Abstract
Background
Accurate diagnosis of urogenital schistosomiasis is vital for surveillance and control programmes. While a number of diagnostic techniques are available there is a need for simple, rapid and highly sensitive point-of-need (PON) tests in areas where infection prevalence and intensity are low. Recombinase Polymerase Amplification (RPA) is a sensitive isothermal molecular diagnostic technology that is rapid, portable and has been used at the PON for several pathogens.
Results
A real time fluorescence RPA assay (RT-ShDra1-RPA) targeting the Schistosoma haematobium Dra1 genomic repeat region was developed and was able to detect 1 fg of S. haematobium gDNA. Results were obtained within 10 minutes using a small portable battery powered tube scanner device that incubated reactions at 40 °C, whilst detecting DNA amplification and fluorescence over time. The assay’s performance was evaluated using 20 urine samples, with varying S. haematobium egg counts, from school children from Pemba Island, Zanzibar Archipelago, Tanzania. Prior to RPA analysis, samples were prepared using a quick crude field DNA extraction method, the Speed Extract Kit (Qiagen, Manchester, UK). Positive assay results were obtained from urine samples with egg counts of 1–926 eggs/10 ml, except for two samples, which had inconclusive results. These two samples had egg counts of two and three eggs/10 ml of urine.
Conclusions
The RT-ShDra1-RPA assay proved robust for S. haematobium gDNA detection and was able to amplify and detect S. haematobium DNA in urine samples from infected patients. The assay’s speed and portability, together with the use of crude sample preparation methods, could advance the rapid molecular diagnosis of urogenital schistosomiasis at the PON within endemic countries.
Similar content being viewed by others
Background
Schistosomiasis is a neglected tropical disease (NTD) caused by parasitic trematodes called schistosomes. Schistosoma haematobium is one of three main human-infecting schistosome species; with > 110 million cases of urogenital schistosomiasis, causing haematuria, bladder wall pathology, hydronephrosis leading to severe kidney disease [1, 2], and bladder cancer [3], with female and male genital schistosomiasis also being linked to infertility and HIV transmission [4]. It is the most commonly occurring schistosome species and is transmitted by various intermediate snail hosts of the genus Bulinus throughout Africa, parts of the Middle East, Madagascar and the Indian Ocean Islands [5] with a recent outbreak on the Mediterranean island of Corsica [6].
Sensitive and specific diagnostic tests are critical for the development, implementation and success of schistosomiasis control and elimination programmes [7,8,9,10,11]. They not only enable the accurate diagnosis and treatment of individual patients but also support the monitoring of control interventions [11,12,13,14,15,16,17]. Moreover, as a control programme achieves success, low infection intensity is common within the population, with a high proportion of those infected excreting low numbers of schistosome eggs that may escape detection by routine methods, namely urine filtration and haematuria detection strips [16, 17]. This increases the need for diagnostic sensitivity and specificity to prevent false negative diagnoses [7, 17]. The recently developed, and very promising, circulating anodic antigen (CAA) based test offers high sensitivity and is currently being optimized and evaluated for PON testing [18, 19].
Molecular diagnostics can be highly sensitive and specific [15]. Polymerase chain reaction (PCR) and quantitative PCR (qPCR) methods, that target and amplify schistosome DNA from urine and stool samples, have been shown to be sensitive (0.01–10 fg) and specific [20,21,22,23]. However, these methods are costly, take time, require a significant laboratory infrastructure and training, hampering their current use in endemic field settings [12, 15, 24]. Loop-mediated isothermal amplification (LAMP), overcomes some of these obstacles and has been successfully used in the field to diagnose human S. haematobium infections [25, 26].
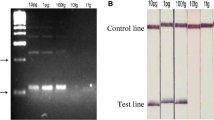
Recombinase polymerase amplification (RPA) is an isothermal DNA amplification technology that can be performed in the field due to its low resource requirements. Reactions are rapid and take place at a low constant temperature using small portable devices and lyophilized reagents. DNA amplification can be detected either by gel electrophoresis, oligo chromatographic lateral flow (LF) strips or real time fluorescence, offering detection flexibility of use in endemic field settings [27,28,29,30]. A lateral flow RPA assay targeting the Dra1 repeat region (Dra1 LF-RPA) has already been developed for S. haematobium [31], and its specificity tested against other urinary pathogens. Here, we advance this research with the first laboratory development and testing of a novel fluorescence real time (RT) Dra1-RPA assay for S. haematobium (RT-ShDra1-RPA).
Methods
Schistosoma haematobium template DNA
For the assay development S. haematobium adult worm genomic DNA (gDNA) originating from the Zanzibar island Unguja was provided by the Schistosomiasis Collection at the Natural History Museum (SCAN) [32]. DNA was quantified using a Qubit 2.0 Fluorimeter (Invitrogen, California, USA) and diluted to a working concentration of 1 ng/µl in ddH20.
RPA assay development
RT-ShDra1-RPA primer design
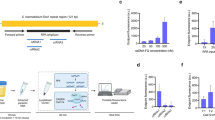
The Dra1 lateral flow RPA primers and internal probe (Dra1 LF-RPA) designed by Rosser et al. [31] were further adapted for RT-ShDra1-RPA amplification and detection. The internal probe was modified for the RT fluorescence-based detection as shown in Table 1 and Fig 1. RPA reactions were performed using the TwistAmp exo kit (TwistDX, UK) in 50 µl reactions containing 29.5 μl rehydration buffer, 2.1 μl of each of the forward and the reverse primers (10 pmol), 0.6 μl of the internal lateral flow probe (10 pmol), 12.2 μl of ddH2o and the DNA template (1 ng), which were added to the lyophilized RPA pellet. The reactions were initiated by the addition of 2.5 μl (280 mM) of magnesium acetate. The reactions were run at 40 °C for 20 minutes in an Axxin T-16 isothermal device (T-16 ISO) (http://www.axxin.com/Molecular-T16), which measures the increase in fluorescence, due to DNA amplification, over time. The reactions were manually mixed after 4 minutes.
RT-ShDra1-RPA sensitivity testing
Sensitivity was determined by running the RPA assay, as described above, using dilutions (1 ng, 1 pg, 1 fg and 0.5 fg) of the S. haematobium gDNA. Negative (no template) and positive (gDNA 1 ng) controls were run with the reactions.
Pilot urine testing
As part of the Zanzibar Elimination of Schistosomiasis Transmission (ZEST) project (2011–2017) [33], 1.5 ml aliquots of urine samples from schoolchildren that participated in the annual surveys, were frozen at −20 °C, and kept at the Public Health Laboratory-Ivo de Carneri (PHL-IdC) on Pemba Island, Zanzibar. All urine samples included in the study presented here were collected in 2013 and were positive for S. haematobium eggs, identified by urine filtration (10 ml) during the ZEST parasitological surveys [33]. Twenty egg-positive urine samples were selected with a range of egg counts classified as very low, low, medium and high (Table 2). The samples were randomly selected from multiple Shehias, so as not to introduce any geographical biases, but they were stratified by egg count.
At PHL-IdC, urine samples were defrosted, mixed and a 100 μl aliquot taken from each sample for DNA extraction. DNA was extracted, in country at PHL, from individual samples using the Qiagen Speed Extract kit (Qiagen). This is a fast, field-friendly and low-cost method for crude extraction of DNA using basic equipment. The protocol followed the manufacturerʼs recommendation with slight modification. All reagents were supplied within the Qiagen Speed Extract kit (Qiagen). 200 μl of EN buffer and 15 μl of the magnet bead mix was added to each urine sample, which was then mixed and incubated at room temperature for 3 min before being placed on a magnetic separation rack (New England Biolabs, Massachusetts, USA) for 1 min. During this time, DNA binds to the magnetic beads forming a pellet, allowing the supernatant to be removed and the pellet to be re-suspended in 100 μl of SL buffer to release the DNA from the beads. Samples were then heated at 95 °C for 5 min before returning to the magnetic rack for 1 min to pellet the magnetic beads. The supernatant, now containing extracted DNA, was removed and stored at room temperature. DNA samples were transported, at ambient temperature, to the Natural History Museum, London, UK, for RPA testing. Samples were analysed using the RT-ShDra1-RPA assay as described above using 5 μl of the DNA preparation in the 50 μl RPA reaction. A negative urine control (from a NHM laboratory staff member) and a positive control (donor urine spiked with S. haematobium gDNA) were also prepared using the Speed Extraction protocol and run alongside the urine samples from PHL-IdC.
Results
RT-ShDra1-RPA assay development and limit of detection
As shown in Fig. 2a, using 1ng of gDNA, the developed RT-ShDra1-RPA assay gave a positive fluorescent sigmoid curve which appeared after ~5 min of reaction time. There was a lower detection limit of 1 fg of gDNA with the final results obtained within 10 min.
RPA fluorescent curves for the S. haematobium gDNA dilutions and the urine samples tested from PHL-IDC, Pemba. a S. haematobium gDNA dilutions. b Urine samples U3-15. c Urine samples U1+2 (high egg counts) and U16-20 (very low egg counts). d Urine samples U1+2 (high egg counts) and negative donor urine (U-ve) and negative donor urine spiked with S. haematobium gDNA (USp). sDNA corresponds to a DNA standard positive control. −ve corresponds to negative controls
Urine sample examination
Fourteen urine samples were analysed from the medium and high egg count categories (Table 2), all of which gave a strong positive sigmoid fluorescent curve obtained within 8 min (Fig. 2b, c). As indicated in Table 2, for the six urine samples in the very low egg count category (1–10 eggs per 10 ml), the results were less easy to interpret. Four of the six samples produced strong positive sigmoid curves but one (U18, 2 eggs/10 ml) showed an inconclusive result (Fig. 2b, c) and one (U19, 3 eggs/10 ml) did not show any significant rise in fluorescence and was therefore considered negative. Positive and negative results were obtained from the spiked (USp) and non-spiked (U-ve) donor urine samples, respectively (Fig. 2d).
Discussion
Schistosomiasis control programs aim to reduce disease morbidity and significant impact has been observed in certain areas, including the Zanzibar islands [8, 34,35,36,37]. However, progress in control and eventual elimination, of schistosomiasis, needs high performance diagnostic tests that can detect low levels of infection [7, 17]. The molecular detection of S. haematobium DNA in urine has been shown to be a highly sensitive indicator of infection [15, 22, 23]. RPA is a DNA amplification and detection technology that is particularly suitable for PON use [24, 31,32,33,34,35,36,37], as all reagents are readily available lyophilized with the main RPA reagents provided in a single dried pellet, simplifying assay preparation, reducing contamination and allowing easy transportation and long-term storage at room temperature. Additionally, DNA amplification occurs at low constant temperatures between 25–40 °C and real time fluorescence amplification detection can be performed using small portable tube-scanner devices, with results obtained within 20 minutes.
Here, a real time fluorescence-based RPA assay was successfully developed to amplify the Dra1 DNA repeat region of S. haematobium (RT-ShDra1-RPA). The present RT-ShDra1-RPA lower limit of detection was 1 fg of S. haematobium gDNA, lower than the existing Dra1 end point PCR diagnostic assay, that has a detection limit of 10 fg of S. haematobium gDNA [38], and also lower than the detection limit of the developed lateral flow RPA Dra1 assay (LF-ShDra1-RPA) [31], that had a detection limit of 100 fg of gDNA. However, the Dra1 qPCR and LAMP assays have been reported to display higher sensitivities detecting as low as 1 fg and 0.1 fg of S. haematobium gDNA, respectively [15, 20,21,22,23, 25, 26]. Theoretically, RPA can detect a single copy of a DNA target thus there is scope for further assay optimization and development, particularly focusing upon different target regions, probe and primer combinations and concentrations to increase assay sensitivity.
RPA has been shown to offer degrees of tolerance to inhibitors found in urine, and works well on crudely prepared samples [30] reducing the need for the resources needed for sophisticated sample preparation and purification methods, further enhancing its feasibility for PON use. The RT-ShDra1-RPA developed here gave positive results when tested on crudely prepared urine samples from infected school children from the endemic Pemba Island, Zanzibar. The DNA preparations took ~15 minutes for 12 samples and required only portable low powered equipment. The assay produced strong positive results for samples that had medium to high egg counts (11–926 eggs per 10 ml), performing as well as microscopy but further analysis of samples with lower egg counts is warranted, particularly as this is the main application for this type of diagnostic. Results varied between samples that had between 1–10 eggs per 10 ml. Four among the 6 samples, within this egg count range, gave positive results but one was inconclusive and one was negative. This maybe due to there not being eggs present in the aliquot of urine analysed, false positive egg identification by microscopy and/or sample degradation. It is worth mentioning that two of the three urine filtration reads (each sample was read by three technicians) performed on these samples were recorded as egg-negative. Further testing of low egg count samples and egg-negative samples is now needed to further assess the performance of the developed assay. It is also necessary to determine if this method is detecting DNA from the S. haematobium eggs or Cell-Free-Parasite DNA (CFPD), which has been reported as a source of DNA in PCR and qPCR assays [15, 39]. RPA reactions are only semi-quantitative with the time to onset of amplification being longer for low and quicker for high DNA concentrations (Fig. 2a) and hence, could currently be used preferentially to identify infected individuals, but not to evaluate infection intensities.
The extraction method used for this study was chosen due to its field applicability and in particular its low resource needs, speed and simplicity. The SpeedXtract method works well for RPA assays as high quality DNA and purity is not needed. However, comparisons with other standard extraction methods would be beneficial as this may increase sensitivity. Particularly, as laboratories in endemic countries grow their infrastructure, more high resource sample preparation methods may become feasible.
Although the ShDra1-RPA assayʼs primer and probe combinations have proved negative with regard to their cross-reactivity with other pathogens found in urine samples and also other schistosome species, this has only been fully tested in the lateral flow form [31]. Here the developed RT-ShDra1-RPA assay is predicted to show the same specificity as the primer and probes used are identical except that they are modified for fluorescent detection. However, full validation on reference pathogen samples and also testing on a range of well-defined negative clinical samples is warranted to further validate the RT-ShDra1-RPA assayʼs clinical specificity.
Defining the gold standard for the diagnosis of schistosomiasis is not easy and there is no real current consensus. Currently egg detection in clinical samples is commonly used but this is known to not have the clinical sensitivity needed in low prevalence and intensity settings. Diagnosis by qPCR [20,21,22,23] is also used as a gold standard in some laboratories with CAA testing also coming on to the agenda however, comparisons between all tests are warranted. In this study the RT-ShDra1-RPA assay has been tested with regard to egg count in a small set of samples but the assay certainly needs testing on a wider range, and on a number of well-defined samples together with direct comparisons to qPCR and CAA [18, 19]. These comparisons are needed to define the diagnostic use case, for such an assay as RPA, in relation to its possible utility at different stages of schistosomiasis control [10,11,12,13]. Certainly, the characteristics of RPA could support diagnosis and possible test and treat scenarios at the elimination stage.
Conclusions
The developed RT-ShDra1-RPA assay offers an alternative S. haematobium DNA amplification system that has the potential use as a molecular diagnostic tool for urogenital schistosomiasis in endemic settings. Results can be rapidly obtained from crudely prepared samples using small portable devices requiring minimal infrastructure. Further development is needed to test its sensitivity and performance in low prevalence settings, where infection intensities and egg counts are mostly low, and in comparison to existing diagnostic tests.
Availability of data and materials
All data involved and arising from the study are included in the publication. Samples are available upon request.
Abbreviations
- RPA:
-
recombinase polymerase amplification
- PCR:
-
polymerase chain reaction
- qPCR:
-
quantitative polymerase chain reaction
- Sh:
-
S. haematobium
- CFPD:
-
cell-free parasite DNA
- gDNA:
-
genomic DNA
- U:
-
urine
- sDNA:
-
synthetic DNA standard
- -ve:
-
negative control
References
van der Werf MJ, de Vlas SJ, Brooker S, Looman CW, Nagelkerke NJ, Habbema JD, Engels D. Quantification of clinical morbidity associated with schistosome infection in sub-Saharan Africa. Acta Trop. 2003;86:125–39.
King CH, Dickman K, Tisch DJ. Reassessment of the cost of chronic helmintic infection: a meta-analysis of disability-related outcomes in endemic schistosomiasis. Lancet. 2005;365:1561–9.
Shiff C, Veltri R, Naples J, Quartey J, Anyan W, Marlow C, et al. Ultrasound verification of bladder damage is associated with known biomarkers of bladder cancer in adults chronically infected with Schistosoma haematobium in Ghana. Trans Roy Soc Trop Med Hyg. 2006;100:847–54.
Kjetland EF, Norseth HM, Taylor M, Lillebo K, Kleppa E, Holmen SD, et al. Classification of the lesions observed in female genital schistosomiasis. Int J Gynecol Obstet. 2014;127:227–8.
Rollinson D. A wake up call for urinary schistosomiasis: reconciling research effort and public health importance. Parasitology. 2009;136:1593–610.
Boissier J, Grech-Angelini S, Webster BL, Allienne JF, Huyse T, Mas Coma S, et al. Outbreak of urogenital schistosomiasis in Corsica (France): an epidemiological case study. Lancet Infect Dis. 2016;16:971–9.
McCarthy JS, Lustigman S, Yang GJ, Barakat RM, Garcia HH, Sripa B, et al. A research agenda for helminth diseases of humans: diagnostics for control and elimination programmes. PLoS Negl Trop Dis. 2012;6:e1601.
Rollinson D, Knopp S, Levitz S, Stothard JR, Tchuel Tchuente LA, Garba A, et al. Time to set the agenda for schistosomiasis elimination. Acta Trop. 2013;128:423–40.
Stothard JR, Stanton MC, Bustinduy AL, Sousa-Figueiredo JC, Van Dam GJ, Betson M, et al. Diagnostics for schistosomiasis in Africa and Arabia: a review of present options in control and future needs for elimination. Parasitology. 2014;141:1947–61.
Utzinger J, Becker SL, van Lieshout L, van Dam GJ, Knopp S. New diagnostic tools in schistosomiasis. Clin Microbiol Infect. 2015;21:529–42.
Bergquist R, Johansen MV, Utzinger J. Diagnostic dilemmas in helminthology: what tools to use and when? Trends Parasitol. 2009;25:151–6.
Knopp S, Becker SL, Ingram KJ, Keiser J, Utzinger J. Diagnosis and treatment of schistosomiasis in children in the era of intensified control. Expert Rev Anti Infect Ther. 2013;11:1237–58.
Cavalcanti MG, Silva LF, Peralta RH, Barreto MG, Peralta JM. Schistosomiasis in areas of low endemicity: a new era in diagnosis. Trends Parasitol. 2013;29:75–82.
Loc L, Hsieh MH. Diagnosing urogenital schistosomiasis: dealing with diminishing returns. Trends Parasitol. 2017;33:378–87.
Weerakoon K, Gordon C, McManus DP. DNA diagnostics for schistosomiasis control. Trop Med Infect Dis. 2018;3:81.
Knopp S, Ame SM, Hattendorf J, Ali SM, Khamis IS, Bakar F, et al. Urogenital schistosomiasis elimination in Zanzibar: accuracy of urine filtration and haematuria reagent strips for diagnosing light intensity Schistosoma haematobium infections. Parasit Vectors. 2018;11:552.
King CH, Bertsch D. Meta-analysis of urine heme dipstick diagnosis of Schistosoma haematobium infection, including low-prevalence and previously-treated populations. PLoS Negl Trop Dis. 2013;7:e2431.
Corstjens PL, Nyakundi RK, de Dood CJ, Kariuki TM, Ochola EA, Karanja DM, et al. Improved sensitivity of the urine CAA lateral-flow assay for diagnosing active Schistosoma infections by using larger sample volumes. Parasit Vectors. 2015;8:241.
Knopp S, Corstjens PL, Koukounari A, Ceramondi CI, Ame SM, Ali SM, et al. Sensitivity and specificity of a urine circulating anodic antigen test for the diagnosis of Schistosoma haematobium in low endemic settings. PLoS Negl Trop Dis. 2015;9:e3752.
Lodh N, Naples JM, Bosompem KM, Quartey J, Shiff CJ. Detection of parasite-specific DNA in urine sediment obtained by filtration differentiates between single and mixed infections of Schistosoma mansoni and S. haematobium from endemic areas in Ghana. PLoS One. 2014;9:e91144.
Obeng BB, Aryeetey YA, de Dood CJ, Amoah AS, Larbi IA, Deelder AM, et al. Application of a circulating-cathodic-antigen (CCA) strip test and real-time PCR, in comparison with microscopy, for the detection of Schistosoma haematobium in urine samples from Ghana. Ann Trop Med Parasitol. 2008;102:625–33.
Cnops L, Soentjens P, Clerinx J, Van Esbroeck M. A Schistosoma haematobium-specific real-time PCR for diagnosis of urogenital schistosomiasis in serum samples of international travelers and migrants. PLoS Negl Trop Dis. 2013;7:e2413.
Vinkeles Melchers NVS, van Dam GJ, Shaproski D, Kahama AI, Brienen EA, Vennervald BJ, van Lieshout L. Diagnostic performance of Schistosoma real-time PCR in urine samples from kenyan children infected with Schistosoma haematobium: day-to-day variation and follow-up after Praziquantel treatment. PLoS Negl Trop Dis. 2014;8:e2807.
Pai NP, Vadnais C, Denkinger C, Engel N, Pai M. Point-of-care testing for infectious diseases: diversity, complexity, and barriers in low- and middle-income countries. PLoS Med. 2012;9:e1001306.
Abbasi I, King CH, Muchirir EM, Hamburger J. Detection of Schistosoma mansoni and Schistosoma haematobium DNA by loop-mediated isothermal amplification: identification of infected snails from early prepatency. Am J Trop Med Hyg. 2010;83:427–32.
Hamburger J, Abbasi I, Kariuki C, Wanjala A, Mzungu E, Mungai P, et al. Evaluation of loop-mediated isothermal amplification suitable for molecular monitoring of schistosome-infected snails in field laboratories. Am J Trop Med Hyg. 2013;88:344–51.
Escadafal C, Faye O, Sall AA, Faye O, Weidmann M, Strohmeier O, et al. Rapid molecular assays for the detection of yellow fever virus in low-resource settings. PLoS Negl Trop Dis. 2014;8:e2730.
Kersting S, Rausch V, Bier FF, von Nickisch-Rosenegk M. Rapid detection of Plasmodium falciparum with isothermal recombinase polymerase amplification and lateral flow analysis. Malaria J. 2014;13:99.
Ahmed A, Linden H, Hartskeerl RA. Development of a recombinase polymerase amplification assay for the detection of pathogenic Leptospira. Int J Environ Res Pub Health. 2014;11:4953–64.
Krolov K, Frolova J, Tudoran O, Suhorutsenko J, Lehto T, Sibul H, et al. Sensitive and rapid detection of Chlamydia trachomatis by recombinase polymerase amplification directly from urine samples. J Mol Diag. 2014;16:127–35.
Rosser A, Rollinson D, Forrest M, Webster BL. Isothermal recombinase polymerase amplification (RPA) of Schistosoma haematobium DNA and oligochromatographic lateral flow detection. Parasit Vectors. 2015;8:446.
Emery AM, Allan FE, Rabone ME, Rollinson D. Schistosomiasis collection at NHM (SCAN). Parasit Vectors. 2012;5:185.
Knopp S, Mohammed KA, Ali SM, Khamis IS, Ame SM, Albonico M, et al. Study and implementation of urogenital schistosomiasis elimination in Zanzibar (Unguja and Pemba islands) using an integrated multidisciplinary approach. BMC Public Health. 2012;12:930.
Stothard JR, French MD, Khamis IS, Basáñez MG, Rollinson D. The epidemiology and control of urinary schistosomiasis and soil-transmitted helminthiasis in schoolchildren on Unguja island, Zanzibar. Trans R Soc Trop Med Hyg. 2009;103:1031–44.
Guidi A, Andolina C, Ame SM, Albonico M, Cioli D, Juma HH. Praziquantel efficacy and long-term appraisal of schistosomiasis control in Pemba island. Trop Med Int Health. 2010;15:614–8.
Knopp S, Ame SM, Person B, Hattendorf J, Rabone M, Juma S, et al. A 5-year intervention study on elimination of urogenital schistosomiasis in Zanzibar: parasitological results of annual cross-sectional surveys. PLoS Negl Trop Dis. 2019;13:e0007268.
Knopp S, Person B, Ame SM, Ali SM, Hattendorf J, Juma S, et al. Evaluation of integrated interventions layered on mass drug administration for urogenital schistosomiasis elimination: a cluster randomised trial. Lancet Glob Health. 2019;7:1118–29.
Hamburger J, Abbasi I, Ramzy RM, Jourdane J, Ruppel A. Polymerase chain reaction assay based on a highly repeated sequence of Schistosoma haematobium: a potential tool for monitoring schistosome-infested water. Am J Trop Med Hyg. 2001;65:907–11.
Wichmann D, Panning M, Quack T, Kramme S, Burchard GD, Grevelding C, Drosten C. Diagnosing schistosomiasis by detection of cell-free parasite DNA in human plasma. PLoS Negl Trop Dis. 2009;3:e422.
Acknowledgements
We thank the ZEST project, funded through the University of Georgia Research Foundation for the Schistosomiasis Consortium for Operational Research and Evaluation (SCORE) project (https://score.uga.edu) awarded by the Bill & Melinda Gates Foundation (prime award no. 50816, sub-award no. RR374-053/4893206), for the provision of the urine samples used in this study. We also acknowledge Dr Andrew Rosser (former LSHTM Masters student) for his RPA Dra1 primer and probe design.
Funding
The study received financial support through a Royal Society Research Grant (awarded to BW, RG160395) and the London School of Hygiene and Tropical Medicine (LSHTM) masters project funds awarded to PR. Also, funds were received from the Natural History Museum Life Sciences Department Investment Funds (DIF).
Author information
Authors and Affiliations
Contributions
BW designed the study, supervised PR, performed the urine sample and control RPA reactions, extracted the DNA from the urine samples in Pemba and wrote the paper. PR performed the laboratory development of the RPA assay and wrote the paper. TP extracted the samples in Pemba and supported the writing of the paper. FB, DR, SK, FA, FB, SA and SA were all involved in the ZEST and the storage and access to the urine samples used within the study. All authors supported the writing of the paper and commented on all versions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The ZEST study protocol was reviewed and obtained ethical approval by (i) the Zanzibar Medical Research Ethical Committee (ZAMREC); (ii) the “Ethikkomission beider Basel” (EKBB) in Switzerland; and (iii) the institutional review board of the University of Georgia (IRB UGA). The trial has been registered at the International Standard Randomised Controlled Trial Number Register (ISRCTN48837681; http://www.controlled-trials.com/ISRCTN48837681).
Written informed consent from parents or legal guardians of participating children was obtained for urine collection and storage, including examination for S. haematobium infections with newly developed diagnostic techniques at a later point in time. School-aged children were offered praziquantel (40 mg/kg) against schistosomiasis free of charge in the frame of the biannual island-wide mass drug administration campaigns.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Rostron, P., Pennance, T., Bakar, F. et al. Development of a recombinase polymerase amplification (RPA) fluorescence assay for the detection of Schistosoma haematobium. Parasites Vectors 12, 514 (2019). https://doi.org/10.1186/s13071-019-3755-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13071-019-3755-6