Abstract
Considering a large class of muscle contraction models accounting for actin–myosin interaction, we present a mathematical setting in which solution properties can be established, including fundamental thermodynamic balances. Moreover, we propose a complete discretization strategy for which we are also able to obtain discrete versions of the thermodynamic balances and other properties. Our major objective is to show how the thermodynamics of such models can be tracked after discretization, including when they are coupled to a macroscopic muscle formulation in the realm of continuum mechanics. Our approach allows to carefully identify the sources of energy and entropy in the system, and to follow them up to the numerical applications.
Similar content being viewed by others
Introduction
The modeling of the active mechanical behavior of muscles has been the object of intense research since the seminal work of Huxley [12] modeling the attachment-detachment process in the actin–myosin interaction responsible for sarcomere contraction. Then, numerous extensions—mostly based on refinements of the chemical process introduced by Huxley—of the previous model have been proposed in order to take into account different time scales of the actin–myosin interaction. In particular several models have been developed to account for the power stroke phenomenon [4, 5, 13, 20]. In parallel, the question of the thermodynamic balances associated with the chemical machinery was intensively studied, notably with the fundamental contributions of Hill [10, 11]. Note that these models are specific cases of molecular motors models without the natural diffusion introduced by the Fokker–Plank equation [2, 3, 14, 18]. In this paper, our objective is to develop a formalism allowing to derive these thermodynamic balances for Huxley’s model and its extensions with an additional tracking of these balances at the discrete level after time-discretizing the model dynamics. Moreover, we present how these microscopic models can be incorporated into a macroscopic model of muscle fibers in the spirit of [2] with the aim of following these thermodynamic balances at the macroscopic level for the continuous-time dynamics but also after adequate time discretization. This last part is general with respect to the chemical microscopic model of interest and could also be extended to similar types of models [3, 19], or those mixing mechanical and chemical modeling elements, for instance [1, 17, 22].
The outline of the paper is as follows. The first section presents the modeling ingredients of the microscopic models of actin–myosin interaction and we derive in a second section the fundamental properties of these models with the associated thermodynamic balances, up to the coupling with the macroscopic mechanical formulation. The third section then describes the discretization scheme and justifies its thermodynamic compatibility. Finally, the last section illustrates our results with numerical investigations.
Modeling of muscle contraction
Physiology of muscle contraction
Muscles are multi-scale structures in which motion is initiated at the cellular level by the relative sliding between two types of filaments: actin filaments and myosin filaments. At the surface of the myosin filament, myosin heads can bind to the actin filament. The actin filament has a periodic structure with regularly spaced attachment sites. The interaction between myosin heads and actin sites occurs in a cyclic manner [16], see Fig. 1. The cycle includes attachment and detachment of the myosin head to and from an actin site and a conformation change of the attached myosin head called the power stroke. The detachment stage requires an energy input obtained from ATP molecules buffered inside the cell.
Different levels of description of the actin–myosin interaction can be considered benefiting from the fact that the power stroke occurs much faster than the attachment and detachment processes.
Huxley’57 model
In his seminal work [12], Huxley describes the myosin head with two chemical states representing the attached and detached configurations. Each myosin can interact with its closest actin site only. The transition rates—for attachment and detachment—depend only on the distance from the myosin head rest position to its nearest attachment site denoted by \( s \). We denote by \( d_a \) the distance between two consecutive attachment sites. The distance \( s \) thus lies in an interval of width \( d_a \), not necessarily symmetric but containing 0, that we denote by \( [s^- , s^+] \)—see Fig. 2.
Huxley’57 model representation. Top: definition of the transitions between the attached state (A) and the detached state (D). Bottom left: model parametrization, representation in the detached state. Bottom right: model parametrization, representation in the attached state. The position of the actin site corresponding to \( s = 0 \) is represented by a thick dashed line
Considering, in a population of myosin heads, the subset of heads with rest position located at distance \( s \) from their nearest attachment site, we define by \( { a(t,s) } \) the ratio of actually attached heads at time \( t \). Equivalently, the ratio of detached heads is denoted by \( d(s,t) = 1 - a(s,t)\), due to the assumption that both types of filaments are rigid. The sliding velocity \( v_c \) between the filaments is a macroscopic variable, hence independent of s and often quasi-static with respect to the microscopic time scales. We refer to “Coupling with a macroscopic model of muscle fiber” section for an illustrating example of coupling between a macroscopic formulation and such a microscopic model.
The detached state is associated with a constant energy level \( w_0 \) and the attached state with an energy \( w_1 \) that depends on the distance \( s \)—the myosin head bound to actin is modeled as an elastic spring. This is where mechanics enters the model, and we point out that we extend here the original Huxley’57 model by allowing the spring to have a non-linear behavior. The myosin head is brought back to the initial detached energy level by the ATP energy input \( \mu _T \).
Transition rates between the states satisfy the detailed balance, i.e. for a transition \( i \) (\( i = 1,2 \), see Fig. 2) from a state of energy \( { w_j(s)} \) to a state of energy \( { w_k(s)} \), the forward and reverse rates—respectively denoted by \( k_{i} \) and \( k_{-i} \)—must satisfy the relation
where \( k_B \) is the Boltzmann constant and \( T \) is the absolute temperature. A schematic of the model is presented in Fig. 2.
The conservation of matter, assuming that there is no coupling between the myosin heads, leads to the following dynamical system, for all \( t > 0 \) and all \( s \in [s^-, s^+] \)
from adequate initial conditions \( a(s, 0) = a^{0}(s) \) and \( d(s, 0) = d^{\,0}(s) \), to be specified later. The assumption that the myosin head can only interact with its nearest actin site imposes that the probability of being attached on the boundaries of the interval \( [s^-, s^+] \) must be zero. Physically, the property \( a(s^-, t) = a(s^+, t) = 0 \) appears when the attachment rates \( k_1(s) \) and \( k_{-2}(s) \) vanish, while the detachment rates \( k_{-1}(s) \) and \( k_{2}(s) \) go to infinity on the boundaries of the interval \( [s^-, s^+] \). Note that the energy levels and the transition rates are linked by the detailed balance (1), which implies that the energy of the attached level goes to infinity on the boundaries of the interval \( [s^-, s^+] \). In a nutshell, the parameter functions must satisfy
the second line enforcing that all heads are detached at the boundaries, see “Model properties based on thermodynamics principles” section [Eqs. (9) and (10)]. This implies energetically that
Actin sites and myosin heads are located at discrete locations separated by regular intervals along their respective filaments. The spatial periodicities are, however, different on each filament. Therefore, for a large population of heads, the distribution of their distance to the nearest actin site can be assumed to be uniform in the interval \( [s^-, s^+] \), and the average tension developed per myosin head is given by
This force can then typically lead to a macroscopic active stress tensor and link to macro-scale models of muscle tissue as presented in [2] or in “Coupling with a macroscopic model of muscle fiber” section.
Extension of Huxley’57 model
To obtain a behavior closer to physiology, and in particular to capture the power stroke, various extensions of Huxley’57 model have been proposed [4, 5, 13, 20]. These extensions can use more than two states to describe the myosin head and allow interactions with an arbitrary number of attachment sites. In this section, our objective is to present these models in a general form, albeit close to the initial 2-state Huxley’s model, in particular concerning their general mathematical and mechanical properties.
Multi-state models
A general formulation of these models considers \( N_s \) chemical states \( \{ X_p \}_{1 \le p \le N_s} \), that we can separate into two categories: attached states and detached states. The states are involved in \( N_r \) reactions between the states in the form
The collection of reactions between the states can be represented by a complete directed graph \( G \). A complete directed graph is a set of vertices connected by edges, in which: edges have a direction; for each edge of the graph, the edge connecting the same vertices in the inverse direction also belongs to the graph; no vertex is connected to itself. We respectively denote by \( V \) and \( E \) the sets of vertices and edges of the complete directed graph, and we write \( G = (V,E) \). This graph is made of \( N_s \) vertices and \( 2 N_r \) edges. A transition \( X_{p_i} \rightarrow X_{q_i} \) is associated with the edge \( E_{p_i q_i} \), whereas the reverse transition \( X_{q_i} \rightarrow X_{p_i} \) is associated with the edge \( E_{q_i p_i} \). The reaction presented in (5), which is bidirectional, is associated with the edges \( E_{p_i q_i} \) and \( E_{q_i p_i} \). The subsets of vertices corresponding to attached and detached states are respectively denoted by \( V_{\mathrm {a}} \) and \( V_{\mathrm {d}} \).
The ratio of heads in state \( X_q \) located at \( s \) at time \( t \) is denoted by \( x_q(s,t) \). We define the chemical flux between states \( X_{p_i} \) and \( X_{q_i} \) through transition \( i \) by
Note that we have
The system dynamics is then governed by
The Huxley’57 model presented in “Huxley’57 model” section can naturally be seen as a particular case of multi-state models with only one attached state and one detached state. The graph \( G \) associated with the Huxley’57 model is given by
Here, we use superscripts in the edges definition to denote that there are two reactions between the same vertices.
Multi-site models
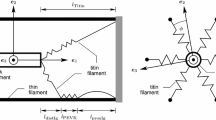
In this further generalization, it is assumed that a myosin head can interact not only with its nearest actin site —located by definition at distance \( s \)—but also with all other actin sites located at distance \( \{s + j d_a\}_{j \in \mathbb {Z}^*} \). A myosin head can thus be detached in state \( q \)—with a probability \( x_q(s) \)—or attached in state \( q \) at a distance \( s + jd_a \) of its rest position—with a probability \( x_q(s + j d_a) \). We extend the definition of the ratio of heads in detached states by periodicity, i.e. if state \( q \) is a detached state \( x_q(s + j d_a) = x_q(s), \, \forall j\). We also refine the description of the graph defined for multi-site models, by splitting the set of edges \( E \) between the edges linking two detached states \( \hat{E} \) and the remaining edges \( \overline{E} \). We define the associated complete directed graphs \( G_d = (V, \hat{E})\) and \( G_a = (V, \overline{E})\). The system dynamics is governed by
The ratios of attached head \( x_q \) for \( V_q \in V_a \) are defined on \( \mathbb {R}\) and must vanish at infinity. By contrast, the ratios of detached head \( x_q \) for \( V_q \in V_d \) are defined on \( [s^-, s^+] \) with periodic boundary conditions.
Example of a multi-state, multi-site model:Piazzesi–Lombardi’95
A specific representative—denoted PL95—of this family of models has been derived in [20] with the aim of accounting for the energetics of muscle contraction. It describes the myosin head with five states arranged in two cycles of chemical reactions, see Fig. 3. The five states are composed of three attached states \( A_1 \), \( A_2 \) and \( A_3 \) and two detached states \( D_1 \) and \( D_2 \). A first long cycle (cycle a) is meant to represent a complete power stroke, while a short cycle (cycle b) allows the myosin head to cycle at small or zero sliding velocity with incomplete power stroke.
An energy \( \mu _T \) is brought to the myosin head by ATP in the transitions \( 2 \rightarrow 5 \) and \( 3 \rightarrow 4 \).
The graph \( G \) associated with this model is given by
Moreover, it is assumed in this model that the myosin can attach to an arbitrary number of actin sites, hence it is also multi-site.
We denote by \( w_q \) the energy associated with the state of vertex \( q \).
Model properties based on thermodynamics principles
From conservation of matter to boundary conditions and monotonicity properties
Let us consider the Huxley’57 model and derive its fundamental properties. System (2) was derived from the conservation of matter, hence we directly verify that for all (s, t), \( a(s,t) + d(s,t) = 1\) as soon as we choose our initial condition \(\forall s \in [s^-, s^+],\,a^0(s) \in [0,1]\) and \( d^0(s) = 1-a^0(s)\), since
where we defined the total derivative by \( \mathrm {d}/\mathrm {d}t(\bullet ) = \partial /\partial t(\bullet ) + v_c \partial /\partial s(\bullet ) \). Therefore, we can rewrite the system (2) in the form of a single equation
where we denote the aggregated transition rates \( k_{+}(s) = k_1(s) + k_{-2}(s) \) and \( k_{-}(s) = k_{2}(s) + k_{-1}(s) \).
Boundary values
As explained in our model presentation, we expect the myosin head to only interact with the nearest actin site, which imposes that the probability of being attached must vanish on the boundaries of the interval \( [s^-, s^+] \). However, the dynamics (2) is a first-order transport equation associated with only one boundary condition. Therefore, we can either consider one single Dirichlet boundary condition at one end of the interval—i.e., in \( s^- \) if \( v_c > 0 \) and \( s^+ \) if \( v_c < 0 \)—and then rely on the conditions (3) to obtain the proper value of the solution at the other end—as a property—or alternatively consider periodic boundary conditions. As the first option yields a periodic solution, it is clear that the two options are equivalent. However, they differ at the discrete level, in which case we will have to make a choice, see “A numerical scheme for Huxley’57 model” section.
In fact, closed-form expressions can be obtained for the solution. To fix the ideas in this derivation, we consider the case \( v_c \ge 0 \), although the same result can be obtained similarly for \( v_c < 0 \). As \(v_c\) is assumed to be constant, the method of characteristic lines gives regular \(C^1\) solutions from regular enough initial condition \(a^0\). Considering \( a(s,t) \) solution of (7), we define the function \( \tilde{a} \) by
which satisfies the equation
where we define \( h(s) = \int _0^s (k_+(\xi ) + k_{-}(\xi )) \, \mathrm {d}\xi \). Solving (8) along a characteristic line and pulling back the result to \( a(s,t) \) we obtain, defining \( t_0 = \max (0, t - \frac{s - s^-}{v_c}) \),
We know that the aggregated attachment rate \( k_+ \) is a continuous function on \( [s^-, s^+] \) and goes to zero on the boundaries of \( [s^-, s^+] \). Therefore, under the condition on the aggregated detachment rate \( \int _{0}^{s^+} k_{-}(\xi ) \, \mathrm {d}\xi = \infty \), we deduce that
using the dominated convergence theorem for the second term of (9). Likewise, the property \( \lim _{s \rightarrow s^-} a(s,t) = 0 \) is obtained for \( v_c < 0 \), and a similar result can be obtained in a similar manner with periodic boundary conditions.
Positivity and boundedness properties
We want to check that the solution has values consistent with ratio quantities. More specifically, we want that, with an initial condition \( a^0(s) \in [0,1] \), the property \( a(s,t) \in [0,1] \) holds. Again, we rely on the solution obtained by the method of characteristic lines (9). As the transition rates and the initial condition are positive, we find that \( a(s,t) \ge 0 \). Then, noting that \( 1 - a(s,t) \) is governed by an equation of the same form as (7) with the initial condition \( 1 - a^0(s) \ge 0 \), we similarly deduce that \( a(s,t) \le 1 \).
First principle
We now want to establish a first thermodynamic property of the Huxley’57 system (2), namely, a first principle, and in this respect we follow the approach proposed by [10]. We consider a system made of a population of myosin heads and define the average energy per myosin head, namely
We then define the chemical fluxes
We will henceforth make the natural assumption that the reaction rates are chosen in order for \(w_1 a\), \(k_{2} a\) and \(k_{-1} a\) to tend to zero when s tends to \(s^{-}\) and \(s^{+}\), with the physical interpretation that no finite energy (\(w_1 a\)) is stored and no detachment flux (\(k_{2} a\) and \(k_{-1} a\)) occurs at the ends of the interval. We will see in “Numerical results and discussion” section that this assumption is easily satisfied in practice when (3) holds. Then, computing the time derivative, we obtain
Using integrations by parts for the transport terms, the boundary properties of the solution—\( w_1(s) a(t,s^-) = w_1(s) a(t,s^+) = 0 \) and \( d(t,s^-) = d(t,s^+)\)—and considering that detachment is associated with the consumption of one ATP, we obtain
where we denoted by
the mean net influx of ATP. Finally, we derive the following formulation of the first principle
where
with the active force \( \tau _c \) defined in (4). The quantity \( \dot{\mathcal {W}} \) is the rate of work given to the system and \(\dot{\mathcal {E}}(t) = \mu _T \overline{J_{2}}(t)\) corresponds to the input flux in energy brought by ATP hydrolysis. The remaining term \( \dot{\mathcal {Q}} \) can be identified with a heat flux. In steady-state shortening (\(\tau _c >0\) and \(v_c < 0\)), the work is negative and in physiological conditions, we expect the energy input term to be positive, and the heat transfer to be negative (see numerical illustrations in “Numerical results and discussion” section). The energy balance can be interpreted as follows: the energy brought by ATP is for one part converted into work, the other part being dissipated as heat production.
Second principle
Let us now derive a second principle thermodynamic balance. We introduce the system entropy as
where \(k_{B}\) is the Bolztmann constant. The system remains at a constant temperature, the outside environment playing the role of a thermostat. We introduce the Helmholtz free energy
which therefore corresponds to
where
are the chemical potentials. Computing the time derivative, we get
Using again integrations by part for the transport terms and the boundary properties of the solution, we obtain
Independently, we also have
Combining Eqs. (14) and (16) and the first principle, we can write
The second principle then reads
where we naturally associate the second term of (17) with the entropy production \( \dot{\mathcal {S}}_{\mathrm {prod}}(t) \).
The model will be compatible with the second principle if this entropy production is always positive. Using the relation (1) deduced from the detailed balance, we recall that
Thus, when introducing the ratio of the one-way fluxes for transition 1, \(J_{1+}\) and \(J_{1-}\), defined by \(J_{1+}(s,t) = k_1(s,t) d(s,t) \) and \(J_{1-}(s,t) = k_{- 1}(s,t) a(s,t) \), we find
As a consequence, we have two cases. If \(\mu _1(s,t) \ge \mu _0(s,t)\), we find that
Conversely, \(\mu _1(s,t) \le \mu _0(s,t)\) implies that \(J_{1}(s,t) \ge 0\). Proceeding in the same way for the second reaction, we finally have
We thus obtain the conclusion that the entropy production term
hence that the model is compatible with the second principle. We can summarize this property using (15) by the free energy balance
Extension to multi-state, multi-site models
Let us now consider the Piazzesi–Lombardi’95 model. We want to establish the thermodynamic balances associated with this model.
Fundamental properties of the solution—in particular the monotonicity properties of the results established for the Huxley’57 model, see “From conservation of matter to boundary conditions and monotonicity properties” section—can be extended to this model. In particular, conservation of matter here reads
First principle
We define the energy as
The time derivative reads, after integrating by parts the transport term
with \(W_{pq} = w_q - w_p\) and the active force defined as
We then obtain
where the mean fluxes are given by
and \(\tilde{W}_{pq} = W_{pq} - \mu _T \delta _{pq=\{25,34\}}\) as an energy \( \mu _T \) is brought to the myosin head by ATP during the transitions \( 2 \rightarrow 5 \) and \( 3 \rightarrow 4 \). Note that the introduction of the modified energy increments \( \tilde{W} \) brings out the input energy fluxes \(\mu _T \big (\overline{J_{25}}(t) + \overline{J_{34}}(t) \big )\). The first principle then naturally reads
with
Second principle
Following the work done for the Huxley’57 model, we here define the entropy of the system as
Then, we define the Helmholtz free energy as \(\mathcal {F}(t) = \mathcal {U}(t) - T \mathcal {S}(t)\), which can be written as
with the definitions
We here define the entropy production as
where \(\tilde{M}_{pq} = \mu _q -\mu _p - \mu _T \delta _{pq=\{34,25\}}\). Then, combining the first principle (25) with the identity \(\dot{F} = \dot{U} - T \dot{S}\), we finally obtain the second principle
Similarly as in the Huxley’57 model, the detailed balance ensures that the entropy production is always positive and the model is thus thermodynamically compatible. We have, as in (23), the free energy balance
Coupling with a macroscopic model of muscle fiber
The thermodynamic properties of these classes of models are very useful when coupling them with a macroscopic model, typically to represent a muscle fiber, as it will ensure a global consistent thermodynamic balance between macroscopic and microscopic contributions. Let us consider, indeed, a macroscopic model of muscle fiber modeled in the realm of non-linear continuum mechanics, as large deformations frequently occur in muscle fibers. The material points coordinates are denoted by \(\underline{\mathrm {x}} \in \varOmega ^0\) in the reference configuration. The displacement field associated with the deformation map is denoted by \(\underline{y}\). We denote by \(\underline{\underline{e}}\) the Green-Lagrange strain tensor, i.e.
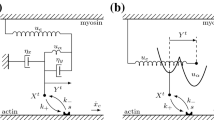
and the second Piola-Kirchhoff stress tensor is denoted by \(\underline{\underline{\varSigma }}\). The fiber as shown in Fig. 4 is subjected to a boundary force \(\underline{t}_N\) on a boundary \(\varGamma ^0_{\!\!N}\). The principle of virtual work (PVW) then reads: for any admissible virtual displacement field \(\underline{w} \in \mathscr {V}_{\text {ad}}\),
where the differential of the Green-Lagrange strain tensor with respect to the displacement field is given by
In this formulation, we want to associate with each material point an active microscopic model based on the Huxley’57 model or its extensions. Typically, we want to incorporate the microscopic model into a 3D visco-hyperelastic constitutive behavior of hyperelastic potential \(\varPsi \) and viscous pseudo-potential \(\varPsi ^v\)—taken here as \(\varPsi ^v(\dot{\underline{\underline{e}}}) = \frac{\eta }{2} \mathrm {tr}(\dot{\underline{\underline{e}}}^2)\) —where \(\eta \) denotes a viscosity modulus—to simplify the presentation. Following [2], which extends the classical Hill-Maxwell scheme [9] to nonlinear behavior, we gather all the constitutive ingredients by defining an adequate rheological scheme—presented in Fig. 5—valid for large deformations. The upper branch represents the sarcomere—including the above active behavior visualized by the collection of myosin heads in the figure—namely, constituents acting in the muscle fibre direction \(\underline{\tau }\). The lower branch represents a 3D passive matrix, associated with the cellular envelope and the extracellular matrix. Each branch contains elastic and viscous constituents, respectively visualized by springs and dashpots, with specific constitutive equations given below.
We consider the following natural rheological rule for the parallel branch
where \(\underline{\underline{e}},\, \underline{\underline{e}}_a,\, \underline{\underline{e}}_p\) denote Green-Lagrange tensors (global, active and passive) and \(\underline{\underline{\varSigma }}\), \(\underline{\underline{\varSigma }}_a\), \(\underline{\underline{\varSigma }}_p\) second Piola-Kirchhoff stress tensors (global, active and passive). However, we will depart from [2] for the series branch.
In fact, the natural view of muscle fibers made of a succession of active and passive segments points to a one-dimensional homogenization type of rheological interpretation. Let us denote by \(e_\text {fib}\) the total (local) extension of a fiber, i.e the ratio of length change over initial length
where we will take for \(\ell _{\text {fib}}\) the length of a half-sarcomere at rest and \(\delta \ell _{\text {fib}}\) the variation thereof. Then, the length change of the half-sarcomere can be decomposed into
where \(\delta \ell _c\) and \(\delta \ell _s\) respectively denote the contributions of the active (rigid filaments) and passive parts in this length change. Note that \(\delta \ell _c\) then represents the relative displacement of the actin and myosin filaments considered above, indeed. We also introduce the corresponding dimensionless extension quantities \(e_c=\delta \ell _c/\ell _{\text {fib}}\) and \(e_s=\delta \ell _c/\ell _{\text {fib}}\), so that \(e_\text {fib} = e_c + e_s\). Of course, the two components carry the same tension, which we denote by \(T_{\text {fib}}\) and define as the force—in the fiber direction—per unit area of transverse cross-section of tissue considered in the reference configuration. Therefore, we can summarize as
Note that the reason why such a simple additive rule holds for \(e_\text {fib}\) in this nonlinear framework is that we are considering extension quantities—scaled in an ad hoc manner —and not Green-Lagrange strains. Moreover, (28) must be complemented by relationships between 3D and 1D quantities. Considering the component of the Green-Lagrange strain tensor in the fiber direction, we directly have
Then, the tension \(T_{\text {fib}}\) corresponds to a contribution in the first Piola-Kirchhoff stress tensor given by
where \(\underline{\underline{F}}=\underline{\underline{1}}+\underline{\underline{\nabla }}\,\underline{y}\) is the classical deformation gradient tensor, as can be easily verified by computing the resulting traction in the fiber direction \(\underline{\underline{T}}_a\cdot \underline{\tau }\). Hence, the associated contribution in the second Piola-Kirchhoff stress tensor reads
since \(\Vert \underline{\underline{F}}\cdot \underline{\tau }\Vert =1+e_\text {fib}\).
Finally, the constitutive equations considered are
where \(\check{T}_c\) represents the aggregation of forces contributed by actin-myosin cross-bridges as described above, i.e. \(\check{T}_c=\rho _{\mathrm {surf}}\tau _c\) with \( \rho _{\mathrm {surf}} \) the number of myosin heads in a layer of thickness \(\ell _{\text {fib}}\) per unit of cross-section area. The series elastic element is here assumed to have a linear constitutive equation of elasticity modulus \(E_s\). Note that a nonlinear hyperelastic behavior could be considered, at the price of having to deal with the dimensionless extension \( e_s \) as an additional internal variable. Nevertheless, in physiological conditions the extension \( e_s \) remains small and a linear behavior is adequate [1]. As regards viscosity, we here incorporate a simple component of viscous modulus \(\nu \) in parallel with the active part in the sarcomere branch, and we recall that viscosity is also present in the parallel branch as provided by the term \(\frac{\partial \varPsi ^v}{\partial \dot{\underline{\underline{e}}}} = \eta \dot{\underline{\underline{e}}}\).
We can now summarize the 3D equations as

where (33c) is based on the Huxley’57 model with sliding velocity \(v_c = \ell _{\text {fib}} \dot{e}_c\). This velocity is independent of the microscopic variable s, which justifies our above study. Note, however, the dependency of \(a(\underline{\mathrm {x}},s,t)\) on \(\underline{\mathrm {x}}\), which means that the microscopic model must be solved everywhere in the domain, i.e. at all numerical quadrature points in numerical simulations.
In order to establish a macroscopic energy balance for the system (33), we take the velocity field \(\dot{\underline{y}}\) as an admissible virtual displacement field in (33a). We thus get
where \(\mathcal {K} = \frac{1}{2}\int _{\varOmega ^0} \rho _0 |\dot{\underline{y}}|^2 \, \mathrm {d}\varOmega \) stands for the kinetic energy and \(\mathcal {P}_{\text {ext}} =\int _{\varGamma ^0_{\!\!N}} \underline{t}_N \cdot \dot{\underline{y}}\, \mathrm {d}\varGamma \) is the power of external forces. Then, we decompose
where the first term is associated with the stored hyperelastic energy, the second term is a macroscopic viscous dissipation, and the last term—denoted \(\mathcal {P}_{\text {fib}}\)—is the power of internal forces in the sarcomere. Then, using the rheological rules we find
where we recognize (1) an elastic energy stored in the series element of the sarcomere, (2) a viscous dissipation term in the sarcomere, and (3) the mechanical work of the actin-myosin bridges. Therefore, combining this energy balance with the free energy balance (23) computed from the Huxley’57 model—or identically from (26) for the extensions—we finally obtain a form of macroscopic Clausius-Duhem relation
where \( \rho _v = \rho _{\mathrm {surf}}/\ell _{\text {fib}}\) is the density of myosin head per unit volume in the reference configuration, and where we recall that \(\mathcal {F}\) is the internal free energy of the bridges introduced in (13), \(\dot{\mathcal {S}}_{\text {prod}}\) is the entropy production term defined in (22) corresponding to energy dissipation associated with chemical transitions, and \(\dot{\mathcal {E}} = \mu _T \bar{J}_2\) as defined in (12b) corresponds to the input flux in energy provided by ATP hydrolysis.
Discretization and thermodynamic principles at discrete level
We now present the proposed discretization scheme for the muscle contraction models. Classical schemes are sufficient for our purposes, and the main originality of this work is to show their compatibility with discrete versions of the thermodynamical principles. Nevertheless, for the sake of completeness, some basic properties of the schemes are quickly re-established before focusing on thermodynamics.
A numerical scheme for Huxley’57 model
To discretize the dynamics (2), we consider a regular grid for the simulation range \([s^-, s^+]\) of discretization length \(\delta s\) and with the convention \(s_0 = s^-\) and \(s_\ell = s^+\). We then choose an upwind implicit scheme, that for the sake of simplicity we only present for a positive sliding velocity \(v_c\), with a natural extension to negative sliding velocities \(v_c\) by inverting the shift in space for the transport term to keep an upwind scheme. We initiate the discretization from an initial condition such that \( a_0^n = a_\ell ^n = 0 \) and with the natural condition \( a_0^n \in [0,1] \). The discretization scheme then reads
with the definition
Note that the exact aggregated detachment rate goes to infinity on the boundary of the interval \( [s^-, s^+] \). Numerically, we use a finite value defined as given in (36c), and we prove in the following section that this choice does not affect the convergence of the scheme.
For the numerical scheme (35), we also need to prescribe adequate boundary conditions. As the analytical solution of (7) vanishes on the boundaries of the interval \( [s^-, s^+] \), we here again can choose: either a Dirichlet condition on one side, and check the consistency on the other side, or choose periodic boundary conditions and again ensure the consistency on the boundary of the interval \( [s^-, s^+] \). From a numerical point of view, it is in fact more convenient for energy estimates to choose periodic boundary conditions for \( a \), i.e. \( a_0^n = a_\ell ^n \). Note that, with this choice, we do not strictly have \( a_0^n = a_\ell ^n = 0 \). This property is only satisfied approximately, or asymptotically when the spatial discretization length goes to zero.
Defining \( \alpha = v_c \delta t/ \delta s\) and \( k_i = k_{+,i} + k_{-,i} \), the scheme can be written in a matrix form on the state vector \( \varvec{a}^n = \left[ \begin{array}{lll} a_1^{n}&\ldots&a_\ell ^{n} \end{array}\right] ^{\mathsf {T}}\)
where

Some fundamentals properties
We first present the basic—but essential—properties of the proposed scheme. This is done using classical strategies for the analysis of transport equations schemes (see for instance [21]).
Uniform positivity and boundedness
One first important property that must be satisfied by the discretization is that the natural bounds for ratio quantities be preserved at the discrete level, namely \(\forall n, \forall i \in [\![1 , \ell ]\!], a^{n}_i \in [0,1] \). To obtain this property, we need \( \mathbb {D} \) to preserve the positivity—i.e. for \( \varvec{a}\in \mathbb {R}^\ell , \mathbb {D} \varvec{a}\ge 0 \Rightarrow \varvec{a}\ge 0 \) (where we use the convention that a vector is positive if all its coefficients \( a_i \) are positive). Let us then take \( \varvec{a}\in \mathbb {R}^\ell \) such that \( \mathbb {D} \varvec{a}\ge 0 \). We have \( \forall i \in [\![1 , \ell ]\!] \)
with the boundary condition \( a_{0} = a_\ell \). Multiplying (37) by \( \alpha ^{l-1} \) for \( i=1 \) and by \( \alpha ^{\ell -i} \prod _{j=1}^{i-1} \big ( 1 +~k_j +~\alpha \big ) \) for \( i \in [\![2 , \ell ]\!] \), and summing, we obtain
Noting that the middle term is a telescoping series, we obtain as expected
Then, recursively from (37), we get \(\forall i \in [\![1 , \ell ]\!],\ a_i \ge 0 \), which shows that the matrix operator \( \mathbb {D} \) preserves the positivity. Knowing that the initial condition and the transition rates are positive, we obtain \(\forall n \ge 0, \forall i \in [\![1 , \ell ]\!], a^{n}_i \ge 0 \). Writing the numerical scheme for the variable \( 1 - a_i^n \) from (35), we similarly obtain that \(\forall n \ge 0, \forall i \in [\![1 , \ell ]\!], a^{n}_i \le 1 \). Therefore, the proposed numerical scheme preserves the adequate positivity and boundedness.
Consistency
Let us here denote \( \overline{a}\) a sufficiently regular solution of (7). Note that this solution satisfies a Dirichlet boundary condition on one side of the simulation interval (i.e. \( \overline{a}(s^-, t) = 0 \)) and that we showed in (10) that \( \lim _{s \rightarrow s^+} \overline{a}(s,t) = 0 \). We denote by \( \overline{\varvec{a}}\) the vector of the values of \( \overline{a}\) at the spatial discretization points at time \( n \, \delta t\), \( \overline{\varvec{a}}^{\, n} = \begin{bmatrix} \overline{a}(\delta s, n \, \delta t)&\ldots&\overline{a}(\ell \, \delta s, n \, \delta t) \end{bmatrix}^\mathsf {T}\) . We define, as usual, the convergence error by
We have \(\mathbb {D} e^{n+1} = e^{n} - \delta t\, \eta ^{n}\), where the consistency error \( \eta ^{n} \) is given by
Evaluating the continuous equation (7) at \(s=s^- + i \delta s\) and \(t=(n+1)\delta t\), we obtain
and this also holds for \(i=\ell \) with the finite numerical value chosen for \(k_{\ell }\), due to the fact that \(\bar{a}(s^{+})=0\) and \(k_{-}(s) \bar{a}(s)\) tends to zero in \(s^{+}\). We directly infer, \(\forall i \in [\![1 , \ell ]\!]\),
The numerical scheme is thus consistent at the first order with the continuous Eq. (7), although different—but compatible—boundary conditions are used. Note that a well-known result by Godunov states that we cannot have more than first-order convergence in time with a discrete scheme that satisfies the positivity and boundedness property [6].
\(L^{2}\) -Stability
We now analyze the \( L^2 \)-stability by justifying the \(\ell ^2\)-stability of the operator \( \mathbb {D}^{-1} \). Multiplying (35) without the source term by \(a^{n+1}_i\), we have
Using the identity \(-ab = \frac{1}{2} \big ( a - b \big )^2 - \frac{1}{2} a^2 - \frac{1}{2} b^2\), we obtain
Summing over \( i \), and using the periodic boundary conditions \( (a^{n+1}_{\ell })^2 = (a^{n+1}_{0})^2 \), we find
Note that this stability property is just a mathematical property.
Convergence
The stability analysis coupled to the consistency analysis gives directly the convergence error. Indeed, we find
so that there exist \(C,T > 0\) such that
First principle
Our objective is more ambitious than numerical convergence, as we want in fine to establish thermodynamic balances at the discrete level. In this respect, let us first consider the energy balance. We recall that the average energy of a myosin head is given at the continuous level by (11). Similarly to (36), we assign a finite value to the energy of the attached state on the boundary of the interval \( [s^-, s^+] \). With the notation
the energy is discretized as
Then, defining the fluxes as
we have
With the definition of the discrete force
where we define \( w_{1,\ell +1} = w_{1,1} \), and using the periodicity of the solution, (38) becomes
We thus obtain the discretized version of the first principle, namely
with
Second principle
Let us now establish a discrete entropy balance. In this respect, we introduce the discrete entropy
and the free energy \(\mathcal {F}^{n} = \mathcal {U}^{n} - T \mathcal {S}^n\), which can be rewritten as
by introducing the discrete chemical potentials
We then rewrite the previous calculation in a manner that closely follows the calculation in the continuous case. We have
Hence,

Developing the expression of the chemical potentials, performing Abel transformations and using the periodic boundary conditions, we obtain
Since \(x \mapsto \ln x\) is a concave function, we have \( \ln a_j - \ln a_i \le \ln '(a_i) \big (a_j - a_i \big )\) so that
Using the fact that the scheme imposes \(\forall n,\ \forall i \in [\![1 , \ell ]\!],\ a_i^n + d_i^n = 1 \), the sums vanish two by two. We finally find
As a point-wise evaluation of the continuous expression (21), we have \(\forall i \in [\![1 , \ell ]\!]\)
Hence, in our case where \( v_c > 0 \), we finally obtain
To relate the decrease of the free energy to the creation of entropy, we first note
Comparing (41) with the formal expression of the second principle (18), we define the discrete entropy creation by
Note that the entropy creation is formally given by
and finally combining (40), (41) and (42) we have the time-discrete counterpart of the free energy balance (23)
Extension to multi-state, multi-site models
The case of multiple states and sites derives from the same principles, hence justifying that we developed precisely the computations for the Huxley’57 model. The developments are, however, not straightforward because the multi-site assumption implies an infinite number of attachment and detachment fluxes, which has to be properly integrated into the discrete thermodynamical balances. Indeed, in the case of a positive sliding velocity \( v_c \), we discretize the system (6) with the following implicit upwind numerical scheme
with the notation \( x^{n}_{q,i+j\ell } = x_q(s^- + (i + j \ell )\delta s, n \delta t) \) for \( V_q \in V_a \), \( x^{n}_{q,i} = x_q(s^- + i \delta s, n \delta t) \) for \( V_q \in V_d \) and \( J_{pq,i+j \ell }^{n} = J_{pq}(s^- + (i + j \ell )\delta s, n \delta t) \). Note that \( x_q \) for \( V_q \in V_d \) has periodic boundary conditions and thus \( x_{q,i+j\ell } = x_{q,i},\ \forall j \in \mathbb {Z}\), and we keep \(j \in \mathbb {Z}\), albeit in practice we will bound the attachment zone, introducing a boundary consistency error.
Mass conservation
Defining the total quantity of matter
which has the periodicity \( m_{0}^{n} = m_{\ell }^{n} \)—and using the scheme (44), we find the classical implicit transport equation
hence, we retrieve, as in the Huxley’57 model, the conservation of matter
First principle
Concerning the energy balance, the internal energy is now defined as
and we find this time, with the notation \( \tilde{W}_{pq,i+j\ell } = \tilde{W}_{pq}(s^- + (i + j \ell )\delta s) \), that
The last term vanishes with the periodic boundary conditions. Performing an Abel transformation on the penultimate term and defining the discrete force as
we obtain the discrete first principle
with
Second principle
We now define the discrete entropy as
and the free energy \(\mathcal {F}^n = \mathcal {U}^n - T \mathcal {S}^n\), that we rewrite as
using discrete chemical potentials
The discrete time derivative of the free energy is
Using the notation \(\tilde{M}_{pq, i+j\ell }^{n} = \mu _{q,i + j \ell }^n - \mu _{p,i + j \ell }^n - \mu _T \delta _{pq=\{25,34\}} \) and the results of the first principle, we get
Performing an Abel transformation on the transport terms, we obtain
Using again that \(\ln x_j - \ln x_i \le \ln '(x_i) \big (x_j - x_i \big )\), the detailed balance and the conservation of matter (45), we finally obtain
As in “Second principle” section this property is equivalent to
Moreover, the entropy production is in fact given by
and, by recombining (46) and (47), we finally get
Discretization of the macroscopic model coupling
We can derive a full discretized version of the macroscopic model presented in Fig. 4 and modeled by the dynamics (33). Here, we will rely—as in [2]—on mid-point rules for the discretization of the PVW, with additional corrections in order to guarantee the energy balance. Therefore, we will typically denote \(\underline{v}^{n+\frac{1}{2}} = \frac{\underline{v}^{n+1}+\underline{v}^n}{2} \) for any variable \(\underline{v}\) and the notation \(n+\frac{1}{2} \sharp \) will allow to indicate when we depart from this classical rule. First, as recommended in [2, 7, 8, 15], we will consider the following non-standard—albeit classical—mid-point quantities
and a passive hyperelastic stress law discretization that includes an energy correction term, namely
Then, we propose the following discretization of (33)

Note here that (50c) and (50d) are defined at each quadrature point \(\underline{\mathrm {x}}_m\), albeit we omit—for the sake of brevity—this explicit dependence in the equations. If the 1D elastic element is chosen nonlinear hyperelastic, the corresponding term in (50c) has to be treated as proposed for the 3D elastic element in (49).
To obtain a complete energy balance, we now proceed as in [2] by considering the mid-point velocity \(\underline{v}^{n+\frac{1}{2}}\) as an admissible displacement field and recalling that \(\mathrm {d}_{\underline{y}}\,\underline{\underline{e}}^{n+\frac{1}{2}\sharp }\cdot \underline{v}^{n+\frac{1}{2}} = \frac{\underline{\underline{e}}^{n+1}-\underline{\underline{e}}^n}{\delta t}\). The balance associated with the hyperelastic contribution is handled by our choice in (49), and the viscous part directly gives a negative contribution, so that we have
The function \(x \mapsto (1+2x)^{\frac{1}{2}}\) is concave of derivative \((1+2x)^{-\frac{1}{2}}\), hence
Therefore we have, recalling that we have defined \(e_s\) such that \(e_{\text {fib}} = e_s + e_c\),
We now incorporate the Huxley’57 discrete-time free energy balance (43)—and we could proceed identically with the other models using (48)—to get
which is the discrete-time counterpart of the Clausius-Duhem relation (34), with here an inequality only due to numerical dissipation and consistent in \(\delta t\). Note finally that in the case of models capturing the power stroke dynamics such as Piazzesi–Lombardi’95, time sub-iterations may be required. In this case it can be shown that the energy balance is preserved provided the active tension \(\check{T}_c^{n+1}\) is redefined by weighing the intermediate states over all sub-iterations.
Numerical results and discussion
In this section, our goal is to illustrate the analysis of the discrete system presented in the previous section for the Huxley’57 model and the Piazzesi–Lombardi’95 model, which we chose as a representative of the multi-site, multi-state models. These illustrations serve several purposes. We first want to demonstrate that the thermodynamics identities established at the discrete level are satisfied in the numerical simulations. Then, we want to show that the ability to compute the thermodynamical balances numerically allows to gain additional insight into the physiology of muscle contraction. Additionally, for the Piazzesi–Lombardi’95 model, we compare our simulation results with that obtained in the original paper as a further validation of our approach.
Huxley’57 model
The choice of model parameters must satisfy the conditions (3) and the assumption that \( w_1 a \) and \( k_{-} a \) tend to zero when \( s \) tends to \( s^- \) and \( s^+ \). We choose the energy levels and transition rates as follows
We choose here to prescribe \( k_1 \) and \( k_2 \) in addition to the energies \( w_1 \) and \( w_0 \). The reverse rates \( k_{-1} \) and \( k_{-2} \) are then derived from the detailed balance (1). The energy levels parametrization is shown in Fig. 6. The transition rates are depicted in Fig. 7. The models parameters used in the following simulations are presented in Table 1.
The asymptotic properties of the chosen transition rates and of the associated solutions can be easily obtained using the analytical solution (9) and the theorem of dominated convergence.
We consider two simulation cases. First, we simulate the tension rise in isometric conditions (\(v_c = 0\)). Then, we compute the muscle response in contraction at constant shortening velocity (\(v_c < 0 \)) starting from the isometric steady-state solution.
Validation of the thermodynamical identities at discrete level
We first want to verify that the discrete versions of the first principle (39) and the second principles (43) are satisfied numerically. To do so, we compute respectively the expressions
and
The results for both simulation cases is presented in Fig. 8. We notice that the first expression is ten orders of magnitude smaller that the individual terms that compose it (see Figs. 9 and 10), showing that the first principle is satisfied at discrete level. The second expression is always negative showing the validity of the discrete second principle.
Tension rise
In our first illustration of the results obtained for the Huxley’57 model, we simulate the tension rise in isometric conditions (\(v_c = 0\)). We initialize all heads in the detached state and let the myosin heads evolve. Along the usual tension evolution, our scheme allows us to compute the thermodynamic fluxes associated with muscle contraction—see Fig. 9. In the steady-state regime, the energy input remains positive and heat is dissipated. The force is sustained through the continuous cycling of the myosin heads in interaction with the actin filament. This process is fueled by the energy brought by ATP. We see here the active nature of muscle contraction. Force is produced when the muscle is supplied with energy. Naturally, as the velocity is zero no work is produced.
Constant velocity contraction
We now show a second illustrative example with a contraction at constant shortening velocity (\(v_c < 0 \)) starting from the isometric steady-state solution. The simulation results are presented in Fig. 10. After a transient phase, the system reaches a permanent regime in which the classical force-velocity curve is measured [9] (note that in the original experimental protocol force and not length is controlled). In this regime, we observe the energy mechano-transduction performed by the molecular motors: the energy input brought by ATP is for one part converted into work produced by the system (\( \mathcal {W} < 0 \)), and for the other part dissipated by entropy production.
Piazzesi–Lombardi’95 model
The Piazzesi–Lombardi’95 model reproduces the physiology of muscle contraction more precisely. In particular, it is able to capture the power stroke fast dynamics observed in length step experiments.
We simulate such an experiment starting from the isometric steady state with a length step of \(8\hbox { nm}\). As in the experimental conditions, the length step is made by a ramp of duration \(100 \upmu \hbox {s}\). Note that, here, the compliance of the myosin and actin filaments is neglected as in the original paper. We choose the energy levels as defined in [20]. We use modified transition rates to ensure that detachment rates diverge at infinity. The energy brought by ATP is set to \(50\hbox { zJ}\) following the model assumption that an ATP molecule can be used for the detachment of several myosin heads.
The results are presented in Fig. 11. They match the results presented in the original paper [20], which shows the consistency of our approach with the original model, hence completed with thermodynamic balances.
Concluding remarks
Considering a large class of muscle contraction models based on actin-myosin interaction—i.e. the Huxley’57 model and various extensions thereof, including the Piazzesi–Lombardi’95 model—we have presented a mathematical setting in which solution properties can be established, including fundamental thermodynamic balances. Moreover, we have proposed a complete discretization strategy for which we were also able to obtain discrete versions of the thermodynamic balances and other properties. In addition, we have also shown how these models can be coupled with a macroscopic continuum mechanics formulation in such a way that these balances carry over to the macroscopic level, including for the discrete versions of the models. As muscle energetics are of major relevance in physiology, this is an important achievement, both from a fundamental and numerical point of view. This paves the way, indeed, for detailed numerical studies of energy exchanges in various applications, such as with a complete realistic heart model.
Abbreviations
- ATP:
-
Adenosine triphosphate
- PL95:
-
Piazzesi–Lombardi’95
- PVW:
-
Principle of virtual work
References
Caruel M, Moireau P, Chapelle D. Stochastic modeling of chemical–mechanical coupling in striated muscles. Biomech Model Mechanobiol. 2018;1:1–25.
Chapelle D, Le Tallec P, Moireau P, Sorine M. Energy-preserving muscle tissue model: formulation and compatible discretizations. J Multiscale Comput Eng. 2012;10:189–211.
Chipot M, Hastings S, Kinderlehrer D. Transport in a molecular motor system. ESAIM: M2AN. 2004;38(6):1011–34.
Eisenberg E, Hill TL. A cross-bridge model of muscle contraction. Progr Biophys Mol Biol. 1978;33(1):55–82.
Eisenberg E, Hill TL, Chen Y. Cross-bridge model of muscle contraction. Quantitative analysis. Biophys J. 1980;29(2):195–227.
Godunov SK. A difference method for numerical calculation of discontinuous solutions of the equations of hydrodynamics. Matematicheskii Sbornik. 1959;89(3):271–306.
Gonzales O. Exact energy and momentum conserving algorithm for general models in nonlinear elasticity. Comput Meth Appl Mech Eng. 2000;190(13–14):1763–83.
Hauret P, Le Tallec P. Energy controlling time integration methods for nonlinear elastodynamics and low velocity impact. Comput Meth Appl Mech Eng. 2006;195:4890–916.
Hill AV. The heat of shortening and the dynamic constants of muscle. In: Proceedings of the Royal Society of London, 1938.
Hill TL. Free energy transduction in biology. Cambridge: Academic Press; 1977.
Hill TL. Free energy transduction and biochemical cycle kinetics. New York: Dover; 2004.
Huxley AF. Muscle structure and theories of contraction. Progr Biophys Chem. 1957;7:255–318.
Huxley AF, Simmons RM. Proposed mechanism of force generation in striated muscle. Nature. 1971;233:533.
Julicher F, Ajdari A, Prost J. Modeling molecular motors. Rev Mod Phys. 1997;69(4):1269–81.
Le Tallec P, Hauret P. Energy conservation in fluid structure interactions. In: Kuznetsov Y, Neittanmaki P, Pironneau O, eds. Numerical methods for scientific computing—variational problems and applications, 2003.
Lymn RW, Taylor EW. Mechanism of adenosine triphosphate hydrolysis by actomyosin. Biochemistry. 1971;10(25):4617–24.
Marcucci L, Washio T, Yanagida T. Including thermal fluctuations in actomyosin stable states increases the predicted force per motor and macroscopic efficiency in muscle modelling. PLoS Comput Biol. 2016;12(9):e1005083.
Mirrahimi S, Souganidis PE. A homogenization approach for the motion of motor proteins. Nonlinear Differ Equ Appl. 2012;20(1):129–47.
Peskin CS. Mathematical aspects of heart physiology. Courant Institute of Mathematical Sciences—NYU, 1975.
Piazzesi G, Lombardi V. A cross-bridge model that is able to explain mechanical and energetic properties of shortening muscle. Biophys J. 1995;68:1966–79.
Richtmyer RD, Morton KW. Difference methods for initial-value problems. Interscience tracts in pure and applied mathematics. Geneva: Interscience Publishers; 1967.
Sheshka R, Truskinovsky L. Power-stroke-driven actomyosin contractility. Phys Rev. 2014;89:1.
Author's contributions
FK was the main investigator in this work; he also performed the numerical implementation and simulations, and initiated the writing. DC was in a position of secondary supervision, and is responsible for some detailed technical aspects and writing. PM was in a position of primary supervision, and proposed initial ideas on topic and discretization strategy; he’s also responsible for some detailed technical aspects and writing. In addition, DC and PM devised the proposed integration of muscle models in continuum mechanics and the associated discretization ingredients. All authors read and approved the final manuscript.
Acknowledgements
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
Not applicable.
Funding
Not applicable.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Kimmig, F., Chapelle, D. & Moireau, P. Thermodynamic properties of muscle contraction models and associated discrete-time principles. Adv. Model. and Simul. in Eng. Sci. 6, 6 (2019). https://doi.org/10.1186/s40323-019-0128-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40323-019-0128-9
Keywords
- Muscle contraction
- Sliding filaments
- Thermodynamically consistent time-discretization
- Clausius–Duhem inequality