (PAH) was demonstrated (15,36). Microfluidic kidney models could advance lead optimization at early stages of drug development and are an emerging technology to study drug-transporter interactions and toxicity (37). Future efforts demonstrating the improvements of such microfluidic 3D PTEC models as compared to regular 2D static culture should include a validation study with a compound library focusing on sensitivity and specificity using nephrotoxic and non-nephrotoxic compounds. Complexity and lack of compatibility with HTS of current kidney-on-a-chip models are a major drawback as opposed to 2D static cell culture in a multiwell setting required for drug screening purposes. Most published kidney-on-a-chip models use complex pump-driven single chip systems that frustrate experiments relying on multiple test conditions (15,34,36,38,39,40). The 3D microfluidic PTEC model presented here consisted of 96 chips on one plate and physiological FSS that is induced by leveling, without the need for external components. Therefore, it has high potential for HTS settings and could relatively easily be implemented in current disposition studies in drug development.
In vitro toxicity screening at early phases of drug discovery benefits from its compatibility for HTS and low costs. Freshly isolated human primary PTEC are the golden standard for in vitro screenings when it comes to intact drug transporter functionality and resemblance to the human kidney. However, expression levels decrease within days after seeding and such studies are limited for their low reproducibility (41,42). Vesicles and over-expression of drug transporters in immortalized cell lines are powerful tools as well to evaluate such drug-transporter interactions, but often lack human PTEC characteristics that could link drug-transporter interactions to clinically relevant toxicological read-outs (43,44,45,46,47). Human-derived immortalized renal cell models such as human embryonic kidney 293 (HEK 293) and human kidney 2 (HK 2) cells have shown to be a valuable tool in screenings in drug development, but have low or no endogenous expression of relevant drug transporters (48,49). To overcome these limitations, we have developed ciPTEC-OAT1 with proven suitability for nephrotoxicity and drug-transporter interactions studies (10,11,12,13,14). The broad range of drug transporter expression in ciPTEC-OAT1 provides a strong basis for highly predictive renal drug interaction studies. The inter-complexity and overlap in substrates of renal drug transporters, rather than through one drug transporter, is incorporated because of this broad range of drug transporter expression. This resembles a more physiological, in vivo-like, mechanism of renal drug handling. Gene expression levels of drug transporters in ciPTEC-OAT1 assessed here were lower compared to levels observed in samples of the human renal cortex, in accordance with other renal cell lines (48). Functionality of OAT1, OCT2, BCRP, P-gp, and MRP2/4 in ciPTEC in 2D cell culture has been demonstrated in multiple studies (9,10,12,13,14).
Establishing highly predictive renal in vitro models for nephrotoxicity and renal drug-transporter interactions could refine and, ultimately, replace animal experiments. Besides the ethical issues, animal testing is expensive and not always translatable because of species differences. Following this study, future research should be directed to develop a method that allows in vitro-in vivo extrapolation of data, using experimental input from the 3D microfluidic PTEC model in accordance with Kunze et al. (50). Such extrapolation could demonstrate the added value and potential of our model to replace current animal studies now used for renal drug handling screenings.
Influx drug transporters should be evaluated in parallel to drug transporter efflux activity to obtain a reliable prediction on renal excretion. Uptake via OCT2 and OAT1/3 has been demonstrated in ciPTEC using uptake of fluorescent substrates (9,10). Using ciPTEC-OAT1, transepithelial transport of the uremic toxins indoxyl sulfate and kynurenic acid could be indirectly demonstrated when cells were cultured on hollow fiber membranes, thereby forming a renal epithelial tubule (13). Expanding the platform with a third lane that includes a perfusable basolateral compartment together with a perfusable apical compartment in a 3D microfluidic platform could be used to study transepithelial transport or organic cations and anion in detail, as shown by Vormann et al. (under revision at AAPSJ) Other potential applications for the three-compartmental system are the incorporation of other segments of the renal tubules or co-culture with vascular endothelial cells, as recently was demonstrated (40,51).
In the current study, fluorescent accumulation in the chips was evaluated from five Z stacks with a 10-μm interval per chip. This approach resulted in a variation of image focus per chip and per experiment. To reduce variability per stack, an image QC was introduced. We used a visually determination of in-focus and out-of-focus images. In our study, image QC metrics tested, such as PLLS, resulted in low precision and corresponding F scores because of overlapping means of image QC metric between in-focus and out-of-focus images assessed. Optimizing imaging in the OrganoPlate by using autofocus could reduce the number of out-of-focus images and improve image QC metric outcome. In addition, by acquiring more Z-stacks per chip, chances of obtaining in-focus images increase. A proper image QC is key in reliably screening larger data sets typical for HT and should therefore be optimized for each setting. Measuring fluorescence intensity in the entire image as compared to using thresholding algorithms provided the most optimal semi-quantification of efflux activity. Corresponding Z′ scores where within an acceptable range (0.5 > Z > 0) where standard deviation bands of positive and negative controls were separated (27). Aforementioned improvements in confocal imaging could further increase separation of standard deviations from controls.
CONCLUSION
To conclude, we present a 3D microfluidic PTEC model including most important renal drug transporters in a platform consisting of 96 chips. Multiplexing of different endpoint read-outs in this platform and testing multiple conditions in one plate allows this platform to be suitable for medium to high-throughput screenings and compatible with most high-content imaging platforms.
References
Morrissey KM, Stocker SL, Wittwer MB, Xu L, Giacomini KM. Renal transporters in drug development. Annu Rev Pharmacol Toxicol. 2013;53:503–29.
Varma MV, Feng B, Obach RS, Troutman MD, Chupka J, Miller HR, et al. Physicochemical determinants of human renal clearance. J Med Chem 2009;52(15):4844–4852.
Ivanyuk A, Livio F, Biollaz J, Buclin T. Renal drug transporters and drug interactions. Clin Pharmacokinet. 2017;56(8):825–92.
EMA. Guideline on the investigation of drug interactions (CPMP/EWP/560/95/Rev. 1). EMA; 2012.
FDA In vitro metabolism- and transporter-mediated drug-drug interaction studies. Guidance for industry (draft). FDA; 2017. p. 13–19.
Ding R, Tayrouz Y, Riedel KD, Burhenne J, Weiss J, Mikus G, et al. Substantial pharmacokinetic interaction between digoxin and ritonavir in healthy volunteers. Clin Pharmacol Ther. 2004;76(1):73–84.
Kovarik JM, Rigaudy L, Guerret M, Gerbeau C, Rost KL. Longitudinal assessment of a P-glycoprotein-mediated drug interaction of valspodar on digoxin. Clin Pharmacol Ther. 1999;66(4):391–400.
Likanonsakul S, Suntisuklappon B, Nitiyanontakij R, Prasithsirikul W, Nakayama EE, Shioda T, et al. A single-nucleotide polymorphism in ABCC4 is associated with tenofovir-related beta2-microglobulinuria in Thai patients with HIV-1 infection. PLoS One. 2016;11(1):e0147724.
Wilmer MJ, Saleem MA, Masereeuw R, Ni L, van der Velden TJ, Russel FG, et al. Novel conditionally immortalized human proximal tubule cell line expressing functional influx and efflux transporters. Cell Tissue Res 2010;339(2):449–457.
Nieskens TT, Peters JG, Schreurs MJ, Smits N, Woestenenk R, Jansen K, et al. A human renal proximal tubule cell line with stable organic anion transporter 1 and 3 expression predictive for antiviral-induced toxicity. AAPS J 2016;18(2):465–475.
Caetano-Pinto P, Jamalpoor A, Ham J, Goumenou A, Mommersteeg M, Pijnenburg D, et al. Cetuximab prevents methotrexate-induced cytotoxicity in vitro through epidermal growth factor dependent regulation of renal drug transporters. Mol Pharm. 2017;14(6):2147–57.
Caetano-Pinto P, Janssen MJ, Gijzen L, Verscheijden L, Wilmer MJ, Masereeuw R. Fluorescence-based transport assays revisited in a human renal proximal tubule cell line. Mol Pharm. 2016;13(3):933–44.
Jansen J, Fedecostante M, Wilmer MJ, Peters JG, Kreuser UM, van den Broek PH, et al. Bioengineered kidney tubules efficiently excrete uremic toxins. Sci Rep. 2016;6:26715.
Jansen J, De Napoli IE, Fedecostante M, Schophuizen CM, Chevtchik NV, Wilmer MJ, et al. Human proximal tubule epithelial cells cultured on hollow fibers: living membranes that actively transport organic cations. Sci Rep. 2015;5:16702.
Jang KJ, Mehr AP, Hamilton GA, McPartlin LA, Chung S, Suh KY, et al. Human kidney proximal tubule-on-a-chip for drug transport and nephrotoxicity assessment. Integr Biol. 2013;5(9):1119–29.
Trietsch SJ, Naumovska E, Kurek D, Setyawati MC, Vormann MK, Wilschut KJ, et al. Membrane-free culture and real-time barrier integrity assessment of perfused intestinal epithelium tubes. Nat Commun. 2017;8(1):262.
Wevers NR, van Vught R, Wilschut KJ, Nicolas A, Chiang C, Lanz HL, et al. High-throughput compound evaluation on 3D networks of neurons and glia in a microfluidic platform. Sci Rep. 2016;6:38856.
Schophuizen CM, De Napoli IE, Jansen J, Teixeira S, Wilmer MJ, Hoenderop JG, et al. Development of a living membrane comprising a functional human renal proximal tubule cell monolayer on polyethersulfone polymeric membrane. Acta Biomater. 2015;14:22–32.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9(7):676–82.
Tutakhel OA, Jelen S, Valdez-Flores M, Dimke H, Piersma SR, Jimenez CR, et al. Alternative splice variant of the thiazide-sensitive NaCl cotransporter: a novel player in renal salt handling. Am J Physiol Ren Physiol. 2016;310(3):F204–16.
Sun Y, Duthaler S, Nelson BJ. Autofocusing in computer microscopy: selecting the optimal focus algorithm. Microsc Res Tech. 2004;65(3):139–49.
Groen FCA, Young IT, Ligthart G. A comparison of different focus functions for use in autofocus algorithms. Cytometry. 1985;6(2):81–91.
Haralick RM. Statistical and structural approaches to texture. P Ieee. 1979;67(5):786–804.
Field DJ, Brady N. Visual sensitivity, blur and the sources of variability in the amplitude spectra of natural scenes. Vis Res. 1997;37(23):3367–83.
Carpenter AE, Jones TR, Lamprecht MR, Clarke C, Kang IH, Friman O, et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 2006;7(10):R100.
Kamentsky L, Jones TR, Fraser A, Bray MA, Logan DJ, Madden KL, et al. Improved structure, function and compatibility for CellProfiler: modular high-throughput image analysis software. Bioinformatics. 2011;27(8):1179–80.
Zhang JH, Chung TD, Oldenburg KR. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J Biomol Screen. 1999;4(2):67–73.
Aller SG, Yu J, Ward A, Weng Y, Chittaboina S, Zhuo R, et al. Structure of P-glycoprotein reveals a molecular basis for poly-specific drug binding. Science. 2009;323(5922):1718–22.
Cascorbi I. P-glycoprotein: tissue distribution, substrates, and functional consequences of genetic variations. Handb Exp Pharmacol. 2011;201:261–83.
Chen ZS, Tiwari AK. Multidrug resistance proteins (MRPs/ABCCs) in cancer chemotherapy and genetic diseases. FEBS J. 2011;278(18):3226–45.
Masereeuw R, Russel FG. Therapeutic implications of renal anionic drug transporters. Pharmacol Ther. 2010;126(2):200–16.
van Aubel RA, Smeets PH, Peters JG, Bindels RJ, Russel FG. The MRP4/ABCC4 gene encodes a novel apical organic anion transporter in human kidney proximal tubules: putative efflux pump for urinary cAMP and cGMP. J Am Soc Nephrol: JASN. 2002;13(3):595–603.
Smeets PH, van Aubel RA, Wouterse AC, van den Heuvel JJ, Russel FG. Contribution of multidrug resistance protein 2 (MRP2/ABCC2) to the renal excretion of p-aminohippurate (PAH) and identification of MRP4 (ABCC4) as a novel PAH transporter. J Am Soc Nephrol: JASN. 2004;15(11):2828–35.
Homan KA, Kolesky DB, Skylar-Scott MA, Herrmann J, Obuobi H, Moisan A, et al. Bioprinting of 3D convoluted renal proximal tubules on perfusable chips. Sci Rep. 2016;6:34845.
Secker PF, Luks L, Schlichenmaier N, Dietrich DR. RPTEC/TERT1 cells form highly differentiated tubules when cultured in a 3D matrix. ALTEX. 2017;
Weber EJ, Chapron A, Chapron BD, Voellinger JL, Lidberg KA, Yeung CK, et al. Development of a microphysiological model of human kidney proximal tubule function. Kidney Int. 2016;90(3):627–37.
Wilmer MJ, Ng CP, Lanz HL, Vulto P, Suter-Dick L, Masereeuw R. Kidney-on-a-chip technology for drug-induced nephrotoxicity screening. Trends Biotechnol. 2016;34(2):156–70.
Frohlich EM, Alonso JL, Borenstein JT, Zhang X, Arnaout MA, Charest JL. Topographically-patterned porous membranes in a microfluidic device as an in vitro model of renal reabsorptive barriers. Lab Chip. 2013;13(12):2311–9.
Gao X, Tanaka Y, Sugii Y, Mawatari K, Kitamori T. Basic structure and cell culture condition of a bioartificial renal tubule on chip towards a cell-based separation microdevice. Anal Sci. 2011;27(9):907–12.
Vedula EM, Alonso JL, Arnaout MA, Charest JL. A microfluidic renal proximal tubule with active reabsorptive function. PLoS One. 2017;12(10):e0184330.
Lash LH, Putt DA, Cai H. Membrane transport function in primary cultures of human proximal tubular cells. Toxicology. 2006;228(2–3):200–18.
Brown CD, Sayer R, Windass AS, Haslam IS, De Broe ME, D'Haese PC, et al. Characterisation of human tubular cell monolayers as a model of proximal tubular xenobiotic handling. Toxicol Appl Pharmacol. 2008;233(3):428–38.
Karlsson JE, Heddle C, Rozkov A, Rotticci-Mulder J, Tuvesson O, Hilgendorf C, et al. High-activity p-glycoprotein, multidrug resistance protein 2, and breast cancer resistance protein membrane vesicles prepared from transiently transfected human embryonic kidney 293-Epstein-Barr virus nuclear antigen cells. Drug Metab Dispos. 2010;38(4):705–14.
El-Sheikh AA, Greupink R, Wortelboer HM, van den Heuvel JJ, Schreurs M, Koenderink JB, et al. Interaction of immunosuppressive drugs with human organic anion transporter (OAT) 1 and OAT3, and multidrug resistance-associated protein (MRP) 2 and MRP4. Transgenic Res. 2013;162(6):398–409.
Wittgen HG, van den Heuvel JJ, van den Broek PH, Dinter-Heidorn H, Koenderink JB, Russel FG. Cannabinoid type 1 receptor antagonists modulate transport activity of multidrug resistance-associated proteins MRP1, MRP2, MRP3, and MRP4. Drug Metab Dispos. 2011;39(7):1294–302.
Fahrmayr C, Konig J, Auge D, Mieth M, Munch K, Segrestaa J, et al. Phase I and II metabolism and MRP2-mediated export of bosentan in a MDCKII-OATP1B1-CYP3A4-UGT1A1-MRP2 quadruple-transfected cell line. Br J Pharmacol 2013;169(1):21–33.
Gartzke D, Fricker G. Establishment of optimized MDCK cell lines for reliable efflux transport studies. J Pharm Sci. 2014;103(4):1298–304.
Jenkinson SE, Chung GW, van Loon E, Bakar NS, Dalzell AM, Brown CD. The limitations of renal epithelial cell line HK-2 as a model of drug transporter expression and function in the proximal tubule. Pflugers Arch. 2012;464(6):601–11.
Astashkina AI, Mann BK, Prestwich GD, Grainger DW. Comparing predictive drug nephrotoxicity biomarkers in kidney 3-D primary organoid culture and immortalized cell lines. Biomaterials. 2012;33(18):4712–21.
Kunze A, Huwyler J, Poller B, Gutmann H, Camenisch G. In vitro-in vivo extrapolation method to predict human renal clearance of drugs. J Pharm Sci. 2014;103(3):994–1001.
Musah S, Mammoto A, Ferrante TC, Jeanty SSF, Hirano-Kobayashi M, Mammoto T, et al. Mature induced-pluripotent-stem-cell-derived human podocytes reconstitute kidney glomerular-capillary-wall function on a chip. Nat Biomed Eng 2017;1.
Acknowledgements
The authors thank Johan van der Vlag (department Nephrology, Radboud University Medical Center, Nijmegen, the Netherlands) for supplying the human kidney cortex tissues. The authors thank Dr. Miriam Schmidts for kindly providing antibodies for acetylated-tubulin and pericentrin. The authors also thank Professor Dr. Hans Tanke from the department of Molecular Cell Biology at the Leiden University Medical Center (LUMC) and the microscopic imaging center at Radboud University Medical Center for using the imaging facilities. The authors thank Renata Škovroňová and Christiaan Stuut for their assistance with imaging processing.
Funding
This project was supported by the NephroTube project funded by the National Center for the Replacement, Refinement and Reduction of Animals in Research (NC3Rs), under the Crack-it challenge 15 (Nephrotube) project no. 37497-25920.
Author information
Authors and Affiliations
Contributions
JV, TTGN, MKV, FGMR, LSD, HLL, PV, RM, and MW participated in research design. JV, BTvdB, and AvdH conducted experiments. JV and BTvdB performed data analysis. JV, TTGN, MKV, BTvdB, FGMR, LSD, HLL, PV, RM, and MW wrote or contributed to the writing of the manuscript.
Additional information
All experiments described in this article comply with the Radboud Institutional Review Board. OrganoPlate® and PhaseGuide™ are a registered trademark of Mimetas BV. The authors MKV, AvdH, HLL, and PV are employees of Mimetas BV.
Corresponding author
Additional information
Guest Editors: Rosalinde Masereeuw and Raymond Lai
Electronic Supplementary Material
ESM 1
(DOCX 22 kb)
Fig. S2
Long-term culture of 20 days of ciPTEC-OAT1. Immunofluorescence staining of F-actin (red), tight-junction protein zonula occludens (ZO-1, green), nuclei (DRAQ5, blue) in ciPTEC-OAT1 in the OrganoPlate, stained at day 20 after seeding. Scale bar represents 100 μm. (PNG 781 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Vriend, J., Nieskens, T.T.G., Vormann, M.K. et al. Screening of Drug-Transporter Interactions in a 3D Microfluidic Renal Proximal Tubule on a Chip. AAPS J 20, 87 (2018). https://doi.org/10.1208/s12248-018-0247-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-018-0247-0