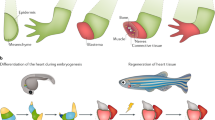
Abstract
Activation of regeneration upon tissue damages requires the activation of many developmental genes responsible for cell proliferation, migration, differentiation, and tissue patterning. Ample evidence revealed that the regulation of chromatin organization functions as a crucial mechanism for establishing and maintaining cellular identity through precise control of gene transcription. The alteration of chromatin organization can lead to changes in chromatin accessibility and/or enhancer-promoter interactions. Like embryogenesis, each stage of tissue regeneration is accompanied by dynamic changes of chromatin organization in regeneration-responsive cells. In the past decade, many studies have been conducted to investigate the contribution of chromatin organization during regeneration in various tissues, organs, and organisms. A collection of chromatin regulators were demonstrated to play critical roles in regeneration. In this review, we will summarize the progress in the understanding of chromatin organization during regeneration in different research organisms and discuss potential common mechanisms responsible for the activation of regeneration response program.
Similar content being viewed by others
Background
Regeneration is a fascinating phenomenon in biology which is depicted as the restoration of damaged body parts to their original state in response to injury or diseases. Unlike mammals including humans that usually have very limited regenerative capacities, lower vertebrates such as fishes and salamanders are good at regenerating various appendages and organs. Ample evidence from regeneration-competent animals revealed that tissues or organs with high regenerative capacities tend to retain high proliferative potential (Chen et al. 2020a; Iismaa et al. 2018; Jopling et al. 2010; Ryoo and Bergmann 2012). Like organ development, cell proliferation and differentiation are essential processes for successful regeneration of damaged organs (Tanaka and Reddien 2011). Upon tissue damage, the cell source for regeneration can vary from organ to organ. For example, progenitor cells, reserve stem cells, or terminally differentiated cells that can undergo de-differentiation or trans-differentiation are common cell sources involved in regeneration (Merrell and Stanger 2016). In classic epimorphic regeneration (e.g., limb regeneration, fin regeneration, and planarian head regeneration), a series of key steps including inflammation response, re-epithelialization (wound healing), blastema formation, regenerative outgrowth, and re-patterning occur to restore the original tissue function (Londono et al. 2018; Pfefferli and Jazwinska 2015; Reddien 2018; Yokoyama 2008). The progression of each phase of regeneration requires precise regulation of gene expression.
Protein-coding and non-coding genomic DNA of each cell is well-organized inside microscopic nuclei as chromatins. The unique structure of the chromatin efficiently packages the genome without compromising DNA accessibility for proper gene expression and replication of the genetic material during cell division. The three-dimensional (3D) genome organization can be defined and characterized at different levels: the chromosomal (distinct distribution of chromosomes in the nucleus) and sub-chromosomal levels (the compartmentalization of chromatin) (Dekker and Mirny 2016; Van Driel et al. 2003; Woodcock and Ghosh 2010). A fundamental unit of the 3D-chromatin organization is the topologically associating domains (TADs) (McArthur and Capra 2021). TADs and their corresponding TAD boundaries within a given cell participate in gene regulation by facilitating or restraining interactions between regulatory sequences and targets. A lot of studies have demonstrated that the organization of accessible chromatin in a genome encodes a network of potential physical interactions that involve promoters, enhancers, insulators, and chromatin-binding factors (Klemm et al. 2019). Precise regulation of chromatin organization is essential for establishing and maintaining cellular identity. For instance, the Polycomb repressive complex PRC1 functions as a master regulator of genome architecture in mouse embryonic stem cells by constraining developmental transcription factor genes (e.g., Hox genes) and their enhancers in three-dimensional interaction networks (Schoenfelder et al. 2015). It was proposed that the selective activation of genes from such a network controls cell fate specification during early embryonic development. In contrast, abnormal regulation of chromatin organization can cause developmental defects and pathogenesis (Anania and Lupianez 2020; Schoenfelder and Fraser 2019; Ushiki et al. 2021; Zheng and Xie 2019).
The regulation of organ regeneration and development share common features in many aspects (Efroni et al. 2016; Goldman and Poss 2020; Malloch et al. 2009). Dynamic changes of chromatin accessibility, epigenetic modification, and the activation of gene promoters and cis-regulatory elements that are known to be critical during development also play essential roles in the activation and progression of regeneration. In the past decades, fruitful new knowledge has been accumulated in the understanding of regeneration due to fast advancement in genetic tools and technologies of various fields, which allows for an easier, faster, and deeper examination of fundamental questions. In this review, we will focus on new findings regarding the regulation of chromatin organization during regeneration in different organisms and discuss potential common mechanisms underpinning the activation of the regeneration program.
Clues from development and diseases
The initial features of genome organization were first observed by Emil Heitz through cytological staining and phase-contrasting to identify heterochromatin and euchromatin in 1928s (Heitz 1928). The development of chromosome-conformation-capture technologies (3C, 4C, 5C, Hi-C, and Micro-C) and their variants have made it possible to examine finer and more comprehensive genomic organizations from territories to compartments, TADs, and even interactive loops (Dekker et al. 2002; Han et al. 2018) (Fig. 1). Previous studies indicated the active transcriptional regions in each chromosomal territory tend to be positioned at the periphery of nuclear speckles, while inactive regions are close to the nuclear envelope (Geyer et al. 2011). Accordingly, the intrachromosomal self-interacting regions can be divided into two types of compartments based on the biochemical marks or activities: A compartment (active marks) and B compartment (inactive marks) (Hildebrand and Dekker 2020). On a finer scale, the active and inactive genomic regions are insulated to form highly interactive TADs with the cooperation of the insulator-binding protein CTCF, cohesin, and others (Beagan and Phillips-Cremins 2020).
The discovery of main features of chromatin organization
Diverse forms of chromatin organization have been identified ranging from the 100 bp scale to more than 100 Mb scale through a combination of different technologies over one century. The association between histone modification and gene transcription was identified through the incorporation of labeled chemical groups into histone structures in 1964 (Allfrey et al. 1964). DNA looping was first discovered using the helical-twist assay (Dunn et al. 1984). With the development of technologies, large chromatin interaction domains called topologically associating domains (TADs) and chromosome compartments indicating the spatial segregation of open and closed chromatin were identified with Hi-C (Dixon et al. 2012; Lieberman-Aiden et al. 2009). Thomas Cremer et al. carried out experiments using the laser to confirm the existence of chromosome territories which help to distinguish one chromosome from its neighbors (Cremer et al. 1982). In 1928, Emil Heitz improved cytological staining to define euchromatin and denser heterochromatin (Heitz 1928)
Integrated with epigenetic and transcriptomic analyses, the chromatin organization from base pairs to territories has been recognized as an increasingly fundamental and sophisticated aspect of embryogenesis, gametogenesis, lineage commitment, and cell differentiation (Bhattacharya et al. 2009; Dixon et al. 2015; Phillips-Cremins et al. 2013; Sati and Cavalli 2017; Zheng and Xie 2019). The plasticity of chromatin organization is a critical mechanism for the transcriptional regulation of gene expression (Vos 2021). A new study in Drosophila revealed that dedicated tethering elements in the genome are critical for fast transcriptional activation by facilitating appropriate enhancer-promoter interactions, while insulators avert unveracious interactions and regulatory interference between neighboring TADs (Batut et al. 2022). The enhancer-promoter interactions are commonly detected during spatiotemporal activation of gene expression. Taking the sonic hedgehog (shh) limb-bud-specific enhancer MFCS1 as an example, long-range interactions between the shh promoter and the MFCS1 enhancer located 1 Mb away were detected in both the anterior and posterior limb buds using 3D-FISH and 3C assays (Amano et al. 2009). The dynamic chromatin conformation of the shh locus drives the pulses of shh activation. Deletion of MFCS1 eliminates the long-range enhancer-promoter interaction, leading to a loss of limb-specific shh expression and truncation of the mouse limb (Amano et al. 2009; Sagai et al. 2005). Further, Hoxd genes have been shown to regulate the induction of shh expression in the mouse limb bud (Kmita et al. 2005; Zakany et al. 2004). The precise regulation of Hoxd gene transcription during early mouse limb development was controlled by the opposite and successive actions of two gene deserts flanking this Hoxd cluster on either side (Andrey et al. 2013). In the early phase, the telomeric domain regulates transcription in the proximal limb until a functional and conformational switch occurs toward the opposite topological domain to take over the regulation in the developing distal limb structures (Andrey et al. 2013).
Similarly, genetic mutations that cause alteration of chromatin organization have been found to contribute to the occurrence and progression of various diseases. Previous studies reported that disease-associated enhancer deletion, relocation, and duplication can lead to aberrant rewiring of gene regulatory circuitry between enhancers and their target genes, and consequently lead to pathogenesis (Krijger and de Laat 2016; Nasser et al. 2021). One such example is deletion-, inversion-, or duplication-induced changes in the structure of the TAD-spanning WNT6/IHH/EPHA4/PAX3 locus give rise to pathogenic rewiring of gene-enhancer interactions and eventually limb malformations in humans (Lupianez et al. 2015). Additionally, oncogenes can be activated by genetic mutations that disrupt chromosome neighborhoods in cancer cells (Hnisz et al. 2016). Together, all these important discoveries on chromatin organization suggest that mapping the spatial TADs, their loop interactions, and TAD boundaries can be extremely informative in deciphering the genetic basis of fundamental biological processes. It is broadly recognized that many regulatory mechanisms by which gene transcription is controlled are shared among development, diseases, and regeneration (Bhatt et al. 2014; Wang et al. 2017). Therefore, adopting concepts and methodologies learned from development and diseases should expedite the understanding of how animal regeneration is achieved.
Genome evolution and the regenerative capacities
The capacity of animal regeneration is unevenly distributed in different animal phyla (Alvarado and Tsonis 2006; Bely 2010; Poss 2010). Animal diversities in nature that involve phenotypic traits, behaviors, and physiology are coded in the genome of each species. Genome evolution can occur at different levels including point mutations, insertion/deletion, genomic recombination, gene duplication, chromosome duplication, and whole genome duplication, and is the driving force for the formation of new features in animals (Dehal and Boore 2005; Henderson and Bomblies 2021; Lin et al. 2019; Lynch and Conery 2000; Tenaillon et al. 2016; Van de Peer et al. 2009). It has been repeatedly observed that evolutionary changes in animal genomes are frequently accompanied by gain or loss of genome size and gene number, expansion or reduction of gene families, and alteration of regulatory complexity (Lynch and Conery 2000; Olson 1999; Petrov 2001; Wittkopp and Kalay 2012). The first analysis of the relationship between genomic features and tissue regeneration was carried out in 1987 by Stanley K. Sessions and Allan Larson (Sessions and Larson 1987). They observed an inverse evolutionary correlation between the genome size and the rate of limb regeneration in salamanders of the family Plethodontidae. Within this largest salamander family, the genome sizes of the group members can range appropriately nine-fold. Interestingly, species with small and large genome sizes in the same phylogenetic group display little differences in the number and shape of the karyotypes (Sessions and Wake 2021). One hypothesis on the evolution of limb regeneration in salamanders is that these animals evolved the ability to regenerate through genome expansion which was mainly driven by the enlargement and dispersion of transposable elements, particularly the LTR retrotransposons (Sessions and Wake 2021). The assembly of the giant axolotl genome (32 Gb, ten times the size of the human genome) was completed recently (Nowoshilow et al. 2018), which provides abundant resources for analyzing the potential genetic regulation of vertebrate regeneration. Massive repetitive sequences (65.6%) were found in the genome and contributed to a dramatic size expansion of introns and intergenic regions compared with those in humans, mice, and frogs (Nowoshilow et al. 2018). Notably, multiple lines of evidence suggest that certain species-restricted coding (e.g., the Ly6 family member Prod1) and non-coding sequences that have been lost or undergone rapid diversification in amniotes contribute to axolotl limb regeneration (Garza-Garcia et al. 2010; Nowoshilow et al. 2018; Silva et al. 2002). In addition to axolotl, genome assembly of other phylogenetically representative species with remarkable regenerative capacities such as the freshwater cnidarian hydra (Chapman et al. 2010), planarians (Grohme et al. 2018), frogs (Kakebeen et al. 2020), zebrafish (Woods et al. 2000), and African killifish (Reichwald et al. 2015; Valenzano et al. 2015) have rendered versatile genomic and transcriptomic analysis for unveiling the mystery of regeneration. Consistently, the presence of a large proportion of noncoding DNA was observed in these regeneration-competent organisms including transposable elements, sequence for the transcription of non-coding RNAs, and other repetitive sequence (Azpiazu and Morata 2022; Harris et al. 2020; Kang et al. 2016; Sen and Ghatak 2015; Shao et al. 2020; Wang et al. 2020c). These observations highlighted that a high percentage of non-coding DNA may be an important source for the generation of new gene regulatory elements that contribute to gene activation upon injury.
A diploid genome is popular in most animals. Polyploidy, a special condition of possessing more than two complete sets of chromosomes, has been observed to participate during injury response in a variety of tissues like hearts, livers, skeletal, and bone marrows (Fig. 2) (Dornen et al. 2020; Matsumoto et al. 2020; Ovrebo and Edgar 2018). Polyploidization (developmentally programmed or stress-induced) can be achieved through either cell–cell fusion or endoreplication (Ovrebo and Edgar 2018). Such a process can bring certain benefits to cells like enlarged cell size and biomass, which confers enhanced cell longevity due to better tolerance to stress (Anatskaya and Vinogradov 2022). Previous studies demonstrated the transition of diploid cardiomyocytes to polyploid cardiomyocytes attenuates the capacity of cardiac regeneration in neonatal mice and zebrafish due to reduced proliferative potential (Alkass et al. 2015; Gonzalez-Rosa et al. 2018; Kadow and Martin 2018; Kirillova et al. 2021; Yahalom-Ronen et al. 2015). Therefore, it was proposed that polyploidization of cardiomyocytes may underlie the failure of heart regeneration in adult mice. However, polyploid hepatocytes are still capable of cell division and do not weaken the regenerative capacity in mouse liver (Miyaoka et al. 2012). In fruit flies, polyploid cells appear in response to injury in diverse tissues such as intestines and abdominal cuticles, and contribute to the restoration of tissue mass, the maintenance of organ size, the protection against oncogenic insults and genomic stress, and the formation of new diploid cells in regeneration (Bailey et al. 2021; Lucchetta and Ohlstein 2017). The distinct observations in different organs or species suggest further investigations are required for elucidating the contribution of polyploid cells upon tissue damages. Currently, few studies have been reported to characterize the chromatin organization between polyploid and diploid cells during regeneration. However, it has been implicated in plants that polyploidization dramatically enhances the complexity of chromatin structures including changes of A/B compartments and the reorganization of TADs (Garcia-Lozano et al. 2021; Wang et al. 2018).
Genome organization and the regulation of tissue regeneration
Summary of current understanding on genome organization and the activation of regeneration in different organisms that have been investigated. Genomic elements, genome features, chromatin modifications, and chromatin regulators all contribute to the regulation of animal regeneration
Further, a recent study using comparative genomics investigated the antler regeneration in ruminants by genome sequencing of pecoran lineages that convergently lack headgear and the collection of hundreds of transcriptomes from bovids and cervids (Lin et al. 2019). This comprehensive analysis indicated that bovid horns and cervid antlers share similar signatures of gene expression and a common neural crest cell origin during development. The rapid regeneration of antlers engages the deployment of oncogenic pathways and a positive selection of certain tumor suppressor genes in deer. Since the first day that multicellular organisms are present on earth, the genome of each extant species is the only information that has been passed from generation to generation during the hundreds of million years of evolution. Therefore, systematic exploration of genome evolution in animals with different regenerative capacities should provide insights into the understanding of the genomic constraints on regeneration.
Regeneration and the remodeling of chromatin organization
Regeneration requires the activation of a regeneration response program to initiate cell proliferation, migration, differentiation, and other biological processes to restore the lost or damaged organs. All these key processes are accompanied by dynamic remodeling of chromatin organization and transcriptomes of cell populations involved in regeneration (Goldman and Poss 2020; van Steensel and Furlong 2019; Vitulo et al. 2017; Wang et al. 2020a). In addition to the classic regulation of transcription and translation, the epigenetic code has been elucidated to be a key mechanism by which the reconfiguration of genome is achieved to allow the turn-on and turnoff of genes essential for cells to acquire new fates or states (Macchi and Sadler 2020; Moris et al. 2016). Plentiful studies indicated that such epigenetic code involves a complex combination of histone variants, histone modifications, DNA modification, and other factors (Fig. 2) (Rothbart and Strahl 2014; Turner 2007). For instance, histone modifications on certain lysine residues that are conserved from yeast to humans are associated with specific regions of the genome, representing different transcriptional states (Jones 2015; Truong and Boeke 2017). In most cases, active promoters are marked by H3K4me3 and H3K9ac, while the gene body tends to have a higher level of H3K36me3 and H3K79me2/3 for an actively transcribing gene (Gates et al. 2017; Sharakhov and Sharakhova 2015; Slotkin and Martienssen 2007). The cis-regulatory elements or enhancers can be defined by H3K27ac (active enhancers) and H3K4me1 (active and primed enhancers) (Nakato and Sakata 2021). Compared with active genes, repressed genes have much higher levels of H3K9me3, H3K27me3, and H4K20me3 (Becker et al. 2016; Peters et al. 2003; Schotta et al. 2004). Whether these histone modifications are functioning as causal effectors is still in debate, such histone marks have been extensively used in the characterization of chromatin accessibility and transcriptional regulation of gene expression under distinct biological contexts.
By taking advantage of these epigenetic markers and tools, a recent breakthrough in understanding the genetic basis of organ regeneration is the discovery of regeneration-responsive enhancers (RREs), also called tissue regeneration enhancer elements or other similar terms (RRE will be used hereafter) (Guenther et al. 2015; Harris et al. 2020; Kang et al. 2016; Sun et al. 2022; Suzuki et al. 2019; Vizcaya-Molina et al. 2018; Wang et al. 2020c; Yang and Kang 2019). Enhancers are DNA-regulatory elements that contain transcription factor binding sites sufficient to activate or boost gene expression by interacting with the gene promoters. The interaction of enhancer-promoter can be achieved by local contacts of DNA-binding transcription factors or by loop formation that mediates the long-range contacts (Higgs 2020). Although it has been suggested that regeneration and development share a similar gene regulatory network (Birnbaum and Sanchez Alvarado 2008; Efroni et al. 2016; Suzuki and Ochi 2020), the mechanisms by which genes are activated can be different. The initial identification of RREs, such as the lepb-linked enhancer (LEN) responsible for the regeneration-dependent expression of lepb in zebrafish (Kang et al. 2016), the WNT gene cluster BRV-B enhancer that directs damaged-induced wingless expression in the fruit fly Drosophila melanogaster (Harris et al. 2016), a Bmp5 enhancer that activates the endogenous bmp5 expression following a bone fracture or soft tissue injury in adult mice (Guenther et al. 2015), and the K-IEN enhancer that controls the regeneration-induced transcription of inhba in African killifish (Wang et al. 2020c), supports the argument that regeneration and development could use distinct regulatory elements to control gene activation. As expected, the regulatory activities of these RREs are turned off or return to a basal level upon the completion of regeneration. Besides, Disruption of certain essential RRE, such as the K-IEN, hampers the progression of regeneration (Wang et al. 2020c). Interestingly, a great deal of redundancy seems to be present for RREs because the deletion of multiple RREs identified from different organs or species did not completely block regeneration (Kang et al. 2016; Thompson et al. 2020). This is consistent with what has been observed for many typical enhancers. The regulatory redundancy may function as a strategy to ensure the robustness of regeneration after tissue injury.
The presence of regeneration-dependent regulatory elements in the genome supports that regeneration requires the alteration of chromatin organization to facilitate accessibility for the transcription of regeneration-responsive genes upon injury (Fig. 2). Systematic identification of RREs across different organs and species has revealed a common regulatory mechanism for injury-induced gene expression (Table 1) (Guenther et al. 2015; Harris et al. 2020; Kang et al. 2016; Murad et al. 2021; Sun et al. 2022; Suzuki et al. 2019; Vizcaya-Molina et al. 2018; Wang et al. 2020c; Yang and Kang 2019). Particularly, a side-by-side comparison between zebrafish and African killifish early fin regeneration not only highlighted a conserved regeneration response program (RRP), but also revealed the activation of many previously overlooked species-specific RREs (Wang et al. 2020c). Comparative single-cell RNA-seq analysis confirmed that blastema cells are the major cell population that employs the RRP during regeneration. This teleost defined RRP only contains 49 genes and is triggered by RREs. The widespread activation of species-specific RREs upon injury is quite surprising because zebrafish and African killifish both belong to the teleost fish, share highly comparable cell types in the caudal fin, and demand a similar amount of time for the completion of regeneration after injury. One vivid difference between the two species with ~ 230 million years of evolutionary distance is their life history. Zebrafish is native to freshwater habitats in Southern Asia and inhabit moderately flowing to stagnant water with shallow depth. In contrast, African killifish is found in ephemeral ponds in semi-arid areas subjected to seasonal desiccation and has adapted to a routine drying of living environment by entering diapause as developmentally arrested and desiccation-resistant embryos that remain dormant in the mud (Hu and Brunet 2018). The adaptation of African killifish genome to such a harsh environment has shaped its unique developmental program and may also directly or indirectly contribute to the evolution of the regeneration program (Valenzano et al. 2015; Wang et al. 2020c). As a result, the African killifish Nothobranchius furzeri has been implicated as a simpler genetic system for regeneration studies due to the reduced complexity of injury response (Wang et al. 2020c).
Comparative studies of RREs using transgenic reporter assays showed that changes in RREs (e.g., enhancer repurposing and epigenetic silencing) with essential functions during regeneration are an important source for the evolution of regenerative capacities in vertebrates (Harris et al. 2016; Wang et al. 2020c). It was implied that the limited regenerative capacity in Xenopus adult limbs is strongly correlated with the DNA methylation status of a limb-specific shh enhancer region during limb regeneration (Yakushiji et al. 2007). This enhancer region is highly methylated in regeneration-incompetent froglets, while is hypomethylated in regeneration-competent tadpoles. Similarly, region-specific epigenetic silencing of a RRE associated with WNT genes limits the regenerative capacity of mature Drosophila imaginal discs (Harris et al. 2016). In sum, the activation of species-specific RRE and epigenetic modification of RREs at different development stages argues that certain regeneration-responsive loci in the genome can be subjected to heritable changes in chromatin organization.
Chromatin regulators and regeneration
The dynamic and strictly controlled regulation of chromatin organization is essential for spatiotemporal and appropriately coordinated gene expression in tissues. Currently, the most characterized chromatin regulators that mediate alterations in the chromatin configuration are DNA modifiers (e.g., methylation and demethylation), histone-modifiers (e.g., methylation, acetylation, ubiquitination, and phosphorylation), ATP-dependent chromatin remodeling complexes (CRCs), and chromatin organizers (e.g., CTCF and cohesin) (Chen and Dent 2014; Clapier et al. 2017; Valencia and Kadoch 2019; Zuin et al. 2014). DNA methylation is a heritable epigenetic mark that cells used to turn off gene expression and this process is directed by DNA methyltransferases (DNMTs). There are five DNMTs encoded in the human genome including DNMT1, DNMT2, DNMT3A, DNMT3B, and DNMT3L. Among those, DNMT1, DNMT3A, and DNMT3B are canonical DNMT enzymes responsible for the addition of methylation marks to genomic DNA (Bestor 2000). In contrast, DNMT2 and DNMT3L lack DNA catalytic activity and are considered non-canonical DNMT members. Genome-wide changes in the pattern of DNA methylation have been observed in different tissues and organs upon injury (Arechederra et al. 2020; Garriga et al. 2018; Gornikiewicz et al. 2016; Lee et al. 2020; Planques et al. 2021; Puttagunta et al. 2014; Yakushiji et al. 2007). Importantly, DNMT1 and the Ubiquitin Like With PHD And Ring Finger Domains 1 (UHRF1) required for localization and stability of DNMTs are key players in the maintenance of methylation during cell proliferation (Bronner et al. 2019). It is not unexpected that these proteins are involved in liver, axon, and muscle regeneration (Lee et al. 2020). Nevertheless, a study in zebrafish revealed that the patterns of lineage-specific DNA methylation are stably maintained during fin regeneration and RREs are preset as hypomethylated before tissue damage (Lee et al. 2020). This result suggested that regeneration responsive genes under the control of RREs are likely activated independent of the DNA demethylation during fin regeneration (Lee et al. 2020; Wang et al. 2020c).
Histone modifiers, such as histone acetyltransferases (HATs) and deacetylases (HDACs) are widely implicated in the regulation of regeneration (Friedrich et al. 2019; Gordon et al. 2015; Huynh et al. 2017). For example, the HAT p300 could acetylate histone H3, pro-regenerative transcription factor p53, and CCAAT-enhancer binding proteins to activate a silent gene regulatory program that is sufficient to promote rat intrinsic axonal regeneration (Gaub et al. 2011). The inhibition of HDACs appears to be a powerful strategy for promoting regeneration in different systems (Ahmad Ganai et al. 2016; Flici and Frank 2018; Huynh et al. 2017). Remarkably, Müller glia specific overexpression of a pro-neural transcription factor Ascl1, in combination with a histone deacetylase inhibitor, enables adult mice to regenerate neurons from Müller glia upon retinal injury (Jorstad et al. 2017). In addition, the motif of the histone demethylase ARID3A was commonly found in RREs (also named regeneration signal-response enhancers) identified in regenerating Xenopus nephric tubules (Suzuki et al. 2019). ARID3A was recruited to reduce the repressive H3K9me3 levels on RREs to promote cell cycle progression and the outgrowth of nephric tubules (Suzuki et al. 2019).
ATP-dependent chromatin remodelers control chromatin architecture by directly mobilizing nucleosomes to enhance local chromatin accessibility (Piatti et al. 2011). Four important families of chromatin remodelers that are conserved from yeast to humans were identified including SWI/SNF, imitation switch (ISWI), nucleosome remodeling and deacetylase (NuRD), and inositol requiring 80 (INO80) complexes (Clapier and Cairns 2009; Tyagi et al. 2016). A recent in vivo CRISPR screening identifies the ISWI component BAZ2 as a druggable suppressor of mammalian liver regeneration (Jia et al. 2022). Inhibition of BAZ2 accelerates liver regeneration through increased ribosomal components and protein synthesis, indicating that targeting chromatin remodelers can be permissive to promote cell growth (Jia et al. 2022). Further, two SWI/SNF complexes, PBAP and BAP, control two distinct aspects (growth and cell fate) of regeneration, respectively (Tian and Smith-Bolton 2021). The PBAP complex is responsible for regenerative growth and developmental timing, while the BAP complex is in charge of correct patterning and cell fate. Additionally, Brg1, another member of the SWI/SNF complex, was reported to regulate myocardial proliferation and regeneration in zebrafish by repressing cyclin-dependent kinase inhibitors (Xiao et al. 2016).
Chromatin organizers, such as CTCF and cohesin, function as physic hinges in organizing TADs into architectural loops (Chien et al. 2011; Han et al. 2008). These proteins participate in higher levels of regulation in chromatin organization. Cohesin catalyzes genome folding through loop extrusion which stops at the CTCF binding sites with a convergent orientation (Tang et al. 2015). The dynamic loop formation increases spatial cis-tethering over long distances and promotes transcriptional regulation. CTCF was identified with increased expression during cell reprogramming, which helps repress somatic genes and maintain chromatin accessibilities for partial enhancer-promoter interactions in cooperation with an ISWI chromatin remodeler SMARCA5 (Song et al. 2022). Moreover, a dual role of CTCF-dependent chromatin organization in controlling myelinogenic programs and recruiting chromatin-repressive complexes was reported in Schwann cells (Wang et al. 2020b). Deletion of CTCF blocks the interaction between promoter and enhancers of the locus of Egr2, leading to a strong reduction in the expression of the pro-myelinogenic factor EGR2 and the suppression of Schwann cell differentiation during nerve repair. Therefore, global changes of chromatin organization caused by aberrant regulation of chromatin organizers can cause unpredicted transcriptional and functional alterations in cells. In summary, chromatin regulators play a significant role in remodeling the architecture of chromatin and are actively involved in the regulation of regeneration (Fig. 2). Understanding how these chromatin regulators establish accessible chromatin and select enhancer-promoter interactions after injury should be informative in developing new strategies for re-activating regeneration in damaged human organs.
The AP-1 complex is a potential master regulator of the regeneration response program
Despite RRP is subjected to evolutionary changes and contains species-specific components, its regulation seems to share common features (Wang et al. 2020c). Identification of the master regulator of RRP is one of the key tasks in the field of regeneration. In adult mice, activation of the neuregulin1 (Nrg1) pathway induces cell-cycle reentry for cardiomyocytes and promotes myocardial regeneration, resulting in improved heart function post myocardial infarction (Bersell et al. 2009). Furthermore, transgenic reactivation of Nrg1 expression in intact zebrafish hearts turns on many hallmarks of cardiac regeneration and significantly enhances ventricular size (Gemberling et al. 2015). Thus, Nrg1 was considered an injury-induced mitogen of cardiomyocytes with the power to induce endogenous heart regeneration program in zebrafish (Gemberling et al. 2015). Because regeneration-responsive genes are restricted at the basal transcriptional levels during homeostasis, activation of this class of genes requires the remodeling of chromatin. Recently, the transcription factor binding motif of the Activator Protein 1 (AP-1) complex was found to be present in reported RREs including LEN, BRV-B, K-IEN, and others (Goldman and Poss 2020; Harris et al. 2016; Harris et al. 2020; Kang et al. 2016; Tamaki et al. 2023; Thompson et al. 2020; Wang et al. 2020c). The AP-1 complex, assembled through the dimerization of the bZIP domain between the Fos and Jun subunits, mediates many cellular and physiological functions in development and diseases. Remarkably, comparative motif enrichment analysis identified the presence of AP-1 binding motif as a common feature in RREs identified from both African killifish and zebrafish fin regeneration (Wang et al. 2020c). Deletion of the AP-1 motifs in tested RREs led to a complete loss of enhancer activities upon tissue damage (Wang et al. 2020c). Moreover, the AP-1 motifs recognized by the Jun family proteins (Jun, JunB, and JunD) exhibit a higher frequency in highly regenerative fish genomes than in human and mouse genomes (Wang et al. 2020c). To date, the AP-1 complex has been shown to play essential roles in regulating fin regeneration, Xenopus tail regeneration, zebrafish heart regeneration, axolotl spinal cord regeneration, mice liver regeneration, peripheral nerve regeneration, and skin repair (Angel et al. 2001; Beisaw et al. 2020; Ishida et al. 2010; Nakamura et al. 2020; Patodia and Raivich 2012; Sabin et al. 2019; Stepniak et al. 2006). Interestingly, the AP-1 complex directs enhancer selection to govern precise gene expression so that cells can differentiate and acquire specialized functions (Bejjani et al. 2019). Such enhancer selection is determined by a collaborative binding of FOS/JUN and cell-type-specific transcription factors to enhancers and the recruitment of the SWI/SNF (BAF) complex to create accessible chromatin (Vierbuchen et al. 2017). This is consistent with the observation that AP-1 transcription factors control the cardiomyocyte response to cryoinjury by regulating chromatin accessibility (Beisaw et al. 2020). These injury-induced open chromatin regions with AP-1 binding motifs allow the activation of a regeneration program that facilitates cardiomyocyte dedifferentiation, proliferation, and protrusion into the damaged area. All these data point out that AP-1 complex may function as a master regulator in the activation of RRP by controlling enhancer selection and chromatin accessibility (Fig. 3).
AP-1 complex and the activation of the regeneration response program
A model for the activation of the regeneration response program (RRP). The AP-1 complex is a potential master regulator of the RRP. Upon tissue damage, alteration of chromatin organization is initiated to establish accessible chromatin. Combined with other binding factors, the AP-1 complex selects enhancer-promoter interactions to turn on the RRP, which leads to the initiation and progression of regeneration
Whether AP-1 complex alone is sufficient to activate RRP needs further investigation. It has been noted that regeneration of different organs involves both shared and organ-specific regulators (Hui et al. 2017; Iismaa et al. 2018). Therefore, AP-1 complex may function with tissue-specific transcription factors and chromatin regulators to initiate regeneration upon tissue damage. Additionally, in highly regenerative invertebrates, such as acoel, the gene early growth response (egr) was identified using ATAC-seq as a pioneer factor to regulate early wound-induced genes (Gehrke et al. 2019). In vertebrates, egr-1 has been considered a critical mediator of fibroblast activation and fibrotic response triggered by diverse stimuli (Bhattacharyya et al. 2013). Abnormal activation of egr-1 has been linked to fibrosis and human fibrotic disorders (Bhattacharyya et al. 2011). Such differences may suggest basal animals with whole-body regeneration and vertebrates that only retain regenerative capacities in certain tissues or organs deploy distinct master regulators to initiate RRP. It would be interesting to investigate the major regulatory differences between animals with unlimited regenerative capacities and others with limited regeneration.
Conclusions
As a long-standing question in biology, regeneration has been widely investigated at different levels including cell regeneration, tissue/organ regeneration, appendage or structure regeneration, and whole-body regeneration. Numerous studies derived from distinct organs and species indicate that mechanisms established from a single species do not ensure successful application in humans. Therefore, there is an urgent need for the identification and characterization of conserved and species-specific regeneration response programs. Synthesizing information collected from different organisms ranging from the basal animals, such as sponges and hydra, to mammals such as deer and African spiny mice is critical for establishing evolutionarily conserved mechanisms underlying regeneration. Further, increasing attention has been paid to develop tools for the manipulation of gene expression in damaged tissues to reactivate regeneration. One such promising tool is the RRE or tissue-regeneration enhancer elements that can confer spatial or temporal control of key regeneration genes (Kang et al. 2016). A new study from the Poss group demonstrated that zebrafish RREs were sufficient to stimulate or suppress endogenous gene expression after ischemic myocardial infarction in mice (Yan et al. 2023). Interestingly, a constitutively active YAP factor driven by such tool was sufficient to promote cardiac regeneration in mice, resulting in improved function of the injured heart (Yan et al. 2023). In addition, other tools (such as chromatin-modifying drugs and metabolites and other small molecules) that are lack of context specificity have also been used to stimulate regeneration in mouse model. Further development and optimization of these tools will pave the way for establishing reliable strategies to restore regeneration in regenerative medicine.
Chromatin organization dependent gene regulation is a highly conserved regulatory mechanism that can be applied in development, regeneration, and diseases. Although several regulators controlling chromatin accessibility during regeneration have been identified, many fundamental questions remain elusive. For example, we still don’t know 1) what kind of chromatin organization underlies regenerative competency? 2) what factors are sufficient to establish such chromatin organization upon tissue damage? 3) what are the differences in chromatin organization between animals with unlimited regenerative capacities and others with limited regeneration? and 4) how do epigenetic modifications and regulations fine-tune regeneration in different species? New methodologies and technologies that have been developed for examining chromatin organization with high resolution will facilitate addressing these questions in future studies. Particularly, the combination of single-cell chromatin profiling techniques (eg., scATAC) and single-cell omics (eg., scRNAseq) provides new opportunities for identifying differences in gene expression and chromatin accessibility in each cell population involved in tissue regeneration (Chen et al. 2020b; Sinha et al. 2022; Wang et al. 2020d). For example, a recent integrated single-cell RNA-seq and ATAC-Seq analysis systematically mapped cell state transitions in more than 10,000 hepatocytes during liver regeneration and identified injury-associated signaling pathways involved in transitioning hepatocytes (Chen et al. 2020b). Similarly, another analysis helped generate open chromatin landscapes and regeneration-associated gene regulatory networks of distinct cardiac cell types following myocardial infarction (Wang et al. 2020d). Mapping cell type specific chromatin organization during regeneration is a critical step toward understanding the genetic basis of regeneration and the uneven distribution of this feature in animals.
Availability of data and materials
Not applicable.
Abbreviations
- 3C:
-
Chromosome Conformation Capture
- CRCs:
-
ATP-dependent chromatin remodeling complexes
- DNMT:
-
DNA methyltransferase
- HAT:
-
Histone acetyltransferase
- HDAC:
-
Histone deacetylase
- HDM:
-
Histone demethylase
- HMT:
-
Histone methyltransferase
- UHRF1:
-
Ubiquitin Like With PHD And Ring Finger Domains 1
- INO80:
-
Inositol requiring 80
- NuRD:
-
Nucleosome remodeling and deacetylase
- ISWI:
-
Imitation Switch
- RRE:
-
Regeneration-Responsive Enhancer
- SWI/SNF:
-
Switching defective/Sucrose nonfermenting
- TADs:
-
Topologically Associating Domains
References
Adam RC, Yang H, Rockowitz S, Larsen SB, Nikolova M, Oristian DS, et al. Pioneer factors govern super-enhancer dynamics in stem cell plasticity and lineage choice. Nature. 2015;521(7552):366–70. https://doi.org/10.1038/nature14289.
Aguilar CA, Pop R, Shcherbina A, Watts A, Matheny RW Jr, Cacchiarelli D, et al. Transcriptional and chromatin dynamics of muscle regeneration after severe trauma. Stem Cell Reports. 2016;7(5):983–97. https://doi.org/10.1016/j.stemcr.2016.09.009.
Ahmad Ganai S, Ramadoss M, Mahadevan V. Histone Deacetylase (HDAC) Inhibitors-emerging roles in neuronal memory, learning, synaptic plasticity and neural regeneration. Curr Neuropharmacol. 2016;14(1):55–71. https://doi.org/10.2174/1570159x13666151021111609.
Alkass K, Panula J, Westman M, Wu TD, Guerquin-Kern JL, Bergmann O. No evidence for cardiomyocyte number expansion in preadolescent mice. Cell. 2015;163(4):1026–36. https://doi.org/10.1016/j.cell.2015.10.035.
Allfrey VG, Faulkner R, Mirsky A. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci. 1964;51(5):786–94. https://doi.org/10.1073/pnas.51.5.786.
Alvarado AS, Tsonis PA. Bridging the regeneration gap: genetic insights from diverse animal models. Nat Rev Genet. 2006;7(11):873–84. https://doi.org/10.1038/nrg1923.
Amano T, Sagai T, Tanabe H, Mizushina Y, Nakazawa H, Shiroishi T. Chromosomal dynamics at the Shh locus: limb bud-specific differential regulation of competence and active transcription. Dev Cell. 2009;16(1):47–57. https://doi.org/10.1016/j.devcel.2008.11.011.
Anania C, Lupianez DG. Order and disorder: abnormal 3D chromatin organization in human disease. Brief Funct Genomics. 2020;19(2):128–38. https://doi.org/10.1093/bfgp/elz028.
Anatskaya OV, Vinogradov AE. Polyploidy as a fundamental phenomenon in evolution, development, adaptation and diseases. Int J Mol Sci. 2022;23(7):3542. https://doi.org/10.3390/ijms23073542.
Andrey G, Montavon T, Mascrez B, Gonzalez F, Noordermeer D, Leleu M, et al. A switch between topological domains underlies HoxD genes collinearity in mouse limbs. Science. 2013;340(6137):1234167. https://doi.org/10.1126/science.1234167.
Angel P, Szabowski A, Schorpp-Kistner M. Function and regulation of AP-1 subunits in skin physiology and pathology. Oncogene. 2001;20(19):2413–23. https://doi.org/10.1038/sj.onc.1204380.
Arechederra M, Berasain C, Avila MA, Fernandez-Barrena MG. Chromatin dynamics during liver regeneration. Semin Cell Dev Biol. 2020;97:38–46. https://doi.org/10.1016/j.semcdb.2019.03.004.
Azpiazu N, Morata G. Chromatin remodelling and retrotransposons activities during regeneration in Drosophila. Dev Biol. 2022;482:7–16. https://doi.org/10.1016/j.ydbio.2021.11.005.
Bailey EC, Kobielski S, Park J, Losick VP. Polyploidy in Tissue Repair and Regeneration. Cold Spring Harb Perspect Biol. 2021;13(10):040881. https://doi.org/10.1101/cshperspect.a040881.
Batut PJ, Bing XY, Sisco Z, Raimundo J, Levo M, Levine MS. Genome organization controls transcriptional dynamics during development. Science. 2022;375(6580):566–70. https://doi.org/10.1126/science.abi7178.
Beagan JA, Phillips-Cremins JE. On the existence and functionality of topologically associating domains. Nat Genet. 2020;52(1):8–16. https://doi.org/10.1038/s41588-019-0561-1.
Becker JS, Nicetto D, Zaret KS. H3K9me3-Dependent Heterochromatin: Barrier to Cell Fate Changes. Trends Genet. 2016;32(1):29–41. https://doi.org/10.1016/j.tig.2015.11.001.
Beisaw A, Kuenne C, Guenther S, Dallmann J, Wu CC, Bentsen M, et al. AP-1 contributes to chromatin accessibility to promote sarcomere disassembly and cardiomyocyte protrusion during zebrafish heart regeneration. Circ Res. 2020;126(12):1760–78. https://doi.org/10.1161/CIRCRESAHA.119.316167.
Bejjani F, Evanno E, Zibara K, Piechaczyk M, Jariel-Encontre I. The AP-1 transcriptional complex: Local switch or remote command? Biochim Biophys Acta Rev Cancer. 2019;1872(1):11–23. https://doi.org/10.1016/j.bbcan.2019.04.003.
Bely AE. Evolutionary loss of animal regeneration: pattern and process. Integr Comp Biol. 2010;50(4):515–27. https://doi.org/10.1093/icb/icq118.
Bersell K, Arab S, Haring B, Kuhn B. Neuregulin1/ErbB4 signaling induces cardiomyocyte proliferation and repair of heart injury. Cell. 2009;138(2):257–70. https://doi.org/10.1016/j.cell.2009.04.060.
Bestor TH. The DNA methyltransferases of mammals. Hum Mol Genet. 2000;9(16):2395–402. https://doi.org/10.1093/hmg/9.16.2395.
Bhatt A, Fan LW, Pang Y. Strategies for myelin regeneration: lessons learned from development. Neural Regen Res. 2014;9(14):1347–50. https://doi.org/10.4103/1673-5374.137586.
Bhattacharya D, Talwar S, Mazumder A, Shivashankar GV. Spatio-temporal plasticity in chromatin organization in mouse cell differentiation and during Drosophila embryogenesis. Biophys J. 2009;96(9):3832–9. https://doi.org/10.1016/j.bpj.2008.11.075.
Bhattacharyya S, Wu M, Fang F, Tourtellotte W, Feghali-Bostwick C, Varga J. Early growth response transcription factors: key mediators of fibrosis and novel targets for anti-fibrotic therapy. Matrix Biol. 2011;30(4):235–42. https://doi.org/10.1016/j.matbio.2011.03.005.
Bhattacharyya S, Fang F, Tourtellotte W, Varga J. Egr-1: new conductor for the tissue repair orchestra directs harmony (regeneration) or cacophony (fibrosis). J Pathol. 2013;229(2):286–97. https://doi.org/10.1002/path.4131.
Birnbaum KD, Sanchez AA. Slicing across kingdoms: regeneration in plants and animals. Cell. 2008;132(4):697–710. https://doi.org/10.1016/j.cell.2008.01.040.
Bronner C, Alhosin M, Hamiche A, Mousli M. Coordinated dialogue between UHRF1 and DNMT1 to ensure faithful inheritance of methylated DNA patterns. Genes. 2019;10(1):65. https://doi.org/10.3390/genes10010065.
Chapman JA, Kirkness EF, Simakov O, Hampson SE, Mitros T, Weinmaier T, et al. The dynamic genome of Hydra. Nature. 2010;464(7288):592–6. https://doi.org/10.1038/nature08830.
Chen T, Dent SY. Chromatin modifiers and remodellers: regulators of cellular differentiation. Nat Rev Genet. 2014;15(2):93–106. https://doi.org/10.1038/nrg3607.
Chen F, Jimenez RJ, Sharma K, Luu HY, Hsu BY, Ravindranathan A, et al. Broad distribution of hepatocyte proliferation in liver homeostasis and regeneration. Cell Stem Cell. 2020;26(1):27–334. https://doi.org/10.1016/j.stem.2019.11.001.
Chen T, Oh S, Gregory S, Shen X, Diehl AM. Single-cell omics analysis reveals functional diversification of hepatocytes during liver regeneration. JCI Insight. 2020;5(22):141024. https://doi.org/10.1172/jci.insight.141024.
Chien R, Zeng W, Ball AR, Yokomori K. Cohesin: a critical chromatin organizer in mammalian gene regulation. Biochem Cell Biol. 2011;89(5):445–58. https://doi.org/10.1139/o11-039.
Clapier CR, Cairns BR. The biology of chromatin remodeling complexes. Annu Rev Biochem. 2009;78:273–304. https://doi.org/10.1146/annurev.biochem.77.062706.153223.
Clapier CR, Iwasa J, Cairns BR, Peterson CL. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat Rev Mol Cell Biol. 2017;18(7):407–22. https://doi.org/10.1038/nrm.2017.26.
Cremer T, Cremer C, Baumann H, Luedtke EK, Sperling K, Teuber V, et al. Rabl’s model of the interphase chromosome arrangement tested in Chinese hamster cells by premature chromosome condensation and laser-UV-microbeam experiments. Hum Genet. 1982;60(1):46–56. https://doi.org/10.1007/BF00281263.
Dehal P, Boore JL. Two rounds of whole genome duplication in the ancestral vertebrate. PLoS Biol. 2005;3(10):314. https://doi.org/10.1371/journal.pbio.0030314.
Dekker J, Mirny L. The 3D Genome as Moderator of Chromosomal Communication. Cell. 2016;164(6):1110–21. https://doi.org/10.1016/j.cell.2016.02.007.
Dekker J, Rippe K, Dekker M, Kleckner N. Capturing chromosome conformation. Science. 2002;295(5558):1306–11. https://doi.org/10.1126/science.1067799.
Denk F, Crow M, Didangelos A, Lopes DM, McMahon SB. Persistent alterations in microglial enhancers in a model of chronic pain. Cell Rep. 2016;15(8):1771–81. https://doi.org/10.1016/j.celrep.2016.04.063.
Díaz-Castillo C. Regeneration: Why junk DNA might matter. PeerJ Preprints. 2018;e27255v1. https://doi.org/10.7287/peerj.preprints.27255v1.
Dixon JR, Selvaraj S, Yue F, Kim A, Li Y, Shen Y, et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature. 2012;485(7398):376–80. https://doi.org/10.1038/nature11082.
Dixon JR, Jung I, Selvaraj S, Shen Y, Antosiewicz-Bourget JE, Lee AY, et al. Chromatin architecture reorganization during stem cell differentiation. Nature. 2015;518(7539):331–6. https://doi.org/10.1038/nature14222.
Dornen J, Sieler M, Weiler J, Keil S, Dittmar T. Cell fusion-mediated tissue regeneration as an inducer of polyploidy and aneuploidy. Int J Mol Sci. 2020;21(5):1811. https://doi.org/10.3390/ijms21051811.
Dunn TM, Hahn S, Ogden S, Schleif RF. An operator at-280 base pairs that is required for repression of araBAD operon promoter: addition of DNA helical turns between the operator and promoter cyclically hinders repression. Proc Natl Acad Sci. 1984;81(16):5017–20. https://doi.org/10.1073/pnas.81.16.5017.
Efroni I, Mello A, Nawy T, Ip PL, Rahni R, DelRose N, et al. Root Regeneration Triggers an Embryo-like Sequence Guided by Hormonal Interactions. Cell. 2016;165(7):1721–33. https://doi.org/10.1016/j.cell.2016.04.046.
Flici H, Frank U. Inhibition of SoxB2 or HDACs suppresses Hydractinia head regeneration by affecting blastema formation. Commun Integr Biol. 2018;11(2):1–5. https://doi.org/10.1080/19420889.2018.1450032.
Friedrich M, Gerbeth L, Gerling M, Rosenthal R, Steiger K, Weidinger C, et al. HDAC inhibitors promote intestinal epithelial regeneration via autocrine TGFbeta1 signalling in inflammation. Mucosal Immunol. 2019;12(3):656–67. https://doi.org/10.1038/s41385-019-0135-7.
Garcia-Lozano M, Natarajan P, Levi A, Katam R, Lopez-Ortiz C, Nimmakayala P, et al. Altered chromatin conformation and transcriptional regulation in watermelon following genome doubling. Plant J. 2021;106(3):588–600. https://doi.org/10.1111/tpj.15256.
Garriga J, Laumet G, Chen S-R, Zhang Y, Madzo J, Issa JPJ, et al. Nerve injury-induced chronic pain is associated with persistent DNA methylation reprogramming in dorsal root ganglion. J Neurosci. 2018;38(27):6090–101. https://doi.org/10.1523/JNEUROSCI.2616-17.2018.10.1523/JNEUROSCI.2616-17.2018.
Garza-Garcia AA, Driscoll PC, Brockes JP. Evidence for the local evolution of mechanisms underlying limb regeneration in salamanders. Integr Comp Biol. 2010;50(4):528–35. https://doi.org/10.1093/icb/icq022.
Gates LA, Foulds CE, O’Malley BW. Histone Marks in the “driver’s seat”: functional roles in steering the transcription cycle. Trends Biochem Sci. 2017;42(12):977–89. https://doi.org/10.1016/j.tibs.2017.10.004.
Gaub P, Joshi Y, Wuttke A, Naumann U, Schnichels S, Heiduschka P, et al. The histone acetyltransferase p300 promotes intrinsic axonal regeneration. Brain. 2011;134(Pt 7):2134–48. https://doi.org/10.1093/brain/awr142.
Ge Y, Gomez NC, Adam RC, Nikolova M, Yang H, Verma A, et al. Stem cell lineage infidelity drives wound repair and cancer. Cell. 2017;169(4):636-50e14. https://doi.org/10.1016/j.cell.2017.03.042.
Gehrke AR, Neverett E, Luo YJ, Brandt A, Ricci L, Hulett RE, et al. Acoel genome reveals the regulatory landscape of whole-body regeneration. Science. 2019;363(6432):eaau6173. https://doi.org/10.1126/science.aau6173.
Gemberling M, Karra R, Dickson AL, Poss KD. Nrg1 is an injury-induced cardiomyocyte mitogen for the endogenous heart regeneration program in zebrafish. Elife. 2015;4:e05871. https://doi.org/10.7554/eLife.05871.
Geyer PK, Vitalini MW, Wallrath LL. Nuclear organization: taking a position on gene expression. Curr Opin Cell Biol. 2011;23(3):354–9. https://doi.org/10.1016/j.ceb.2011.03.002.
Goldman JA, Poss KD. Gene regulatory programmes of tissue regeneration. Nat Rev Genet. 2020;21(9):511–25. https://doi.org/10.1038/s41576-020-0239-7.
Gonzalez-Rosa JM, Sharpe M, Field D, Soonpaa MH, Field LJ, Burns CE, et al. Myocardial Polyploidization Creates a Barrier to Heart Regeneration in Zebrafish. Dev Cell. 2018;44(4):433-46e7. https://doi.org/10.1016/j.devcel.2018.01.021.
Gordon JAR, Stein JL, Westendorf JJ, van Wijnen AJ. Chromatin modifiers and histone modifications in bone formation, regeneration, and therapeutic intervention for bone-related disease. Bone. 2015;81:739–45. https://doi.org/10.1016/j.bone.2015.03.011.
Gornikiewicz B, Ronowicz A, Krzeminski M, Sachadyn P. Changes in gene methylation patterns in neonatal murine hearts: Implications for the regenerative potential. BMC Genomics. 2016;17(1):231. https://doi.org/10.1186/s12864-016-2545-1.
Grohme MA, Schloissnig S, Rozanski A, Pippel M, Young GR, Winkler S, et al. The genome of Schmidtea mediterranea and the evolution of core cellular mechanisms. Nature. 2018;554(7690):56–61. https://doi.org/10.1038/nature25473.
Guenther CA, Wang Z, Li E, Tran MC, Logan CY, Nusse R, et al. A distinct regulatory region of the Bmp5 locus activates gene expression following adult bone fracture or soft tissue injury. Bone. 2015;77:31–41. https://doi.org/10.1016/j.bone.2015.04.010.
Han L, Lee D-H, Szabó PE. CTCF is the master organizer of domain-wide allele-specific chromatin at the H19/Igf2 imprinted region. Mol Cell Biol. 2008;28(3):1124–35. https://doi.org/10.1128/MCB.01361-07.
Han J, Zhang Z, Wang K. 3C and 3C-based techniques: the powerful tools for spatial genome organization deciphering. Mol Cytogenet. 2018;11:21. https://doi.org/10.1186/s13039-018-0368-2.
Harris RE, Setiawan L, Saul J, Hariharan IK. Localized epigenetic silencing of a damage-activated WNT enhancer limits regeneration in mature Drosophila imaginal discs. Elife. 2016;5:e11588. https://doi.org/10.7554/eLife.11588.
Harris RE, Stinchfield MJ, Nystrom SL, McKay DJ, Hariharan IK. Damage-responsive, maturity-silenced enhancers regulate multiple genes that direct regeneration in Drosophila. Elife. 2020;9:e58305. https://doi.org/10.7554/eLife.58305.
Heitz E. Das heterochromatin der Moose. Jahrb Wiss Botanik. 1928.
Henderson IR, Bomblies K. Evolution and plasticity of genome-wide meiotic recombination rates. Annu Rev Genet. 2021;55:23–43. https://doi.org/10.1146/annurev-genet-021721-033821.
Higgs DR. Enhancer-promoter interactions and transcription. Nat Genet. 2020;52(5):470–1. https://doi.org/10.1038/s41588-020-0620-7.
Hildebrand EM, Dekker J. Mechanisms and functions of chromosome compartmentalization. Trends Biochem Sci. 2020;45(5):385–96. https://doi.org/10.1016/j.tibs.2020.01.002.
Hnisz D, Weintraub AS, Day DS, Valton AL, Bak RO, Li CH, et al. Activation of proto-oncogenes by disruption of chromosome neighborhoods. Science. 2016;351(6280):1454–8. https://doi.org/10.1126/science.aad9024.
Hoang T, Wang J, Boyd P, Wang F, Santiago C, Jiang L, et al. Gene regulatory networks controlling vertebrate retinal regeneration. Science. 2020;370(6519). https://doi.org/10.1126/science.abb8598
Hu CK, Brunet A. The African turquoise killifish: A research organism to study vertebrate aging and diapause. Aging cell. 2018;17(3):e12757. https://doi.org/10.1111/acel.12757.
Hui SP, Sheng DZ, Sugimoto K, Gonzalez-Rajal A, Nakagawa S, Hesselson D, et al. Zebrafish regulatory T cells mediate organ-specific regenerative programs. Dev Cell. 2017;43(6):659-72e5. https://doi.org/10.1016/j.devcel.2017.11.010.
Hung HA, Sun G, Keles S, Svaren J. Dynamic regulation of Schwann cell enhancers after peripheral nerve injury. J Biol Chem. 2015;290(11):6937–50. https://doi.org/10.1074/jbc.M114.622878.
Huynh CN, Everts V, Ampornaramveth RS. Histone deacetylases and their roles in mineralized tissue regeneration. Bone Rep. 2017;7:33–40. https://doi.org/10.1016/j.bonr.2017.08.001.
Iismaa SE, Kaidonis X, Nicks AM, Bogush N, Kikuchi K, Naqvi N, et al. Comparative regenerative mechanisms across different mammalian tissues. NPJ Regen Med. 2018;3:6. https://doi.org/10.1038/s41536-018-0044-5.
Ishida T, Nakajima T, Kudo A, Kawakami A. Phosphorylation of Junb family proteins by the Jun N-terminal kinase supports tissue regeneration in zebrafish. Dev Biol. 2010;340(2):468–79. https://doi.org/10.1016/j.ydbio.2010.01.036.
Jia Y, Li L, Lin YH, Gopal P, Shen S, Zhou K, et al. In vivo CRISPR screening identifies BAZ2 chromatin remodelers as druggable regulators of mammalian liver regeneration. Cell Stem Cell. 2022;29(3):372-85e8. https://doi.org/10.1016/j.stem.2022.01.001.
Jones B. Epigenetics: Histones pass the message on. Nat Rev Genet. 2015;16(1):3. https://doi.org/10.1038/nrg3876.
Jopling C, Sleep E, Raya M, Marti M, Raya A, Izpisua Belmonte JC. Zebrafish heart regeneration occurs by cardiomyocyte dedifferentiation and proliferation. Nature. 2010;464(7288):606–9. https://doi.org/10.1038/nature08899.
Jorstad NL, Wilken MS, Grimes WN, Wohl SG, VandenBosch LS, Yoshimatsu T, et al. Stimulation of functional neuronal regeneration from Muller glia in adult mice. Nature. 2017;548(7665):103–7. https://doi.org/10.1038/nature23283.
Kadow ZA, Martin JF. A Role for Ploidy in Heart Regeneration. Dev Cell. 2018;44(4):403–4. https://doi.org/10.1016/j.devcel.2018.02.004.
Kakebeen AD, Chitsazan AD, Williams MC, Saunders LM, Wills AE. Chromatin accessibility dynamics and single cell RNA-Seq reveal new regulators of regeneration in neural progenitors. Elife. 2020;9:e52648. https://doi.org/10.7554/eLife.52648.
Kang J, Hu J, Karra R, Dickson AL, Tornini VA, Nachtrab G, et al. Modulation of tissue repair by regeneration enhancer elements. Nature. 2016;532(7598):201–6. https://doi.org/10.1038/nature17644.
Kirillova A, Han L, Liu H, Kuhn B. Polyploid cardiomyocytes: implications for heart regeneration. Development. 2021;148(14):dev199401. https://doi.org/10.1242/dev.199401.10.1242/dev.199401.
Klemm SL, Shipony Z, Greenleaf WJ. Chromatin accessibility and the regulatory epigenome. Nat Rev Genet. 2019;20(4):207–20. https://doi.org/10.1038/s41576-018-0089-8.
Kmita M, Tarchini B, Zakany J, Logan M, Tabin CJ, Duboule D. Early developmental arrest of mammalian limbs lacking HoxA/HoxD gene function. Nature. 2005;435(7045):1113–6. https://doi.org/10.1038/nature03648.
Krijger PH, de Laat W. Regulation of disease-associated gene expression in the 3D genome. Nat Rev Mol Cell Biol. 2016;17(12):771–82. https://doi.org/10.1038/nrm.2016.138.
Lee HJ, Hou Y, Chen Y, Dailey ZZ, Riddihough A, Jang HS, et al. Regenerating zebrafish fin epigenome is characterized by stable lineage-specific DNA methylation and dynamic chromatin accessibility. Genome Biol. 2020;21(1):52. https://doi.org/10.1186/s13059-020-1948-0.
Leone M, Magadum A, Engel FB. Cardiomyocyte proliferation in cardiac development and regeneration: a guide to methodologies and interpretations. Am J Physiol Heart Circ Physiol. 2015;309(8):H1237–50. https://doi.org/10.1152/ajpheart.00559.2015.
Lieberman-Aiden E, van Berkum NL, Williams L, Imakaev M, Ragoczy T, Telling A, et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. 2009;326(5950):289–93. https://doi.org/10.1126/science.1181369.
Lin Z, Chen L, Chen X, Zhong Y, Yang Y, Xia W, et al. Biological adaptations in the Arctic cervid, the reindeer (Rangifer tarandus). Science. 2019;364:eaav6312. https://doi.org/10.1126/science.aav6312.
Londono R, Sun AX, Tuan RS, Lozito TP. Tissue repair and epimorphic regeneration: an overview. Curr Pathobiol Rep. 2018;6(1):61–9. https://doi.org/10.1007/s40139-018-0161-2.
Lucchetta EM, Ohlstein B. Amitosis of Polyploid Cells Regenerates Functional Stem Cells in the Drosophila Intestine. Cell Stem Cell. 2017;20(5):609-20e6. https://doi.org/10.1016/j.stem.2017.02.012.10.1016/j.stem.2017.02.012.
Lupianez DG, Kraft K, Heinrich V, Krawitz P, Brancati F, Klopocki E, et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell. 2015;161(5):1012–25. https://doi.org/10.1016/j.cell.2015.04.004.
Lynch M, Conery JS. The evolutionary fate and consequences of duplicate genes. Science. 2000;290(5494):1151–5. https://doi.org/10.1126/science.290.5494.1151.
Macchi F, Sadler KC. Unraveling the epigenetic basis of liver development, regeneration and disease. Trends Genet. 2020;36(8):587–97. https://doi.org/10.1016/j.tig.2020.05.002.
Malloch EL, Perry KJ, Fukui L, Johnson VR, Wever J, Beck CW, et al. Gene expression profiles of lens regeneration and development in Xenopus laevis. Dev Dyn. 2009;238(9):2340–56. https://doi.org/10.1002/dvdy.21998.
Matsumoto T, Wakefield L, Tarlow BD, Grompe M. In vivo lineage tracing of polyploid hepatocytes reveals extensive proliferation during liver regeneration. Cell Stem Cell. 2020;26(1):34-47e3. https://doi.org/10.1016/j.stem.2019.11.014.10.1016/j.stem.2019.11.014.
McArthur E, Capra JA. Topologically associating domain boundaries that are stable across diverse cell types are evolutionarily constrained and enriched for heritability. Am J Human Genet. 2021;108(2):269–83. https://doi.org/10.1016/j.ajhg.2021.01.001.
Merrell AJ, Stanger BZ. Adult cell plasticity in vivo: de-differentiation and transdifferentiation are back in style. Nat Rev Mol Cell Biol. 2016;17(7):413–25. https://doi.org/10.1038/nrm.2016.24.
Miyaoka Y, Ebato K, Kato H, Arakawa S, Shimizu S, Miyajima A. Hypertrophy and unconventional cell division of hepatocytes underlie liver regeneration. Curr Biol. 2012;22(13):1166–75. https://doi.org/10.1016/j.cub.2012.05.016.
Moris N, Pina C, Arias AM. Transition states and cell fate decisions in epigenetic landscapes. Nat Rev Genet. 2016;17(11):693–703. https://doi.org/10.1038/nrg.2016.98.
Murad R, Macias-Munoz A, Wong A, Ma X, Mortazavi A. Coordinated Gene Expression and Chromatin Regulation during Hydra Head Regeneration. Genome Biol Evol. 2021;13(12):evab221. https://doi.org/10.1093/gbe/evab221.10.1093/gbe/evab221.
Naik S, Larsen SB, Gomez NC, Alaverdyan K, Sendoel A, Yuan S, et al. Inflammatory memory sensitizes skin epithelial stem cells to tissue damage. Nature. 2017;550(7677):475–80. https://doi.org/10.1038/nature24271.
Nakamura M, Yoshida H, Takahashi E, Wlizla M, Takebayashi-Suzuki K, Horb ME, et al. The AP-1 transcription factor JunB functions in Xenopus tail regeneration by positively regulating cell proliferation. Biochem Biophys Res Commun. 2020;522(4):990–5. https://doi.org/10.1016/j.bbrc.2019.11.060.
Nakato R, Sakata T. Methods for ChIP-seq analysis: A practical workflow and advanced applications. Methods. 2021;187:44–53. https://doi.org/10.1016/j.ymeth.2020.03.005.
Nasser J, Bergman DT, Fulco CP, Guckelberger P, Doughty BR, Patwardhan TA, et al. Genome-wide enhancer maps link risk variants to disease genes. Nature. 2021;593(7858):238–43. https://doi.org/10.1038/s41586-021-03446-x.
Nowoshilow S, Schloissnig S, Fei JF, Dahl A, Pang AWC, Pippel M, et al. The axolotl genome and the evolution of key tissue formation regulators. Nature. 2018;554(7690):50–5. https://doi.org/10.1038/nature25458.
Olson MV. When less is more: gene loss as an engine of evolutionary change. Am J Human Genet. 1999;64(1):18–23. https://doi.org/10.1086/302219.
Ovrebo JI, Edgar BA. Polyploidy in tissue homeostasis and regeneration. Development. 2018;145(14):dev156034. https://doi.org/10.1242/dev.156034.10.1242/dev.156034.
Patodia S, Raivich G. Role of transcription factors in peripheral nerve regeneration. Front Mol Neurosci. 2012;5:8. https://doi.org/10.3389/fnmol.2012.00008.
Peters AH, Kubicek S, Mechtler K, O’Sullivan RJ, Derijck AA, Perez-Burgos L, et al. Partitioning and plasticity of repressive histone methylation states in mammalian chromatin. Mol Cell. 2003;12(6):1577–89. https://doi.org/10.1016/s1097-2765(03)00477-5.
Petrov DA. Evolution of genome size: new approaches to an old problem. Trends Genet. 2001;17(1):23–8. https://doi.org/10.1016/s0168-9525(00)02157-0.
Pfefferli C, Jazwinska A. The art of fin regeneration in zebrafish. Regeneration (Oxf). 2015;2(2):72–83. https://doi.org/10.1002/reg2.33.
Phillips-Cremins JE, Sauria ME, Sanyal A, Gerasimova TI, Lajoie BR, Bell JS, et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell. 2013;153(6):1281–95. https://doi.org/10.1016/j.cell.2013.04.053.
Piatti P, Zeilner A, Lusser A. ATP-dependent chromatin remodeling factors and their roles in affecting nucleosome fiber composition. Int J Mol Sci. 2011;12(10):6544–65. https://doi.org/10.3390/ijms12106544.
Planques A, Kerner P, Ferry L, Grunau C, Gazave E, Vervoort M. DNA methylation atlas and machinery in the developing and regenerating annelid Platynereis dumerilii. BMC Biol. 2021;19(1):148. https://doi.org/10.1186/s12915-021-01074-5.
Poss KD. Advances in understanding tissue regenerative capacity and mechanisms in animals. Nat Rev Genet. 2010;11(10):710–22. https://doi.org/10.1038/nrg2879.
Potts HG, Stockdale WT, Mommersteeg MTM. Unlocking the secrets of the regenerating fish heart: comparing regenerative models to shed light on successful regeneration. J Cardiovasc Dev Dis. 2021;8(1):4. https://doi.org/10.3390/jcdd8010004.
Puttagunta R, Tedeschi A, Soria MG, Hervera A, Lindner R, Rathore KI, et al. PCAF-dependent epigenetic changes promote axonal regeneration in the central nervous system. Nat Commun. 2014;5(1):3527. https://doi.org/10.1038/ncomms4527.
Reddien PW. The Cellular and Molecular Basis for Planarian Regeneration. Cell. 2018;175(2):327–45. https://doi.org/10.1016/j.cell.2018.09.021.
Reichwald K, Petzold A, Koch P, Downie BR, Hartmann N, Pietsch S, et al. Insights into sex chromosome evolution and aging from the genome of a short-lived fish. Cell. 2015;163(6):1527–38. https://doi.org/10.1016/j.cell.2015.10.071.
Rothbart SB, Strahl BD. Interpreting the language of histone and DNA modifications. Biochim Biophys Acta. 2014;1839(8):627–43. https://doi.org/10.1016/j.bbagrm.2014.03.001.
Ryoo HD, Bergmann A. The role of apoptosis-induced proliferation for regeneration and cancer. Cold Spring Harb Perspect Biol. 2012;4(8):a008797. https://doi.org/10.1101/cshperspect.a008797.
Sabin KZ, Jiang P, Gearhart MD, Stewart R, Echeverri K. AP-1(cFos/JunB)/miR-200a regulate the pro-regenerative glial cell response during axolotl spinal cord regeneration. Commun Biol. 2019;2(1):91. https://doi.org/10.1038/s42003-019-0335-4.
Sagai T, Hosoya M, Mizushina Y, Tamura M, Shiroishi T. Elimination of a long-range cis-regulatory module causes complete loss of limb-specific Shh expression and truncation of the mouse limb. Development. 2005;132(4):797–803. https://doi.org/10.1242/dev.01613.
Sati S, Cavalli G. Chromosome conformation capture technologies and their impact in understanding genome function. Chromosoma. 2017;126(1):33–44. https://doi.org/10.1007/s00412-016-0593-6.
Schoenfelder S, Fraser P. Long-range enhancer-promoter contacts in gene expression control. Nat Rev Genet. 2019;20(8):437–55. https://doi.org/10.1038/s41576-019-0128-0.
Schoenfelder S, Sugar R, Dimond A, Javierre BM, Armstrong H, Mifsud B, et al. Polycomb repressive complex PRC1 spatially constrains the mouse embryonic stem cell genome. Nat Genet. 2015;47(10):1179–86. https://doi.org/10.1038/ng.3393.
Schotta G, Lachner M, Sarma K, Ebert A, Sengupta R, Reuter G, et al. A silencing pathway to induce H3–K9 and H4–K20 trimethylation at constitutive heterochromatin. Genes Dev. 2004;18(11):1251–62. https://doi.org/10.1101/gad.300704.
Sen CK, Ghatak S. miRNA control of tissue repair and regeneration. Am J Pathol. 2015;185(10):2629–40. https://doi.org/10.1016/j.ajpath.2015.04.001.
Sessions SK, Larson A. Developmental correlates of genome size in plethodontid salamanders and their implications for genome evolution. Evolution. 1987;41(6):1239–51. https://doi.org/10.1111/j.1558-5646.1987.tb02463.x.
Sessions SK, Wake DB. Forever young: Linking regeneration and genome size in salamanders. Dev Dyn. 2021;250(6):768–78. https://doi.org/10.1002/dvdy.279.
Shao Y, Wang XB, Zhang JJ, Li ML, Wu SS, Ma XY, et al. Genome and single-cell RNA-sequencing of the earthworm Eisenia andrei identifies cellular mechanisms underlying regeneration. Nat Commun. 2020;11(1):2656. https://doi.org/10.1038/s41467-020-16454-8.
Sharakhov IV, Sharakhova MV. Heterochromatin, histone modifications, and nuclear architecture in disease vectors. Curr Opin Insect Sci. 2015;10:110–7. https://doi.org/10.1016/j.cois.2015.05.003.
Silva SMD, Gates PB, Brockes JP. The Newt Ortholog of CD59 Is Implicated in Proximodistal Identity during Amphibian Limb Regeneration. Dev Cell. 2002;3(4):547–55. https://doi.org/10.1016/s1534-5807(02)00288-5.
Sinha S, Sparks HD, Labit E, Robbins HN, Gowing K, Jaffer A, et al. Fibroblast inflammatory priming determines regenerative versus fibrotic skin repair in reindeer. Cell. 2022;185(25):4717-36e25. https://doi.org/10.1016/j.cell.2022.11.004.10.1016/j.cell.2022.11.004.
Slotkin RK, Martienssen R. Transposable elements and the epigenetic regulation of the genome. Nat Rev Genet. 2007;8(4):272–85. https://doi.org/10.1038/nrg2072.
Song Y, Liang Z, Zhang J, Hu G, Wang J, Li Y, et al. CTCF functions as an insulator for somatic genes and a chromatin remodeler for pluripotency genes during reprogramming. Cell Rep. 2022;39(1):110626. https://doi.org/10.1016/j.celrep.2022.110626.
Stepniak E, Ricci R, Eferl R, Sumara G, Sumara I, Rath M, et al. c-Jun/AP-1 controls liver regeneration by repressing p53/p21 and p38 MAPK activity. Genes Dev. 2006;20(16):2306–14. https://doi.org/10.1101/gad.390506.
Sun F, Ou J, Shoffner AR, Luan Y, Yang H, Song L, et al. Enhancer selection dictates gene expression responses in remote organs during tissue regeneration. Nat Cell Biol. 2022;24(5):685–96. https://doi.org/10.1038/s41556-022-00906-y.
Suzuki N, Ochi H. Regeneration enhancers: A clue to reactivation of developmental genes. Dev Growth Differ. 2020;62(5):343–54. https://doi.org/10.1111/dgd.12654.
Suzuki N, Hirano K, Ogino H, Ochi H. Arid3a regulates nephric tubule regeneration via evolutionarily conserved regeneration signal-response enhancers. Elife. 2019;8:e43186. https://doi.org/10.7554/eLife.43186.
Suzuki N, Kanai A, Suzuki Y, Ogino H, Ochi H. Adrenergic receptor signaling induced by Klf15, a regulator of regeneration enhancer, promotes kidney reconstruction. Proc Natl Acad Sci U S A. 2022;119(33):e2204338119. https://doi.org/10.1073/pnas.2204338119.
Tamaki T, Yoshida T, Shibata E, Nishihara H, Ochi H, Kawakami A. Splashed E-box and AP-1 motifs cooperatively drive regeneration response and shape regeneration abilities. Biol Open. 2023;12(2):bio059810. https://doi.org/10.1242/bio.059810.10.1242/bio.059810.
Tanaka EM, Reddien PW. The cellular basis for animal regeneration. Dev Cell. 2011;21(1):172–85. https://doi.org/10.1016/j.devcel.2011.06.016.
Tang Z, Luo OJ, Li X, Zheng M, Zhu JJ, Szalaj P, et al. CTCF-Mediated Human 3D Genome Architecture Reveals Chromatin Topology for Transcription. Cell. 2015;163(7):1611–27. https://doi.org/10.1016/j.cell.2015.11.024.
Tenaillon O, Barrick JE, Ribeck N, Deatherage DE, Blanchard JL, Dasgupta A, et al. Tempo and mode of genome evolution in a 50,000-generation experiment. Nature. 2016;536(7615):165–70. https://doi.org/10.1038/nature18959.
Thompson JD, Ou J, Lee N, Shin K, Cigliola V, Song L, et al. Identification and requirements of enhancers that direct gene expression during zebrafish fin regeneration. Development. 2020;147(14):dev191262. https://doi.org/10.1242/dev.191262.10.1242/dev.191262.
Tian Y, Smith-Bolton RK. Regulation of growth and cell fate during tissue regeneration by the two SWI/SNF chromatin-remodeling complexes of Drosophila. Genetics. 2021;217(1):1–16. https://doi.org/10.1093/genetics/iyaa028.
Truong DM, Boeke JD. Resetting the Yeast Epigenome with Human Nucleosomes. Cell. 2017;171(7):1508-19e13. https://doi.org/10.1016/j.cell.2017.10.043.
Turner BM. Defining an epigenetic code. Nat Cell Biol. 2007;9(1):2–6. https://doi.org/10.1038/ncb0107-2.
Tyagi M, Imam N, Verma K, Patel AK. Chromatin remodelers: We are the drivers!! Nucleus. 2016;7(4):388–404. https://doi.org/10.1080/19491034.2016.1211217.
Ushiki A, Zhang Y, Xiong C, Zhao J, Georgakopoulos-Soares I, Kane L, et al. Deletion of CTCF sites in the SHH locus alters enhancer-promoter interactions and leads to acheiropodia. Nat Commun. 2021;12(1):2282. https://doi.org/10.1038/s41467-021-22470-z.
Valencia AM, Kadoch C. Chromatin regulatory mechanisms and therapeutic opportunities in cancer. Nat Cell Biol. 2019;21(2):152–61. https://doi.org/10.1038/s41556-018-0258-1.
Valenzano DR, Benayoun BA, Singh PP, Zhang E, Etter PD, Hu C-K, et al. The African turquoise killifish genome provides insights into evolution and genetic architecture of lifespan. Cell. 2015;163(6):1539–54. https://doi.org/10.1016/j.cell.2015.11.008.
Van de Peer Y, Maere S, Meyer A. The evolutionary significance of ancient genome duplications. Nat Rev Genet. 2009;10(10):725–32. https://doi.org/10.1038/nrg2600.
Van Driel R, Fransz PF, Verschure PJ. The eukaryotic genome: a system regulated at different hierarchical levels. J Cell Sci. 2003;116(20):4067–75. https://doi.org/10.1242/jcs.00779.
van Steensel B, Furlong EEM. The role of transcription in shaping the spatial organization of the genome. Nat Rev Mol Cell Biol. 2019;20(6):327–37. https://doi.org/10.1038/s41580-019-0114-6.
Vierbuchen T, Ling E, Cowley CJ, Couch CH, Wang X, Harmin DA, et al. AP-1 transcription factors and the BAF complex mediate signal-dependent enhancer selection. Mol Cell. 2017;68(6):1067-82e12. https://doi.org/10.1016/j.molcel.2017.11.026.10.1016/j.molcel.2017.11.026.
Vitulo N, Dalla Valle L, Skobo T, Valle G, Alibardi L. Transcriptome analysis of the regenerating tail vs. the scarring limb in lizard reveals pathways leading to successful vs. unsuccessful organ regeneration in amniotes. Dev Dyn. 2017;246(2):116–34. https://doi.org/10.1002/dvdy.24474.
Vizcaya-Molina E, Klein CC, Serras F, Mishra RK, Guigo R, Corominas M. Damage-responsive elements in Drosophila regeneration. Genome Res. 2018;28(12):1852–66. https://doi.org/10.1101/gr.233098.117.
Vos SM. Understanding transcription across scales: From base pairs to chromosomes. Mol Cell. 2021;81(8):1601–16. https://doi.org/10.1016/j.molcel.2021.03.002.
Wang S, Miller SR, Ober EA, Sadler KC. Making it new again: insight into liver development, regeneration, and disease from zebrafish research. Curr Top Dev Biol. 2017;124:161–95. https://doi.org/10.1016/bs.ctdb.2016.11.012.
Wang M, Wang P, Lin M, Ye Z, Li G, Tu L, et al. Evolutionary dynamics of 3D genome architecture following polyploidization in cotton. Nat Plants. 2018;4(2):90–7. https://doi.org/10.1038/s41477-017-0096-3.
Wang AW, Wang YJ, Zahm AM, Morgan AR, Wangensteen KJ, Kaestner KH. The Dynamic Chromatin Architecture of the Regenerating Liver. Cell Mol Gastroenterol Hepatol. 2020a;9(1):121–43. https://doi.org/10.1016/j.jcmgh.2019.09.006.
Wang J, Wang J, Yang L, Zhao C, Wu LN, Xu L, et al. CTCF-mediated chromatin looping in EGR2 regulation and SUZ12 recruitment critical for peripheral myelination and repair. Nat Commun. 2020b;11(1):4133. https://doi.org/10.1038/s41467-020-17955-2.
Wang W, Hu CK, Zeng A, Alegre D, Hu D, Gotting K, et al. Changes in regeneration-responsive enhancers shape regenerative capacities in vertebrates. Science. 2020;369(6508):eaaz3090. https://doi.org/10.1126/science.aaz3090.
Wang Z, Cui M, Shah AM, Tan W, Liu N, Bassel-Duby R, et al. Cell-type-specific gene regulatory networks underlying murine neonatal heart regeneration at single-cell resolution. Cell reports. 2020;33(10):108472. https://doi.org/10.1016/j.celrep.2020.108472.
Wang X, Guo H, Yu F, Zhang H, Peng Y, Wang C, et al. Dataset for transcriptomic, H3K9ac and H3K9me3 profiles during cardiac regeneration. Data Brief. 2022;45:108569. https://doi.org/10.1016/j.dib.2022.108569.
Wei X, Li H, Guo Y, Zhao X, Liu Y, Zou X, et al. An ATAC-seq Dataset Uncovers the Regulatory Landscape During Axolotl Limb Regeneration. Front Cell Dev Biol. 2021;9:651145. https://doi.org/10.3389/fcell.2021.651145.
Wittkopp PJ, Kalay G. Cis-regulatory elements: molecular mechanisms and evolutionary processes underlying divergence. Nat Rev Genet. 2012;13(1):59–69. https://doi.org/10.1038/nrg3095.
Woodcock CL, Ghosh RP. Chromatin higher-order structure and dynamics. Cold Spring Harb Perspect Biol. 2010;2(5):a000596. https://doi.org/10.1101/cshperspect.a000596.
Woods IG, Kelly PD, Chu F, Ngo-Hazelett P, Yan YL, Huang H, et al. A comparative map of the zebrafish genome. Genome Res. 2000;10(12):1903–14. https://doi.org/10.1101/gr.10.12.1903.
Xiao C, Gao L, Hou Y, Xu C, Chang N, Wang F, et al. Chromatin-remodelling factor Brg1 regulates myocardial proliferation and regeneration in zebrafish. Nat Commun. 2016;7:13787. https://doi.org/10.1038/ncomms13787.
Yahalom-Ronen Y, Rajchman D, Sarig R, Geiger B, Tzahor E. Reduced matrix rigidity promotes neonatal cardiomyocyte dedifferentiation, proliferation and clonal expansion. Elife. 2015;4:e07455. https://doi.org/10.7554/eLife.07455.
Yakushiji N, Suzuki M, Satoh A, Sagai T, Shiroishi T, Kobayashi H, et al. Correlation between Shh expression and DNA methylation status of the limb-specific Shh enhancer region during limb regeneration in amphibians. Dev Biol. 2007;312(1):171–82. https://doi.org/10.1016/j.ydbio.2007.09.022.
Yan R, Cigliola V, Oonk KA, Petrover Z, DeLuca S, Wolfson DW, et al. An enhancer-based gene-therapy strategy for spatiotemporal control of cargoes during tissue repair. Cell Stem Cell. 2023;30(1):96–1116. https://doi.org/10.1016/j.stem.2022.11.012.10.1016/j.stem.2022.11.012.
Yang K, Kang J. Tissue regeneration enhancer elements: a way to unlock endogenous healing power. Dev Dyn. 2019;248(1):34–42. https://doi.org/10.1002/dvdy.24676.
Yokoyama H. Initiation of limb regeneration: the critical steps for regenerative capacity. Dev Growth Differ. 2008;50(1):13–22. https://doi.org/10.1111/j.1440-169X.2007.00973.x.
Zakany J, Kmita M, Duboule D. A dual role for Hox genes in limb anterior-posterior asymmetry. Science. 2004;304(5677):1669–72. https://doi.org/10.1126/science.1096049.
Zheng H, Xie W. The role of 3D genome organization in development and cell differentiation. Nat Rev Mol Cell Biol. 2019;20(9):535–50. https://doi.org/10.1038/s41580-019-0132-4.
Zuin J, Dixon JR, van der Reijden MI, Ye Z, Kolovos P, Brouwer RW, et al. Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc Natl Acad Sci U S A. 2014;111(3):996–1001. https://doi.org/10.1073/pnas.1317788111.
Acknowledgements
We thank all the anonymous reviewers for insightful comments. We also apologize that we cannot cite all published work in this field due to the limited length of the manuscript.
Funding
This work was supported by National Natural Science Foundation of China (W.W., 32270891).
Author information
Authors and Affiliations
Contributions
XH. Jia and W. Wang wrote the manuscript. WF. Lin summarized the literatures and generated the Table 1. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Jia, X., Lin, W. & Wang, W. Regulation of chromatin organization during animal regeneration. Cell Regen 12, 19 (2023). https://doi.org/10.1186/s13619-023-00162-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13619-023-00162-x