Abstract
Vaccines targeting enterohaemorrhagic Escherichia coli (EHEC) O157:H7 shedding in cattle are only partially protective. The correlates of protection of these vaccines are unknown, but it is probable that they reduce bacterial adherence at the mucosal surface via the induction of blocking antibodies. Recent studies have indicated a role for cellular immunity in cattle during colonisation, providing an impetus to understand the bacterial epitopes recognised during this response. This study mapped the epitopes of 16 EHEC O157:H7 proteins recognised by rectal lymph node CD4+ T-cells from calves colonised with Shiga toxin producing EHEC O157:H7 strains. 20 CD4+ T-cell epitopes specific to E. coli from 7 of the proteins were identified. The highly conserved N-terminal region of Intimin, including the signal peptide, was consistently recognised by mucosal CD4+ T-cell populations from multiple animals of different major histocompatibility complex class II haplotypes. These T-cell epitopes are missing from many Intimin constructs used in published vaccine trials, but are relatively conserved across a range of EHEC serotypes, offering the potential to develop cross protective vaccines. Antibodies recognising H7 flagellin have been consistently identified in colonised calves; however CD4+ T-cell epitopes from H7 flagellin were not identified in this study, suggesting that H7 flagellin may act as a T-cell independent antigen. This is the first time that the epitopes recognised by CD4+ T-cells following colonisation with an attaching and effacing pathogen have been characterised in any species. The findings have implications for the design of antigens used in the next generation of EHEC O157:H7 vaccines.
Similar content being viewed by others
Introduction
Enterohaemorrhagic Escherichia coli (EHEC) O157:H7 causes diarrhoea and potentially fatal renal failure in humans as a result of Shiga toxin (Stx) activity [1]. Cattle are the primary reservoir for EHEC O157:H7 [2] and this has led to the development of two licensed vaccines aimed at reducing EHEC O157 carriage in cattle. Econiche (Econiche Corp, Belleville, Canada), is manufactured from culture supernatants containing type III secreted proteins (T3SPs), whilst the other is manufactured from cell membrane extracts (Epitopix, Willmar, USA) prepared from iron restricted cultures. Both vaccines are only partially protective [3] and the correlates of protection have not yet been determined.
The primary site of colonisation in cattle is the terminal rectum where the bacteria form attaching and effacing (A/E) lesions on the apical epithelial surface [4]. We have recently shown that CD4+ T-cells infiltrate the rectal mucosa during experimental colonisation of cattle with EHEC O157:H7 and that CD4+ T-cells isolated from the rectal lymph nodes of colonised calves proliferate in response to T3SPs [5]. Furthermore, transcriptional profiling of the rectal mucosa during colonisation reveals a bias towards a T-helper type 1 (TH1) response, with colonisation inducing increased levels of interferon-gamma (IFN-γ) and T-bet transcripts within rectal mucosal tissues [5]. This suggests that cellular immunity may play an important role in controlling EHEC O157:H7 in cattle, particularly as cattle clear EHEC O157:H7 despite only generating low and highly variable mucosal antibody titres to key bacterial antigens [6]. Thus we hypothesise that while antibody production following vaccination may block binding to the epithelium, as suggested by passive immunisation studies [7], vaccines that induce a cellular response may be more effective in clearance once bacteria have formed A/E lesions with epithelial cells.
Given the importance of CD4+ T-cells in coordinating humoral and cellular immunity, an understanding of the epitopes that drive this response is essential to the design of protective vaccines, specifically with respect to establishing a CD4+ T-cell memory pool that can respond to epitopes presented during natural colonisation. Additionally, as CD4+ T-cells recognise linear epitopes presented on major histocompatibility complex (MHC) Class II molecules, studying the epitope repertoire recognised following colonisation allows basic bioinformatics tools to be used to compare these epitopes to the sequences of other EHEC serotypes of public health concern (O26, O111, O103, O121, O45 and O145). Epitopes that are conserved between these serotypes may provide cross-protection between EHEC serotypes.
The aim of this study was to characterise the epitopes recognised by mucosal CD4+ T-cells following experimental colonisation with Stx producing EHEC O157:H7. To achieve this, we measured CD4+ T-cell responses to overlapping peptides from 16 EHEC proteins that have previously been shown to play a role in colonisation or immunity to EHEC O157:H7 in cattle and/or have been previously used in experimental EHEC vaccines.
Materials and methods
The experimental approach is summarised in Figure 1A. CD4+ T-cells from the rectal lymph node (RLN) of experimentally colonised calves were expanded using an approach adapted from [8]. Autologous APCs were generated by immortalisation of peripheral blood mononuclear cells (PBMC) following Theileria annulata infection. These cells express MHCII and efficiently present exogenous antigen to CD4+ T-cells [9]. Cryopreserved PBMC and RLN cells were available from eight calves experimentally colonised with EHEC O157:H7 in a previous study [5], which was performed at the Moredun Research Institute (MRI) in accordance with the UK. Animals (Scientific Procedures) Act, 1986 under Home Office license 60/3179. The study was approved by the MRI Animal Experiments and Ethical Review Committee. All cultures were at 37 °C in a humidified 5% CO2 atmosphere.
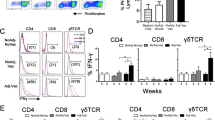
Experimental approach. A Graphical representation of APC and CD4+ T-cell line generation. B, C Immunofluorescent staining of T. annulata immortalised PBMC. Nuclei stained using DAPI (blue) and macroschizonts stained using monoclonal antibody 1C12 (green). D Flow cytometric analysis following staining of T. annulata immortalised PBMC for MHCII, CD80 and CD86. Filled histograms denote stained samples, open histograms denote the no primary control.
Peptide library
The length of peptides presented by MHCII in cattle has not been defined. In humans, the optimum MHCII binding affinity occurs with peptides 18–20 amino acids in length [10]. Lyophilised N-terminus amine, C-terminus glycine acid 18mer peptides overlapping by 12 amino acids were supplied by JPT Peptide Technologies (Berlin, Germany) for 16 bacterial proteins (see Additional file 1). 330 µM stock solutions were prepared in 5% dimethyl sulfoxide (DMSO) in phosphate buffered saline (PBS). Sequential peptides from each protein were pooled in groups of 10 (details in Additional file 2). Theileria parva peptides (Additional file 3) were prepared as a negative control to the same final concentration in 5% DMSO in PBS.
APC line generation
1 × 107 PBMC were resuscitated and infected with Theileria annulata sporozoites in lymphocyte culture medium (LCM) containing: Roswell Park Memorial Institute (RPMI) 1640 medium (Gibco), 10% heat inactivated (HI) fetal calf serum (FCS), 200 mM l-glutamine, 100 IU/mL penicillin and 100 µg/mL streptomycin (Invitrogen). Cultures were incubated in 24 well plates and inspected daily. Once growing rapidly, cells were transferred to flasks. APC lines were considered established after 2 weeks of culture.
Flow cytometric analysis of APC lines
Cells were incubated with anti-bovine CD80 (ILA159 [11]), CD86 (ILA190 [11]) or MHCII (CC158 [12]) antibodies, washed and incubated with goat anti-mouse IgG fluorescein isothiocyanate (FITC) conjugated antibody (Molecular Probes, Paisley, UK); unstained and no primary antibody controls were included. Incubations were for 30 min at 4 °C and 5% FCS/0.02% sodium azide/PBS was used for washes and antibody dilutions. Cells were analysed using a FACS Calibur flow cytometer (BD Biosciences, San Jose, USA) and FlowJo v10.0.7 (FlowJo, Ashland, USA).
Immunofluorescent microscopic assessment of APC lines
Cells were pelleted on to glass slides using the Cytospin system (Thermo Scientific, Paisley, UK) and fixed in acetone. Slides were stained using the Sequenza Immunostaining Center (Thermo Scientific) using 50 µL reaction volumes with reagents diluted in PBS as follows: cells were incubated with anti-T. annulata p104 vb (1C12 [13]) followed by goat anti-mouse IgG FITC conjugated antibody (Molecular Probes) then 3 nM DAPI. Slides were mounted using Fluorescent Mounting Media (Agilent Technologies, Wokingham, UK) and examined using a DMLB Fluorescent Microscope (Leica, Wetzlar, Germany).
CD4+ T-cell line generation
1 × 107 RLN cells were resuscitated into LCM medium containing 1 µg/mL phytohaemagglutinin (PHA) (Thermo Scientific) and 100 IU/mL rhIL-2 (Novartis, Basel, Switzerland) and seeded into round bottomed 96 well plates at 35 000 cells/well. After 4 days, wells were pooled and cells stained using anti-bovine CD4 AlexaFluor 647 conjugated antibody (CC8, AbD Serotec, Oxford, UK) diluted in 0.5% bovine serum albumin (BSA) (Sigma) in PBS. CD4+ cells were sorted at 4 °C using an Aria IIIu FACS machine (BD Biosciences). Cells in LCM containing 100 IU/mL rhIL-2 were seeded into 48 well plates at 1 × 106 cells/well, incubated for 10 days.
Peptide stimulations
Fourteen days after PHA stimulation, 250 000 CD4+ T-cells/well in LCM were seeded into 96 well plates and 50 000 γ-irradiated (60 Gy) APCs in LCM added to each well. Test and control peptide pools were added to a final concentration of 3 µM per peptide; approximately 1 µg/well (average MW 1932 Da). One µg/mL PHA was used as a positive control. Plates were incubated for 3 days. Interferon-γ (IFN-γ) release into the supernatant was measured using a commercial bovine IFN-γ ELISA kit (Mabtech AB, Nacka Strand, Sweden) and a microplate reader (Tecan, Männedorf, Switzerland). Individual peptides from up to eight positive pools were then tested individually. The process was as per the pools, except that peptide TC4 (Additional file 3) was used as a negative control and the final concentration of DMSO was 0.05 versus 0.5% for the pools.
MHC II genotyping of calves
DNA was extracted from the APC lines using the DNeasy Blood and Tissue DNA extraction kit (Qiagen, Hilden, Germany). Exon 2 of the bovine MHC Class II DRB3 gene was amplified using GoTaq DNA Polymerase (Promega, Southampton, UK) as per the manufacturer’s instructions. The forward (F455: TATCCCGTCTCTGCAGCACATTTC) and reverse (R329: CACCCCCGCGCTCACCTCGCCGC) primers were a modification of those previously published for genotyping the ovine DRB1 locus [14]. All reactions yielded a single amplicon of the predicted size (301 bp) which was purified and sequenced. Alleles were assigned as detailed in [14].
Peptide alignments
Peptide alignments were performed using the translated BLAST (tblastn, National Center for Biotechnology Information (NCBI)) and exported for manual inspection in Excel v14.0.7128 (Microsoft, Seattle, USA). Where necessary, annotations were obtained for accession numbers using Batch Entrez (NCBI). R package seqinr was used to append annotations to tblastn query results. Intimin alignments were performed using Clustal Omega and formatted using MView (EBI).
Data analysis
R v3.1.0 and packages ggplot2, GGally, scales, grid and gridExtra was used for analysis and plots. IFN-γ release in response to non-reactive peptides was assumed to be normally distributed around the mean IFN-γ release from the negative control wells, with reactive peptides skewing the overall distribution of the data. The standard deviation was therefore calculated manually where the mean was the IFN-γ release from the negative control wells and the deviance from this mean calculated using the wells where IFN-γ release was less than this mean. Positive wells were defined as wells where IFN-γ release was >3 standard deviations from the mean of the negative control wells.
Results
APC cell line phenotypes
Autologous Theileria annulata infected cell lines were established for all eight animals to be utilised as APC. Subsets of lines were stained to confirm T. annulata infection (Figures 1B and C). All eight lines exhibited MHCII and CD80 expression (Figure 1D). CD86 expression was undetectable.
Peptide pool screening
Twelve proteins were chosen because they had been shown to be targets of the antibody response in cattle colonised with EHEC O157:H7 (Intimin, Tir, EspA, EspB, EspD, EspM2, NleA, TccP, FliC, FliD and Stx2a and Sxt2c B-subunits [15–18]) and would therefore be expected to contain CD4+ T-cell epitopes. A further four secreted effector proteins were selected (EspF, EspK, NleC and NleD) as our previous studies suggested that T3SP preparations containing increased secreted effector proteins but minimal translocon-associated proteins stimulate stronger lymphoproliferative responses in rectal lymph node cells compared to T3SP preparations predominantly consisting of translocon-associated proteins [5]. IFN-γ release was used as a readout as this study also demonstrated the presence of TH1 responses in colonised cattle, and rectal lymph node cells released IFN-γ following re-stimulation with EHEC O157 T3SPs.
The CD4+ T-cell line from animal 101 053 exhibited high background IFN-γ release (mean of 30 439 pg/mL in the negative control wells compared to 120 326 pg/mL in PHA controls; ~25% of the PHA positive control) and the CD4+ T-cell expansion for animal 101 398 was less efficient, yielding only enough cells to seed 110 000 cells/well. Consequently the assay lacked sensitivity and specificity respectively for these animals (Additional file 4) and they were excluded from subsequent analysis. Mean IFN-γ release in response to PHA by the lines from the other 6 animals ranged from 14 070 pg/mL to 42 682 pg/mL. Background IFN-γ release by the negative control wells ranged from 1.9 to 7.5% of the PHA positive control.
IFN-γ release by each cell line in response to the 84 peptide pools is shown in Figure 2, with positive pools inset next to each graph. Positive pools are also summarised for each animal in Table 1 alongside the DRB3 MHCII haplotypes. None of the animals were of the same haplotype, demonstrating the outbred nature of the calves. Thirty-six positive pools were identified, representing 10/16 proteins tested. Ten of these pools were from Intimin, seven from EspK and five from NleD. Lines from 5/6 animals demonstrated responsiveness to at least one Intimin and NleD pool and 4/6 to at least one EspK pool.
IFN-γ production by rectal CD4 + T-cells of EHEC O157 colonised calves in response to pooled bacterial peptides. CD4+ T-cell lines prepared from the rectal lymph nodes of animals challenged with EHEC O157:H7 were stimulated with pools of peptides from 16 EHEC O157:H7 proteins and subsequent IFN-γ release quantified by ELISA. Each panel represents results from a single animal. Peptide pools are ordered on the Y axis by the amount of IFN-γ produced. Peptide pool names are given for positive pools only. Solid lines denote the mean of two negative control wells. Dotted lines denote three standard deviations from the negative control as detailed in the "Materials and methods" section. Positive pools are shown in the inset next to each graph.
Peptide pool deconvolution
Where sufficient CD4+ T-cells were available, individual peptides were assessed from up to eight positive pools. Overall IFN-γ release was higher, likely due to the reduced DMSO concentration in each well. IFN-γ release in response to PHA ranged from 31 360 to 525 433 pg/mL. Negative control wells ranged from 0.1 to 10.8% of the PHA positive control. IFN-γ release in response to the individually deconvolved peptides is shown in Figure 3. From a total of five calves, 20 unique positive peptides were identified. Intimin was the most commonly recognised protein, with 7/20 positive peptides representing sequences from the highly conserved N-terminal region of Intimin (Figure 4).
IFN-γ production by rectal CD4 + T-cells of EHEC O157 colonised calves in response to individual bacterial peptides. CD4+ T-cell lines prepared from the rectal lymph nodes of animals challenged with EHEC O157:H7 were stimulated with individual peptides from the top eight positive pools for each animal in Table 1 and subsequent IFN-γ release quantified by ELISA. Each panel represents results from a single animal. Peptides are ordered on the Y axis by the amount of IFN-γ produced. Peptide names are given for positive peptides only. Solid lines denote the mean of four negative control wells. Dotted lines denote three standard deviations from the negative control as detailed in the "Materials and methods" section. Positive peptides are shown in the inset next to each graph.
Location and conservation of bovine CD4 + T-cell Intimin epitopes after experimental colonisation with EHEC O157:H7. A Schematic of EHEC Intimin-γ, labelled with structural domain coordinates (Uniprot accession: P43261). Location of CD4+ T-cell epitopes is indicated in red. B Sequence alignment of CD4+ T-cell epitopes against the parental EHEC O157:H7 Intimin-γ sequence (accession BAB37982.1) and an Intimin consensus sequence. Conservation of this consensus is indicated on a scale of (*)-0, where * = 100% identity, 9 is highly conserved, down to 0, which is not conserved. Structural domains shown in A are also highlighted in this alignment. The consensus sequence was generated by a MUSCLE [32] multiple sequence alignment of representative Intimin sequences from EHEC of public health concern: O157:H7 (γ, accession BAB37982.1); O145:H28 (γ, accession AHY73165.1); O26:H11 (β, accession BAI28422.1); O103:H2 (ε, accession BAI32332.1); O121:H19 (ε, accession KDV53260.1); O45:H2 (ε, accession EZA22403.1); O111:H- (τ, accession BAI37528.1) and a prototypical EPEC O127:H6 comparator (α, accession YP_002331401.1). Intimin types are indicated in brackets. Conservation levels of this consensus sequence were generated using the PRALINE consistency output [33].
A tblastn alignment of the amino acid sequence for each peptide was performed using the nr/nt database and 1714 hits with 100% coverage and 100% identity selected. After excluding synthetic sequences, the only alignments not attributed to E. coli were hits for Intimin from E. albertii and Shigella boydii and NleD and EspK from E. albertii.
To investigate the degree of conservation between EHEC serotypes, a tblastn alignment for each peptide using the 16 complete EHEC genomes was performed. The percent identity for each peptide is summarised in Table 2. Peptide identity was 100% across the six O157:H7 and four O145:H28 genomes, with the exception of the NleD peptides, where no alignments were returned for the O145:H28 genomes. NleD is encoded by cryptic prophage CP-933 K and a tblastn alignment of the Sakai protein sequence (accession NP308877.1) against the O145:H28, O103:H2, O111:H- and O26:H11 genomes yielded no hits. There were no hits for peptides EspD1.5 and Tir9.10 against the O103:H2, O111:H- and O26:H11 genomes. Tblastn alignments of the EspD and Tir Sakai protein sequences (accessions BAB37978.1 and BAB37984.1 respectively) against the genomes of these three serotypes resulted in single hits of 72–74% and 60–66% identity respectively, indicating a lower level of homology for these proteins. The alignments for the remaining peptides against the whole genome sequences of the O103:H2, O111:H- and O26:H11 strains varied from 72 to 100% identity. As anticipated, no hits were identified for any of the peptides in the three O104:H4 genomes, which do not express a T3SS [19].
Intimin alignments
As the most commonly recognised epitopes in colonised cattle were from the N-terminal region of Intimin, these sequences were aligned with the β, γ, ε and θ Intimin sequences from the six other major EHEC serotypes (O145:H28, O26:H1, O103:H2, O121:H19, O45:H2 and O111:H-) and α Intimin from enteropathogenic E. coli (EPEC) O127:H6 using Intimin from EHEC O157:H7 as a scaffold (Additional file 5). All 7 Intimin peptides originate from a region of the molecule that is highly conserved between EHEC serotypes. A tblastn alignment of Sakai Intimin (accession BAB37982.1) against all deposited O157:H7 sequences (excluding Sakai), identified 11/12 sequences with 100% identity and 1/12 sequences with 99% identity. The same alignment for Intimin from the O145:H28 strain RM12581 (accession AHY73165.1) for all deposited O145:H28 sequences (excluding RM12581) resulted in 3/3 sequences with 100% identity.
Discussion
A rational screening approach was undertaken to investigate CD4+ T-cell epitopes recognised by cattle following EHEC O157:H7 colonisation. T. annulata transformed PBMC were used as APCs. These lines have been shown to efficiently process and present peptides in vitro [9], although potential impacts of T. annulata transformation on antigen processing and presentation cannot be excluded. Positive peptides were not identified from all of the positive peptide pools identified in the initial screening experiment, probably as the frequency of peptide-specific CD4+ T-cells within these cell lines was too low for detection. Epitopes were identified from EspA, EspB, EspD, EspK, Intimin, NleD and Tir. Serological responses have been detected to all of these proteins with the exception of EspK and NleD. As these are injected effector proteins, it is possible that these epitopes are more efficient at driving cellular rather than humoral responses.
All six animals from which CD4 epitope data were analysed in this study demonstrated rising anti-H7 flagellin IgA and/or IgG1 serum titres during colonisation (data not shown) and H7 flagellin is known to induce strong antibody responses in colonised calves [20, 21]. The failure to identify a convincing CD4+ T-cell response against either FliC or FliD peptides suggest that anti-flagellin antibody production during E. coli O157 colonisation may largely occur independently of T-cell help as flagellins are known to activate B cells in a T-cell independent manner [22].
Intimin epitopes were identified in 5/6 animals, with all the peptides originating from the intracellular N-terminal region of the molecule, over regions that are identical between sequenced strains of the same serotype and highly conserved between different serotypes. Intimin contains an N-terminal signal peptide, which in EPEC has historically been thought to be cleaved following translocation of the protein into the periplasm [23]. Experimental evidence of signal peptide cleavage in either EPEC or EHEC has not been published, whilst recent models for the biogenesis and secretion of Intimin do not propose cleavage of the signal peptide [24]. The peptides identified in this study sit predominantly within the signal peptide and are adjacent to a LysM domain (Figure 4) which is thought to enable peptidoglycan tethering in the periplasmic space [25]. The Intimin1.6 peptide identified in this study spans the putative signal peptide cleavage site between residues 39 and 40. If the Intimin signal peptide is cleaved, this would suggest that antigen presenting cells in vivo are processing cytoplasmic Intimin.
Intimin is 934 amino acids long. Due to its size and technical challenges associated with expressing full-length Intimin, C-terminal Intimin fragments of varying sizes have been used in vaccine studies. The longest C899 [7] and shortest C280 [26] fragments have been used in studies of passive transfer of immunity in pigs and cattle respectively where memory recall responses were not relevant. C531 [27] and C280 [28] fragments have been trialled in cattle and a C280 [29] fragment has also been trialled in goats as part of a fusion protein. These shorter Intimin fragments excluded all the epitopes identified in this study. This study suggests that the N-terminus of Intimin is an important immunological target and whilst C-terminal fragments may result in protective antibody titres, the use of full length Intimin or shorter constructs that include the N-terminal amino acid sequences may be required to induce a memory response that can be recalled during colonisation.
It is interesting to note that the Intimin epitopes identified in this study sit in a region of the molecule that is unlikely to be accessible to host antibodies. Mathematical modelling has identified signal peptides and transmembrane domains as molecule regions that are particularly dense in CD8+ T-cell epitopes [30]. This study suggests that this may also be the case for CD4+ T-cell epitopes. Whether this is due to their inherent hydrophobicity or part of a regulatory mechanism to ensure that B cell responses only receive T-cell help if they are directed against epitopes that originate from a full length pathogen molecule is unclear.
Antibodies raised to the EHEC O157:H7 recombinant C-terminal fragment Intimin-531 cross react with whole cell preparations from six different EHEC serotypes: O111, O103:H2, O145, O26:H11, O26:H- and O26 [31], despite the variability in the extra-cellular domains between the different Intimin types represented by these strains. Taken together, both the CD4+ T-cell epitopes identified in this study and antibody epitopes identified in previous studies suggest that immunisation with Intimin has the potential to provide cross-protection against different EHEC serotypes.
In conclusion, this study demonstrates that CD4+ T-cells from cattle colonised with EHEC O157:H7 recognise and respond to peptide sequences from 7/16 EHEC O157:H7 proteins tested. The most commonly recognised molecule was the N-terminal region of Intimin, highlighting important considerations with respect to the use of C-terminal Intimin fragments in EHEC O157:H7 vaccine development. The failure to identify CD4+ T-cell responses against H7 flagellin raises questions as to the mechanism of anti-flagellin antibody generation in cattle by EHEC O157:H7.
References
Karmali MA (2004) Infection by Shiga toxin-producing Escherichia coli: an overview. Mol Biotechnol 26:117–122
Armstrong GL, Hollingsworth J, Morris JG Jr (1996) Emerging foodborne pathogens: Escherichia coli O157:H7 as a model of entry of a new pathogen into the food supply of the developed world. Epidemiol Rev 18:29–51
Snedeker KG, Campbell M, Sargeant JM (2012) A systematic review of vaccinations to reduce the shedding of Escherichia coli O157 in the faeces of domestic ruminants. Zoonoses Public Health 59:126–138
Naylor SN, Low JC, Besser TE, Mahajan A, Gunn GJ, Pearce MC, McKendrick IJ, Smith DG, Gally DL (2003) Lymphoid follicle-dense mucosa at the terminal rectum is the principal site of colonization of enterohemorrhagic Escherichia coli O157:H7 in the bovine host. Infect Immun 71:1505–1512
Corbishley A, Ahmad NI, Hughes K, Hutchings MR, McAteer SP, Connelley TK, Brown H, Gally DL, McNeilly TN (2014) Strain dependent cellular immune responses in cattle following Escherichia coli O157:H7 colonisation. Infect Immun 82:5117–5131
Bretschneider G, Berberov EM, Moxley RA (2008) Enteric mucosal antibodies to Escherichia coli O157:H7 in adult cattle. Vet Rec 163:218–219
Dean-Nystrom EA, Gansheroff LJ, Mills M, Moon HW, O’Brien AD (2002) Vaccination of pregnant dams with intimin(O157) protects suckling piglets from Escherichia coli O157:H7 infection. Infect Immun 70:2414–2418
Geiger R, Duhen T, Lanzavecchia A, Sallusto F (2009) Human naive and memory CD4+ T cell repertoires specific for naturally processed antigens analyzed using libraries of amplified T cells. J Exp Med 206:1525–1534
Glass EJ, Spooner RL (1990) Parasite-accessory cell interactions in theileriosis. Antigen presentation by Theileria annulata-infected macrophages and production of continuously growing antigen-presenting cell lines. Eur J Immunol 20:2491–2497
O’Brien C, Flower DR, Feighery C (2008) Peptide length significantly influences in vitro affinity for MHC class II molecules. Immunome Res 4:6
Glew EJ (2003) Differential effects of bovine viral diarrhoea virus on monocytes and dendritic cells. J Gen Virol 84:1771–1780
Howard CJ, Sopp P, Brownlie J, Kwong LS, Parsons KR, Taylor G (1997) Identification of two distinct populations of dendritic cells in afferent lymph that vary in their ability to stimulate T cells. J Immunol 159:5372–5382
Woods KL, Theiler R, Mühlemann M, Segiser A, Huber S, Ansari HR, Pain A, Dobbelaere DAE (2013) Recruitment of EB1, a master regulator of microtubule dynamics, to the surface of the Theileria annulata schizont. PLoS Pathog 9:e1003346
Ballingall KT, Tassi R (2010) Sequence-based genotyping of the sheep MHC class II DRB1 locus. Immunogenetics 62:31–39
Asper DJ, Karmali MA, Townsend H, Rogan D, Potter AA (2011) Serological response of Shiga toxin-producing Escherichia coli type III secreted proteins in sera from vaccinated rabbits, naturally infected cattle and humans. Clin Vaccine Immunol 18:1052–1057
Bretschneider G, Berberov EM, Moxley RA (2007) Isotype-specific antibody responses against Escherichia coli O157:H7 locus of enterocyte effacement proteins in adult beef cattle following experimental infection. Vet Immunol Immunopathol 118:229–238
Vilte DA, Larzabal M, Cataldi AA, Mercado EC (2008) Bovine colostrum contains immunoglobulin G antibodies against intimin, EspA, EspB and inhibits hemolytic activity mediated by the type three secretion system of attaching and effacing Escherichia coli. Clin Vaccine Immunol 15:1208–1213
Naylor SW, Flockhart A, Nart P, Smith DG, Huntley J, Gally DL, Low JC (2007) Shedding of Escherichia coli O157:H7 in calves is reduced by prior colonization with the homologous strain. Appl Env Microbiol 73:3765–3767
Pierard D, De Greve H, Haesebrouck F, Mainil J (2012) O157:H7 and O104:H4 Vero/Shiga toxin-producing Escherichia coli outbreaks: respective role of cattle and humans. Vet Res 43:13
McNeilly TN, Naylor SW, Mitchell MC, McAteer SP, Mahajan A, Smith DG, Gally DL, Low JC, Huntley JF (2007) Simple methods for measurement of bovine mucosal antibody responses in vivo. Vet Immunol Immunopathol 118:160–167
McNeilly TN, Naylor SW, Mahajan A, Mitchell MC, McAteer SP, Deane D, Smith DG, Low JC, Gally DL, Huntley JF (2008) Escherichia coli O157:H7 colonization in cattle following systemic and mucosal immunization with purified H7 flagellin. Infect Immun 76:2594–2602
Pike BL, Alderson MR, Nossal GJ (1987) T-independent activation of single B cells: an orderly analysis of overlapping stages in the activation pathway. Immunol Rev 99:119–152
Gomez-Duarte O, Kaper J (1995) A plasmid-encoded regulatory region activates chromosomal eaeA expression in enteropathogenic Escherichia coli. Infect Immun 63:1767–1776
Leo JC, Oberhettinger P, Schütz M, Linke D (2015) The inverse autotransporter family: intimin, invasin and related proteins. Int J Med Microbiol 305:276–282
Bodelón G, Marín E, Fernández LA (2009) Role of periplasmic chaperones and BamA (YaeT/Omp85) in folding and secretion of intimin from enteropathogenic Escherichia coli strains. J Bacteriol 191:5169–5179
Rabinovitz BC, Vilte DA, Larzábal M, Abdala A, Galarza R, Zotta E, Ibarra C, Mercado EC, Cataldi A (2014) Physiopathological effects of Escherichia coli O157:H7 inoculation in weaned calves fed with colostrum containing antibodies to EspB and Intimin. Vaccine 32:3823–3829
McNeilly TN, Mitchell MC, Rosser T, McAteer SP, Low JC, Smith DG, Huntley JF, Mahajan A, Gally DL (2010) Immunization of cattle with a combination of purified intimin-531, EspA and Tir significantly reduces shedding of Escherichia coli O157:H7 following oral challenge. Vaccine 28:1422–1428
Vilte DA, Larzábal M, Garbaccio S, Gammella M, Rabinovitz BC, Elizondo AM, Cantet RJC, Delgado F, Meikle V, Cataldi A, Mercado EC (2011) Reduced faecal shedding of Escherichia coli O157:H7 in cattle following systemic vaccination with γ-intimin C280 and EspB proteins. Vaccine 29:3962–3968
Zhang X, Yu Z, Zhang S, He K (2014) Immunization with H7-HCP-tir-intimin significantly reduces colonization and shedding of Escherichia coli O157:H7 in goats. PLoS One 9:e91632
Kovjazin R, Volovitz I, Daon Y, Vider-Shalit T, Azran R, Tsaban L, Carmon L, Louzoun Y (2011) Signal peptides and trans-membrane regions are broadly immunogenic and have high CD8+ T cell epitope densities: Implications for vaccine development. Mol Immunol 48:1009–1018
McNeilly TN, Mitchell MC, Corbishley A, Nath M, Simmonds H, McAteer SP, Mahajan A, Low JC, Smith DG, Huntley JF, Gally DL (2015) Optimizing the protection of cattle against Escherichia coli O157:H7 colonization through immunization with different combinations of H7 flagellin, Tir, intimin-531 or EspA. PLoS One 10:e0128391
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Simossis VA, Heringa J (2005) PRALINE: a multiple sequence alignment toolbox that integrates homology-extended and secondary structure information. Nucleic Acids Res 33:289–294
Competing interests
AC received financial support from Bioniche Life Sciences Inc., who played no role in the study design, performance of experimental work, analysis and interpretation of the data, nor the writing of the report. Approval was sought from Bioniche Life Sciences Inc. prior to submission of the manuscript for publication.
Authors’ contributions
AC, TM and DG conceived and designed the experiments. AC, TM, TC and AB performed the experimental work. Structural modelling of Intimin was performed by EW. Input into the MHCII typing was provided by KB. All authors read and approved the final manuscript.
Acknowledgements
MRI receives funding from the Scottish Government. DLG is supported at the Roslin Institute by BBSRC institute programme funding (BB/J004227/1). TM and KB receive funding from the Scottish Government. DLG and TM were also supported by BBSRC project grant BB/I011625/1. AC was supported by a Bioniche Life Sciences Inc. PhD studentship and BBSRC funding. AB is supported by a Food Standards Agency Grant FS101055.
Author information
Authors and Affiliations
Corresponding author
Additional files
13567_2016_374_MOESM1_ESM.xlsx
Additional file 1. Details of EHEC O157 protein sequences for epitope mapping. Names and accession numbers of 16 EHEC O157 protein sequences used to synthesise peptides for T-cell epitope mapping.
13567_2016_374_MOESM2_ESM.xlsx
Additional file 2. Pooling strategy for EHEC O157 peptides. Details of peptide pools used to stimulate CD4+ T-cell lines from rectal lymph nodes of EHEC O157 challenged calves.
13567_2016_374_MOESM3_ESM.xlsx
Additional file 3. Theileria parva peptide sequences. Amino acid sequences of T. parva peptides used as negative controls for CD4+ T-cell stimulation assays.
13567_2016_374_MOESM4_ESM.pdf
Additional file 4. IFN-γ production by CD4 + T-cells from calves 101398 and 101053 in response to pooled EHEC O157 peptides. Graphs represent IFN-γ released from CD4+ T-cell lines generated from the rectal lymph nodes of calves 101398 and 101053 in response to EHEC O157 peptide stimulation. Peptide pools are ordered on the Y axis by the amount of IFN-γ produced. Solid lines denote the mean of two negative control wells. Dotted lines denote three standard deviations from the negative control.
13567_2016_374_MOESM5_ESM.pdf
Additional file 5. Sequence alignment of Intimin epitopes against Intimin sequences from non-O157 EHEC serotypes. Alignment of Intimin CD4+ T-cell epitope sequences with representative Intimin sequences from EHEC serotypes O145, O127, O26, O103, O121, O45 and O111. Percentage values indicate % similarity to the EHEC O157:H7 reference sequence.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Corbishley, A., Connelley, T.K., Wolfson, E.B. et al. Identification of epitopes recognised by mucosal CD4+ T-cell populations from cattle experimentally colonised with Escherichia coli O157:H7. Vet Res 47, 90 (2016). https://doi.org/10.1186/s13567-016-0374-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13567-016-0374-5