Abstract
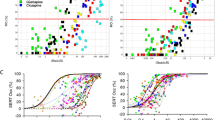
The physiologically based pharmacokinetic (PBPK) model for liver transporter substrates has been established previously and used for predicting drug–drug interactions (DDI) and for clinical practice guidance. So far, nearly all the published PBPK models for liver transporter substrates have one or more hepatic clearance processes (i.e., active uptake, passive diffusion, metabolism, and biliary excretion) estimated by fitting observed systemic data. The estimated hepatic clearance processes are then used to predict liver concentrations and DDI involving either systemic or liver concentration. However, the accuracy and precision of such predictions are unclear. In this study, we try to address this question by using the PBPK model to generate simulated compounds for which we know both systemic and liver profiles. We then developed an approach to assess the accuracy and precision of predicted liver concentration. With hepatic clearance processes estimated using plasma data, model predictions of liver are typically accurate (i.e., true value is bounded by predicted maximum and minimum); however, only for a few compounds are predictions also precise. The results of the current study indicate that extra attention is required when using the current PBPK approach to predict liver concentration and DDI for transporter substrates dependent upon liver concentrations.






Similar content being viewed by others
References
Peters SA. Physiologically based pharmacokinetic (PBPK) modeling and simulations : principles, methods, and applications in the pharmaceutical industry. Hoboken: Wiley; 2011. xvii, 430 p. p.
Rose RH, Neuhoff S, Abduljalil K, Chetty M, Rostami-Hodjegan A, Jamei M. Application of a physiologically based pharmacokinetic model to predict OATP1B1-related variability in pharmacodynamics of rosuvastatin. CPT Pharmacometrics Syst Pharmacol. 2014;3, e124.
Watanabe T, Kusuhara H, Maeda K, Shitara Y, Sugiyama Y. Physiologically based pharmacokinetic modeling to predict transporter-mediated clearance and distribution of pravastatin in humans. J Pharmacol Exp Ther. 2009;328(2):652–62.
Li R, Barton HA, Varma MV. Prediction of pharmacokinetics and drug-drug interactions when hepatic transporters are involved. Clin Pharmacokinet. 2014;53(8):659–78.
Li R, Barton HA, Yates PD, Ghosh A, Wolford AC, Riccardi KA, et al. A “middle-out” approach to human pharmacokinetic predictions for OATP substrates using physiologically-based pharmacokinetic modeling. J Pharmacokinet Pharmacodyn. 2014;41(3):197–209.
Li R, Barton HA, Maurer TS. Toward prospective prediction of pharmacokinetics in OATP1B1 genetic variant populations. CPT Pharmacometrics Syst Pharmacol. 2014;3, e151.
Shah DK, Betts AM. Towards a platform PBPK model to characterize the plasma and tissue disposition of monoclonal antibodies in preclinical species and human. J Pharmacokinet Pharmacodyn. 2012;39(1):67–86.
Davies B, Morris T. Physiological parameters in laboratory animals and humans. Pharm Res. 1993;10(7):1093–5.
Rodgers T, Leahy D, Rowland M. Physiologically based pharmacokinetic modeling 1: predicting the tissue distribution of moderate-to-strong bases. J Pharm Sci. 2005;94(6):1259–76.
Rodgers T, Rowland M. Physiologically based pharmacokinetic modelling 2: predicting the tissue distribution of acids, very weak bases, neutrals and zwitterions. J Pharm Sci. 2006;95(6):1238–57.
Poirier A, Funk C, Scherrmann JM, Lave T. Mechanistic modeling of hepatic transport from cells to whole body: application to napsagatran and fexofenadine. Mol Pharm. 2009;6(6):1716–33.
Bellu G, Saccomani MP, Audoly S, D’Angio L. DAISY: a new software tool to test global identifiability of biological and physiological systems. Comput Methods Prog Biomed. 2007;88(1):52–61.
Spear RC, Bois FY. Parameter variability and the interpretation of physiologically based pharmacokinetic modeling results. Environ Health Perspect. 1994;102 Suppl 11:61–6.
Martin PD, Warwick MJ, Dane AL, Brindley C, Short T. Absolute oral bioavailability of rosuvastatin in healthy white adult male volunteers. Clin Ther. 2003;25(10):2553–63.
DiStefano JJ. Dynamic systems biology modeling and simulation. First edition. ed. Amsterdam: Elsevier, Academic Press; 2013. xxii, 859 pages p.
Acknowledgments
The authors greatly appreciate the sincere help on structural identifiability analysis from Dr. Nicolette Meshkat at Santa Clara University and Dr. Joe DiStefano III at University of California, Los Angeles.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 267 kb)
Rights and permissions
About this article
Cite this article
Li, R., Maurer, T.S., Sweeney, K. et al. Does the Systemic Plasma Profile Inform the Liver Profile? Analysis Using a Physiologically Based Pharmacokinetic Model and Individual Compounds. AAPS J 18, 746–756 (2016). https://doi.org/10.1208/s12248-016-9895-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1208/s12248-016-9895-0