Abstract
Geobacillus sp. Y412MC52 was isolated from Obsidian Hot Spring, Yellowstone National Park, Montana, USA under permit from the National Park Service. The genome was sequenced, assembled, and annotated by the DOE Joint Genome Institute and deposited at the NCBI in December 2011 (CP002835). Based on 16S rRNA genes and average nucleotide identity, Geobacillus sp. Y412MC52 and the related Geobacillus sp. Y412MC61 appear to be members of a new species of Geobacillus. The genome of Geobacillus sp. Y412MC52 consists of one circular chromosome of 3,628,883 bp, an average G + C content of 52 % and one circular plasmid of 45,057 bp and an average G + C content of 45 %. Y412MC52 possesses arabinan, arabinoglucuronoxylan, and aromatic acid degradation clusters for degradation of hemicellulose from biomass. Transport and utilization clusters are also present for other carbohydrates including starch, cellobiose, and α- and β-galactooligosaccharides.
Similar content being viewed by others
Introduction
Identification of new organisms that produce biomass-degrading enzymes is of considerable interest. Commercial uses for these enzymes include paper manufacturing, brewing, biomass deconstruction and the production of animal feeds [1–3]. Hot springs, especially those at Yellowstone National Park, have been a source of many new organisms including Thermus aquaticus [4, 5], Thermus brockianus [6], and Acidothermus cellulolyticus [7] that possess enzymes with significant potential in biotechnological applications [8]. As part of a project in conjunction with the Great Lakes Bioenergy Research Center, Dept. of Energy, C5–6 Technologies and Lucigen Corp. isolated, characterized, and sequenced a number of new enzyme-producing aerobic organisms from Yellowstone hot springs.
Geobacillus species were the most common aerobic organisms isolated during the cultivation of most hot springs samples. Geobacillus species were originally classified as members of the genus Bacillus , but were subsequently reclassified as a separate genus based on 16S rRNA gene sequence analysis, lipid and fatty acid analysis, phenotypic characterization, and DNA—DNA hybridization experiments [9]. Geobacillus species have been isolated from a number of extreme environments including high-temperature oilfields [10], a corroded pipeline in an extremely deep well [11], African [12] and Russian [13] hot springs, marine vents [14], and the Mariana Trench [15], yet they can also be found in garden soils [16] and hay composts [17]., The ability of Geobacillus species to thrive in these varied and often hostile environments suggests that these species possess enzymes suitable for applications in challenging industrial environments. We therefore sequenced a number of these Geobacillus isolates including strains Y41MC52, Y41MC61, C56-T3, and Y4.1MC1 [18] to identify new enzymes suitable for use in biomass conversion into fuels and chemicals.
Organism information
Classification and features
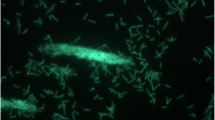
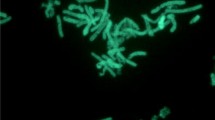
Geobacillus sp. Y412MC52 and Geobacillus sp. Y412MC61 are two thermophilic organisms isolated from Obsidian Hot Spring, Yellowstone National Park, Montana, USA (44.6100594° latitude and −110.4388217° longitude) under a sampling permit from the National Park Service. The hot spring possesses a pH of 6.37 and a temperature range of 42–90 °C. The organisms were isolated from a sample of hot spring water by enrichment and plating on YTP-2 medium [19] at 70 °C. The cultures are available from the Bacillus Genetic Stock Center as GSCID: 96A11 (MC52) and GSCID: 96A12 (MC61). Both cultures are routinely grown in YTP-2 medium media and maintained on YTP-2 agar plates. MC52, is a Gram-positive, rod-shaped facultative anaerobe (Table 1 and Additional file 1: Table S1), with optimum growth temperature of 65 °C and maximum growth temperature of 75 °C. MC52 appears to grow as a mixture of single cells and occasional large clumps of cells in liquid culture (Fig. 1). Growth is not observed on minimal medium supplemented with glucose, xylose or other sugars. Excellent growth is seen in Luria Broth, Terrific Broth, Tryptic Soy Broth and other common lab media with and without additional carbohydrate, indicating potential growth requirements for both vitamins and amino acids. Growth in YTP-2 medium is stimulated by addition of monosaccharides, disaccharides, soluble starch, xylan, arabinan, and arabinogalactan. Growth in YTP-2 medium is not stimulated by addition of cellulose, mannan, glucomannan, galactomannan, chitin, or pectin. MC52 produces extracellular xylanase when grown in YTP-2 medium supplemented with pyruvate, xylose, xylooligosaccharides and arabinogalactan. No secreted xylanase is detected when MC52 is grown in YTP-2 medium supplemented with glucose or arabinose. Extracellular arabinase is detected only in cultures grown in YTP-2 medium supplemented with arabinogalactan. Extracellular amylase is detected in cultures grown in YTP-2 medium supplemented with soluble starch or pullulan. Blue (positive) colonies of MC52 are observed on plates containing either 5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside or 5-bromo-4-chloro-3-indolyl-α-D-galactopyranoside, indicating production of α-galactosidase and β-galactosidase. Fluorescent colonies are observed on plates containing 4-methylumbelliferyl-β-D-cellobioside, 4-methylumbelliferyl-β-D-xylopyranoside, and 4-methylumbelliferyl-β-D-glucoyranoside indicating production of β-glucosidase and β-xylosidase.
Micrograph of Geobacillus sp. Y412MC52 cells showing individual cells and clumps of cells. Cells were grown in TSB plus 0.4 % glucose for 18 h. at 70 °C. A 1.0 ml aliquot was removed, centrifuged, re-suspended in 0.2 ml of sterile water, and stained using a 50 μM solution of SYTO® 9 fluorescent stain in sterile water (Molecular Probes). Dark field fluorescence microscopy was performed using a Nikon Eclipse TE2000-S epifluorescence microscope at 2000× magnification using a high-pressure Hg light source and a 500 nm emission filter
A phylogenetic tree was constructed to identify the relationship of Geobacillus sp. Y412MC52 and Geobacillus sp. Y412MC61 to other members of the Geobacillus family. MC52 and MC61 both contain eight annotated 16S rRNA genes. The 16S rRNA genes located at MC52 genome coordinates 11,820 through 13,365 and MC61 genome coordinates 10,516 through 12,061 were used for tree construction. Trees constructed with the remaining seven MC52 16S rRNA genes were identical to the tree shown here. The phylogeny was determined using the described 16S rRNA gene sequences, 16S rRNA gene sequences of the type strains of all validly described Geobacillus species and full-length 16S rRNA gene sequences of Geobacillus species present in GenBank. The 16S rRNA gene sequences were aligned using MUSCLE [20], pairwise distances were estimated using the Maximum Composite Likelihood approach, and initial trees for heuristic search were obtained automatically by applying the Neighbour-Joining method in MEGA 5 [21]. The alignment and heuristic trees were then used to infer the phylogeny using the Maximum Likelihood method based on Tamura-Nei [22]. The phylogenetic tree (Fig. 2) indicates that MC52, MC61 and Geobacillus sp. C56-T3 cluster separately from other validly named species.
The evolutionary history was inferred by using the Maximum Likelihood method based on the Tamura-Nei model [22]. The bootstrap consensus tree inferred from 500 replicates [45] is taken to represent the evolutionary history of the taxa analyzed [45]. Branches corresponding to partitions reproduced in less than 50 % bootstrap replicates are collapsed. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (500 replicates) are shown next to the branches [45]. Initial tree(s) for the heuristic search were obtained automatically by applying Neighbor-Join and BioNJ algorithms to a matrix of pairwise distances estimated using the Maximum Composite Likelihood (MCL) approach, and then selecting the topology with superior log likelihood value. The analysis involved 26 nucleotide sequences. All positions containing gaps and missing data were eliminated. There were a total of 1271 positions in the final dataset. Evolutionary analyses were conducted in MEGA5 [21]. The type strains of all validly described species are included (NCBI accession numbers): G. caldoxylosilyticus ATCC700356T (AF067651), G. galactosidasius CF1BT (AM408559), G. jurassicus DS1T (FN428697), G. kaustophilus NCIMB8547T (X60618), G. lituanicus N-3T (AY044055), G. stearothermophilus R-35646T (FN428694), G. subterraneus 34T (AF276306), G. thermantarcticus DSM9572T (FR749957), G. thermocatenulatus BGSC93A1T (AY608935), G. thermodenitrificans R-35647T (FN538993), G. thermoglucosidasius BGSC95A1T (FN428685), G. thermoleovorans DSM5366T (Z26923), G. toebii BK-1T (FN428690), G. uzenensis UT (AF276304) and G. vulcani 3S-1T (AJ293805). Additional16S rRNA sequences of G. thermoleovorans strain NP54 (JN871595G. thermoleovorans strain NP33 (JQ343209), G. thermoleovorans strain LEH-1 (NR_036985), G. thermocatenulatus strain DSM 730 (NR_119305), G. vulcani 3S-1 (NR_025426), G. strain C56-T3 (NC_014206), G. strain GHH01 (NC_020210), G. strain C56-YS93 (CP002835), and G. strain G11MC16 (CP002835)
Genome sequencing and annotation
Genome project history
Y412MC52 was selected for sequencing on the basis of its biotechnological potential as part of the U.S. Department of Energy Genomic Science program (formerly Genomics:GTL). The genome sequence is deposited in the Genomes On Line Database [23, 24] (GOLD ID = Gc01757), and in GenBank (NCBI Reference Sequence = CP002442.1). Sequencing, finishing and annotation were performed by the DOE Joint Genome Institute. A summary of the project information and its association with MIGS identifiers is shown in Table 2.
Growth conditions and genomic DNA preparation
For preparation of genomic DNA, cultures of Y51MC23 were grown from a single colony in YTP-2 in 1000 ml medium in a 2000 ml Erlenmeyer flask at 70 °C, 200 rpm for 18 h. Cells were collected by centrifugation at 4 °C and stored frozen until used for DNA preparation. The cell concentrate was lysed using a combination of SDS and proteinase K, and genomic DNA was isolated using a phenol/chloroform extraction method [25]. The genomic DNA was precipitated, and treated with RNase to remove residual contaminating RNA.
Genome sequencing and assembly
The genome of Geobacillus sp. Y412MC52 was sequenced at the Joint Genome Institute (JGI) using a combination of Sanger, Illumina and 454 technologies [26]. An Illumina GAii shotgun library with reads of 664 Mb, a 454 Titanium draft library with average read length of 250 bp, and two Sanger libraries with average insert size of 3 and 8 Kb were generated for this genome. Illumina sequencing data was assembled with VELVET [27], and the consensus sequences were shredded into 1.5 Kb overlapped fake reads and assembled together with the 454 data. Draft assemblies were based on 95.5 MB 454 draft data. Newbler parameters are - consed -a 50–1 350 -g -m -ml 20. The initial Newbler assembly contained 40 contigs in 18 scaffolds. We converted the initial 454 assembly into a phrap assembly by making fake reads from the consensus, collecting the read pairs in the 454 paired end library. The Phred/Phrap/Consed software package was used for sequence assembly and quality assessment [28–30] in the following finishing process. Illumina data was used to correct potential base errors and increase consensus quality using a software Polisher developed at JGI (Alla Lapidus, unpublished). After the shotgun stage, reads were assembled with parallel phrap (High Performance Software, LLC). Possible mis-assemblies were corrected with gapResolutioin (Cliff Han, unpublished), Dupfinisher, or sequencing cloned bridging PCR fragments with subcloning. Gaps between contigs were closed by editing in Consed, by PCR and by Bubble PCR primer walks. A total of 1069 additional reactions and 9 shatter libraries were necessary to close gaps and to raise the quality of the finished sequence. The overall average error rate achieved was 0.01 errors/10 Kb.
Genome annotation
Genes were identified using Prodigal [31] as part of the Oak Ridge National Laboratory genome annotation pipeline, followed by a round of manual curation using the JGI GenePRIMP pipeline [32]. The predicted CDSs were translated and used to search the National Center for Biotechnology Information (NCBI) nonredundant database, UniProt, TIGRFam, Pfam, PRIAM, KEGG, COG, and InterPro databases. These data sources were combined to assert a product description for each predicted protein. Non-coding genes and miscellaneous features were predicted using tRNAscan-SE [32], RNAMMer [33], Rfam [34], TMHMM [35], and signalP [35].
Genome properties
The genome of Geobacillus sp. Y412MC52 consists of one circular chromosome of 3,628,883 bp (Table 3 and Fig. 3) and an average G + C content of 52 % and one circular plasmid of 45,057 bp and an average G + C content of 45 % (Table 4). There are 88 tRNA genes, 25 rRNA genes and 3 “other” identified RNA genes. There are 3634 predicted protein-coding regions and 175 pseudogenes in the genome. A total of 2569 genes (68.51 %) have been assigned a predicted function while the rest have been designated as hypothetical proteins (Table 4). The numbers of genes assigned to each COG functional category are listed in Table 5. About 35 % of the annotated genes were not assigned to a COG or have an unknown function.
Insights from the genome sequence
Average Nucleotide Identity (ANI) calculations [36] were used to compare the genomes of MC52 and other sequenced Geobacillus species. The comparison of the MC52 genome to the other genomes (Table 6) confirms the phylogenetic tree obtained using 16S rRNA genes. MC52 is most closely related to MC61 (100 % identity) followed by Geobacillus sp. C56-T3 (98.3 %). These values are above the species cutoff value of 98.2 % to 99.0 % [37] indicating that these are most likely strains of the same species. The ANI values for all other sequenced strains are below 98 %, suggesting that MC52, MC61, and C56-T3 represent members of a new species. Comparison of genes shows MC52 and MC61 share 3329 genes (Fig. 4). MC52 has 52 unique genes and MC61 has 48. These unique genes code mostly for hypothetical proteins and are randomly distributed throughout both genomes. Alignment of the MC52 and M61 genomes using progressiveMauve [38] shows one predominant, four medium, and two small Locally Collinear Blocks of conserved genes (Fig. 5). In Y412MC61, two of the medium blocks precede the predominant block, while these blocks follow the predominant block in Y412MC52. In addition to having alternate locations within these genomes, these two blocks reverse their orientation between the two genomes. Taken together, these results indicate that MC52 and M61 are not two different isolates of the same strain, but are two closely related strains of the same species with a unique relationship to each other.
Venn Diagram of Y412MC52 and Y412MC61 determined using software at https://edgar.computational.bio.uni-giessen.de
MC52 possesses a 45-gene arabinan and xylan degradation cluster that allows degradation of hemicellulose components of biomass (GYMC52_1817 through GYMC52_1867). The cluster contains one secreted xylanase (GYMC52_1825) and one secreted arabinase (GYMC52_1858), in agreement with the experimental results. The organization of the xylan degradation portion of the cluster matches the glucuronic acid utilization cluster described for G. stearothermophilus [39]. The arabinan degradation part of the cluster is smaller than the arabinan cluster of G. stearothermophilus [40], lacking araP, araS, araT, araE, araG and araH genes. MC52 also possesses three clusters annotated for degradation of aromatic acid molecules, GYMC52_1956 through GYMC52_1962, GYMC52_1990 through GYMC52_2001, and GYMC52_3134 through GYMC52_3141. Geobacillus species utilize xylan by transporting large xylooligosaccharides into the cell and then degrading these xylooligosaccharides intracellularly [39]. These aromatic acid degradation clusters may allow degradation and utilization of lignin fragments such as ferulic, sinapic, and cinnamic acids that are attached to the xylooligosaccharides. Utilization of these aromatic acids increases the metabolic energy obtained from the fragments and eliminates potential toxicity of these aromatic acids. Transport and metabolic clusters for utilization of cellobiose and related oligosaccharides (GYMC52_1797 through GYMC52_1801), α- and β-galactooligosaccharides (GYMC52_12121 through GYMC52_2132), and α-1,4-linked glucooligosaccharides (GYMC52_06321 through GYMC52_0637) were identified, confirming the experimental observations of the corresponding enzymatic activities.
The smaller arabinan cluster in MC52 is the result of an 11-gene insert (GYMC52_1870 through GYMC52_1880) coding for a peptide utilization cluster that replaces part of the arabinan cluster. This peptide utilization cluster is found in only a few Geobacillus strains, including Geobacillus sp. Y412MC61 (GYMC61_2740 through GYMC52_2750), Geobacillus sp. Y4.1MC1 (GY4MC1_2192 through GY4MC1_2202), and Geobacillus sp. C56-YS93 (Geoth_2276 through Geoth_2288). The cluster does not code for a secreted protease or peptidase, but contains an annotated five-gene ABC peptide transporter system and two intracellular peptidases.
Geobacillus strain Y412MC52 possesses a 54.4 Kb, 73-gene insert that codes for 47 phage genes identified using phast [41, 42] phage identification software (Fig. 5), an identical insert is present in Y412MC61. The prophage insert has 39 % coverage and 83 % identity to Geobacillus phage E2 (GenBank NC_009552) [43], isolated from a deep sea location. The phage is not present in Geobacillus strain C56-YS93 also isolated from Obsidian Hot Spring, indicating the phage may have a limited range of hosts in the hot spring.
Conclusions
Obsidian Hot Spring is home to a wide variety of organisms, including Paenibacillus lautus Y412MC10 [19], Geobacillus thermoglucosidans C56-YS93 (manuscript submitted) and Geobacillus sp. Y412MC52 and Y412MC61. Especially of interest is the isolation of both low G + C (C56-YS93, 43.9 % G + C) and high G + C (Y412MC52 and Y412MC61, 52.3 % G + C) xylanolytic Geobacillus species from the same hot spring sample. This suggests that the high and low G + C Geobacillus species may occupy separate ecological niches that allow each strain to thrive in the same site. Based on the genomic analysis, Geobacillus sp. Y412MC52 appears to utilize only some biomass components such as xylan, arabinoglucuronoxylan, and the arabinan component of arabinogalactan. MC52 shows no genes coding for utilization of other biomass components such as cellulose, mannan, glucomannan, galactomannan, chitin, or pectin, confirming experimental observations. The limited range of substrates suggests MC52 functions as part of a microbial consortium in degrading biomass. The presence of aromatic acid metabolic clusters and the lack of mannan-utilization clusters suggest the organism has a preference for utilization of hemicellulose derived from grassy plants rather than woody plants.
Based on 16S rRNA genes and average nucleotide identity, Geobacillus sp. Y412MC52 and the related Geobacillus sp. Y412MC61 appear to be members of a new species of Geobacillus . The presence of multiple 16S rRNA genes in Geobacillus species as well as the small differences observed in 16S rRNA gene sequences makes assignment of strains to new or existing species difficult. Utilization of recN sequences [44] has been proposed as an alternative to 16S rRNA gene sequences, but it is unclear if this leads to a more accurate description of the distinct species. Sequencing of additional genomes and in-depth microbiological characterizations are needed to clarify the relationships among Geobacillus species.
Abbreviations
- MC52:
-
Geobacillus sp. Y412MC52
- MC61:
-
Geobacillus sp. Y412MC61
References
Valls C, Gallardo O, Vidal T, Pastor FI, Diaz P, Roncero MB. New xylanases to obtain modified eucalypt fibres with high-cellulose content. Bioresour Technol. 2010;101(19):7439–45.
Valls C, Roncero MB. Using both xylanase and laccase enzymes for pulp bleaching. Bioresour Technol. 2009;100(6):2032–9.
Tricarico JM, Dawson KA. Influence of supplemental endoglucanase or xylanase on volatile fatty acid production from ruminant feed by ruminal in vitro cultures. Arch Anim Nutr. 2005;59(5):325–34.
Brock TD, Freeze H. Thermus aquaticus gen. n. and sp. n., a nonsporulating extreme thermophile. J Bacteriol. 1969;98(1):289–97.
Brock TD, Edwards MR. Fine structure of Thermus aquaticus, an extreme thermophile. J Bacteriol. 1970;104(1):509–17.
Williams RA, Smith KE, Welch SG, Micallef J, Sharp RJ. DNA relatedness of Thermus strains, description of Thermus brockianus sp. nov., and proposal to reestablish Thermus thermophilus (Oshima and Imahori). Int J Syst Bacteriol. 1995;45(3):495–9.
Mohagheghi A, Grohmann K, Himmel M, Leighton L, Updegraff DM. Isolation and characterization of Acidothermus cellulolyticus gen. nov., sp. nov., a new genus of thermophilic, acidophilic, cellulolytic bacteria. Int J Syst Bacteriol. 1986;36:435–43.
Brock TD. The value of basic research: discovery of Thermus aquaticus and other extreme thermophiles. Genetics. 1997;146(4):1207–10.
Nazina TN, Tourova TP, Poltaraus AB, Novikova EV, Grigoryan AA, Ivanova AE, et al. Taxonomic study of aerobic thermophilic bacilli: descriptions of Geobacillus subterraneus gen. nov., sp. nov. and Geobacillus uzenensis sp. nov. from petroleum reservoirs and transfer of Bacillus stearothermophilus, Bacillus thermocatenulatus, Bacillus thermoleovorans, Bacillus kaustophilus, Bacillus thermodenitrificans to Geobacillus as the new combinations G. stearothermophilus, G. thermocatenulatus, G. thermoleovorans, G. kaustophilus, G. thermoglucosidasius and G. thermodenitrificans. Int J Syst Evol Microbiol. 2001;51(Pt 2):433–46.
Kuisiene N, Raugalas J, Chitavichius D. Geobacillus lituanicus sp. nov. Int J Syst Evol Microbiol. 2004;54(Pt 6):1991–5.
Popova NA, Nikolaev Iu A, Turova TP, Lysenko AM, Osipov GA, Verkhovtseva NV, et al. Geobacillus uralicus, a new species of thermophilic bacteria. Mikrobiologiia. 2002;71(3):391–8.
Hawumba JF, Theron J, Brozel VS. Thermophilic protease-producing Geobacillus from Buranga hot springs in Western Uganda. Curr Microbiol. 2002;45(2):144–50.
Nazina TN, Lebedeva EV, Poltaraus AB, Tourova TP, Grigoryan AA, Sokolova D, et al. Geobacillus gargensis sp. nov., a novel thermophile from a hot spring, and the reclassification of Bacillus vulcani as Geobacillus vulcani comb. nov. Int J Syst Evol Microbiol. 2004;54(Pt 6):2019–24.
Maugeri TL, Gugliandolo C, Caccamo D, Stackebrandt E. Three novel halotolerant and thermophilic Geobacillus strains from shallow marine vents. Syst Appl Microbiol. 2002;25(3):450–5.
Takami H, Nishi S, Lu J, Shimamura S, Takaki Y. Genomic characterization of thermophilic Geobacillus species isolated from the deepest sea mud of the Mariana Trench. Extremophiles. 2004;8(5):351–6.
Wiegand S, Rabausch U, Chow J, Daniel R, Streit WR, Liesegang H. Complete genome sequence of Geobacillus sp. Strain GHH01, a thermophilic lipase-secreting bacterium. Genome Announc. 2013;1(2):e0009213.
Sung MH, Kim H, Bae JW, Rhee SK, Jeon CO, Kim K, et al. Geobacillus toebii sp. nov., a novel thermophilic bacterium isolated from hay compost. Int J Syst Evol Microbiol. 2002;52(Pt 6):2251–5.
Brumm P, Land M, Hauser LJ, Jeffries C, Chang YJ, Mead D. Complete genome sequence of Geobacillus strain Y4.1MC1, a novel co-utilizing geobacillus thermoglucosidasius strain isolated from bath hot spring in Yellowstone National Park. BioEnerg Res. 2015;8(3):1039–45.
Mead DA, Lucas S, Copeland A, Lapidus A, Cheng JF, Bruce DC, et al. Complete genome sequence of Paenibacillus strain Y4.12MC10, a novel Paenibacillus lautus strain isolated from Obsidian Hot Spring in Yellowstone National Park. Stand Genomic Sci. 2012;6(3):381–400.
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32(5):1792–7.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 2011;28(10):2731–9.
Tamura K, Nei M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol. 1993;10(3):512–26.
Liolios K, Tavernarakis N, Hugenholtz P, Kyrpides NC. The Genomes On Line Database (GOLD) v. 2: a monitor of genome projects worldwide. Nucleic Acids Res. 2006;34(Database issue):D332–4.
Liolios K, Chen IM, Mavromatis K, Tavernarakis N, Hugenholtz P, Markowitz VM, et al. The Genomes On Line Database (GOLD) in 2009: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res. 2010;38(Database issue):D346–54.
Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: a laboratory manual. NY: Cold Spring Harbor Laboratory Press; 1989.
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature. 2005;437(7057):376–80.
Zerbino DR. Using the Velvet de novo assembler for short-read sequencing technologies. Curr Protoc Bioinformatics. 2010;Chapter 11:Unit 11.5.
Ewing B, Green P. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 1998;8(3):186–94.
Ewing B, Hillier L, Wendl MC, Green P. Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 1998;8(3):175–85.
Gordon D, Abajian C, Green P. Consed: a graphical tool for sequence finishing. Genome Res. 1998;8(3):195–202.
Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal prokaryotic dynamic programming genefinding algorithm. BMC Bioinformatics. 2010;11:119.
Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997;25(5):955–64.
Lagesen K, Hallin P, Rodland EA, Staerfeldt HH, Rognes T, Ussery DW. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 2007;35(9):3100–8.
Griffiths-Jones S, Bateman A, Marshall M, Khanna A, Eddy SR. Rfam: an RNA family database. Nucleic Acids Res. 2003;31(1):439–41.
Krogh A, Larsson B, von Heijne G, Sonnhammer EL. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol. 2001;305(3):567–80.
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol. 2007;57(Pt 1):81–91.
Kim M, Oh HS, Park SC, Chun J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol. 2014;64(Pt 2):346–51.
Darling AE, Mau B, Perna NT. ProgressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One. 2010;5(6):e11147.
Shulami S, Gat O, Sonenshein AL, Shoham Y. The glucuronic acid utilization gene cluster from Bacillus stearothermophilus T-6. J Bacteriol. 1999;181(12):3695–704.
Shulami S, Raz-Pasteur A, Tabachnikov O, Gilead-Gropper S, Shner I, Shoham Y. The L-Arabinan utilization system of Geobacillus stearothermophilus. J Bacteriol. 2011;193(11):2838–50.
Hubisz MJ, Pollard KS, Siepel A. PHAST and RPHAST: phylogenetic analysis with space/time models. Brief Bioinform. 2011;12(1):41–51.
Zhou Y, Liang Y, Lynch KH, Dennis JJ, Wishart DS. PHAST: a fast phage search tool. Nucleic Acids Res. 2011;39(Web Server issue):W347–52.
Wang Y, Zhang X. Characterization of a novel portal protein from deep-sea thermophilic bacteriophage GVE2. Gene. 2008;421(1–2):61–6.
Zeigler DR. Application of a recN sequence similarity analysis to the identification of species within the bacterial genus Geobacillus. Int J Syst Evol Microbiol. 2005;55(Pt 3):1171–9.
Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985;10:512–26.
Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol. 2008;26(5):541–7.
Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci U S A. 1990;87(12):4576–9.
Priest FG, Goodfellow M, Todd C. A numerical classification of the genus Bacillus. J Gen Microbiol. 1988;134(7):1847–82.
Ash C, Priest FG, Collins MD. Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Proposal for the creation of a new genus Paenibacillus. Antonie Van Leeuwenhoek. 1993;64(3–4):253–60.
Camon E, Magrane M, Barrell D, Lee V, Dimmer E, Maslen J, et al. The Gene Ontology Annotation (GOA) database: sharing knowledge in uniprot with gene ontology. Nucleic Acids Res. 2004;32(Database issue):D262–6.
Markowitz VM, Chen IM, Palaniappan K, Chu K, Szeto E, Pillay M, et al. IMG 4 version of the integrated microbial genomes comparative analysis system. Nucleic Acids Res. 2014;42(Database issue):D560–7.
Acknowledgements
This work was funded by the DOE Great Lakes Bioenergy Research Center (DOE Office of Science BER DE-FC02-07ER64494). Sequencing work was performed under the auspices of the US Department of Energy’s Office of Science, Biological and Environmental Research Program, and by the University of California, Lawrence Berkeley National Laboratory under contract No. DE-AC02-05CH11231, Lawrence Livermore National Laboratory under Contract No. DE-AC52-07NA27344, and Los Alamos National Laboratory under contract No. DE-AC02-06NA25396.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
This work was funded by the DOE Great Lakes Bioenergy Research Center (DOE Office of Science BER DE-FC02-07ER64494, but did not have any additional role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript, DM is an employee and shareholder of Lucigen Corporation and PB is an employee and owner of C5•6 Technologies LLC. Neither company had any role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Authors’ contributions
PB isolated the organism, prepared genomic DNA, performed microbial characterization and drafted the manuscript. ML, LH, CJ and YC performed the genome sequencing, assembly, and annotation. DM conceived of the study, and participated in its design and coordination and helped to draft the manuscript. All authors read and approved the final manuscript.
Additional file
Additional file 1: Table S1.
Associated MIGS record. (DOCX 33 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Brumm, P., Land, M.L., Hauser, L.J. et al. Complete genome sequences of Geobacillus sp. Y412MC52, a xylan-degrading strain isolated from obsidian hot spring in Yellowstone National Park. Stand in Genomic Sci 10, 81 (2015). https://doi.org/10.1186/s40793-015-0075-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40793-015-0075-0