Abstract
Diverse neuropeptides participate in cell–cell communication to coordinate neuronal and endocrine regulation of physiological processes in health and disease. Neuropeptides are short peptides ranging in length from ~3 to 40 amino acid residues that are involved in biological functions of pain, stress, obesity, hypertension, mental disorders, cancer, and numerous health conditions. The unique neuropeptide sequences define their specific biological actions. Significantly, this review article discusses how the neuropeptide field is at the crest of expanding knowledge gained from mass-spectrometry-based neuropeptidomic studies, combined with proteomic analyses for understanding the biosynthesis of neuropeptidomes. The ongoing expansion in neuropeptide diversity lies in the unbiased and global mass-spectrometry-based approaches for identification and quantitation of peptides. Current mass spectrometry technology allows definition of neuropeptide amino acid sequence structures, profiling of multiple neuropeptides in normal and disease conditions, and quantitative peptide measures in biomarker applications to monitor therapeutic drug efficacies. Complementary proteomic studies of neuropeptide secretory vesicles provide valuable insight into the protein processes utilized for neuropeptide production, storage, and secretion. Furthermore, ongoing research in developing new computational tools will facilitate advancements in mass-spectrometry-based identification of small peptides. Knowledge of the entire repertoire of neuropeptides that regulate physiological systems will provide novel insight into regulatory mechanisms in health, disease, and therapeutics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION NEUROPEPTIDES FOR CELL–CELL COMMUNICATION
Neuropeptides Regulate Neuronal and Endocrine Functions
Neuropeptides are required for physiological functions as neurotransmitters in the nervous system and are essential as peptide hormones for endocrine regulation of target biological systems (Fig. 1). Thus, neuropeptides are critical mediators of cell–cell communication in neuroendocrine systems. Such neuropeptides are composed of diverse peptide sequences typically consisting of about 3–40 residues. It is estimated that hundreds to thousands of different neuropeptides may exist, with many yet to be discovered.
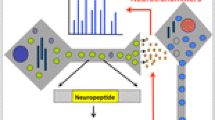
Neuropeptides for neuronal and endocrine cell–cell communication. a Neuropeptides, peptide neurotransmitters, in the central nervous system of brain. Brain neuropeptides function as peptide neurotransmitters to mediate chemical cell–cell communications among neurons. Neuropeptides are synthesized within secretory vesicles that are transported from the neuronal cell body via the axon to nerve terminals. The prohormone (also known as proneuropeptide) is packaged within the newly formed secretory vesicle in the cell body, and proteolytic processing of the precursor protein occurs during axonal transport and maturation of the secretory vesicle. Mature processed neuropeptides are contained within secretory vesicles at the synapse where activity-dependent, regulated secretion of neuropeptides occurs to mediate neurotransmission via neuropeptide activation of peptidergic receptors. b Neuropeptides, peptide neurotransmitters and peptide hormones, in the peripheral nervous system and endocrine systems for regulation of physiological organ functions. The peripheral nervous system regulates all organ systems, linking the central nervous system of the brain with peripheral neuronal control of physiological functions. In the body, neuropeptides also function as hormones that mediate endocrine cell–cell communication
Neuropeptides represent one of two main classes of neurotransmitters. Prior to the discovery of the neuropeptides, classical neurotransmitters are known as key mediators of cell–cell communication in the nervous system (1). The “classical” neurotransmitters consist of small molecules such as, for example, norepinephrine, serotonin, GABA, acetylcholine, and many others (1). The classical neurotransmitters are synthesized by (a) modifications of single amino acids such as, for example, norepinephrine synthesized from tyrosine and serotonin synthesized from tryptophan or (b) synthesized by enzymatic reactions such as, for example, acetylcholine generated from choline and acetyl-CoA by the choline acetyl transferase enzyme. The “peptide” and “classical” neurotransmitters together mediate neuronal cell–cell communication.
Among the neuropeptides, each of their unique primary sequences defines selective and potent biological actions. The same neuropeptide may participate in multiple regulatory systems in the nervous system and endocrine systems (Table I). For example, enkephalin neuropeptides function as neurotransmitters in the brain for regulation of behavior and pain and are also involved in peripheral actions including regulation of intestinal motility and immune cell functions (1,2). Adrenocorticotropin hormone (ACTH) is present in brain where it functions as a neuromodulator; furthermore, ACTH is a prominent peptide hormone released from the pituitary gland for control of glucocorticoid production in the adrenal cortex (3). Neuropeptides such as ß-endorphin, neuropeptide Y (NPY), galanin, corticotropin-releasing factor (CRF), vasopressin, insulin, and numerous others mediate diverse physiological functions that include analgesia, feeding behavior and blood pressure regulation, cognition, stress, water balance, and glucose metabolism, respectively (4,5). These and other neuropeptides regulate physiological functions in mammalian systems, as well as in invertebrate and related organisms (6–8). Clearly, neuropeptides possess a wide scope of diverse actions among numerous organisms.
Neuropeptides Generated by Proteolytic Processing of Prohormones
Neuropeptides are derived from larger protein precursors known as prohormones or proneuropeptides (9–11). The term “prohormone” is most commonly used in the field. Such protein precursors undergo proteolytic processing to generate the smaller peptide neurotransmitters and hormones.
Prohormone precursors share distinct and common features. Notably, the small active form of each neuropeptide is a domain present within its prohormone protein. A prohormone may contain one copy of the active neuropeptide, as represented by pro-NPY, progalanin, and pro-vasoactive intestinal polypeptide (VIP; Fig. 2) (12–14). Alternatively, a precursor may contain several related copies of the active neuropeptide. For example, proenkephalin (PE) contains four copies of (Met)enkephalin, one copy of the related (Leu)enkephalin, and one copy each of the ME-Arg-Gly-Leu and ME-Arg-Phe (Fig. 2) (15–17). Proteolysis of these precursors, especially tissue-specific processing is required for biologically active neuropeptides to be generated.
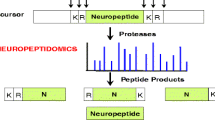
Prohormone precursors undergo proteolysis to generate active neuropeptides. Neuropeptides are synthesized as inactive preprohormone precursors that undergo removal of the N-terminal signal peptide sequence in the rough endoplasmic reticulum to generate the prohormone precursors (9–11). The prohormones, also known as proneuropeptides, undergo proteolytic processing at dibasic and monobasic cleavage sites to liberate the active neuropeptides. The precursor proteins may contain one copy of the active neuropeptide, such as the proneuropeptides for NPY, galanin, CRF, and VIP. Some proneuropeptides such as proenkephalin contains multiple copies of the active neuropeptide; proenkephalin contains four copies of (Met)enkephalin (ME), one copy of (Leu)enkephalin (L), and the related opioid peptides ME-Arg-Phe (H) and ME-Arg-Gly-Cleu (O). Certain precursors contain different peptide hormones within the same precursor, such as the POMC precursor which gives rise to the distinct peptide hormones ACTH, α-MSH, and ß-endorphin. The presence of ACTH in anterior pituitary and the presence of α-MSH and ß-endorphin in intermediate pituitary illustrate that tissue-specific processing of the POMC prohormone occurs
While each prohormone precursor possesses a distinct primary sequence, proteolytic processing occurs at dibasic residue sites that typically flank the NH2 and COOH termini of neuropeptides within their precursors (Fig. 2) (9–11). The dibasic residues Lys-Arg (KR) most often flank the neuropeptides; however, the dibasic sites Lys-Lys, Arg-Arg, and sometimes Arg- Lys also occur. Processing sometimes occurs at monobasic Arg sites as well as at multibasic residue sites. Processing at nonbasic residues occurs occasionally. Significantly, proteolytic processing is a key process required for the biosynthesis of numerous active neuropeptides from inactive precursors.
MASS SPECTROMETRY-BASED NEUROPEPTIDOMICS FOR GLOBAL ANALYSES OF DIVERSE PEPTIDES
Mass Spectrometry for Analyses of Multiple Neuropeptides
Multiple neuropeptides of unique sequences are secreted and utilized for coordinate regulation of physiological functions. A critical unmet need in the field is global knowledge of the neuropeptide profiles utilized in neuroendocrine control of cellular and biological functions. It is likely that previously unidentified peptide products exist that result from proteolytic processing of prohormones. Exciting new approaches utilizing mass spectrometry-based neuropeptidomics (18–22) to simultaneously identify neuropeptide profiles in a single biological event will open our understanding of how multiple neuropeptides, rather than a single neuropeptide, are cosecreted for coregulation of key biological functions.
For example, the stress response to the “fight or flight” condition results in active secretion of multiple neuropeptides and catecholamine neurotransmitters from the adrenal medulla of the sympathetic nervous system. Stress induces the secretion of enkephalins, catestatin, NPY, VIP, galanin, and other neuropeptides (23–26). Pain involves the tachykinins, notably substance P, for afferent transmission of pain to spinal and brain neurons, with modulation of the spinal–brain pain pathway by opioid neuropeptides including enkephalin and dynorphins (27,28). Multiple vasoactive peptide hormones including angiotensin (Ang), vasopressin, bradykinin (BK), and others regulate blood pressure conditions including hypertension (29–31). These examples illustrate that profiles of neuropeptides participate in regulating specific physiological functions.
Advantages of Mass Spectrometry-Based Neuropeptide Analyses Compared to Traditional Radioimmunoassay of Neuropeptides
The analysis of profiles of distinct groups of neuropeptides can be readily accomplished by mass spectrometry-based neuropeptidomics. Liquid chromatography separation of neuropeptides with online tandem mass spectrometry allows identification of hundreds to thousands of peptides in single experiments. In contrast, traditional antibody-based detection of neuropeptides can only obtain information of one neuropeptide in a single assay, resulting in lack of knowledge of profiles of neuropeptides. A key limitation of antibody-based radioimmunoassays (RIA) is that RIA detection of a peptide in the biological sample indicates that it is “related” to the standard peptide, but the peptide sequence structure of the detected peptide(s) is not defined by the RIA method since antibodies can detect several related peptides.
Significantly, directed mass spectrometry of selected neuropeptides provides definitive identification of the peptide of interest. Relative quantitation of the peptide can be achieved by “normalized spectral abundance” analyses or by isotopic labeling approaches (29,32–34). Such directed “multiple reaction monitoring” (MRM) of peptides is useful for designated peptides of biological interest. These neuropeptidomic and MRM mass spectrometry approaches provide identification and quantitation of defined peptide species in a single experiment, which is not possible with antibody-based approaches. The mass spectrometry-based neuropeptidomic approach provides knowledge of neuropeptide profiles that participate in biological regulation.
Importantly, most neuropeptides are active at very low concentrations in vivo (nanomolar range). Therefore, the high sensitivity of the mass spectrometry approach is advantageous for identifying neuropeptides. Application of mass spectrometry will likely lead to identification of numerous peptides generated from respective prohormone precursors. Subsequent biological analyses will then be needed to determine active peptides or inactive peptide products. Smaller inactive peptides may represent degradation of active neuropeptides; the spectrum of degradative pathways to inactivate neuropeptides has yet to be fully defined. Comparison of neuropeptide profiles under different cellular conditions in future studies can indicate how regulation of neuropeptide forms participates in health, disease, and drug treatment conditions.
NEUROPEPTIDOMICS DEMONSTRATE NOVEL PEPTIDES GENERATED FROM PROENKEPHALIN AND CHROMOGRANIN A IN DENSE CORE SECRETORY VESICLES
Proenkephalin-Derived Neuropeptides
The active pentapeptides (Met)enkephalin (ME) and (Leu)enkephalin (LE) are both derived from the proenkephalin (Fig. 2) precursor by proteolytic processing (9–11,15,16). Each proenkephalin precursor yields four copies of ME, one copy of LE, and the related peptides ME-Arg-Gly-Leu and ME-Arg-Phe. Studies of these active peptides have been based on their selection by bioassay and peptide sequencing of purified (Met)enkephalin (35) and deduced enkephalin-related peptides resulting from proteolytic processing of proenkephalin at dibasic residues.
Neuropeptidomic analyses by nano-liquid chromatography tandem mass spectrometry (LC-MS/MS) of neuropeptide-containing secretory vesicles isolated from a human pheochromocytoma revealed new information of how proenkephalin is processed (Fig. 3) (18). Peptides were analyzed in a low molecular weight pool of peptides less than 10 kDa (obtained by passage of the soluble fraction of isolated secretory vesicles through a 10-kDa Millipore membrane). Tandem mass spectrometry identified numerous extended forms of (Met)enkephalin that included “intervening” sequences of non-enkephalin domains. Also, intervening peptide sequences that did not include enkephalin sequences were identified. Yet, some intervening peptide domains were not detected (residues 145–161). These intervening peptide sequences have never been observed in prior studies.
Neuropeptidomic analyses of human proenkephalin-derived peptides in secretory vesicles. Neuropeptidomic studies investigated endogenous peptides derived from human proenkephalin in chromaffin secretory vesicles. Endogenous peptides derived from human PE in human adrenal medullary secretory vesicles (purified from human pheochromocytoma tissue) are illustrated with respect to their location within PE. Peptides were identified by ion-trap and QTOF MS/MS, combined with InsPecT (Ins) and Spectrum Mill (SM) bioinformatic analyses of MS/MS data at 1% false discovery rate (FDR; with the exception of (Leu)enkephalin that was indicated at 5% FDR) (18). Peptides identified under each of these conditions were mapped to PE, illustrated by colored lines: QTOF MS/MS data analyzed by InsPect (Ins, orange) or Spectrum Mill (SM, yellow) and ion-trap (Trap) analyzed by InsPect (Ins, green) or SM (olive). Within PE, the active enkephalin neuropeptides sequences are shown in yellow. Dibasic cleavage sites are highlighted by boxes; in addition, monobasic residues within PE are shown. (Hyphens at the end of some lines indicate peptides that were split between two lines in the figure.)
Chromogranin-A-Derived Neuropeptides
In secretory vesicles isolated from human pheochromocytoma, the chromogranin A (CgA) precursor undergoes proteolysis to generate an extensive group of peptide products (Fig. 4) (18,36). Neuropeptidomic data identified numerous CgA-derived peptide domains of catestatin, vasostatin, parastatin, and related neuropeptides derived by proteolytic processing of CgA (18). Interestingly, the low molecular pool of peptides less than 10 kDa contained several intervening peptides of CgA. But some intervening peptide domains of CgA were not detected. Thus, similar to peptides derived from proenkephalin, the presence of intervening peptide domains indicates products derived from proneuropeptide processing. In addition, the absence of particular intervening peptide products may indicate their presence only with the intact precursor or of high molecular weight intermediate fragments larger than 10 kDa, or as peptide domains that may have undergone extensive proteolysis.
Neuropeptidomic analyses of human chromogranin-A-derived peptides in secretory vesicles. Neuropeptidomics studies investigated endogenous peptides derived from human CgA in chromaffin secretory vesicles. CgA-derived peptides in human adrenal medullary secretory vesicles (purified from human pheochromocytoma tissue) identified in neuropeptidomic studies are illustrated within the CgA precursor (18). Peptides were identified (as described in legend of Fig. 3) by ion-trap and QTOF MS/MS, combined with InsPecT (Ins) bioinformatic analyses of MS/MS data at 1% FDR. Peptides identified under each of these conditions were mapped to CgA, illustrated by colored lines: QTOF MS/MS data analyzed by InsPect (Ins, purple) and ion-trap (Trap) MS/MS data analyzed by InsPect (Ins, olive green). Within CgA, names of known peptide sequences are indicated. Dibasic cleavage sites are highlighted by boxes
These neuropeptidomic results demonstrate that global, unbiased analyses of peptides by nano-LC-MS/MS indicates multiple, diverse peptides generated from proenkephalin. Ongoing (19–21) and future mass spectrometry analyses of prohormone precursors will reveal a multitude of previously unknown peptide products generated from such precursors. The diversity of neuropeptides implicates their broad biological activities.
PROFILING OF VASOACTIVE PEPTIDE HORMONES IN PLASMA BY MASS SPECTROMETRY REVEALS NOVEL ANTIHYPERTENSIVE DRUG MECHANISMS
Secretion of active neuropeptides participates in peptide hormone regulation of physiological systems in health and disease conditions. Profiling changes in the levels of functionally related peptide hormones is critical to understanding the effectiveness of therapeutic strategies. The high potency of low concentrations of plasma peptide hormones indicates the necessity for quantitative detection of low-abundance plasma peptides, with the challenge of their high turnover and regulated levels in physiological functions. Our recent studies demonstrate the use of nano-LC-MS/MS with stable isotope labeling and MRM analysis for measuring selected groups of vasoactive peptides consisting of angiotensin, bradykinin, and related peptide hormones (29). The effects of an inhibitor of angiotensin-converting enzyme (ACE) on the profile of these vasoactive peptides illustrated that the drug affects not only its ACE target for angiotensin production but also results in prominent alteration of the profiles of bradykinin and related peptides that regulate blood pressure (Fig. 5) (29). The ACE inhibitor, captopril, reduced plasma levels of several angiotensin-related peptides, as expected. The drug also resulted in substantial time-dependent increases in bradykinin and kallidin. Thus, the drug has widespread effects on profiles of vasoactive peptide hormones.
Multiple vasoactive peptide hormones regulated by ACE inhibitor drug therapeutics. The effects of an ACE inhibitor, captopril, on levels of plasma vasoactive peptides were analyzed in time course studies by nano-LC-MS/MS with quantitation using stable isotope-labeled internal peptide standards (29). ACE inhibitors are utilized as antihypertensive drugs. Chromatographic separation of target peptides and MRM provided quantitation of Ang I, Ang II, Ang1–7, BK 1–8, BK 2–9, and kallidin (KD). Results show significant reduction by the ACE inhibitor of the angiotensin peptides, with an interesting concomitant increase in plasma bradykinins and kallidin (potent vasodilators). The percent change in plasma concentration at different times after drug administration is shown in the table below the bar graph. Results illustrate the utility of simultaneous profiling of multiple peptides using mass spectrometry analysis to monitor drug-induced changes in vasoactive neuropeptides
Clearly, targeted peptidomic analyses to profile related peptides can enhance elucidation of the events regulating complex and dynamic physiological processes responding to therapeutic agents.
THE SECRETORY VESICLE PROTEOME FOR BIOSYNTHESIS OF ITS NEUROPEPTIDOME
The “neuropeptidome,” representing the profile of cellular neuropeptides, is produced from protein precursors within regulated secretory vesicles of neurons and endocrine cells. The “proteome” of these secretory vesicles is responsible for the biosynthesis, storage, and secretory release of the cellular “neuropeptidome.” The key segments of the secretory vesicle proteome responsible for biosynthesis of neuropeptidomes consist of the protease pathways utilized to convert proneuropeptides into active neuropeptides and protein systems required by secretory vesicles for vesicular trafficking, neuropeptide biosynthesis, signal transduction, and secretion of neuroeffectors.
Protease Pathways for Neuropeptidome Production
Chemical Biology for Activity-Based Identification of Proteases by Mass Spectrometry
Recent achievements in development of active site-directed affinity probes for proteases and other enzyme classes provide direct chemical labeling of proteases of interest in the biological system (37–40). Such activity-based probes that selectively label the main protease subclasses—cysteine, serine, metallo, aspartic, and threonine—provide advantageous chemical approaches for functional protease identification. Such chemical probes directed to cysteine proteases have been instrumental for identification of the new cathepsin L cysteine protease pathway for neuropeptide biosynthesis.
The activity probe DCG-04, the biotinylated form of E64c that inhibits cysteine proteases, was utilized for specific affinity labeling and mass spectrometry identification of the primary proenkephalin cleaving activity as cathepsin L (41–44). Confirmation of the localization of cathepsin L in neuropeptide-containing secretory vesicles was demonstrated by immunofluorescence confocal microscopy and immunoelectron microscopy (44–47). The key role of cathepsin L for neuropeptide biosynthesis has been illustrated by gene knockout, gene expression, and inhibitor studies. Results demonstrate the significant function of cathepsin L for producing neuropeptides including enkephalin, beta-endorphin, dynorphin, ACTH, α-melanocyte-stimulating hormone (α-MSH), cholecystokinin, NPY, and others (41–47). These findings suggested a new biological function for cathepsin L in secretory vesicles (in contrast to its known role in lysosomes) for producing the enkephalin and related neuropeptides.
Together with current knowledge in the field, prohormone processing utilizes two distinct protease pathways (Fig. 6) consisting of the cathepsin L cysteine protease pathway that includes Arg/Lys aminopeptidase (aminopeptidase B) and the well-known proprotein convertase (PC) family of subtilisin-like proteases (9–11) that process proneuropeptides with carboxypeptidase E. These protease pathways, and possibly others, generate neuropeptidomes of neuroendocrine cells.
Protease pathways for neuropeptide production. Distinct cysteine protease and subtilisin-like protease pathways participate in prohormone processing (9–11). The cysteine protease cathepsin L in secretory vesicles functions as a processing enzyme for the production of neuropeptides. Preference of cathepsin L to cleave at the NH2-terminal side of dibasic residue processing sites yields peptide intermediates with NH2-terminal residues, which are removed by Arg/Lys aminopeptidase that has been identified as aminopeptidase B. Cathepsin L also cleaves between the dibasic residues, resulting in peptide intermediates that then require both aminopeptidase and carboxypeptidase E to remove NH2-terminal and C-terminal basic residues to generate active neuropeptides. The subtilisin-like protease pathway involves the prohormone convertases PC1/3 and PC2. The PC enzymes preferentially cleave at the COOH-terminal side of dibasic processing sites, which results in peptide intermediates with basic residue extensions at their COOH termini that are removed by carboxypeptidase E
Proteomics of Secretory Vesicles for Neuropeptidome Biosynthesis and Secretion
Proteomic studies of neuropeptide-containing secretory vesicles can identify the functional protein categories in secretory vesicles utilized for neuropeptide production and secretion. Recent examination of proteins in model bovine chromaffin secretory vesicles revealed multiple functional protein categories that participate in secretory vesicle production of neuropeptides and catecholamines for cell–cell communication (Fig. 7) (48,49). Protein systems involved in vesicular neuropeptide biosynthesis were examined in proteomic studies of soluble and membrane fractions of dense core secretory vesicles purified from chromaffin cells of the sympathetic nervous system. Proteomic results revealed functional categories of prohormones, proteases, catecholamine neurotransmitter metabolism, protein folding, redox regulation, ATPases, calcium regulation, signaling components, exocytotic mechanisms, and related functions. Interestingly, these secretory vesicles contained an extensive number of GTP nucleotide-binding proteins related to Rab, Rho, and Ras signaling molecules (50,51), together with SNARE-related proteins and annexins that are involved in trafficking and exocytosis of secretory vesicle components (52,53). These vesicles also contain ATPases that regulate proton translocation (54), combined with components for signaling and exocytosis of neuropeptides. It will be of interest to compare the proteomics of neuropeptide secretory vesicles with that of synaptic vesicles that secrete classic small-molecule neurotransmitters (55–57).
The secretory vesicle proteome for neuropeptidome biosynthesis and secretion. Proteins of the secretory vesicle, known as the neuroproteome, participate in the biosynthesis, storage, and regulated secretion of neuropeptides, known as the neuropeptidome. Thus, the neuropeptidome is generated and secreted by the neuroproteome of regulated secretory vesicles. The neuroproteome consists of soluble and membrane proteins that participate in secretory vesicle functions for providing neuropeptides for cell–cell communication in the nervous and endocrine systems. Proteomic studies of the soluble and membrane fractions of neuropeptide secretory vesicles isolated from adrenal medullary chromaffin cells of the sympathetic nervous system (bovine) indicate the protein systems participating in neuropeptide production and secretion (illustrated in the pie charts) that include neuropeptides and neurohumoral factors, proteases, neurotransmitters enzymes and transporters, receptors, enzymes, carbohydrate functions, lipids, reduction oxidation, ATPases and nucleotide metabolism, protein folding, signal transduction and GTP-binding proteins, vesicular trafficking and exocytosis, structural proteins, and cell adhesion proteins (48,49)
Overall, knowledge of secretory vesicle proteomes provides novel insights into the biosynthesis and secretion of neuropeptidomes.
ADVANCES IN COMPUTATIONAL MASS SPECTROMETRY ARE NECESSARY FOR NEUROPEPTIDOMIC AND PROTEOMIC INVESTIGATIONS OF THE SYSTEMS BIOLOGY OF CELL–CELL COMMUNICATION
Computational mass spectrometry for bioinformatic analyses of mass spectrometry data is essential for identification and organization of peptidomic and proteomic components. Bioinformatics comprises the major effort for understanding mass spectrometry information. Importantly, analyses of neuropeptides and proteins subjected to mass spectrometry analyses each requires different bioinformatic approaches, as explained below.
Bioinformatics for Peptidomics
Mass spectrometry data of neuropeptidomes utilize bioinformatics that is distinguished from that used for protein identification. Unique features of peptides, compared to proteins, require distinct and appropriate bioinformatic tools for several reasons. Mass spectrometry analyses of neuropeptides depend entirely on the successful identification of a single peptide, which contrasts with protein identification by several peptide fragments generated by protease (typically trypsin) digestion. The neuropeptides of very short lengths of three to seven residues or of long lengths of more than 15–20 residues will benefit from the design of new algorithms for effective identification. Furthermore, whereas neuropeptides are nontryptic, many bioinformatic programs are designed for analyzing tryptic peptides for protein identification. Peptide identifications by search of protein databases are difficult because a substantial fraction of all short peptide sequences is present in the database, thus essentially reducing the search to de novo sequencing (a search in the space of all possible peptide sequences). In this case, the difficulty is that database search algorithms perform poorly whenever the database contains a substantial fraction of all possible peptides. Therefore, it will be necessary to improve search algorithms for peptide identifications. In addition, if the sequence database used is large, short peptides are likely to be present in the corresponding decoy database simply by chance, thus making peptide identification difficult. Significantly, structural complexity of neuropeptides is represented by posttranslational modifications including C-terminal amidation, acetylation, phosphorylation, sulfation, and other modifications (6,58). Thus, while some neuropeptides can be identified with current bioinformatic tools, complete neuropeptidomics will require the design of novel computational tools for identifying small neuropeptides from mass spectrometry data.
Bioinformatic analyses of neuropeptide sequences include genomic analyses for predictions of neuropeptide genes and peptide products from prohormones (59–64). Design of novel computational tools for predictions of neuropeptides from combined genomic and peptidomic sequences is an active area of algorithmic research in biomedical research. Such new bioinformatics tools will facilitate identification of neuropeptides by mass spectrometry, which will likely lead to discovery of previously unknown neuropeptides.
Bioinformatics for Proteomics
Investigation of proteomic data of secretory vesicles obtained from sympathoadrenal chromaffin cells has been achieved in our studies with bioinformatic analyses of mass spectrometry data by Spectrum Mill and Sequest (48,49). The identified proteins comprising these data were clustered to gain knowledge of the protein functional protein categories present in secretory vesicles for neuropeptide biosynthesis and regulated secretion, as illustrated in Fig. 7. These proteomic data indicate how these protein systems together coordinate secretory vesicle functions for neuropeptide production and release to mediate extracellular cell–cell communication.
CONCLUSION: NEUROPEPTIDOMICS AND PROTEOMICS FOR INVESTIGATION OF THE REGULATION OF NEUROPEPTIDES IN HEALTH, DISEASE, AND THERAPEUTICS
Regulation of brain and neuroendocrine functions utilizes profiles of neuropeptides that together function to mediate the complex cell–cell communication network among organ systems. Changes in physiological functions are represented by alterations in profiles of neuropeptides that can be investigated by neuropeptidomic approaches in health and disease. Furthermore, neuropeptidomics will be used for biomarker applications for monitoring disease status and the effectiveness of therapeutic agents. Significantly, because the secretory vesicle organelle produces neuropeptides, joint proteomic studies of this organelle provide knowledge of the key protein systems required for neuropeptide production. Elucidation of the neuropeptidomic systems, and their biosynthesis by the secretory vesicle proteome, can provide insight into new drug targets for novel disease therapeutics.
References
Holz RW, Fisher SK. Synaptic transmission and cellular signaling: an overview. In: Siegel GJ, editor. Basic neurochemistry. Philadelphia: Lippincott Williams & Wilkins; 1999. p. 191–212.
Law PY, Loh HH. Regulation of opioid receptor activities. J Pharmacol Exp Ther. 1999;289:607–24.
Norris D. Vertebrate endocrinology. San Diego: Academic; 1997.
Vaudry H. Brain peptides. In: Kastin AJ, editor. Handbook of biologically active peptides. Amsterdam: Elsevier; 2006. p. 621–828.
Krieger D, Brownstein MJ, Martin JB. Brain peptides. New York: Wiley-Interscience; 1983.
Siegel GJ, Agranoff BW, Albers RW, Fisher SK, Uhler MD. Peptides. In: Siegel GJ, editor. Basic neurochemistry. Philadelphia: Lippincott Williams & Wilkins; 1999. p. 363–82.
Nacham RJ. Invertebrate peptides. In: Kastin AJ, editor. Handbook of biologically active peptides. Amsterdam: Elsevier; 2006. p. 157–260.
(No authors) Proceedings of invertebrate neuropeptide conference 2009 (INC2009). Jan. 11–15, 2009, Khajuraho, India. 2009. Peptides 2009;31:371–524.
Hook V, Funkelstein L, Lu D, Bark S, Wegrzyn J, Hwang SR. Proteases for processing proneuropeptides into peptide neurotransmitters and hormones. Annu Rev Pharmacol Toxicol. 2008;48:393–423.
Zhou A, Webb G, Zhu X, Steiner DF. Proteolytic processing in the secretory pathway. J Biol Chem. 1999;274:20745–8.
Seidah GN, Chretien M. Proprotein and prohormone convertases: a family of subtilases generating diverse bioactive polypeptides. Brain Res. 1999;8484:45–62.
Higuchi H, Yang HY, Sabol SL. Rat neuropeptide Y precursor gene expression. mRNA structure, tissue distribution, and regulation by glucocorticoids, cyclic AMP, and phorbol ester. J Biol Chem. 1988;263:6288–95.
Rokaeus A, Brownstein MJ. Construction of a porcine adrenal medullary cDNA library and nucleotide sequence analysis of two clones encoding a galanin precursor. Proc Natl Acad Sci USA. 1986;83:6287–91.
Robinson BG, D'Angio Jr LA, Pasieka KB, Majzoub JA. Preprocorticotropin releasing hormone: cDNA sequence and in vitro processing. Mol Cell Endocrinol. 1989;61:175–80.
Yoshikawa K, Williams C, Sabol S. Rat brain preproenkephalin mRNA, cDNA cloning, primary structure, and distribution in the central nervous system. J Biol Chem. 1984;259:14301–8.
Comb M, Rosen H, Seeburg P, Adelman J, Herbert E. Primary structure of the human proenkephalin gene. DNA. 1983;2:213–29.
Roberts JL, Seeburg PH, Shine J, Herbert E, Baxter JD, Goodman HM. Corticotropin and beta-endorphin: construction and analysis of recombinant DNA complementary to mRNA for the common precursor. Proc Natl Acad Sci USA. 1979;76:2153–7.
Gupta N, Bark SJ, Lu WD, Taupenot L, O’Connor DT, Pevzner P, and Hook V. Mass spectrometry-based neuropeptidomics of secretory vesicles reveals novel properties of post-translational processing for neuropeptide biosynthesis. J Proteome Res. 2010 (in press).
Bora A, Annagudi SP, Millet LJ, Rubakhin SS, Forbes AJ, Kelleher NL, et al. Neuropeptidomics of the supraoptic nucleus. J Proteome Res. 2008;7:4992–5003.
Perkel JM. Neuropeptidomics study profiles hypothalamic ‘nucleus’, individual cells. J Proteome Res. 2008;7:4610.
Fricker LD. Neuropeptidomics to study peptide processing in animal models of obesity. Endocrinology. 2007;148:4185–90.
Amare A, Hummon AB, Southey BR, Zimmerman TA, Rodriguez-Zas SL, Sweedler JV. Bridging neuropeptidomics and genomics with bioinformatics: prediction of mammalian neuropeptide prohormone processing. J Proteome Res. 2006;5:1162–7.
Goldstein DS, Kopin IG. Adrenomedullary, adrenocortical, and sympathoneural responses to stressors: a meta-analysis. Endocr Regul. 2008;42:111–9.
Hook V, Toneff T, Baylon S, Sei C. Differential activation of enkephalin, galanin, somatostatin, NPY, and VIP neuropeptide production by stimulators of protein kinases A and C in neuroendocrine chromaffin cells. Neuropeptides. 2008;42:503–11.
Whitworth EJ, Kosti O, Renshaw D, Hinson JP. Adrenal neuropeptides: regulation and interaction with ACTH and other adrenal regulators. Microsc Res Tech. 2003;61:259–67.
Nankova BB, Sabban EL. Multiple signaling pathways exist in the stress-triggered regulation of gene expression for catecholamine biosynthetic enzymes and several neuropeptides in the rat adrenal medulla. Acta Physiol Scand. 1999;167:1–9.
Otsuka M, Yanagisawa M. Pain and neurotransmitters. Cell Mol Neurobiol. 1990;10:293–302.
Wilson PR, Yaksh TL. Pharmacology of pain and analgesia. Anaesth Intensive Care. 1980;8:248–56.
Lortie M, Bark S, Blantz R, Hook V. Detecting low abundance vasoactive peptides in plasma: progress towards absolute quantitation using nano liquid chromatography–mass spectrometry. Anal Biochem. 2009;394:164–70.
Molinaro G, Rouleau JL, Adam A. Vasopeptidase inhibitors: a new class of dual zinc metallopeptidase inhibitors for cardiorenal therapeutics. Curr Opin Pharmacol. 2002;2:131.
Ardaillou R. Active fragments of angiotensin II: enzymatic pathways of synthesis and biological effects. Curr Opin Nephrol Hypertens. 1997;6:28–34.
Florens L, Swanson CMJ, SK FM, Coleman MK, Workman JL, Wahsburn MP. Analyzing chromatin remodeling complexes using shotgun proteomics and normalized spectral abundance factors. Methods. 2006;40:303–11.
Bark SJ, Lu WD, Hook V. Linear and accurate quantitation of proenkephalin-derived peptides by isotopic labeling with internal standards and mass spectrometry. Anal Biochem. 2009;389:18–26.
Bantscheff M, Schirie M, Swwetman G, Rick J, Kuster B. Quantitative mass spectrometry in proteomics: a critical review. Anal Bioanal Chem. 2007;389:1017–31.
Hughes J, Smith TW, Kosterlitz HW, Fothergill LA, Morgan BA, Morris HR. Identification of two related pentapeptides from the brain with potent opiate agonist activity. Nature. 1975;17:91–6.
Lee JC, Hook V. Proteolytic fragments of chromogranins A and B represent major components of chromaffin granules, illustrated by 2-D proteomics with NH2-terminal Edman peptide sequencing and MALDI-TOF MS. Biochemistry. 2009;48:5254–62.
Barglow KT, Cravatt BF. Activity-based protein profiling for the functional annotation of enzymes. Nat Method. 2007;4(10):822–7.
Evans MJ, Cravatt BF. Mechanism-based profiling of enzyme families. Chem Rev. 2006;106:3279–301.
Yuan F, Verhelst SH, Blum G, Coussens LM, Bogyo M. A selective activity-based probe for the papain family cysteine protease dipeptidyl peptidase I/cathepsin C. J Am Chem Soc. 2006;128(17):5616–7.
Kato D, Boatright KM, Berger AB, Nazif T, Blum G, Ryan C, et al. Activity-based probes that target diverse cysteine protease families. Nat Chem Biol. 2005;1(1):33–8.
Hook V, Yasothornsrikul S, Greenbaum D, Medzihradszky KF, Troutner K, Toneff T, et al. Cathepsin L and Arg/Lys aminopeptidase: a distinct prohormone processing pathway for the biosynthesis of peptide neurotransmitters and hormones. Biol Chem. 2004;385:473–80.
Schiller MR, Mende-Mueller L, Moran K, Meng M, Miller KW, Hook VY. “Prohormone thiol protease” (PTP) processing of recombinant proenkephalin. Biochemistry. 1995;34:7988–95.
Yasothornsrikul S, Aaron W, Toneff T, Hook VY. Evidence for the proenkephalin processing enzyme prohormone thiol protease (PTP) as a multicatalytic cysteine protease complex: activation by glutathione localized to secretory vesicles. Biochemistry. 1999;38:7421–30.
Yasothornsrikul S, Greenbaum D, Medzihradszky KF, Toneff T, Bundey R, et al. Cathepsin L in secretory vesicles functions as a prohormone-processing enzyme for production of the enkephalin peptide neurotransmitter. Proc Natl Acad Sci USA. 2003;100:9590–5.
Funkelstein L, Toneff T, Hwang SR, Beuschlein F, Lichtenauer UD, Reinheckel T, et al. Major role of cathepsin L for producing the peptide hormones ACTH, β-endorphin, and α-MSH, illustrated by protease gene knockout and expression. J Biol Chem. 2008;83:35652–9.
Minokadeh A, Funklestein L, Toneff T, Hwang SR, Reinheckel T, Peters C, et al. Cathepsin L participates in dynorphin neuropeptide production in brain cortex, illustrated by protease gene knockout and expression. Mol Cell Neurosci. 2010;43:98–107.
Funkelstein L, Toneff T, Hwang SR, Reinheckel T, Peters C, Hook V. Cathepsin L participates in the production of neuropeptide Y in secretory vesicles, demonstrated by protease gene knockout and expression. J Neurochem. 2008;106:384–91.
Wegrzyn J, Lee J, Neveu JM, Lane WS, Hook V. Proteomics of neuroendocrine secretory vesicles reveal distinct functional systems for biosynthesis and exocytosis of peptide hormones and neurotransmitters. J Proteome Res. 2007;6(5):1652–65.
Wegrzyn JL, Bark SJ, Yap A, Hook V. Proteomics of dense core secretory vesicles reveal distinct protein categories for secretion of neuroeffectors for cell-cell communication: relevance to health and disease. J Proteome Res. 2010 (in press).
Pfeffer S, Aivazian D. Targeting RAB GTPases to distinct membrane compartments. Nat Rev Mol Cell Biol. 2004;5:886–96.
Colicelli J. Human RAS superfamily proteins and related GTPases. Sci STKE. 2004;250:RE13.
Ungar D, Hughson FM. SNARE protein structure and function. Annu Rev Cell Dev Biol. 2003;19:493–517.
Gerke V, Moss SE. Annexins: from structure to function. Physiol Rev. 2002;82:331–71.
Taupenot L, Harper KL, O’Connor DT. Role of H+-ATPase-mediated acidification in sorting and release of the regulated secretory protein chromogranin A: evidence for a vesiculogenic function. J Biol Chem. 2005;280:3885–97.
Brunner Y, Schvartz D, Couté Y, Sanchez JC. Proteomics of regulated secretory organelles. Mass Spectrom Rev. 2009;28:844–67. Review.
Gilchrist A, Au CE, Hiding J, Bell AW, Fernandez-Rodriguez J, Lesimple S, et al. Quantitative proteomics analysis of the secretory pathway. Cell. 2006;127:1265–81.
Chen X, Walker AK, Strahler JR, Simon ES, Tomanicek-Volk SL, Nelson BB, et al. Organellar proteomics: analysis of pancreatic zymogen granule membranes. Mol Cell Proteomics. 2006;5:306–12.
Kastin AJ. Handbook of biologically active peptides. Amsterdam: Elsevier; 2006. p. 1–1479.
Clynenm E, Liu F, Husson SJ, Landuyt B, Hayakawa E, Baggerman G, et al. Bioinformatic approaches to the identification of novel neuropeptide precursors. Methods Mol Biol. 2010;615:357–74.
Burbach JP. Neuropeptides from concept to online database www.neuropeptides.nl. Eur J Pharmacol. 2010;626:27–48.
Southey BR, Rodriguez-Zas SL, Sweedler JV. Characterization of the prohormone complement in cattle using genomic libraries and cleavage prediction approaches. BMC Genomics. 2009;10:228.
Sonmez K, Zaveri NT, Kerman IA, Burke S, Neal CR, Xie X, et al. Evolutionary sequence modeling for discovery of peptide hormones. PLoS Comput Biol. 2009;5:e1000258.
Castellana NE, Payne SH, Shen Z, Stanke M, Bafna V, Briggs SP. Discovery and revision of Arabidopsis genes by proteogenomics. Proc Natl Acad Sci USA. 2008;105:21034–8.
Gupta N, Benhamida J, Bhargava V, Goodman D, Kain E, Nguyen N, et al. Comparative proteogenomics: combining mass spectrometry and comparative genomics to analyze multiple genomes. Genome Res. 2008;18:1133–42.
Acknowledgments
This work was supported by grants from the National Institute of Health (NIH grants R01DA04271, R01MH077305, R01NS24553, and P01HL58120) and the Alzheimer’s Association to V. Hook, NIH grants (R01RR016522 and P41RR024851) to P. Pevzner, and NIH grant P01 HL58120 to D. O’Connor. The authors thank Thomas Toneff for technical assistance.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Guest Editors: Rao S. Rapaka, Lloyd D. Fricker, and Jonathan V. Sweedler
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Hook, V., Bark, S., Gupta, N. et al. Neuropeptidomic Components Generated by Proteomic Functions in Secretory Vesicles for Cell–Cell Communication. AAPS J 12, 635–645 (2010). https://doi.org/10.1208/s12248-010-9223-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1208/s12248-010-9223-z