Abstract
Today, we are experiencing unprecedented growth and innovation within the pharmaceutical industry. Established protein therapeutic modalities, such as recombinant human proteins, monoclonal antibodies (mAbs), and fusion proteins, are being used to treat previously unmet medical needs. Novel therapies such as bispecific T cell engagers (BiTEs), chimeric antigen T cell receptors (CARTs), siRNA, and gene therapies are paving the path towards increasingly personalized medicine. This advancement of new indications and therapeutic modalities is paralleled by development of new analytical technologies and methods that provide enhanced information content in a more efficient manner. Recently, a liquid chromatography-mass spectrometry (LC-MS) multi-attribute method (MAM) has been developed and designed for improved simultaneous detection, identification, quantitation, and quality control (monitoring) of molecular attributes (Rogers et al. MAbs 7(5):881–90, 2015). Based on peptide mapping principles, this powerful tool represents a true advancement in testing methodology that can be utilized not only during product characterization, formulation development, stability testing, and development of the manufacturing process, but also as a platform quality control method in dispositioning clinical materials for both innovative biotherapeutics and biosimilars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
To date, multi-attribute method (MAM) has been applied to several early stage clinical programs with different classes of protein therapeutics and compared to current practices. This experience shows that the MAM technology can be utilized for different types of protein therapeutics delivering highly specific and quantitative information, which is invaluable during process development and essential for molecular characterization. Data has also been generated to support its use for release and stability in alignment with Quality by Design (QbD) principles. Utilizing recent advances in the technology, research, and development labs, in industry, has generated convincing data that MAM would be highly beneficial in a cGMP environment. This article will summarize the advantages of MAM relative to conventional product testing approaches. We recommend that MAM, unlike conventional purity methods, be used to specifically monitor and quantify molecular product quality attributes and product/process-related impurities. This increased specificity for attributes with increased relevance to safety and efficacy can lead to better product/process understanding, shorter process and product development timelines, and an improved control strategy by improving specificity of the measurement.
MAM OVERVIEW
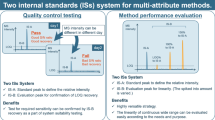
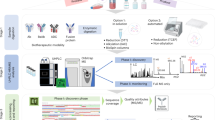
Reduced LC-MS-based peptide mapping methods have been used in academia and the industry for several decades. Biopharmaceutical companies are increasingly using quantitative LC-MS-based peptide mapping methods for clinical material characterization, as well as release and stability testing (1,2,3,4,5,6,7). Quantitative peptide mapping practices for assessing the relative extent of modification at specific amino acids have been shifting away from UV chromatographic peak integration to MS-based extracted-ion chromatograms (EICs) for increased specificity. Even though potential differences in ionization efficiencies between modified vs unmodified peptides are acknowledged, they appear to have minimal effect to date (8,9,10,11,12,13,14,15). For protein therapeutics, MAM involves a large-scale, targeted search of the peptide mapping data using commercially available software and predefined EIC parameters that originate based on the in-depth, site-specific characterization work performed during product development (Fig. 1). Furthermore, MAM employs site-specific quantitation, where for a particular amino acid modification, the readout is percent modified (relative to the unmodified peptide, summing all related forms of the modified peptide). An additional data processing function called new peak detection (NPD) is an essential component of the MAM. NPD automatically identifies new peaks in samples tested in comparison with a reference, with some pre-set peak selection criteria implemented with the intention of minimizing false-positive peak detection. NPD functions as an impurity/purity method in potentially detecting any new impurities in the samples present above the pre-set detection limit. Sample preparation protocols have been optimized and improved (16), which makes it possible to generate consistent, rapid, and low artifact peptide digestions for MAM analysis. The invention and evolution of high resolution/high mass accuracy mass spectrometers has drastically improved the sensitivity and specificity, which increases the confidence of identification and quantification of post-translational modifications. The robustness and bench-top footprint of modern mass spectrometers makes it practical for cGMP lab use. Instrument vendors now offer Title 21 CFR Part 11 compliant software that can be used for MAM data acquisition and automated data analysis in a quality laboratory.
MAM workflow. The MAM begins with a digestion method that limits the introduction of artificial PTMs. Initially, the digested biotherapeutic is characterized using a mass spectrometer capable of LC-MS/MS analysis. The resulting data are interrogated with an automated search algorithm. The search results yield sequence coverage, PTM identifications, chemical modifications, and clip identifications. These results are used to create a processing method to monitor PQAs by relative quantification (dividing the area for a modified PQA peptide by the summed area of the parent peptide and the modified PQA peptide). The monitoring stage of the MAM can be accomplished with LC/MS only data. Here, experimental samples are compared to the characterized reference standard. The processing method developed in the characterization stage is applied to quantify the relative amounts of PQAs, and NPD is applied to identify impurities and potential changes in PTMs. If new species are identified, the experimental sample(s) will be rerun on an MS/MS capable mass spectrometer for identification. Finally, the relative PQA values and NPD results can be exported to either a developmental report for supporting product and process understanding or a compliant report for releasing the biotherapeutic from a QC laboratory
REGULATORY LANDSCAPE AND MAM
For nearly two decades, regulatory agencies have suggested manufacturers leverage QbD and Design Space Models in order to provide operational flexibility while reducing product life cycle management burden. One crucial element of QbD is the generation of a Quality Target Product Profile (QTPP) and enhanced process control strategies to assure the product quality (17,18,19,20,21,22,23). These concepts have been debated, discussed, piloted, and reflected in regulatory guidance documents (17,18,19,20,24); however, the implementation and transition to such manufacturing models has been difficult to attain and has not been fully realized. To date, we have not fully leveraged the capabilities, flexibilities, and opportunities that these principles potentially provide.
Another key concept described in ICH Q8 and Q11 is that of the critical quality attribute (CQA). A CQA is defined as “A physical, chemical, biological, or microbiological property or characteristic that should be within an appropriate limit, range, or distribution to ensure the desired product quality.” For purposes of this discussion, a product quality attribute (PQA) can be considered any attribute of the product—either critical or noncritical. Those PQAs that should be within an appropriate limit, range, or distribution to ensure the desired product quality throughout the product’s life cycle are, by definition, CQAs. CQAs are generally associated with the drug substance, excipients, intermediates (in-process materials), and drug product. For biopharmaceutical products, CQAs may include items such as the nature and quantity of product-related substances, product-related impurities, and process-related impurities.
As defined in ICH Q6B, product-related impurities are “Molecular variants of the desired product (e.g., precursors, certain degradation products arising during manufacture and/or storage) which do not have properties comparable to those of the desired product with respect to activity, efficacy, and safety.” CQAs in the category of product-related impurities can be related to the post-translational modifications (PTMs) and chemical modifications of various amino acid residues of the drug product, amino acid misincorporations (sequence variants), or cleavage of its primary sequence due to hydrolysis during production and storage (25).
Conventional purity assays such as CEX-HPLC, cIEF, and CE-SDS used for lot release and stability testing are profile based and are often not capable of identifying and quantifying residue-specific CQAs. Each conventional assay is designed to primarily monitor one type of CQA and, in general, has limited resolution and specificity. Therefore, orthogonal methods, such as MAM, should be implemented to compensate for these limitations. MAM has the capability and sensitivity for pinpointing the location of the PTMs or cleavages, affording better understanding and monitoring of these attributes. If deemed critical, this method could then allow for specific monitoring in release and stability in a cGMP lab environment as part of overall control over these attributes.
Taken together, LC-MS-based MAM represents a significant opportunity to enable the pharmaceutical industry to quickly implement and leverage the QbD principles outlined above:
-
MAM provides residue-specific identification, direct quantitative monitoring, and enhanced overall understanding and trending of PTMs and chemical modifications, as compared to conventional profile-based purity methods
-
MAM is a key analytical tool in efforts to classify and differentiate CQAs from PQAs in conjunction with structure-function studies, as well as identify the relationship between one or more individual attributes and the product profile and performance
-
Specificity and quantitative ability of MAM provides control of CQAs at the appropriate point in the process, as well as consolidation and removal of less informative and redundant testing methods
-
MAM can simplify the life cycle management activities providing regulatory relief where a single method is maintained in place of several conventional purity methods
HISTORICAL APPROACHES FOR THE ASSESSMENT OF PRODUCT AND PROCESS UNDERSTANDING
Product-related purity assessments and controls have historically related to the degree of process understanding and analytical technologies available at the time. For example, the first monoclonal antibodies (mAb) commercialized in 1985, OKT3, included SDS-PAGE (reduced and non-reduced), gel permeation chromatography, and ion-exchange chromatography as part of its quality control (26). As technology and process understanding improved, quality control methods evolved as exemplified in the A-mAb case study (27) where proposed QC testing for product-related impurities only included HPSEC.
Recent years have seen great advancement made for analytical technologies. Conventional slab-gel SDS-PAGE used for purity and impurity tests of biologics have been largely replaced by capillary or chip-based electrophoretic separation methods. Gel IEF methods are mostly replaced by user-friendly imaging capillary isoelectric focusing (iCIEF) methods in QC laboratories. Advancement of analytical technologies has also led to more in-depth understanding of overall product quality. Conventional analytical methods, such as the chromatography or electrophoresis methods, when applied to heterogeneous biologics, report CQAs as acidic and basic charge isoforms or total fragmented impurities, respectively. These CQAs are good indicators of the overall stability of biologics. Any shifts in the charge or size distributions and/or peak profiles (i.e., new species) signal a change in the product; however, the root causes of the charge or size distribution change are usually not clearly understood or directly monitored using these methods.
The biotechnology industry has used conventional assays, including capillary electrophoresis (CE), ion exchange chromatography, reversed phase high performance liquid chromatography (RP-HPLC), and normal phase (NP) or hydrophilic interaction chromatography (HILIC), to monitor modifications like clipping, N-terminal signal peptides, C-terminal lysine, deamidation, isomerization, oxidation, and glycosylation. These conventional methods are still essential for early development and characterization, but at best offer limited information on specific product-related species and/or product-related impurities. Reducing CE is used to detect polypeptide clips in the final purified product as well as during upstream/downstream processing, formulation development, and stability studies; however, the reducing CE data does not provide any sequence information about where the clip is occurring in the protein. Ion exchange chromatography is used to show the charge distribution on a molecule based on surface charge at a specific pH with readout of main peak, acidic peaks, and basic peaks, but does not give any site specificity for where a modification is located in the amino acid sequence. The released N-glycan HILIC assay gives the overall distribution of the combined N-linked glycans on a molecule, but if a molecule has more than one asparagine site that is subject to glycosylation, the glycan distribution shown by the HILIC glycan assay does not represent the glycan profile at either specific asparagine, neither does the HILIC glycan map provide details around the percent occupancy at the potential glycosylation sites (12). Historically, in the absence of MAM, these advanced methods, prior to the characterization with orthogonal assays, have served as surrogate measures, often capable of only assessing global rather than site-specific chemical modification at the amino acid level.
MAM IMPLEMENTATION
A significant development in analytical technology for characterization and control testing of complex biopharmaceutical products is the wide-spread application of mass spectrometry. Peptide mapping coupled with mass spectrometric analysis is uniquely suited for the identification of a large number of PQAs in mAb therapeutics (28), including many potential CQAs. Peptide mapping is widely accepted as an identity test for therapeutics in the QC lab. While there are multiple contributing factors such as the use of reversed phase U/HPLC for enhanced peptide separation and the ever-improving detection sensitivity and resolving power, accurate mass determinations of intact and fragmented proteolytic peptides are the key to the extraordinary specificity now routinely available in the identification of a majority of product variants and impurities with unambiguity. As a result of such advancement in the use of mass spectrometry, it is now feasible and common practice that detailed product characterization is performed earlier in the product development cycle, alongside bioprocess and formulation process development.
Chromatography with hyphenated use of mass spectrometry such as MAM, combined with advanced data processing software, provides the opportunities to first identify, then inform and support process development to control, and/or minimize, product variants and impurities that could potentially impact safety and efficacy. MAM is a purity test, but also an “identity” test based on unique product specific sequences such as unique CDR sequences in a mAb. As mentioned above, conventional purity tests include ion exchange chromatography and reducing CE. The analysis is performed by automated integration and visual inspection of the chromatogram or electropherogram. There is a high probability that an impurity or new species could co-elute with an existing peak making it impossible to detect or quantify the impurity. Furthermore, the visible baseline of each assay limits the sensitivity of conventional purity tests. MAM does not suffer from any of the limitations of the conventional purity tests. MAM utilizes software to interrogate a sample for impurities or new species. The purity component of the MAM involves differential analysis of a reference standard compared to a test sample. The software performs a base peak alignment of the reference and test sample. The software is able to search the full mass range and retention time range of the data. The high resolution and mass accuracy of MAM allows the software to examine thousands of peaks for differences between the reference sample and the test sample. A reportable intensity threshold limit can be determined such that new species will be detected while eliminating false positives. By leveraging software to detect impurities, the need to visually inspect chromatograms or electropherograms for new species has been removed. MAM, with NPD, significantly reduces the potential for false negatives and co-eluting peaks, inherent in a visual inspection of the conventional purity tests. Additionally, identification of the impurity can be determined if MS/MS data were acquired for MAM analysis. Direct characterization is not possible with the conventional purity tests. As well, significant effort is still necessary to enrich and identify a particular variant or impurity if it is detected with a conventional purity test.
Many post-translational modifications (PTMs) and chemical modifications contribute to the formation of charge heterogeneity or size variants of biologics product, which can be readily monitored by MAM, often with the convenience of not requiring product-specific method development or optimization. Sequence variants due to either genetic mutation or amino acid misincorporation can be identified by mass spectrometry (29,30,31). When appropriate, cell lines with sequence variants can be rejected early in development, or accepted for further development with careful monitoring and process control of known sequence variants using MAM. Similarly, MAM can facilitate the monitoring of host cell proteins (HCPs) with immunogenicity or safety risks, and for any HCPs at unacceptable levels, MAM can aid the optimization of purification processes for greater clearance (32). In addition to being an excellent formulation screening tool, MAM is now widely used to monitor specific oxidation, deamidation, isomerization, and/or succinimide formation events that can impact product potency and establish appropriate product expiries to mitigate their impact (8,10,12,13,14,32,33).
MAM ENABLING OF IMPROVED CONTROL STRATEGIES AND ADVANCED BIOMANUFACTURING
ICH Q10 defines a control strategy as “A planned set of controls, derived from current product and process understanding, that assures process performance and product quality” (17). Control strategy design, development, implementation, and maintenance is managed across the life cycle of the product. Monitoring and verification activities within the quality systems are an integral part of demonstrating our ability to manage risk and increase the confidence in our control strategy throughout the life cycle. The MAM method monitors specific attributes at the molecular level which provides product quality information at the amino acid level, i.e., chemical modifications, fragmentation, glycosylation, and any new variants including point mutations/sequence variants. If variants were to be observed due to any potential deviation from the cell line and/or process changes, the NPD capability within MAM would be able to detect attribute changes which is not possible using conventional methods. This further improves our ability to control attributes that matter for patient safety at a desired level.
Specifications are one part of an overall control strategy, and, in a traditional control strategy, drug substance/drug product specifications are the primary means of control. The ability to specifically measure attributes at the molecular level in a quantitative manner via MAM enables us to establish a direct link between product quality attributes and clinical performance. The transition from characterization of a CQA in process development to a QC release specification comes from product and process development experience and adequate bridging to conventional methods. MAM can be a critical component to establishing clinically relevant specifications using direct attribute measurements and biological relevance information from preclinical and clinical studies providing an improved control strategy to report the variants (oxidation, deamidation, isomerization, sialylation, high mannose, and clips) for lot release/stability and set biologically relevant specifications.
Furthermore, an enhanced control strategy usually starts from identification of critical control points (CCPs) of the CQAs and the critical process parameters as well as input parameters at the CCPs that drive the formation and level of the CQAs. A critical control point (34) of a CQA refers to a particular unit operation of a production process, after which there is little to no potential for a CQA to change. The enhanced control strategy can involve application of a validated control loop for a CQA at its CCP, utilizing real-time product quality prediction through modeling or direct measurement and allowing self-correction of the process to meet the desired range of the CQA.
Based on its features, MAM is ideally suited for application in enhanced control strategies as it measures and quantifies several CQAs simultaneously in a single analysis. When combined with advanced sampling techniques from the bioreactor followed by sample clean-up, MAM can universally provide “near-real-time measurements” (current turn-around time for MAM data is 3–4 h) of any number of attributes at line on the manufacturing floor as a process analytical technology (PAT) application with direct monitoring of CQAs and product attribute control if linked with the appropriate process levers. Monitoring and control of an array of attributes was previously not achievable for biologics with one single method; furthermore, when MAM is applied at the CCP, it lends itself perfectly to disposition with real-time-release testing (RTRT) without the need for extensive quality control laboratory end-product release testing of drug substance and drug product.
Examples of enhanced control strategies have been demonstrated by a variety of biotechnology companies. Control of the glucose feeding based on real-time monitoring of the culture at the bioreactor stage using a Raman probe allows a more precise control of the glucose concentration in the culture than conventional periodic glucose measurements and therefore a more consistent level of glycation in mAb products manufactured (35). Similarly, others have reported real-time glycosylation monitoring of bioreactor using a micro-sequential injection system coupled with U/HPLC N-glycan analysis (36), which supported the use of the method for process monitoring and control. In another report, Zupke et al. showcased direct control of high-mannose N-glycosylation levels by controlling mannose feed to the bioreactor (37). This procedure was enabled by combining near real-time mass spectrometry measurements and standard bioreactor monitoring with a nonlinear predictive model, developed based on understanding of the relationship between bioreactor mannose concentration with high-mannose level on the mAb and bioprocess performance. Biogen successfully replaced conventional chromatography assays by multivariate models that predict lower pI isoforms and N-glycan properties, e.g., levels of galactosylation and levels of sialylation, based on cell culture process parameters, as part of the drug substance and drug product release of a commercial mAb approved by the European Medicines Agency in 2016 (38). This approach successfully removed the end-product release testing of these product qualities and provided an avenue for a newer paradigm for CQA control using product quality predictive models established based on process understanding. MAM can potentially be the one method that simultaneously reports multiple attributes including levels of N-glycans and/or O-glycans as well as levels of specific PTMs which are correlated with purity or impurity from conventional assays, as part of these advanced process control loops.
CHALLENGES AND SOLUTIONS FOR BROADER IMPLEMENTATION OF MAM
Implementation of any new technology is not achieved without some resistance to change or hurdles that must be overcome for broader implementation. This is no different for the expanded use of MAM for cGMP testing of biotherapeutics and enabling of an advanced control strategy. Common issues and concerns around broader implementation of MAM typically fall into four categories: technical, regulatory/compliance, capability as a replacement (instead of an additional) release test, and a diversified regulatory environment.
The most common concerns for MAM in the technical space include critical reagents for digestion (sample preparation), hardware robustness between instruments (mainly MS focused), method transferability, and software robustness for full automation of data analysis. These technical concerns have been debated at biopharmaceutical conferences since 2014 with the MS hardware robustness and software for new peak detection (NPD) as the most hotly debated topics. Regulatory/compliance concerns include large data volume, method validation, potential for increased out of specification results/investigations, parallel testing (conventional assays plus MAM), lack of proper technical understanding from quality control labs, and overall regulatory acceptance.
There also seems to be varying opinions on the capability of MAM to fully replace conventional assays in a cGMP environment. The notion of having conventional assays in addition to MAM is a barrier to broader implementation for most companies. In order to potentially replace conventional assays with MAM, it has been recommended by a regulatory agency to demonstrate strong correlation between conventional purity/impurity assays such as CE or ion exchange chromatography (IEC) with MAM results to support filing for a replacement of conventional release assays with MAM. It has been reported recently at the AccBio conference in 2017 that a good correlation between the MAM and IEC or N-glycan results was observed with a large number of commercial drug substance batches of a mAb product; however, this type of correlation was not found for a different IgG1 mAb in clinical development (39). The dependence of correlation on the particular protein biopharmaceuticals adds complication in replacing the conventional release tests using MAM agnostic of molecule types. The notion of demonstrating a like for like comparison between the MAM and the conventional assay should not preclude the replacement of the conventional methods with MAM for releasing biopharmaceuticals. Relative to the progression and acceptance of MAM, efforts have been made for adoption of this technology as part of the CQA control strategy. To date, limited success has been achieved regarding the replacement of conventional release and stability assays with MAM in the QC environment due to regulatory unfamiliarity, experience, and level of comfort with the MAM technology. Subsequently, as the regulatory agencies gain familiarity, dual and repetitive testing is currently being performed, where both MAM and conventional assays are included side by side for product release and stability testing. While dual testing beyond early development is unsustainable, the ultimate goal is to have MAM replace lower-resolving conventional methods in the long term.
The diverse regulatory environment worldwide presents additional challenges to MAM implementation in a cGMP environment. Regulatory agencies may hold different opinions in adoption of this assay as a release test especially in lieu of conventional release tests. This white paper is intending to provide the regulatory agencies a unified industrial view of the MAM and help pave a path forward to minimize the regulatory diversity. The regulatory acceptance and assay replacement topics have also been discussed at industry and scientific conferences, including the sharing of successful experiences at Amgen at 2017 WCBP MAM roundtable (40). In a Bioanalysis editorial earlier this year, Zhang and Guo provided a summary of the current state of MAM highlighting both advantages and challenges (41).
In efforts to expand industry and regulatory acceptance of MAM, numerous research and development studies are ongoing to provide solutions for the most common technical and compliance-related challenges. For critical reagents, multiple enzyme vendors are optimizing their reagent quality and platforms to simplify the process for executing a digestion with their materials. Advancements in liquid handling robotics also provide a means to improve the reproducibility of the sample preparation for MAM. Robotics can help streamline method qualification in addition to providing a resource savings and less variability from sample preparation. Regarding the MS hardware, instrument vendors are aware of the equipment settings critical to performance between systems. Enabling user access to advance instrument control settings was deemed critical for obtaining consistent hardware performance across systems (e.g., vacuum and ion transfer speed) and will help improve mass spectrometer to mass spectrometer performance of the MAM assay in a cGMP laboratory, especially for identical make and models. Software robustness and options for MAM data collection and analysis are also improving (41). Multiple vendors provide 21CFR Part 11 compliant packages for execution of the entire MAM data analysis workflow. Expanding the options in the compliant MAM data analysis space, including new peak detection (NPD), will provide much needed technical alternatives for implementation. These hardware and software advances will enable more successful MAM method transfer and validation. Regardless of the technical advances, it is clear from a regulatory agency perspective that equipment redundancy can assist the implementation of MAM in a cGMP environment.
For regulatory/compliance-related concerns such as large amounts of data and method validation, the hardware and software advances described above will help significantly. Concern for increased investigations and out of specification results can be accounted for early during the process development and specification setting. Acceptance criteria for MAM are quite different than conventional assays because these require sufficient characterization of assay sensitivity for each CQA monitored by the LC-MS MAM approach. Combining phase appropriate acceptance criteria for the CQA will enable a robust overall control strategy leveraging MAM. Remaining GMP/compliance hurdles can be addressed through participation in the MAM consortium (42), discussions/seminar with regulatory agencies, and participation in round robin MAM testing opportunities for protein standards such as the NIST mAb. These activities provide the forum to train and educate laboratory, quality and regulatory colleagues on a unified MAM workflow through focused data and technology discussions. Having been widely applied for antibodies and other proteins, MAM has been accepted as a characterization assay. An increased understanding, confidence and acceptance of the MAM technology and its application, will allow the industry to fully leverage the benefits of MAM as an advanced and emerging technology across the entire product and process development life cycle.
MAM ADVANTAGES, CURRENT LIMITATIONS, AND FUTURE APPLICATIONS
While there is still a need for conventional assays to assess some product characteristics (e.g., multimers), the application of MAM provides the opportunity to better align a specific critical quality attribute with overall product structure-function relationship, control, and assessment, resulting in a more thorough understanding of the relationship of CQAs with the product profile and performance. As a result of increased performance in mass spectrometry technology, sophisticated data processing software packages and data interpretation, substantial advances have been made in the data analysis of biologics. The goal of introducing MAM technology is to expand the usefulness of this information in order to assess all molecular attributes in one single analysis in a fully automated fashion, while maintaining compliance with ICH and local regional regulatory requirements.
Going forward, replacing conventional assays from the QC environment, as release specifications and stability testing protocols, in favor of MAM with adequate acceptance criteria is desirable. Additionally, where appropriate and justified with data and scientific rationale, future clinical trial application filings can increasingly leverage the capabilities of MAM as a characterization, release assay, and as part of the formal stability assessment. To fully realize the benefits of this emergent technology, MAM implementation should include submission within new market applications to regulatory authorities as a characterization assay, and where appropriate, as a release and stability-indicating test method, in original marketing applications and currently approved commercial life cycle products. Successful implementation of MAM as part of the analytical control strategy in these environments provides an opportunity to fully realize MAM as an advancement in testing methodology.
At present, MAM is not suitable for assessing disulfide bond linkages, disulfide isoform ratios (e.g., IgG2 mAbs), trisulfide levels, cysteine adducts, etc. in mAbs, although a non-reducing version of MAM would likely address this category of PQAs. Likewise, direct monitoring of succinimide is not always possible via trypsin peptide mapping when the digestion pH is above 7.5, as it is in MAM (pH 7.9), due to rapid hydrolysis to isoAsp and Asp. Additional current limitations of MAM include the measurement of DNA levels (or other non-proteinaceous molecules not amenable to proteolytic mapping conditions), as well as self-association and aggregation, and higher-order structure (HOS) and potential conformers (43,44,45). With respect to the latter PQAs, structure-based MS techniques are making big strides, and one can envision a specialized version of MAM for HOS offering site-specific assessment of structure with high sensitivity before and after stress conditions.
Future applications associated with development of MAM could be envisioned by the following:
-
1.
Including additional PQAs for quantitation and control
-
2.
Data standardization across multiple instrument and software platforms
-
3.
On-the-floor real-time testing (in-process and/or release) and product attribute control
-
4.
Smaller instrument footprint, automation, and faster run times
-
5.
Expedited regulatory submissions and life cycle management
-
6.
Raw data submission for regulatory agencies to evaluate MAM data
CONCLUSIONS AND OUTLOOK
MAM, an emerging sensitive, high-resolution, low artifact product testing technique, can enable a future state for biopharmaceutical products of enhanced direct measurement that can facilitate approaches for control of multiple individual CQAs and PQAs. MAM can support QbD implementation by offering in-depth product and process knowledge as well as more efficient product testing by consolidation (and/or replacement) of multiple profile-based release and stability assays. With a greater appreciation of clinically relevant attribute criticality throughout the industry and scientific literature, MAM is poised to contribute immensely to heightened process and product understanding. Improved detection will enable development of processes capable of more consistent product quality, thereby better ensuring patient safety and drug efficacy.
MAM greatly enhances product characterization which in turn leads to improved process understanding and the development of process control levers for specific PTMs, size variants, or impurities, delivering improved performance in product consistency, safety, and efficacy. Combination of the control levers with real-time in-process PQA monitoring using user-friendly and fast on-the-floor analytical methods results in process control loops that can be employed as part of an advanced control strategy for biologics. Replacement of conventional methods with MAM, along with incorporation of process control levers, within an advanced control strategy could enable product attribute control (PAC) for biologics. Furthermore, utilization of MAM can facilitate root cause analysis of raw material variation or manufacturing process deviations and help the industry enhance product knowledge and process understanding towards realizing the QbD goal in product development and manufacturing.
With unprecedented growth and innovation in the biotechnology industry, it is essential for the industry to be able to implement and regulators to accept more advanced, modality independent, analytical technologies. The novel protein therapeutic modalities, such as recombinant human proteins, mAbs, BiTEs, CARTs, antibody drug conjugates, and fusion proteins, require improved techniques for product and process characterization, CQA assessment with structure function studies, and stability testing and release from the QC laboratory. MAM offers the opportunity to platform a global release strategy that can improve the safety and efficacy of these therapeutics for the patient. By implementing MAM, the industry is not only heading towards enhanced control strategies, but is also able to better link CQAs to the clinical performance of a therapeutic to improve safety and efficacy for the patient.
References
Lundell N, Schreitmüller T. Sample preparation for peptide mapping—a pharmaceutical quality-control perspective. Anal Biochem [Internet]. 1999;266(1):31–47. Available from: http://linkinghub.elsevier.com/retrieve/pii/S0003269798929190. https://doi.org/10.1006/abio.1998.2919.
Bongers J, Cummings JJ, Ebert MB, Federici MM, Gledhill L, Gulati D, et al. Validation of a peptide mapping method for a therapeutic monoclonal antibody: what could we possibly learn about a method we have run 100 times? J Pharm Biomed Anal [Internet]. 2000;21(6):1099–128. Available from: http://www.ncbi.nlm.nih.gov/pubmed/10708395
Dougherty J, Mhatre R, Moore S. Using peptide maps as identity and purity tests for lot release testing of recombinant therapeutic proteins. Process Dev [Internet]. 2003;54. Available from: http://alfresco.ubm-us.net/alfresco_images/pharma/2014/08/22/4e9452e7-dfb9-40e7-a143-fd92a0ab6183/article-54510.pdf
Challener CA. Mass spectrometry measures up to analytical challenges. BioPharm Int [Internet]. 2017;30(2):14–7. Available from: http://www.biopharminternational.com/mass-spectrometry-measures-analytical-challenges
Zhang Z. Large-Scale Identification and quantification of covalent modifications in therapeutic proteins. Anal Chem [Internet]. 2009;81(20):8354–64. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19764700
Rogstad S, Faustino A, Ruth A, Keire D, Boyne M, Park J. A Retrospective evaluation of the use of mass spectrometry in FDA biologics license applications. J Am Soc Mass Spectrom [Internet]. 2017;28(5):786–94. [cited 2017 Oct 31], Available from: http://www.ncbi.nlm.nih.gov/pubmed/27873217
Nowak C, Cheung KJ, Dellatore MS, Katiyar A, Bhat R, Sun J, et al. Forced degradation of recombinant monoclonal antibodies: a practical guide. MAbs [Internet]. 2017;9(8):1–14. [cited 2017 Oct 31], Available from:https://www.tandfonline.com/doi/full/10.1080/19420862.2017.1368602
Rogers RS, Nightlinger NS, Livingston B, Campbell P, Bailey R, Balland A, et al. Multi-attribute method for characterization, quality control testing and disposition of biologics. MAbs. 2015;7(5):881–90. https://doi.org/10.1080/19420862.2015.1069454.
Bomans K, Haberger M, Bonnington L, Wagner K, Kailich T, Wegele H, et al. Multi-attribute monitoring of antibody modifications by semi-automated liquid chromatography mass spectrometry peptide mapping|American Pharmaceutical Review—The Review of American Pharmaceutical Business & Technology. Am Pharm Rev [Internet]. 2016; Available from: http://www.americanpharmaceuticalreview.com/Featured-Articles/331529-Multi-Attribute-Monitoring-of-Antibody-Modifications-by-Semi-Automated-Liquid-Chromatography-Mass-Spectrometry-Peptide-Mapping/
Eris T. Application of a mass spectrometry based multi-attribute-method (MAM) for QC release and stability testing of protein therapeutics. In: CASSS CMC Strategy Forum [Internet]. 2016. Available from: http://c.ymcdn.com/sites/www.casss.org/resource/resmgr/2016_CMCS_ErisTamer.pdf
Shah B, Jiang XG, Chen L, Zhang Z. LC-MS/MS peptide mapping with automated data processing for routine profiling of N-glycans in immunoglobulins. J Am Soc Mass Spectrom [Internet]. 2014;25(6):999–1011. Available from: http://link.springer.com/10.1007/s13361-014-0858-3
Wang T, Chu L, Li W, Lawson K, Apostol I, Eris T. Application of a quantitative LC–MS multiattribute method for monitoring site-specific glycan heterogeneity on a monoclonal antibody containing two N-linked glycosylation sites. Anal Chem [Internet]. 2017;89(6):3562–7. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28199092
Xu X, Qiu H, Li N. LC-MS multi-attribute method for characterization of biologics. J Appl Bioanal [Internet]. 2017;3(2):21–25. Available from: http://www.betasciencepress.com/index.php/jab17003. 10.17145/jab.17.003.
Wang Y, Li X, Liu Y-H, Richardson D, Li H, Shameem M, et al. Simultaneous monitoring of oxidation, deamidation, isomerization, and glycosylation of monoclonal antibodies by liquid chromatography-mass spectrometry method with ultrafast tryptic digestion. MAbs [Internet]. 2016;8(8):1477–86. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27598507
Lyubarskaya Y, Kobayashi K, Swann P. Application of mass spectrometry to facilitate advanced process controls of biopharmaceutical manufacture litate tical. Pharm Bioprocess [Internet]. 2015;3(4):313–321. Available from: http://www.openaccessjournals.com/articles/application-of-mass-spectrometry-to-facilitate-advanced-process-controls-of-biopharmaceutical-manufacture.pdf. https://doi.org/10.4155/pbp.15.10.
Ren D, Pipes GD, Liu D, Shih L-Y, Nichols AC, Treuheit MJ, et al. An improved trypsin digestion method minimizes digestion-induced modifications on proteins. Anal Biochem [Internet]. 2009;392(1):12–21. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19457431
ICH Harmonised Tripartite Guideline Pharmaceutical Quality System Q10 Pharmaceutical Quality System. 2008; Available from: http://www.ich.org/fileadmin/Public_Web_Site/ICH_Products/Guidelines/Quality/Q10/Step4/Q10_Guideline.pdf
ICH Harmonised Tripartite Guideline Quality Risk Management Q9 quality risk management. 2005; Available from: http://www.ich.org/fileadmin/Public_Web_Site/ICH_Products/Guidelines/Quality/Q9/Step4/Q9_Guideline.pdf
Ich Harmonised Tripartite Guideline Pharmaceutical Development Q8(R2). 2009; Available from: http://www.ich.org/fileadmin/Public_Web_Site/ICH_Products/Guidelines/Quality/Q8_R1/Step4/Q8_R2_Guideline.pdf
Guidance for Industry Q11 Development and Manufacture of Drug Substances. 2012;301–827. Available from: http://www.fda.gov/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/default.htm
Yu LX, Amidon G, Khan MA, Hoag SW, Polli J, Raju GK, et al. Understanding Pharmaceutical Quality by Design. AAPS J [Internet]. 2014;16(4):771–83. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24854893
Robinson CJ, Jones C. Quality control and analytical techniques for biopharmaceuticals. Bioanalysis [Internet]. 2011;3(1):81–95. Available from: http://www.ncbi.nlm.nih.gov/pubmed/21175369
Luciani F, Galluzzo S, Gaggioli A, Kruse NA, Venneugues P, Schneider CK, et al. Implementing quality by design for biotech products: are regulators on track? MAbs [Internet]. 2015;7(3):451–5. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25853461
Guidance for Industry Immunogenicity Assessment for Therapeutic Protein Products. 2014; Available from: http://www.fda.gov/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/default.htm
ICH Harmonised Tripartite Guideline Specifications: Test procedures and acceptance criteria for Biotechnological/biological products Q6B specifications: test procedures and acceptance criteria for biotechnological/biological products. 1999; Available from: https://www.ich.org/fileadmin/Public_Web_Site/ICH_Products/Guidelines/Quality/Q6B/Step4/Q6B_Guideline.pdf
Beekman MJHI, Turnock G. Biotechnological innovations in health care. [Internet]. Butterworth-Heinemann; 1991. 299 p. Available from: http://www.sciencedirect.com/science/book/9780750614979
The CMC Biotech Working Group. A-Mab: a case study in bioprocess Development. 2009; Available from: http://c.ymcdn.com/sites/www.casss.org/resource/resmgr/imported/A-Mab_Case_Study_Version_2-1.pdf
Beck A, Sanglier-Cianférani S, Van Dorsselaer A. Biosimilar, biobetter, and next generation antibody characterization by mass spectrometry. Anal Chem [Internet]. 2012;84(11):4637–46. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22510259
Yang J, Manolio TA, Pasquale LR, Boerwinkle E, Caporaso N, Cunningham JM, et al. Genome partitioning of genetic variation for complex traits using common SNPs. Nat Genet [Internet]. 2011;43(6):519–25. Available from: http://www.ncbi.nlm.nih.gov/pubmed/21552263
Zhang Z, Shah B, Bondarenko PV. G/U and certain wobble position mismatches as possible main causes of amino acid misincorporations. Biochemistry [Internet]. 2013;52(45):8165–76. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24128183
Zhang T, Huang Y, Chamberlain S, Romeo T, Zhu-Shimoni J, Hewitt D, et al. Identification of a single base-pair mutation of TAA (Stop codon) → GAA (Glu) that causes light chain extension in a CHO cell derived IgG1. MAbs [Internet]. 2012;4(6):694–700. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23018810
Rogers R. Leveraging the MAM to improve biotherapeutic characterization [Internet]. 21st Symposium on the Interface of Regulatory and Analytical Sciences for Biotechnology Health Products. CASSS; 2017 [cited 2017 Oct 31]. Available from: http://c.ymcdn.com/sites/www.casss.org/resource/resmgr/wcbp_speaker_slides/2017_WCBP_RogersRich.pdf
Xu W, Jimenez RB, Mowery R, Luo H, Cao M, Agarwal N, et al. A Quadrupole Dalton-based multi-attribute method for product characterization, process development, and quality control of therapeutic proteins. MAbs [Internet]. 2017;9(7):1186–96. [cited 2017 Oct 31], Available from: https://www.tandfonline.com/doi/full/10.1080/19420862.2017.1364326
Annex 7 Application of Hazard Analysis and Critical Control Point (HACCP) methodology to pharmaceuticals. World Heal Organ WHO Tech Rep Ser [Internet]. 2003;(908). Available from: http://www.who.int/medicines/areas/quality_safety/quality_assurance/ApplicationHACCPMethodologyPharmaceuticalsTRS908Annex7.pdf
Berry BN, Dobrowsky TM, Timson RC, Kshirsagar R, Ryll T, Wiltberger K. Quick generation of Raman spectroscopy based in-process glucose control to influence biopharmaceutical protein product quality during mammalian cell culture. Biotechnol Prog [Internet]. 2016;32(1):224–34. Available from: http://doi.wiley.com/10.1002/btpr.2205
Tharmalingam T, Wu C-H, Callahan S, Goudar TC. A framework for real-time glycosylation monitoring (RT-GM) in mammalian cell culture. Biotechnol Bioeng [Internet]. 2015;112(6):1146–54. Available from: http://doi.wiley.com/10.1002/bit.25520
Zupke C, Brady LJ, Slade PG, Clark P, Caspary RG, Livingston B, et al. Real-time product attribute control to manufacture antibodies with defined N-linked glycan levels. Biotechnol Prog [Internet]. 2015;31(5):1433–41. Available from: http://doi.wiley.com/10.1002/btpr.2136
Zang L (Biogen), Swann P (Biogen). Personal Communication. 2017.
5th International Conference on Accelerating Biopharmaceutical Development [Internet]. 2017. Available from: https://www.aiche.org/sbe/conferences/accelerating-biopharmaceutical-development-abd/2017
Rogers R, Abernathy M. Discussion surrounding the implementation of a mass spectrometry-base method for real-time monitoring of bioreactors and QC release. In: WCBP round table [Internet]. 2017. Available from: http://c.ymcdn.com/sites/www.casss.org/resource/resmgr/wcbp/2017_WCBP_Table_45.pdf
Zhang Y, Guo J. Characterization and QC of biopharmaceuticals by MS-based “multi-attribute method”: advantages and challenges. Bioanalysis [Internet]. 2017;9(6):499–502. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28207284.
MAM Consortium-Google Groups [Internet]. Available from: https://groups.google.com/forum/?hl=en#!forum/mam-consortium
Berkowitz SA, Engen JR, Mazzeo JR, Jones GB. Analytical tools for characterizing biopharmaceuticals and the implications for biosimilars. Nat Rev Drug Discov [Internet]. 2012;11(7):527–40. [cited 2017 Oct 31], Available from: http://www.ncbi.nlm.nih.gov/pubmed/22743980
Beck A, Wagner-Rousset E, Ayoub D, Van Dorsselaer A, Sanglier-Cianférani S. Characterization of therapeutic antibodies and related products. Anal Chem. 2013;85(2):715–36. https://doi.org/10.1021/ac3032355.
Fekete S, Guillarme D, Sandra P, Sandra K. Chromatographic, electrophoretic, and mass spectrometric methods for the analytical characterization of protein biopharmaceuticals. Anal Chem [Internet]. 2016;88(1):480–507. [cited 2017 Oct 31], Available from: http://www.ncbi.nlm.nih.gov/pubmed/26629607
Acknowledgements
We would like to thank all of our colleagues for their critical feedback for this white paper.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Rogers, R.S., Abernathy, M., Richardson, D.D. et al. A View on the Importance of “Multi-Attribute Method” for Measuring Purity of Biopharmaceuticals and Improving Overall Control Strategy. AAPS J 20, 7 (2018). https://doi.org/10.1208/s12248-017-0168-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-017-0168-3