Abstract
Background
Antibiotic resistance is on the rise, and new antibiotic research has slowed in recent years, necessitating the discovery of possibly novel microbial resources capable of producing bioactive compounds. Microbial infections are gaining resistance to existing antibiotics, emphasizing the need for novel medicinal molecules to be discovered as soon as possible. Because the possibilities of isolating undiscovered actinomycetes strains have decreased, the quest for novel products has shifted to rare actinomycetes genera from regular environments or the identification of new species identified in unusual habitats.
Main body of the abstract
The non-streptomyces actinobacteria are known as rare actinomycetes that are extremely difficult to cultivate. Rare actinomycetes are known to produce a variety of secondary metabolites with varying medicinal value. In this review, we reported the diversity of rare actinomycetes in several habitat including soil, plants, aquatic environment, caves, insects and extreme environments. We also reported some isolation methods to easily recover rare Actinobacteria from various sources guided with some procedures to identify the rare Actinobacteria isolates. Finally, we reported the biosynthetic potential of rare actinomycetes and its role in the production of unique secondary metabolites that could be used in medicine, agriculture, and industry. These microbial resources will be of interest to humanity, as antibiotics, insecticides, anticancer, antioxidants, to mention but a few.
Short conclusion
Rare actinomycetes are increasingly being investigated for new medicinal compounds that could help to address existing human health challenges such as newly emerging infectious illnesses, antibiotic resistance, and metabolic disorders. The bioactive secondary metabolites from uncommon actinomycetes are the subject of this review, which focuses on their diversity in different habitats, isolation, identification and biosynthetic potentials.
Similar content being viewed by others
Background
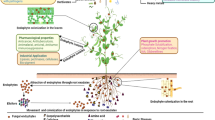
Actinomycetes have long been recognized as a top source of biopharmaceuticals, particularly antibiotics [1, 2]. Gram-positive filamentous bacteria with a high G + C concentration are known as actinomycetes [3]. They are a key part of microbial diversity and have been found in a variety of habitats and unique settings. Rare actinomycetes are a group of actinomycetes whose isolation frequency is significantly lower than that of streptomyces strains obtained using traditional procedures [4]. Isolating and cultivating them is challenging. Due to their ability to produce a large variety of structurally diverse natural compounds with unusual bioactivity, these microbial groups from underexplored habitats are being studied in drug development [5]. They are found in a variety of habitats, including soil, aquatic, mangrove, desert, mountains, and plants, and account for around 10% of all isolated actinomycetes. They have shown to be an excellent and exciting source of novel and potent bioactive compounds [6]. Efforts in the past and present to isolate uncommon actinomycetes from underexplored diverse natural settings have resulted in the isolation of over 220 rare actinomycetes genera, with more than 50 taxa producing 2500 bioactive compounds [5]. This number accounts for more than a quarter of all actinomycetes metabolites, indicating that selective isolation techniques are being developed and widely used. This review updates all selected isolation medium, including pretreatment and enrichment procedures for the isolation of rare actinomycetes, to aid in that discovery. It reveals several processes toward the discovery of novel anti-microbial compounds from rare actinomycetes (Fig. 2). Furthermore, this research reveals that rigorous efforts in isolating and screening rare genera of actinomycetes from new and underexplored habitat can increase the discovery of new compounds with novel scaffolds. To address the rising number of antibiotic-resistant pathogenic bacteria, new antibiotics are critically needed. Natural products continue to be the most potential source of new antimicrobials and bioactive compounds. Actinobacteria are well-known for being prolific makers of natural bioactive substances. Intensive efforts in isolating and screening rare genera of microorganisms are thought to boost the chances of identifying a new drug with a novel chemical structure. One strategy to break into novel bioactive chemical discovery is to screen rare actinomycetes and their hitherto underrepresented genera from unfamiliar settings in natural product screening collections [4]. The importance of unusual actinomycetes in this regard can also be shown in the fact that they produce several of the most effective antibacterial drugs now on the market. We want to refresh our understanding of the potential of rare actinomycetes by focusing on their biodiscovery potential; therefore, we want to give the reader a quick overview of the bioactive compounds from unusual actinomycetes. New compounds identified from these microbes with bioactive potential are the focus. Actinomycete strains that are difficult to identify are of particular interest to researchers. As a result, providing access to rare actinomycete strains with a high potential for producing novel bioactive compounds is of great importance [7].
The so-called "rare actinomycetes" are rather numerous in many habitats, according to molecular tools, and can be retrieved in large numbers using an appropriate isolation procedure [8]. We expect that investigating unusual actinomycetes that are difficult to isolate will yield a variety of beneficial compounds [9]. The distribution of rare actinomycetes is influenced by a variety of parameters such as habitat type, ambient pH, and nutrient content [6]. The following genera are rare actinobacteria: Gordonia, Isoptericola, Jiangella, Knoellia, Kocuria, Krasilnikoviella, Kribbella, Actinocorallia, Actinomadura, Agromyces, Alloactinosynnema, Amycolatopsis, Beutenbergia, Cellulosimicrobium, Gordonia, Isoptericola, Jiangella, Knoellia, Kocuria, Krasilnikoviella Nocardia, Nocardioides, Nocardiopsis, Nonomuraea, Oerskovia, Pseudokineococcus, Pseudonocardia, Rhodococcus, Saccharothrix, Streptosporangium, and Tsukamurella [10].
It is challenging to isolate unusual actinomycetes using traditional dilution plate procedures. Isolation, preservation, and cultivation are all demanding procedures. The reason for this is that they are frequently obscured by fast-growing organisms including bacteria, fungus, and common Streptomyces [11]. Pretreatments such as dry heat, calcium carbonate, phenol, thermal, microwave, and sonication are required for the isolation of uncommon actinobacteria. One or more of these are done before plating the sample on appropriate media such as humic acid agar with vitamins (HVA) and oatmeal agar (ISP3), with 50 mg/L nalidixic acid and 100 mg/L of cycloheximide incubating at 30 °C for at least 7 days [12, 13]. These treatments remove non-filamentous bacteria from samples and restrict fungal growth, allowing slow-growing uncommon actinomycetes to thrive [12]. For fostering the growth of rare actinomycetes while suppressing bacterial and fungal contamination, appropriate selective media containing macromolecules such as casein, chitin, and humic acid are essential.
Diverse habitats for sourcing rare actinomycetes
Soil and plants
Actinomycetes populations have been thoroughly investigated in soil, and the majority of the rare actinomycetes reported so far have come from various types of soil [6]. Table 2 shows that the isolation of several novel and rare taxa mentioned in this analysis came from a variety of soil types. Many unusual actinomycetes are now being isolated from plants [14, 15], often to uncover new microbial resources for screening of potential bioactive compounds [16]. Endophytic habitats were used to isolate Saccharopolyspora, Dietzia, Blastococcus, Dactylosporangium, Promicromonospora, Oerskovia, Actinocorallia, and Jiangella species [17]. Endophytic Actinomycetes, such as the Frankia genera, can fix nitrogen, which is an important function in ecological systems [18]. Rare actinomycetes belonging to the Micromonospora, Microbispora, Actinoplanes, and Streptosporangium genera have been isolated consistently from numerous Korean soils [4].
Extreme environments
High and low temperatures, salt, alkaline and acidic pH, radioactivity, and high pressure are all examples of unique growth conditions found in extreme habitats. Microorganisms from harsh habitats have gotten a lot of attention because of their unique processes for adapting to their extreme surroundings and their ability to create uncommon bioactive compounds [19]. Despite the interest, actinomycetes that live in harsh settings have yet to be extensively studied since the discovery of pioneer Actinopolyspora halophila by chance [5]. Researchers have been looking for unusual actinomycetes in a variety of habitats, including salt soil, alkaline soil, salty seas, and the ocean [20]. Researchers have isolated Naxibacter, Actinopolyspora, Amycolatopsis, Citricoccus, Halomonas, Isoptericola, Jonesia, Kocuria, Kribbella, Liuella, Marinococcus, Massilia, Microbacterium, Nesterenkonia, Nocardia, Nocardiopsis, Prauserella, Rhodococcus, Saccharomonospora, Saccharopolyspora, Sphingomona from extreme environments [19]. Rare halophilic actinomycetes, such as Nocardiopsis strains, have been found to contain a high frequency of non-ribosomal peptide synthase (NRPS) genes, which could be linked to their great capacity for synthesizing huge numbers of physiologically active compounds [19].
Caves
Caves offer low nutrition, temperature, and light intensity as a microbiological environment, but high humidity [21]. These conditions may increase competition, which could boost the development of antibiotics and hydrolytic enzymes that stop other microbes from growing [22]. Spirillospora, Nonomuraea, Catellatospora, Nonomuraea, Micromonospora, isolated members of the Actinomadura, Saccharopolyspora, Actinoplanes, Gordonia, Microbispora, Micromonospora, Nocardia, and Nonomuraea, among others, have been isolated from caves. These findings support the idea that caves could be rich in rare actinomycetes that produce new compounds [22,23,24,25,26].
Insects and birds
The insect kingdom is yet another uncharted territory for discovering unique and unusual actinomycetes [27]. Some insects, such as Pseudonocardia and Amycolatopsis, kill weeds due to their natural ability to produce antimicrobials through a symbiotic interaction with actinomycete bacteria [28]. Insect-associated actinomycetes have been found to produce a few numbers of antifungal compounds. Pseudonocardia species isolated from lower attines Apterostigma dentigerum produced dentigerumycin, whereas Streptomyces species isolated from higher attine ants belonging to the genus Acromyrmex produced candicidin, a well-known antifungal [29, 30]. Antifungal activity was also observed in Pseudonocardia isolated from Acromyrmex octospinosus, although no antifungal compounds have been extracted or identified [29]. A Pseudonocardia species was recently discovered in the ant Acromyrmex octospinosus that produced a unique polyene antifungal metabolite [31]. Switching the search from explored to undiscovered areas could boost the discovery of new bioactive compounds [32]. Streptosporangium, Actinomadura, Saccharopolyspora, Thermoactinomyces, and Nocardia have recently been isolated from soils in the nests of solitary wasps and swallow birds [33]. Insects and birds are quickly becoming important sources for finding unique and novel bioactive compounds in Actinomycetes.
Aquatic habitat
In rivers, lakes, oceans, and marine habitats, rare actinomycetes are common. Actinoplanes with sporangium and zoospores will grow in moist environments and survive in dry environments as spores. Micromonospora spp. is a naturally occurring bacterium found in freshwater lakes and mud that can be isolated from lake sediments. Representatives of Thermoactinomyces, Streptomyces, and Rhodococcus have been found to be predominantly isolated from aquatic habitats, according to researchers [34]. Actinoplanes, Actinomadura, Microbispora, Micropolyspora, Microtetraspora, Mycobacterium, Nocardiopsis, Nocardia, Promicromonospora, Rhodococcus, Saccharomonospora, Saccharopolyspora, Streptosporangium, Thermoactinomyces, Thermomonospora, and Thermopolyspora are examples of rare genera of actinomycetes isolated from aquatic habitat [35].
Pretreatment of samples for isolation of rare actinomycetes
The discovery of humic acid vitamin agar (HVA) was a watershed moment in the isolation of uncommon actinomycetes. It is made entirely of soil humic acid, which is an excellent source of carbon and nitrogen for recovering rare actinomycetes from natural samples. Although humic acid is a highly heterogeneous cross-linked polymer that resists biological degradation and inhibits the formation of non-filamentous bacteria colonies, it is an exceptionally heterogeneous cross-linked polymer [4]. To limit duplication of isolation, different natural samples used for the isolation of unusual actinomycetes are frequently treated before the isolation to remove common actinomycetes like streptomyces and undesirable bacteria. For the isolation of rare actinomycetes from samples, a variety of pre-treatment methods and isolation media (Table 1) are used, including dilution and mixing with sterile natural decoction water from plant samples, seawater [36], artificial seawater, saline solution, and deionized/distilled water supplemented with NaCl for sea or marine sediment samples [37, 38]. A variety of pre-treatment procedures have been used to isolate uncommon actinomycetes selectively. Most researchers use drying and moist heating of sample materials [39], because actinomycetes spores are resistant to desiccation and heating, they can be used to screen against Gram-positive bacteria [39]. Because actinomycetes' spores are resistant to a variety of substances, including benzethonium chloride, chlorhexidine gluconate, phenol, sodium dodecyl sulfate, and antibiotics, they are commonly used to isolate actinomycetes. These compounds can reduce or prevent the growth of aerobic Gram-negative bacteria, endospore-forming bacilli, and pseudomonads when treated with the samples for 30 min, improving the chances of isolating actinomycetes selectively [40]. The following sub-headings are used to discuss these pre-treatment techniques:
Heat treatments
Most researchers propose using these pretreatment processes (wet and dry heat) in combination with selected isolation media for the selective isolation of novel and rare actinomycetes [4]. Most actinomycete genera' airborne spores are resistant to desiccation and have a significantly higher resilience to wet or dry heat than their vegetative hyphae [4]. The growth of Streptosporangium spp. is considerably aided by a dry heat treatment (120 °C for 1 h) of natural samples. Following surface sterilization and continuous drying at 100 °C for 15 min before directly plating on different selective media, numerous strains belonging to the genera Pseudonocardia, Nocardiopsis, Micromonospora, Microbispora, Acitinomadura and Streptosporangium were isolated [17]. Dry heating of samples treated with chemicals like 0.01 percent benzethonium chloride, 0.03 percent chlorhexidine gluconate, 0.05 percent sodium dodecylsulfate (SDS), 6 percent yeast extract, and 1.5 percent phenol and supplemented with different selective antibiotics like leucomycin and nalidixic acid on HVA has greatly increased the selectivity of rare actinomycetes [6, 41]. Pretreatment with moist (50 °C for 6 min) and dry (120 °C for 1 h) heating and 1.5 percent phenol reduced the quantity of unwanted bacteria and improved the selective separation of Actinoplanes, Actinomadura, Saccharopolyspora, Gordonia, Microbispora, Micromonospora, Nocardia, and Nonomuraea [26].
Phenol treatment
Alternative approaches for the selective isolation of uncommon actinomycetes include adding chemicals such as phenol to natural samples [41]. Because 1.5 percent phenol is poisonous to bacteria, fungus, and streptomycetes, it increases the chances of isolating rare actinobacteria. As a result, 1.5 percent phenol treatment reduces the quantity of such organisms by removing sensitive species [42]. By pretreating samples with 1.5 percent phenol and then plating on HVA, several non-streptomycetes, including the rare genera Actinomadura, Microbispora, Micromonospora, Nocardia, Polymorphospora, and Nonomurea, were isolated [41, 43].
Selective antimicrobial agents
Several rare actinomycetes are resistant to a wide spectrum of antibiotics. Thus, several antibiotic molecules have been used in selective media to inhibit the competing bacteria including fast-growing actinomycetes. Selective isolation plates containing novobiocin significantly increased the numbers of Micromonospora-like colonies while gentamicin is also one of the selective agents used to access Micromonospora spp. [44]. Isolating media are mostly modified with nalidixic acid (50 mg liter−1) and nystatin (100 mg liter−1) to suppress the growth of Gram-negative bacteria and fungi [17].
Calcium carbonate treatment
The use of calcium carbonate to treat natural habitat samples enhanced the populations of rare actinomycetes genera [45]. Although the process is unknown, researchers discovered that mixing natural samples with calcium carbonate powder alters the pH in favor of actinomycete propagule growth, and the presence of calcium ions encourages the development of aerial mycelia in actinomycetes [46]. Actinokineospora spp., Saccharopolyspora, Dietzia, Blastococcus, Dactylosporangium, Promicromonospora, Oerskovia, Actinocorallia, and Jiangella species have all been successfully isolated using a combination of calcium carbonate rehydration and centrifugation [46, 47]. For the isolation of rare actinomycetes genera from natural samples, a combination of the calcium carbonate process and additional selective isolation procedures is usually recommended [45]
Microwave irradiation
The usage of microwave energy is commonly used to sterilize soil [48]. Total fungal and total prokaryote counts in soil extracts were lowered after microwave irradiation [49]. Micromonospora, Micropolyspora, Norcardia, Actinomadura, Streptosporangium, and Lentzea spp. are among the rare actinomycetes that have been isolated by microwave irradiation [48, 49]. Other physical agents are used to isolate rare actinomycetes in a selective manner. Electric pulses, electromagnetic radiation, super high frequency radiation, ultrasonic waves, and extremely high frequency radiation are some of the methods used [26, 50, 51]. The use of these techniques has resulted in a large rise in the overall number of isolated uncommon actinomycetes.
Centrifugation method
Another physical method is centrifugation, which removes Streptomycetes and other non-motile Actinomycetes from the liquid phase, allowing for the selective growth of rare motile actinomycetes [46, 52]. Endophytic uncommon actinobacteria Pseudonocardia, Nocardiopsis, Micromonospora, Amycolatopsis, Nocardia, Nonomuraea, Actinomadura, Gordonia, Promicromonospora, and Mycobacterium species were isolated using a combination of enzymatic hydrolysis and differential centrifugation [53]
Chlorination and chemo-attractants
Selective isolation of sporulating actinomycetes known to produce motile spores can be done using xylose, chloride, γ-collidine, bromide and vanillin which act as chemo-attractants for accumulating spores of rare actinomycetes such as Actinoplanes, Dactylosporangium and Catenuloplanes [6]. The use of chloramine treatment has been used to selectively isolate rare genera Herbidospora, Microbispora, Microtetraspora and Streptosporangium. This is because chlorination is believed to suppress growth of contaminant bacteria but promote the growth of rare actinomycetes upon plating on humic acid vitamin media [6, 54]. Generally, rare actinomycetes are selectively isolated from natural habitats using combined physical and chemical treatments [45]. Several new Actinobacteria species are recovered from different sources using various media types (Table 1).
Isolation of rare actinomycetes
Collected samples (soil, marine sediment, plant parts) undergo series of pretreatments to promote the possibility of isolating rare actinomycetes and suppress the growth of often isolated streptomyces [96]. These physical and chemical pretreatments include the use of dry heat, phenol treatments, sucrose gradient centrifugation and sodium dodecyl sulfate treatment [42, 97]. In case of isolating endophytic actinobacteria, plant samples are subjected to surface sterilization and are fragmented (8 × 8 mm) before deposition onto petri dishes containing the isolation media [98, 99]. Starch casein agar (SCA) and humic acid vitamin agar (HVA) supplemented with nalidixic acid (50 μg/mL) and cycloheximide (100 μg/mL) are mostly employed for selective isolation of rare actinomycetes [99]. The media are supplemented with a pinch of nalidixic and cycloheximide to inhibit unwanted bacterial and fungal contamination, respectively. An aliquot of 0.1 ml sample would be serially diluted up to 10–9 and a pour plate technique would be performed and incubated for 30 days at 28 °C and would be examined daily for the presence of colonies. The actinomycetes colonies are mostly identified by their chalky, powdery colonies and leathery texture [100]. These colonies would be sub-cultured and maintained at 4 °C for further characterization. It is well established that several other antimicrobial agents such as anisomycin, gentamicin, kanamycin, novobiocin, nystatin, penicillin, primaricin, polymyxin, rifampicin, streptomycin, tunicamycin and vancomycin can also be added to the isolation media to promote the selective isolation of rare actinobacteria [54, 101].
Morphological identification of actinomycetes
Different culture media are employed to assess the macro-morphological characteristics of actinomycetes. These include: Agar yeast-malt extract (ISP2); Oatmeal Agar (ISP3); Agar Starch and inorganic salts (ISP4); Glycerol Asparagine Agar (ISP5), Soya bean meal agar, Glucose -Yeast Malt extract agar, Czapeks agar, Luria Bertani Agar (LBA), Starch casein agar and nutrient agar [102]. Each media would be sterilized, poured into sterile petri dishes and then left to solidify. Each strain would be aseptically streaked on the media surface and incubated at 28–30 °C for 7–21 days. The morphological characteristics to be examined among isolates include their color or soluble pigment, surface morphology, type of aerial hyphae, formation of aerial and substrate mycelia. These features are observed and compared using colour chart [102].
Microscopic characterization and biochemical tests for identification of actinomycetes
There are several microscopic and biochemical tests that are employed in identification of actinobacteria. They include Gram staining, starch hydrolysis test, casein hydrolysis test, urea hydrolysis test, lipase test, gelatin hydrolysis test, salt tolerance test, oxidase test, milk coagulation and peptonization test [103]. Most biochemical tests investigate the ability of the actinobacteria to produce different enzymes [104,105,106]. For example, coagulation and peptonization of milk test investigate the ability of the actinobacteria to produce protease enzyme, starch hydrolysis investigates their ability to produce certain exoenzymes like α-amylase and oligo-1,6-glucosidase while cellulose hydrolysis test checks the ability of actinobacteria to produce cellulase enzyme [107, 108].
Molecular and species level characterization
Sequel to morphological, microscopic and biochemical characterization, the isolated actinobacterial strains are subjected to species level identification done by 16S rRNA gene sequencing. The genomic DNA would be extracted using DNA extraction kit and the 16S rRNA gene amplified using pair of primers like (27F, 5′-AGAGTTTGATCMTGGCTCAG-3′; 1492R, 5′-GGTTACCTTGTTACGACT T-3′) and 9F(5′GAGTTTGATCCTGGCTCAG3′); 1541R (5′AAGGAGGTGATCCAGCC3′) [109, 110]. The amplified fragment for each strain would be sequenced utilizing the primers (forward and reverse). High-quality sequences would be assembled to produce the partial 16S rRNA contig for each strain. National Center for Biotechnology Information (NCBI) server are used to check the similarity for each contig against the available 16S rRNA genes data to determine the closest homologs. The homology search can be performed by comparing the sequence with thus present in the public database (NCBI) using the standard Basic Local Alignment Search Tool (BLAST) program. The 16S rRNA gene sequence of the selected strains would be submitted in the NCBI database to get GenBank accession numbers. For phylogenetic analysis, a neighbour joining tree based on the 16S rRNA gene sequences of the actinobacterial strains and their closely related type strains would be constructed at 1000 bootstrap replicates using by Molecular Evolutionary Genetic Analysis (MEGA) software [111, 112].
Genomic mining and omic based screening of rare actinomycetes
In rare actinomycete research, genome mining is an important bioprospecting tool. The fast advancement in genome sequencing, followed by mining of the genome using bioinformatic methods, including the identification of secondary metabolite gene clusters, has resulted in the finding of genetic machinery encoding for novel natural compounds from microbes that have yet to be chemically identified [113]. Polyketides (PK), non-ribosomally synthesized peptides (NRP), ribosomally and post-translationally modified peptides (RiPPs), and aminoglycosides are all encoded by most of these gene clusters [113]. Silent secondary metabolite gene clusters can also be discovered via bioinformatic analysis of genomes, which are not expressed under typical laboratory settings [114]. So far, more than 23,000 PK and NRP have been documented, many of which are discovered in actinomycetes and are being evaluated for pharmaceutical purposes [115, 116]. This method has also been utilized to discover novel antibiotic scaffolds in marine sediments from uncommon actinomycetes genera [117]. Due to revolutionary developments in genome- and metagenome-based approaches for drug discovery [118], the number of new biosynthetic gene clusters and corresponding compounds will undoubtedly increase in the near future, and it is likely that omics-based screening for novel bioactive compounds will overtake microbial isolation as the most efficient method for first identification of bioactive compounds [119].
The genes involved in the manufacture of bioactive secondary metabolites are found in the actinobacterial genome in the form of gene clusters, according to the literature [120]. Genome mining tools have made it more convenient to look for innovations in natural product discovery with majority of the bioactive compounds biosynthetic pathway of polyketides governed by a complex enzyme system, called polyketide synthase encoded by PKS gene cluster [121, 122]. Available whole genome draft of endophytic actinobacteria also revealed the presence of PKS and NPRS genes suggesting that these microbes are the possible source for many novel bioactive compounds [123, 124]. Screening for the presence of bioactive secondary metabolites in actinobacteria can be done using a high throughput method based on gene clusters. The antiSMASH (antibiotics & Secondary Metabolite Analysis Shell) pipeline is the first to identify biosynthetic loci across the whole spectrum of known secondary metabolite compound classes (polyketides, non-ribosomal peptides, terpenes, aminoglycosides, aminocoumarins, indolocarbazoles, antibiotics, bacteriocins, nucleosides, beta-lactams, butyrolactones, siderophores, melanins and others). It integrates or cross-links all previously existing secondary-metabolite specific gene analysis methods in one interactive view and aligns the detected regions at the gene cluster level to their nearest relatives from a database including all other known gene clusters [125].
Biopharmaceutical significance of rare actinomycete
Actinomycetes are major members of the soil microbial community, and their ability to create pharmaceutically useful compounds is of great interest to humans. Their interaction with rhizosphere soils has demonstrated their potential use as plant disease biocontrol agents. Their role as bioactive compound producers is well-documented. They are interesting prospects for the development of antimicrobials with medical and pharmaceutical applications [126].
Actinomycetes are known makers of antimicrobial compounds, which are significant medications in health care. Antibiotics could be produced by the genera Streptomyces and Micromonospora have shown to possess powerful therapeutic and acceptable pharmacokinetic qualities for clinical use [3]. Several substances derived from uncommon actinomycetes have been studied for their potential as antibacterial agents. Munumbicins were found to be efficient against Mycobacterium tuberculosis and Bacillus anthracis [127]. Actinomycetes produce peptide antibiotics called kakadumycins, which have shown to be effective against B. anthracis [3]. Actinomycete-produced coronamycin was effective against pythiaceous fungi as well as the human pathogen Cryptococcus neoformans [128]. Maklamycin, an antibacterial polyketide discovered in the culture filtrate of Micromonospora isolated from the Thai medicinal plant Abrus pulcellus, has been proven to be active against Gram-positive pathogens [129].
It is crucial to remember that biodiversity is the key to bioprospecting natural products. The isolation and discovery of new compounds with various chemical structures has frequently resulted from the diversity of microorganisms in unique habitats. When testing a molecule for a certain biological activity, multiple strains are screened against a wide range of targets, and the positive result is referred to as the "lead." Deciphering the pathways involved in secondary metabolite production has proven valuable in determining a strain's metabolite-producing capacity. The polyketide synthase (PKS) and non-ribosomal peptide synthetase (NRPS) enzymes are encoded in the actinomycete genome. The ability of a strain to create secondary metabolites by the identification of these genes is reported using recognized primers [79]. This method eliminates the requirement to test many strains' fermentation products for bioactivities. The positive strains should be subjected to the metabolite-producing potentials in either case, as some of the genes encoding these pathways may not be functional or necessitating different growth conditions [15]. Bioactivities of several secondary metabolites isolated from uncommon actinomycetes have been examined, including:
Antimicrobial effect
Antibacterial activity of actinomycetes strains was significant and varied against Gram-negative and Gram-positive bacteria [130]. Because numerous bioactive compounds were secreted rather than a single inhibitory molecule, many actinomycetes possessed a diverse range of activities including antimicrobial activity [131]. Rare actinomycetes have been shown to have antifungal and antagonistic activities against human pathogens in recent decades [130]. Rare actinomycetes of the genera Nocardia and Micromonospora have been shown to be efficient against a variety of pathogenic yeasts, but the species Nonomuraea has shown only mild antibacterial action [132]. Furthermore, antimicrobial substances produced by uncommon actinomycetes of the genera Micromonospora and Nocardia had previously been discovered to have broad-spectrum activity against both bacterial and fungal infections [133, 134]. The emergence and spread of multi-resistant bacteria have affected practically all antimicrobial agent classes [135]. This necessitates a call for urgency in the quest for novel antimicrobials. Antimicrobial-resistant microorganisms have been identified as a serious global public health problem, resulting in increased morbidity, mortality, and healthcare costs [135]. Antibiotic misuse is frequent in many underdeveloped countries, resulting in large outbreaks of antimicrobial-resistant bacteria and a lack of surveillance and data collection. Antibiotics with novel structures derived from unusual actinomycetes are urgently needed to combat multidrug-resistant pathogenic bacteria. Natural products continue to be the best source of new antibiotics. Rare actinobacteria are known to be prolific producers of natural bioactive chemicals, hence, screening unusual actinomycetes isolates can be used for new antibiotic discovery. We believe that intense efforts in isolating and screening rare genera of microbes can boost the chances of identifying a new drug with a novel chemical structure. One technique to do this is to screen rare actinomycetes and their previously under-represented taxa from unfamiliar settings in natural product screening collections [136]. Several bioactive substances derived from actinomycetes have been shown to suppress multidrug resistant pathogens such as vancomycin resistant Enterococci, methicillin resistant Staphylococcus aureus, Shigella dysenteriae, Klebsiella sp., Escherichia coli, and Pseudomonas aeruginosa [101, 137].
Antioxidant effect
To date, several actinobacterial antioxidants have been identified, including dihydroherbimycin A, N-carbamoyl-2,3-dihydroxybenzamide, 2-acetamido-3-(2,3-dihydroxybenzoylthio) propanoic acid, 2-allyloxyphenol, phenazines, and saccharomonopyrone A [138,139,140,141,142]. The genus Streptomyces has produced most physiologically active antioxidant compounds among actinobacteria [138]. Less prevalent or culturable strains of actinobacteria, such as rare genera, should be targeted for the discovery of new bioactive compounds due to the high likelihood of finding already known antioxidant metabolites (re-isolation of known antioxidant chemicals) [5]. UTMC 537 Saccharothrix ecbatanensis is a valuable source for the development of multipotent antioxidant compounds [143].
Anticancer/cytotoxic effect
Despite major advancements in the treatment of malignant tumors, cancer remains a primary cause of death and a public health issue around the world. The prospect of microbial secondary metabolites represents an effective source for the development of therapeutic leads, among the keyways for the discovery of new bioactive molecules [144]. Many secondary metabolites from rare actinomycetes have been extracted and tested for anticancer activity in a variety of carcinoma cell lines, including K562 (Human acute myelocytic leukemia), HeLa (cervical carcinoma), AGS (Human gastric), MCF-7 (breast adenocarcinoma), and HL-60 (Human acute promyelocytic leukemia). The discovery of taxol, a strong anticancer agent derived from endophytic fungi, sparked an interest in microbes as a source of possible antitumor agents. The anticancer potentials of rare actinomycetes' staurosporine and kigamicin have also been investigated, with promising results [144].
Insecticide/pesticide/herbicide
Pesticides made from natural products have grown in popularity around the world because to their excellent efficacy, environmental friendliness, and positive safety profile. This rise in popularity is reflected on the development of polyketide insecticides derived from actinomycetes in recent decades. Avermectins, spinosyns, polynactins, tetramycin, and analogues of these pesticides have all been used successfully in crop protection [145]. Furthermore, biotechnology's advancement has resulted in ongoing improvements in the research and production procedures. Actinomadura, Nocardiopsis, Dactylosporangium, Kibdelosporangium, Microbispora, Kitasatospora, Planomonospora, Planobispora, Salinispora, Marinispora, Serinicoccus, and Verrucosispora are among the less well-known uncommon taxa. These consequences highlight the importance of continuing study in this domain, and investments in uncommon actinomycetes can be deemed totally justified. PKSI, PKSII, and NRPS gene clusters were found in endophytic actinobacteria isolated from Artemisia annua, which had herbicidal activity against Echinochloa crusgalli [146]. Various antimicrobials and other bioactive compounds are obtained from rare actinomycetes (Table 2).
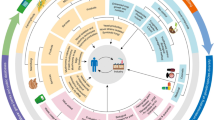
Several newer compounds isolated from rare actinomycetes include but not limited to Neomaclafungi A, Maklamicin, chaxamycin D, Macrolactin AI, Gilvocarin HE, RSP 01, Formicamycin J, Isoikarugamycin, Ageloline A, Arenimycin C, 5-hydroxynovobiocin, citreamycin A, Salinamide F, Arylomycin A6, Kibdelomycin, Kocurin, actinomadurol, Kibdelomycin (Fig. 1). Neomaclafungi A is a metabolite product of Actinoalloteichus sp. with potent antimicrobial activity. Kibdelomycin is got from a rare actinomycete of genus Kibdelosporangium. Chaxamycin is a product of Streptomyces sp. strain C34. Maklamicin, salinamide F, Kocurin, actinomodurol, citreamycin A and Formicamycin J are respectively from Actinomodura sp TP-AO878, Streptomyces sp, Kocuria palustris, Actinomodura sp., S. caelestis and S. formicae [161,162,163,164].
Considerable factors affecting bioactive molecule production in rare actinomycetes
The ability of actinomycete cultures to form these bioactive products is not a fixed trait; it can be considerably enhanced or completely lost depending on nutrition and cultivating conditions [165, 166]. This is because antibiotic biosynthesis is a unique feature of bacteria that is highly dependent on growth conditions. Manipulation of the nutritional and physical characteristics of the culture environment can be used to improve growth and antibiotic production. As a result, media composition is critical to the efficiency and profitability of the final process. Therefore, choosing the right fermentation medium is crucial in the generation of secondary metabolites [165]. Antibiotic biosynthesis in actinomycetes has been shown to be affected by changes in the nature and type of carbon and nitrogen sources [167]. Several culture parameters like as pH, cell density, microbial strain, incubation time, and temperature also play significant roles in the formation of bioactive metabolites [168]. When it comes to getting the best antibacterial output, cell density is crucial [169]. There are many natural products to be discovered from rare actinomycetes. Screening uncommon actinomycetes for novel bioactive metabolites is the first step in the search for useful antibiotics. This is followed by optimization of growth conditions for optimum antimicrobial compound production. Then comes antibiotic assay, chemical characterization, and identification of antibiotic compounds [101]. The amount and kind of actinomycetes present in the niche is influenced by ecological parameters such as environmental temperature and pH, habitat type, culture, organic matter concentration, exposure to air, and moisture content. Alkaliphilic actinomycetes, on the other hand, are extensively spread and easily isolated from their maritime environments [100, 169] (Fig. 2).
Conclusions
Rare actinomycetes have consistently produced a small number of novel bioactive compounds, but their promise in this field has been largely untapped. Due to the difficulty in cultivating most naturally occurring microorganisms, microbiologists have been severely limited in their research of natural microbial communities until recently. The search for unique biosynthetic potential species in unusual settings must be expanded. Microorganisms that are yet to be found or are rare may hold the key to developing new antibiotics to treat multidrug-resistant human infections and emerging fatal diseases. Using selective isolation and enhanced techniques, new rare bioactive producing actinobacteria can be discovered in previously unexplored environments. A combination of pretreatment procedures, appropriate selective isolation media, and enrichment culture supplemented with specific antibiotics allowed the isolation of rare and unique actinomycetes that produced unusual bioactive compounds and new enzymes. Rare actinobacteria have new genomes and structural diversities that are just waiting to be identified and applied in biotechnological and pharmaceutical industries.
Availability of data and materials
Not applicable.
Abbreviations
- ISP:
-
International Streptomyces Project
- HVA:
-
Humic acid Vitamin Agar
- LB:
-
Luria Bertani Agar
- SCA:
-
Starch Casein Agar
- NCBI:
-
National Center for Biotechnology Information
- BLAST:
-
Basic Local Alignment Search Tool
- rRNA:
-
Ribosomal Ribonucleic Acid
- DNA:
-
Deoxyribonucleic Acid
- MEGA:
-
Molecular Evolutionary Genetic Analysis
- MRSA:
-
Methicillin Resistant Staphylococcus aureus
- MDR:
-
Multidrug Resistant
- SDS:
-
Sodium dodecylsulphate
- PK:
-
Polyketides
- NRP:
-
Non-ribosomally synthesized Peptides
- RiPPs:
-
Ribosomally and post-translationally modified peptides
- PKS:
-
Polyketide synthetase
- NPRS:
-
Non-ribosomal peptide synthetase
- AntiSMASH:
-
Antibiotic and Secondary Metabolite Analysis Shell
References
Amin DH, Abolmaaty A, Tolba S, Abdallah NA, Wellington EM (2018) Phylogenic characteristics of a unique antagonistic Micromonospora sp. Rc5 to S. aureus isolated from Sinai desert of Egypt. Curr Res Microbiol Biotechnol 5(6):1295–1306
Amin DH, Abolmaaty A, Borsetto C (2019) In silico genomic mining reveals unexplored bioactive potential of rare actinobacteria isolated from Egyptian soil. Bull Natl Res Cent 43:78
Nalini MS, Prakash HS (2017) Diversity and bioprospecting of actinomycete endophytes from the medicinal plants. Lett Appl Microbiol 64(4):261–270
Seong CN, Choi JH, Baik KS (2001) An improved selective isolation of rare Actinomycetes from forest soil. J Microbiol 39(1):17–23
Subramani R, Aalbersberg W (2013) Culturable rare actinomycetes: diversity, isolation and marine natural product discovery. Appl Microbiol Biotechnol 97(21):9291–9321
Hayakawa M (2008) Studies on the isolation and distribution of rare Actinomycetes in soil. Actinomycetol 22:12–19
Matsumoto A, Takahashi Y (2017) Endophytic actinomycetes: promising source of novel bioactive compounds. J Antibiot 70:514–519
Sosio M, Donadio S (2004) Exploiting and expanding actinomycete diversity for antibiotic discovery. Microbiol Aust 25:32–33
Donadio S, Monciardini P, Alduina R, Mazza P, Chiocchini C, Cavaletti L, Sosio M, Puglia AM (2002) Microbial technologies for the discovery of novel bioactive metabolites. J Biotechnol 99(3):98–187
Fang B, Salam N, Han M, Jiao J, Chang J, Wei D, Xiao M, Li W (2017) Insights on the effects of heat pretreatment, pH and calcium salts on the isolation of rare actinobacteria from karstic caves. Front Microbiol. https://doi.org/10.3389/fmicb.2017.01535
Lazzarini A, Cavaletti L, Toppo G, Marinelli F (2000) Rare genera of actinomycetes as potential producers of new antibiotics. Antonie Van Leeuwenhoek 78(3–4):399–405
Arango C, Acosta-Gonzalez A, Parra-Giraldo CM, Sánchez-Quitian ZA, Kerr R, Díaz LE (2018) Characterization of actinobacterial communities from Arauca river sediments (Colombia) reveals antimicrobial potential presented in low abundant isolates. Open Microbiol J 12:181–194
Zhang J (2011) Improvement of an isolation medium for actinomycetes. Mod Appl Sci 5:124–127
Takahashi Y, Nakashima T (2018) Actinomycetes an inexhaustible source of naturally occurring antibiotics. Antibiotics (Basel) 7(2):45
Janso JE, Carter GT (2010) Biosynthetic potential of phylogenetically unique endophytic Actinomycetes from tropical plants. Appl Environ Microbiol 76:4377–4386
Inahashi Y, Matsumoto A, Omura S, Takahashi Y (2011) Streptosporangium oxazolinicum sp. nov., a novel endophytic Actinomycete producing new antitrypanosomal antibiotics, spoxazomicins. J Antibiot 64:297–302
Qin S, Li J, Chen HH, Zhao GZ, Zhu WY, Jiang CL, Xu LH, Li WJ (2009) Isolation, diversity, and antimicrobial activity of rare Actinobacteria from medicinal plants of tropical rain forests in Xishuangbanna, China. Appl Environ Microbiol 75:6176–6186
Xu LH, Li WJ, Liu ZH, Jiang CL (2007) Actinomycete taxonomy. Acedemic Press, Beijing, pp 202–208
Meklat A, Sabaou N, Zitouni A, Mathieu F, Lebrihi A (2011) Isolation, taxonomy, and antagonistic properties of halophilic Actinomycetes in Saharan soils of Algeria. Appl Environ Microbiol 77:6710–6714
Jiang Y, Li Q, Chen X, Jiang C (2016) Isolation and cultivation methods of Actinobacteria. In: Dhanasekaran D, Jiang Y (eds) Actinobacteria-basics and biotechnological applications. InTech, London, pp 39–57
Schabereiter-Gurtner C, Saiz-Jimenez C, Pinar G, Lubitz W, Rolleke S (2002) Altamira Cave Paleolithic paintings harbour partly unknown bacterial communities. FEMS Microbiol Lett 211:7–11
Nakaew N, Pathom-aree W, Lumyong S (2009) Generic diversity of rare Actinomycetes from Thai cave soils and their possible use as new bioactive compounds. Actinomycetologica 23:21–26
Lee SD (2006) Actinocorallia cavernae sp. nov., isolated from a natural cave in Jeju, Korea. Int J Syst Evol Microbiol 56:1085–1088
Lee SD (2006) Amycolatopsis jejuensis sp. nov. and Amycolatopsis halotolerans sp. nov., novel Actinomycetes isolated from a natural cave. Int J Syst Evol Microbiol 56:549–553
Lee SD (2006) Nocardia jejuensis sp. nov., a novel Actinomycetes isolated from a natural cave on Jeju Island, Republic of Korea. Int J Syst Evol Microbiol 56:559–562
Niyomvong N, Pathom-aree W, Thamchaipenet A, Duangmal K (2012) Actinomycetes from tropical limestone Caves. ChiangMai J Sci 39:373–388
Kaltenpoth M (2009) Actinobacteria asmutualists: general healthcare for insects? Trends Microbiol 17:529–535
Sen R, Ishak HD, Estrada D, Dowd SE, Hong E, Mueller UG (2009) Generalized antifungal activity and 454-screening of Pseudonocardia and Amycolatopsis bacteria in nests of fungus growing ants. Proc Natl Acad Sci 106:17805–17810
Haeder S, Wirth R, Herz H, Spiteller D (2009) Candicidin-producing Streptomyces support leaf-cutting ants to protect their fungus garden against the pathogenic fungus Escovopsis. Proc Natl Acad Sci 106:4742–4746
Oh DC, Poulsen M, Currie CR, Clardy J (2009) Dentigerumycin: a bacterial mediator of an ant–fungus symbiosis. Nat Chem Biol 5:391–393
Barke J, Seipke RF, Grüschow S, Heavens D, Drou N, Bibb MJ, Goss RJM, Yu DW, Hutchings MI (2010) A mixed community of Actinomycetes produce multiple antibiotics for the fungus farming ant Acromyrmex octospinosus. BMC Biol 8:109–118
Clardy J, Fischbach MA, Currie CR (2009) The natural history of antibiotics. Curr Biol 19:R437–R441
Kumar V, Bharti A, Gupta VK, Gusain O, Bisht GS (2012) Actinomycetes from solitary wasp mud nest and swallow bird mud nest: isolation and screening for their antibacterial activity. World J Microbiol Biotechnol 28:871–880
Zhang DF, Pan HQ, He J, Zhang XM, Zhang YG, Klenk HP, Hu JC, Li WJ (2013) Description of Streptomonospora sediminis sp. nov. and Streptomonospora nanhaiensis sp. nov., and reclassification of Nocardiopsis arabia. Int J Syst Evol Microbiol 63:4447–4455
Cho Y, Jang G, Hwang CY, Kim EH, Cho BC (2013) Nocardioides salsibiostraticola sp. Nov., isolated from biofilm formed in coastal seawater. Int J Syst Evol Microbiol 63:3800–3806
Lee SD (2013) Tamlicoccus marinus gen. nov., sp. nov., isolated from seawater. Int J Syst Evol Microbiol 63:1951–1954
Afonso CB, Afonso RS, Souza WR, Parma M, Melo IS, Zucchi TD, Fantinatti GF (2017) Williamsia spongiae sp. nov., an actinomycete isolated from the marine sponge Amphimedon viridis. Int J Syst Evol Microbiol 67:1260–1265
Souza DT, Silva FS, Silva JD, Crevelin EJ, Moraes AB, Zucchi TD, Melo IS (2017) Saccharopolyspora spongiae sp. nov., a novel actinomycete isolated from the marine sponge scopalina ruetzleri. Int J Syst Evol Microbiol 67:2019–2025
Fenical W, Jensen PR (2006) Developing a new resource for drug discovery: marine actinomycete bacteria. Nat Chem Biol 2:666–673
Jiang Y, Wei X, Chen X, Jiang Y, Xue Q, Lai H, Jiang C (2016) Saccharopolyspora griseoalba sp. nov., a novel actinomycete isolated from the dead sea. Antonie Van Leeuwenhoek 109:1635–1641
Khamna S, Yokota A, Lumyong S (2009) Actinomycetes isolated from medicinal plant rhizosphere soils: diversity and screening of antifungal compounds, indole-3-acetic acid and siderophore production. World J Microbiol Biotechnol 25:649–655
Hayakawa M, Yoshida Y, Iimura Y (2004) Selection of bioactive soil Actinomycetes belonging to the Streptomyces violaceus niger phenotypic cluster. J Gen Appl Microbiol 96:973–981
Istianto Y, Koesoemowidodo RSA, Saputra H, Watanabe Y, Pranamuda H, Marwoto B (2012) Application of phenol pretreatment for the isolation of rare Actinomycetes from Indonesian soil. Microbiology (Indonesia) 6:42–47
Qiu FB, Huang Y, Sun L, Zhang XX, Liu ZH, Song W (2007) Leifsonia ginsengi sp. nov., isolated from ginseng root. Int J Syst Evol Microbiol 57:405–408
Tiwari K, Gupta RK (2012) Diversity and isolation of rare Actinomycetes: an overview. Crit Rev Microbiol 39:256–294
Qin S, Chen HH, Klenk HP, Zhao GZ, Li J, Xu LH, Li WJ (2009) Glycomyces scopariae sp. nov. and Glycomyces mayteni sp. nov., isolated from two medicinal plants in China. Int J Syst Evol Microbiol 59:1023–1027
Otoguro M, Hayakawa M, Yamazaki T, Iimura Y (2001) An integrated method for the enrichment and selective isolation of Actiniokineospora spp. in soil and plant litter. J Appl Microbiol 91:118–130
Wang DS, Xue QH, Zhu WJ, Zhao J, Duan JL, Shen GH (2013) Microwave irradiation is a useful tool for improving isolation of Actinomycetes from soil. Microbiology 82:102–110
Xue Q, Dua CM, Wang LN, Lin YB (2010) The influence of microwave irradiation to the isolation effect of soil Actinomycetes. Chin J Microbiol 3:19–24
Jiang Y, Cao Y, Zhao L, Wang Q, Jin R, He W, Xue Q (2010) Ultrasonic treatment of soil samples for Actinomycete isolation. Wei ShengWu Xue Bao 50:1094–1097
Li IV, Terekhova LP, Alferova IV, Galatenko OA, Gapochka MG (2003) The application of succession analysis in combination with extremely high-frequency irradiation to the selective isolation of Actinomycetes from soil. Mikrobiologiia 72:131–135
Hayakawa M, Otoguro M, Takeuchi T, Yamazaki T, Iimura Y (2000) Application of a method incorporating differential centrifugation for selective isolation of motile Actinomycetes in soil and plant litter. Antonie Van Leeuwenhoek 78:171–185
Qin S, Zhu WY, Jiang JH, Klenk HP, Li J, Zhao GZ, Xu LH, Li WJ (2009) Pseudonocardia tropica sp. nov., a novel endophytic actinomycete isolated from the stem of Maytenus austroyunnanensis. Int J Syst Evol Microbiol 60:2524–2528
Hong K, Gao AH, Xie QY, Gao H, Zhuang L, Lin HP, Yu HP, Li J, Yao XS, Goodfellow M, Ruan JS (2009) Actinomycetes for marine drug discovery isolated from mangrove soils and plants in China. Mar Drugs 7(1):24–44
Zhang DF, Chen W, He J, Zhang XM, Xiong ZJ, Sahu MK, Sivakumar K, Li WJ (2013) Saccharomonospora oceani sp. nov. isolated from marine sediments in little Andaman. India Antonie Van Leeuwenhoek 103:1377–1384
Taechowisan T, Peberdy JF, Lumyong S (2003) Isolation of endophytic actinomycetes from selected plants and their antifungal activity. World J Microbiol Biotechnol 19:381–385
Saadi SA, Meklat A, Mokrane S, Achour HY, Holtz MD, Klenk HP, Bouras N (2021) Isolation and characterization of a new Saccharothrix strain AH023 with antimicrobial activity from an unexploited Algerian sahara region. Analele Univ din Oradea Fasc Biol 28(1):71–77
Wang L, Li J, Zhang G (2016) Nocardioides rotundus sp. nov., isolated from deep seawater. Int J Syst Evol Microbiol 66:1932–1936
Supong K, Suriyachadkun C, Pittayakhajonwut P, Suwanborirux K, Thawai C (2013) Micromonospora spongicola sp. nov., an actinomycete isolated from marine sponge in the gulf of Thailand. J Antibiot 66:505–509
Wu JF, Li J, You ZQ, Zhang S (2014) Prauserella coralliicola sp. nov., isolated from the coral Galaxea fascicularis. Int J Syst Evol Microbiol 64:3341–3345
Kaur G, Mual P, Kumar N, Verma A, Kumar A, Krishnamurthi S, Mayilraj S (2016) Microbacterium aureliae sp. nov., a novel actinobacterium isolated from Aurelia aurita, the moon jellyfish. Int J Syst Evol Microbiol 66:4665–4670
De Menezes CB, Tonin MF, Silva LJ, De Souza WR, Parma M, Melo IS, Zucchi TD, Destefano SA, Fantinatti GF (2015) Marmoricola aquaticus sp. nov., an actinomycete isolated from marine sponge. Int J Syst Evol Microbiol 65:2286–2291
Lee JY, Hyun DW, Soo KP, Sik KH, Shin NR, Yun JH, Jung MJ, Kim MS, Woong WT, Bae JW (2016) Arthrobacter echini sp. nov., isolated from the gut of a purple sea urchin, Heliocidaris crassispina. Int J Syst Evol Microbiol 66:1887–1893
Ramaprasad EV, Sasikala C, Ramana CV (2015) Ornithinimicrobium algicola sp. nov., a marine actinobacterium isolated from the green alga of the genus Ulva. Int J Syst Evol Microbiol 65:4627–4631
Thawai C, Rungjindamai N, Klanbut K, Tanasupawat S (2017) Nocardia Xestospongiae sp. nov., isolated from a marine sponge in the Andaman sea. Int J Syst Evol Microbiol 67:1451–1456
Kampfar P, Glaeser SP, Busse H, Abdelmohsen UR, Hentschet U (2014) Rubrobacter aplysinae sp. Nov., isolated from the marine sponge Aplysina aerophoba. Int J Syst Evol Microbiol 3:64
Kampfer P, Glaeser SP, Busse HJ, Abdelmohsen UR, Ahmed S, Hentschd U (2015) Actinokineospora spheciospongiae sp. nov., isolated from the marine sponge Spheciospongia vagabunda. Int J Syst Evol Microbiol 65:879–884
Sarmiento VA, Gonzalez V, Brana AF, Molina A, Acuna JL, Garcia LA, Blanco G (2015) Myceligenerans cantabricum sp. nov., a barotolerant actinobacterium isolated from a deep cold water cord. Int J Syst Evol Microbiol 65:1328–1334
Veyisoglu A, Sazak A, Cetin D, Guven K, Sahin N (2013) Saccharomonospora amisosensis sp. nov., isolated from deep marine sediment. Int J Syst Evol Microbiol 63:3782–3786
Zhang DF, Jiang Z, Zhang XM, Yang LL, Tian XP, Long LJ, Zhang S, Li WJ (2014) Actinophytocola sediminis sp. nov., an actinomycete isolated from a marine sediment. Int J Syst Evol Microbiol 64:2834–2840
Zhang DF, Jiang Z, Li L, Liu BB, Zhang XM, Tian XP, Zhang S, Li WJ (2014) Pseudonocardia sediminis sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol 64:745–750
Wei X, Jiang Y, Chen X, Jiang Y, Lai H (2015) Amycolatopsis flava sp. nov., a halophilic actinomycete isolated from dead sea. Antonie Van Leeuwenhoek 108:879–885
Lee DW, Lee AH, Lee H, Kim JJ, Khim JS, Yim UH, Kim BS (2017) Nocardioides litoris sp. nov., isolated from the Taean seashore. Int J Syst Evol Microbiol 67:2332–2336
Hamada M, Shibata C, Tamura T, Suzuki K (2014) Agromyces marinus sp. nov., a novel actinobacterium isolated from sea sediment. J Antibiot 67:703–706
Mawlankar RR, Mual P, Sonalkar VV, Thorat MN, Verma A, Srinivasan K, Dastager SG (2015) Microbacterium enclense sp. nov., isolated from sediment sample. Int J Syst Evol Microbiol 65:2064–2070
Yan L, Wang J, Chen Z, Guan Y, Li J (2015) Microbacterium nanhaiense sp. nov., an actinobacterium isolated from sea sediment. Int J Syst Evol Microbiol 65:3697–3702
Gu Q, Zheng W, Huang Y (2007) Glycomyces sambucus sp. nov., an endophytic actinomycete isolated from the stem of Sambucus adnata wall. Int J Syst Evol Microbiol 57:1995–1998
Zhang X, Ren K, Du J, Liu H, Zhang L (2014) Glycomyces artemisiae sp. nov., an endophytic actinomycete isolated from the roots of Artemisia argyi. Int J Syst Evol Microbiol 64:3492–3495
Zhao GZ, Li J, Huang HY, Zhu WY, Zhao LX, Tang SK, Xu LH, Li WJ (2011) Pseudonocardia artemisiae sp. nov., a novel actinobacterium isolated from surface-sterilized Artemisia annua L. Int J Syst Evol Microbiol 61:1061–1065
Gu Q, Luo H, Zheng W, Huang Y (2006) Pseudonocardia oroxyli sp. nov., a novel actinomycete isolated from surface sterilized Oroxylum indicum root. Int J Syst Evol Microbiol 56(9):2193–2197
Hamada M, Shibata C, Tamura T, Suzuki K (2013) Zhihengliuella flava sp. nov., an actinobacterium isolated from sea sediment, and emended description of the genus Zhihengliuella. Int J Syst Evol Microbiol 63:4760–4764
Dastager SG, Tang SK, Srinivasan K, Lee JC, Li WJ (2014) Kocuria indica sp. nov., isolated from a sediment sample. Int J Syst Evol Microbiol 64:869–874
Zhang G, Zhang Y, Yin X, Wang S (2015) Nesterenkonia alkaliphila sp. nov., an alkaliphilic, halotolerant actinobacteria isolated from the western Pacific Ocean. Int J Syst Evol Microbiol 65:516–521
Fan X, Zhang Z, Li Z, Zhang XH (2014) Luteococcus sediminum sp. nov., isolated from deep subseafloor sediment of the South Pacific Gyre. Int J Syst Evol Microbiol 64:2522–2527
Zhang DF, Wang HF, Xiong ZJ, Tian XP, Liu L, Zhang XM, Jiang Z, Zhang S, Li WJ (2014) Mariniluteicoccus flavus gen. nov., sp. nov., a new member of the family Propionibacteriaceae, isolated from a deep-sea sediment. Int J Syst Evol Microbiol 64:1051–1056
Bai JL, Wang Y, Qin S, Ding P, Xing K, Yuan B, Cao CL, Huang Y, Zhang YQ, Jiang JH (2016) Nocardia jiangsuensis sp. nov., an actinomycete isolated from coastal soil. Int J Syst Evol Microbiol 66:4633–4638
Hamada M, Shibata C, Saitou S, Tamura T, Komaki H, Ichikawa N, Oguchi A, Hosoyama A, Fujita N, Yamamura H (2015) Proposal of nine novel species of the genus Lysinimicrobium and emended description of the genus Lysinimicrobium. Int J Syst Evol Microbiol 65:4394–4402
Ren J, Li L, Wei B, Tang YL, Deng ZX, Sun M, Hong K (2013) Micromonospora wenchangensis sp. nov., isolated from mangrove soil. Int J Syst Evol Microbiol 63:2389–2395
Tang YL, Lin HP, Xie QY, Li L, Peng F, Deng Z, Hong K (2013) Actinoallomurus acanthiterrae sp. nov., an actinomycete isolated from rhizosphere soil of the mangrove plant Acanthus ilicifolius. Int J Syst Evol Microbiol 63:1874–1879
Lee LH, Azman AS, Zainal N, Yin WF, Mutalib NS, Chan KG (2015) Sinomonas humi sp. nov., an amylolytic actinobacterium isolated from mangrove forest soil. Int J Syst Evol Microbiol 65:996–1002
Huang HQ, Xing SS, Yuan WD, Wang Y, Liu M, Sun QG, Lin XZ, Bao SX (2015) Nocardiopsis mangrovei sp. nov., isolated from mangrove sediment. Antonie Van Leeuwenhoek 107:1541–1556
Hamada M, Shibata C, Tamura T, Nurkanta A, Ratnakomala S, Lisdiyanti P, Suzuki K (2016) Kocuria pelophila sp. nov., an actinobacterium isolated from the rhizosphere of a mangrove. Int J Syst Evol Microbiol 66:9
Lee LH, Zainal N, Azman AS, Mutalib NS, Hong K, Chan KG (2014) Mumia flava gen. nov., sp. nov., an actinobacterium of the family Nocardioidaceae. Int J Syst Evol Microbiol 64:1461–1467
Azman AS, Zainal N, Mutalib NA, Yin WF, Chan KG, Lee LH (2016) Monashia flava gen. nov., sp. nov., an actinobacterium of the family Intrasporangiaceae. Int J Syst Evol Microbiol 66:554–561
Duangmal K, Muangham S, Mingma R, Yimyai T, Srisuk N, Kitpreechavanich V, Matsumoto A, Takahashi Y (2016) Kineococcus mangrovi sp. nov., isolated from mangrove sediment. Int J Syst Evol Microbiol 66:1230–1235
Stanek RJ, Maher MB, Norton NB, Mufson MA (2011) Emergence of a unique penicillin-resistant Streptococcus pneumoniae serogroup 35 Strain. J Clin Microbiol 49(1):400–404
Yamamura H, Hayakawa M, Iimura Y (2003) Application of sucrose-gradient centrifugation for selective isolation of Nocardia spp. from soil. J Appl Microbiol 95(4):677–685
Ezeobiora CE, Igbokwe NH, Amin DH, Mendie UE (2021) Endophytic microbes from Nigerian ethnomedicinal plants: a potential source for bioactive secondary metabolites—a review. Bull Natl Res Cent 45:103
Burgdorf RJ, Laing MD, Morris CD, Jamal-Ally SF (2014) A procedure to evaluate the efficiency of surface sterilization methods in culture-independent fungal endophyte studies. Braz J Microbiol 45(3):977–983
Sivanandhini T, Ramasamy S, Gopinath M, Angrasan JM, Kabilan T, Selvam M (2015) An investigation on morphological characterization of actinomycetes isolated from marine sediments. Res J Pharm Biol Chem Sci 6(2):1234
Khanna M, Solanki R, Lal R (2011) Selective isolation of rare actinomycetes producing novel antimicrobial compounds. Int J Adv Biotechnol Res 2(3):357–375
Li Q, Li G, Chen X, Xu F, Li Y, Xu L, Jiang Y, Jiang C (2015) Kineococcus gypseus sp. Nov., isolated from saline sediment. Int J Syst Evol Microbiol 65(10):3703–3708
Dhananjeyan V, Selvan N, Dhanapal K (2010) Isolation, Characterization, Screening and Antibiotic sensitivity of actinomycetes from locally (Near MCAS) collected soil samples. J Biol Sci 10:514–519
Bhagat N, Virdi JS (2009) Molecular and biochemical characterization of urease and survival of Yersinia enterocolitica biovar 1A in acidic pH in vitro. BMC Microbiol 9:262
Mobarak-Qamsari E, Kasra-Kermanshahi R, Moosavi-Nejad Z (2011) Isolation and identification of a novel, lipase-producing bacterium, Pseudomnas aeruginosa KM110. Iran J Microbiol 3(2):92–98
Tille PM, Forbes BA (2014) Bailey & Scott’s diagnostic microbiology, Thirteenth. Elsevier, St. Louis
Cappuccino JG, Sherman N (2008) Microbiology: a laboratory manual, 8th edn. Pearson Benjamin Cummings, San Francisco
Tille PM (2014) Bailey and Scott’s diagnostic microbiology. Thirteen edition. Mosby, Inc., an affiliate of Elsevier Inc. 3251 Riverport Lane. St. Louis. Missouri 63043
Braesel J, Lee JH, Arnould B, Murphy BT (2019) Diazaquinomycin biosynthetic gene clusters from marine and freshwater actinomycetes. J Nat Prod 82:937–946
Savi DC, Aluizio R, Terasawa L, Kava V, Glienke C (2016) 16S-gyrB-rpoB multilocus sequence analysis for species identification in the genus Microbispora. Antonie Van Leeuwenhoek 109:801–815
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA 6: molecular evolutionary genetic analysis version 6.0. Mol Biol Evol 30(12):2725–2729
Cheema MT, Ponomareva LV, Tao L, Voss RS, Thorson JS, Shaaban KA, Sajid I (2021) Taxonomic and metabolomic profiling of Actinobacteria strains from Himalayan collection sites in parkistan. Curr Microbiol. https://doi.org/10.1007/S00284-021-02557-y
Ziemert N, Alanjaryab M, Weber T (2016) The evolution of genome mining in microbes—a review. Nat Prod Rep 33:988–1005
Hug JJ, Bader CD, Remškar M, Cirnski K, Müller R (2018) Concepts and methods to aaccess novel antibiotics from actinomycetes. Antibiotics 7:44
Wei Y, Zhang L, Zhou Z, Yan X (2018) Diversity of gene clusters for polyketide and nonribosomal peptide biosynthesis revealed by metagenomic analysis of the yellow sea sediment. Front Microbiol 9:295
Katz L, Baltz RH (2016) Natural product discovery: Past, present, and future. J Ind Microbiol Biotechnol 43:155–176
Schwager E, Luo C, Huttenhower C, Morgan XC (2015) Sequencing and other tools for studying microbial communities: Genomics and “meta’omic” tools are enabling us to explore the microbiome from three complementary perspectives—taxonomic, functional and ecological. Microbe 10:419–425
Schorn MA, Alanjary MM, Aguinaldo K, Korobeynikov A, Podell S, Patin N, Lincecum T, Jensen PR, Ziemert N, Moore BS (2016) Sequencing rare marine actinomycete genomes reveals high density of unique natural product biosynthetic gene clusters. Microbiology 162:2075–2086
Loureiro C, Medema MH, Van der Oost J, Sipkema D (2018) Exploration and exploitation of the environment for novel specialized metabolites. Curr Opin Biotechnol 50:206–213
Doroghazi JR, Metcalf WW (2013) Comparative genomics of actinomycetes with a focus on natural product biosynthetic genes. BMC Genomics 14:1–13
Jackson SA, Crossman L, Almeida EL, Margassery LM, Kennedy J, Dobson ADW (2018) Diverse and abundant secondary metabolism biosynthetic gene clusters in the genomes of marine sponge derived Streptomyces spp. isolates. Mar Drugs 16:1–18
Corre C, Challis GL (2007) Heavy tools for genome mining. Chem Biol 14:7–9
Komaki H, Ichikawa N, Hosoyama A, Fujita N, Thamchaipenet A, Igarashi Y (2015) Draft genome sequence of Linfuranone producer Microbispora sp. GMKU 363. Genome Announc 3:e01471-e1515
Angolini CFF, Gonçalves AB, Sigrist R, Paulo BS, Samborskyy M, Cruz PLR (2016) Genome mining of endophytic Streptomyces wadayamensis reveals high antibiotic production capability. J Braz Chem Soc 27:1465–1475
Medema MH, Blin K, Cimermancic P, Jager V, Zakrzewski P, Fischbach MA, Weber T, Takano E, Breitling R (2011) antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res 39:W339–W346
Blin K, Kim HU, Medema MH, Weber T (2019) Recent development of antiSMASH and other computational approaches to mine secondary metabolite biosynthetic gene clusters. Brief Bioinform 20(4):1103–1113
Ek-Ramos MJ, Gomez-Flores R, Orozco-Flores AA, Rodríguez-Padilla C, González-Ochoa G, Tamez-Guerra P (2019) Bioactive products from endophytic gram-positive bacteria. Front Microbiol 10:463
Ezra D, Castillo UF, Strobel GA, Hess WM, Porter H, Jensen JB, Condron MAM, Teplow DB (2004) Coronamycins, peptide antibiotics produced by a verticillate Streptomyces sp. (MSU-2110) endophytic on Monstera sp. Microbiology 150:785–793
Igarashi Y, Ogura H, Furihata K, Oku N, Indananda C, Thamchaipenet A (2011) Maklamicin, an antibacterial polyketide from an endophytic Micromonospora sp. J Nat Prod 74:670–674
Benhadj M, Gacemi-Kirane D, Menasria T, Guebla K, Ahmane Z (2019) Screening of rare actinomycetes isolated from natural wetland ecosystem (Fetzara Lake, northern Algeria) for hydrolytic enzymes and antimicrobial activities. J King Saud Univ Sci 31(4):706–712
Benhadj M, GacemiKirane D, Toussaint M, Hotel L, Bontemps C, Duval RE, Aigle B, Leblond P (2018) Diversity and antimicrobial activities of Streptomyces isolates from Fetzara Lake, northeastern Algeria. Ann Biol Clin 76(1):81–95
Tanvir R, Sajid I, Hasnain S, Kulik A, Grond S (2016) Rare actinomycetes Nocardia caishijiensis and Pseudonocardia carboxydivorans as endophytes, their bioactivity and metabolites evaluation. Microbiol Res 185:22–35
Kavitha A, Prabhakar P, Narasimhulu M, Vijayalakshmi M, Venkateswarlu Y, Rao KV, Raju VBS (2010) Isolation, characterization and biological evaluation of bioactive metabolites from Nocardia levis MK-VL_113. Microbiol Res 165:199–210
Kodani S, Komaki H, Ishimura S, Hemmi H, Ohnishi-Kameyama M (2016) Isolation and structure determination of a new antibiotic cinnamycin B from Actinomadura atramentaria based on genome mining. J Ind Microbiol Biotechnol 43:1159–1165
Serwecinska L (2020) Antimicrobials and antibiotic resistant bacteria: a risk to the environment and to public health. Water 12:3313
Tiwari K, Gupta RK (2011) Rare actinomycetes: a potential storehouse for novel antibiotics. Crit Rev Biotechnol 32(2):108–132
Chaudhary SH, Soni B, Shrivastava AR, Shrivastava S (2013) Diversity and versatility of actinomycetes and its role in antibiotic production. J Appl Pharm Sci 3(8):S83–S94
Chang HB, Kim JH (2007) Antioxidant properties of dihydroherbimycin A from a newly isolated Streptomyces sp. Biotechnol Lett 29:599–603
Kumar S, Krishnan K (2012) Cytotoxicity and antioxidant activity of 5-(2,4-dimethylbenzyl)pyrrolidin-2-one extracted from marine Streptomyces VITSVK5 spp. Saudi J Biol Sci 19(1):81–86
Sugiyama Y, Hirota A (2009) New potent DPPH radical scavengers from a marine-derived Actinomycete strain USF-TC31. Biosci Biotechnol Biochem 73:2731–2734
Arumugam M, Mitra A, Jaisankar P, Dasgupta S, Sen T, Gachhui R, Mukhopadhyay UK, Mukherjee J (2010) Isolation of an unusual metabolite 2-allyloxyphenol from a marine actinobacterium, its biological activities and applications. Appl Microbiol Biotechnol 86:109–117
Abdelmageed WM, Milne BF, Wagner M, Schumacher M, Sandor P, Pathomaree W, Goodfellow M, Bull AT, Horikoshi K, Ebel R (2010) Dermacozines, a new phenazine family from deep-sea dermacocci isolated from a Mariana Trench sediment. Org Biomol Chem 8:2352–2362
Zotchev SB (2012) Marine actinomycetes as an emerging resource for the drug development pipelines. J Biotechnol 158:168–175
Mohammadipanah F, Momenilandi M (2018) Potential of rare actinomycetes in the production of metabolites against multiple oxidant agents. Pharm Biol 56(1):51–59
Davies-Bolorunduro FO, Adeleye AI, Akinleye MO, Wang GP (2019) Anticancer potentials of metabolic compounds from marine actinomycetes isolated from Lagos Lagoon sediments. J Pharm Anal 9(3):201–208
Li S, Yang B, Tan GY, Ouyang LM, Qiu S, Wang W, Xiang W, Zhang L (2021) Polyketide pesticides from actinomycetes. Curr Opin Biotechnol 69:299–307
Li J, Guozhen Z, Huang H, Strobel G (2012) Isolation and characterization of culturable endophytic actinobacteria associated with Artemisia annua L. Antonie Van Leeuwenhoek 101(3):515–527
Yamanaka K, Reynolds KA, Kersten RD, Ryan KS, Gonzalez DJ, Nizet V, Dorrestein PC, Moore BS (2014) Direct cloning and refactoring of a silent lipopeptide biosynthetic gene cluster yields the antibiotic taromycin A. Proc Natl Acad Sci USA 111:1957–1962
Duncan KR, Crüsemann M, Lechner A, Sarkar A, Li J, Ziemert N, Wang M, Bandeira N, Moore BS, Dorrestein PC (2015) Molecular networking and pattern-based genome mining improves discovery of biosynthetic gene clusters and their products from Salinispora species. Chem Biol 22:460–471
Richter TK, Hughes CC, Moore BS (2015) Sioxanthin, a novel glycosylated carotenoid, reveals an unusual subclustered biosynthetic pathway. Environ Microbiol 17:2158–2171
Schulze CJ, Donia MS, Siqueira-Neto JL, Ray D, Raskatov JA, Green RE, McKerrow JH, Fischbach MA, Linington RG (2015) Genome-directed lead discovery: Biosynthesis, structure elucidation, and biological evaluation of two families of polyene macrolactams against Trypanosoma brucei. ACS Chem Biol 10:2373–2381
Tan Y, Hu Y, Wang Q, Zhou H, Wang Y, Gan M (2016) Tetrocarcins N and O, glycosidic spirotetronates from a marine-derived Micromonospora sp. identified by PCR-based screening. RSC Adv 6:91773–91778
Jiang X, Zhang Q, Zhu Y, Nie F, Wu Z, Yang C, Zhang L, Tian X, Zhang C (2017) Isolation, structure elucidation and biosynthesis of benzo[b]fluorine nenestatin A from deep-sea derived Micromonospora echinospora SCSIO 04089. Tetrahedron 73:3585–3590
Kaeberlein T, Lewis K, Epstein SS (2002) Isolating “uncultivable” microorganisms in pure culture in a simulated natural environment. Science 296:1127–1129
Vartoukian SR, Palmer RM, Wade WG (2010) Strategies for culture of ‘unculturable’ bacteria. FEMS Microbiol Lett 309:1–7
Stewart EJ (2012) Growing unculturable bacteria. J Bacteriol 194:4151–4160
Zengler K, Toledo G, Rappé M, Elkins J, Mathur EJ, Short JM, Keller M (2002) Cultivating the uncultured. Proc Natl Acad Sci USA 99:15681–15686
Butler MS, Blaskovich MA, Cooper MA (2017) Antibiotics in the clinical pipeline at the end of 2015. J Antibiot 70:3–24
Feling RH, Buchanan GO, Mincer TJ, Kauffman CA, Jensen PR, Fenical W (2003) Salinosporamide A: a highly cytotoxic proteasome inhibitor from a novel microbial source, a marine bacterium of the new genus Salinospora angew. Chem Int Ed Engl 42:355–357
Asolkar RN, Freel KC, Jensen PR, Fenical W, Kondratyuk TP, Park EJ, Pezzuto JM (2009) Arenamides A-C, cytotoxic NFκB Inhibitors from the marine actinomycete Salinispora arenicola. J Nat Prod 72:396–402
Jang KH, Nam SJ, Locke JB, Kauffman CA, Beatty DS, Paul LA, Fenical W (2013) Anthracimycin, a potent anthrax antibiotic from a marine-derived actinomycete. Angew Chem Int Ed Engl 52:7822–7824
Jakubiec-Krzesniak K, Rajnisz-Mateusiak A, Guspiel A, Ziemska J, Solecka J (2018) Secondary metabolites of actinomycetes and their antibacterial, antifungal and antiviral properties. Proc J Microbiols 67(3):259–272. https://doi.org/10.21307/pjm-2018-048
Hassan HM, Degen D, Jang KH, Ebright RH, Fenical W (2015) Salinamide F, new depsipeptide antibiotic and inhibitor of bacterial RNA polymerase from a marine-derived Streptomyces sp. J Antibiot 68(3):206–209
Martín J, Sousa T, Crespo G, Palomo S, González I, Tormo JR, Cruz M, Anderson M, Hill RT, Vicente F, Genilloud O, Reyes F (2013) Kocurin, the true structure of PM181104, an anti-methicillin-resistant Staphylococcus aureus (MRSA) thiazolyl peptide from the marine-derived bacterium Kocuria palustris. Mar Drugs 11(2):387–398
Phillips JW, Goetz MA, Smith SK, Zink DL, Polishook J, Onishi R, Salowe S, Wiltsie J, Allocco J, Sigmund J, Dorso K, Lee S, Skwish S, de la Cruz M, Martín J, Vicente F, Genilloud O, Lu J, Painter RE, Young K, Overbye K, Donald RG, Singh SB (2011) Discovery of kibdelomycin, a potent new class of bacterial type II topoisomerase inhibitor by chemical-genetic profiling in Staphylococcus aureus. Chem Biol 18(8):955–965
Gao H, Liu M, Liu J, Dai H, Zhou X, Liu X (2009) Medium Optimization for the production of avermectin B1a by Streptomyces Avermitilis 14–12A using response surface methodology. Bioresour Technol 100:4012–4016
Bundale S, Begde D, Nashikkar N, Kadam T, Upadhyay A (2015) Optimization of culture conditions for production of bioactive metabolites by Streptomyces spp. Isolated from. Soil Adv Appl Microbiol 5:441–451
Reddy NG, Ramakrishna D, Rajagopal S (2011) Optimization of culture conditions of Streptomyces rochei (MTCC 10109) for the production of antimicrobial metabolites. Egypt J Biol 13:21–29
Usha KM, Sudhakar P, Sreenivasulu K, Vijayalakshmi M (2011) Optimization of culturing conditions for improved production of bioactive metabolites by Pseudonocardia sp. VUK-10. Mycobiology 39:174–181
Abdelwahed N, Abdallah NA, El-Ghawas DE, El-Din SMB, El-Diwany AL (2012) Isolation, identification and optimization of antimicrobial metabolites produced by soil derived actinomycetes. Egypt J Exp Biol (Bot) 8(2):205–217
Acknowledgements
Not applicable.
Funding
There was no special funding for the research. The authors funded the research.
Author information
Authors and Affiliations
Contributions
ECE conceived the project and was a major contributor in writing the manuscript; CFO helped in writing the manuscript. INH, DHA and MUE supervised the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate.
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ezeobiora, C.E., Igbokwe, N.H., Amin, D.H. et al. Uncovering the biodiversity and biosynthetic potentials of rare actinomycetes. Futur J Pharm Sci 8, 23 (2022). https://doi.org/10.1186/s43094-022-00410-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s43094-022-00410-y