Abstract
Bacterial adaptive immunity hinges on CRISPR-Cas systems that provide DNA-encoded, RNA-mediated targeting of exogenous nucleic acids. A plethora of CRISPR molecular machines occur broadly in prokaryotic genomes, with a diversity of Cas nucleases that can be repurposed for various applications.
Similar content being viewed by others
CRISPR-Cas systems and adaptive immunity
The characterization of biological processes that underlie CRISPR-based adaptive immunity in bacteria and archaea has shaped many crucial aspects of the past decade in the fields of microbiology and genetics, and has enabled the current ‘genome editing’ craze [1]. Clustered regularly interspaced short palindromic repeats (CRISPRs) and their CRISPR-associated (Cas) proteins constitute the CRISPR-Cas immune system (Fig. 1), which provides adaptive immunity against invasive elements such as viruses and plasmids in bacteria and archaea [2–5]. Although CRISPR loci were first observed in the genome of Escherichia coli in 1987 [6], it took 15 years of microbial genomics renaissance to appreciate their widespread occurrence in bacteria and archaea [7, 8]. Actually, it was exactly 10 years ago that the first functional clue emerged, with the observation that CRISPR spacers showed homology to viral sequences [9–11], leading to the hypothesis that they might constitute a prokaryotic equivalent to RNA interference (RNAi) [12]. Shortly thereafter, their biological function as adaptive immune systems was established [13], revealing that CRISPR arrays, together with cas genes, provide acquired immunity against bacteriophages in a sequence-specific manner. The mechanism of action of various CRISPR-Cas systems has since been determined through milestone discoveries establishing that CRISPR-encoded immunity is mediated by CRISPR RNA (crRNAs) [14], and targets invasive DNA [15] and sometimes RNA [16].
CRISPR-Cas systems and adaptive immunity. CRISPR repeats, together with CRISPR spacers, constitute repeat-spacer arrays that define clustered regularly interspaced short palindromic repeats (CRISPRs). These CRISPR arrays are typically flanked by CRISPR-associated sequences (cas) that encode Cas proteins involved in the three stages of CRISPR-encoded immunity, namely adaptation, expression and interference. During adaptation, Cas proteins (including the universal Cas1 and Cas2) sample invasive DNA, leading to the genesis of a new repeat-spacer unit that is inserted in a polarized manner in the CRISPR array. During the second stage — expression — the CRISPR array is transcribed into a full pre-crRNA transcript that is processed into small, mature, interfering CRISPR RNAs (crRNAs). In the third — interference — stage, crRNAs guide Cas effector proteins towards complementary nucleic acids for sequence-specific targeting. Interaction between the interference complex and the target nucleic acid is typically initiated by binding to the protospacer adjacent motif (PAM), which triggers interrogation of flanking DNA by the loaded crRNA. If complementarity extends beyond the seed sequence, an R-loop is formed, and nickase domains within Cas effector proteins cleave the target DNA. dsDNA double-stranded DNA, L leader
Key discoveries quickly established that targeting is generally dependent upon a short DNA sequence known as the protospacer adjacent motif (PAM) [17–19], is driven by seed sequences [20, 21] and is mediated by Cas endonucleases that specifically cleave complementary DNA [22]. For type I systems, early efforts defined the biochemical and structural underpinning of the ‘CRISPR-associated complex for antiviral defence’ (Cascade) [14], and the endonucleolytic and exonucleolytic degradation of DNA by Cas3 [23–29]. For type II systems, early studies defined crRNA biogenesis [30], Cas9-dependent immunity [13] and cleavage [22], and eventually re-programmable targeting [31] and genesis of precise double-stranded DNA (dsDNA) breaks [32–34].
Arguably, it was the turning of native CRISPR-Cas systems into engineered and programmable two-component systems comprising Cas9 and single guide RNAs (sgRNAs) [33] that was the technological tipping point that singlehandedly enabled Cas9-driven genome editing [35–37] and fuelled the CRISPR craze that has unabatedly unfolded since then [1, 38]. The technical tour de force essentially turned the native Cas9–trans-activating CRISPR RNA (tracrRNA)–crRNA–RNase-III four-component system into the streamlined Cas9–sgRNA technology, rendering the challenge of co-opting the system for eukaryotic applications accessible. The synthetic genesis of sgRNAs allowed the repurposing of CRISPR-Cas immune systems into powerful and nimble molecular machines that can yield double-stranded breaks. Indeed, the Cas9 molecular-scalpel-based genome editing craze was foreshadowed in the fall of 2012 [39], following the release of the sgRNA–Cas9 technology, and preceding the publication of proof of concept in human [35, 36] and bacterial cells [37]. Within months, the Church, Zhang and Marraffini labs were able to concurrently establish that the sgRNA–Cas9 technology can be exploited for efficient genome editing, and immediately thereafter, hundreds of studies showed that this approach can be universally implemented in a wide range of cells and model organisms. The avalanche of Cas9-based genome editing studies attests to the potential of this broadly applicable technology.
Mechanistically, CRISPR-Cas immunity hinges on three distinct steps, defined as adaptation, expression and interference (Fig. 1). In the adaptation stage, CRISPR immunization occurs through the uptake and polarized integration of invasive DNA as a novel CRISPR spacer into the CRISPR array, creating a serial record of vaccination events. In the expression stage, the CRISPR array is transcribed into a full pre-CRISPR RNA (pre-crRNA) transcript that is processed into mature crRNAs containing partial CRISPR spacer sequences attached to partial CRISPR repeats, forming CRISPR guide RNAs. In the interference stage, crRNAs guide Cas nucleases towards complementary nucleic acids for sequence-specific targeting and cleavage of invasive genetic elements. Most CRISPR effector proteins initiate targeting by interaction with a particular two-to-four nucleotide sequence motif, the PAM. Once interaction with the PAM has been established, the crRNA guide loaded within the Cas nuclease can then interrogate the flanking target DNA [40, 41]. The strength and duration of the molecular interaction correlates with the level of complementarity between the crRNA and target DNA, which drives conformational changes in Cas effector proteins, such as Cas9 [40, 42, 43] and Cascade [44–46], that eventually lead to a cleavage-competent structural state [40]. If complementarity between the guide RNA and target DNA extends beyond the seed sequence, a DNA R-loop is directionally formed [29, 47, 48], which triggers subsequent nicking by the Cas effector nucleases (i.e., Cas3, Cas9, Cpf1) at particular locations defined by a ruler-anchor mechanism. The literature includes many reviews that cover the history [49–52], biology [3–5, 53–56] and applications [57–63] of CRISPR-Cas systems.
Diversity of CRISPR-Cas systems and mechanisms of action
In general terms, there are two main classes [64] of CRISPR-Cas systems, which encompass five major types and 16 different subtypes based on cas gene content, cas operon architecture, Cas protein sequences, and processes that underlie the aforementioned steps (Fig. 1) [65, 66]. The first class is defined by multiprotein effector complexes (Cascade, Cmr, Csm), and encompasses types I, III and IV. In particular, type I systems are the most frequent and widespread systems, which target DNA in a Cascade-driven and PAM-dependent manner, destroying target nucleic acids by using the signature protein Cas3 [26, 28, 67–71] (Fig. 2). Many studies have led to extensive biochemical and structural characterization of the effector proteins and protein–DNA–RNA complexes implicated in type I CRISPR-Cas systems [20, 23, 24, 46, 72–77]. Likewise, type III systems occur frequently in archaea and are characterized by the multiprotein Csm [78–82] or Cmr [16, 83–95] complexes; they operate in a PAM-independent manner and can cleave DNA or RNA by using the signature Cas10 protein together with effector nucleases such as Cmr4 (the RNase within the Cmr complex for type III-B systems) [85, 95] and Csm3 (the RNase within the Csm complex for type III-A systems) [81, 82]. Interestingly, several recent studies have revealed that type III CRISPR-Cas systems can actually target both nucleic acid types, through co-transcriptional RNA and DNA cleavage [80, 82]. Specifically, distinct active sites within the Cas10–Csm ribonucleoprotein effector complex drive co-transcriptional RNA-guided DNA cleavage and RNA cleavage [80]. Type IV systems are rather rare and still remain to be characterized in terms of their distribution and function.
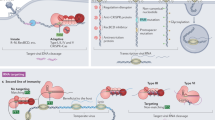
Diversity of CRISPR-Cas molecular machines. Two main classes of CRISPR-Cas systems exist, which are defined by the nature of their Cas effector nucleases, either constituted by multiprotein complexes (class 1), or by a single signature protein (class 2). For class 1 systems, the main types of CRISPR-Cas systems include type I and type III systems. Illustrated here as an example, the Escherichia coli K12 type I-E system (upper left) targets sequences flanked by a 5′-located PAM. Guide RNAs are generated by Cascade, in a Cas6-defined manner and typically contain an eight-nucleotide 5′ handle derived from the CRISPR repeat, a full spacer sequence, and a 3′ hairpin derived from the CRISPR repeat. Following nicking of the target strand, the 3′ to 5′ Cas3 exonuclease destroys the target DNA in a directional manner. In the Pyrococcus furiosus DSM 3638 type III-B system (lower left), a short crRNA guide directs the Cmr complex towards complementary single-stranded RNA in a PAM-independent manner. For the canonical type II-A Streptococcus thermophilus LMD-9 system (upper right), a dual crRNA–tracrRNA guide generated by Cas9 and RNase III targets a 3′-flanked PAM DNA complementary sequence for the genesis of a precise double-stranded break using two nickase domains (RuvC and HNH). For the Francisella novicida U112 type V system (lower right), a single guide RNA targets complementary dsDNA flanked by a 5′-PAM using Cpf1, which generates a staggered dsDNA break. Cascade CRISPR-associated complex for antiviral defense, CRISPR clustered regularly interspaced short palindromic repeat, crRNA CRISPR RNA, dsDNA double-stranded DNA, L leader, nt nucleotide, PAM protospacer adjacent motif, ssRNA single-stranded RNA, tracrRNA trans-activating CRISPR RNA
By contrast, the second class is defined by single effector proteins and encompasses types II and V. Type II systems are defined by the popular Cas9 endonuclease [22], which hinges on dual crRNA–tracrRNA guides [30] that direct the RuvC and HNH nickase domains to generate precise blunt DNA breaks in target DNA sequences flanked by a 3ʹ PAM [22, 31–34, 96, 97]. Type V systems are rare, and characterized by the signature Cpf1 nuclease, which is guided by a single crRNA that directs this RuvC-like endonuclease for staggered dsDNA nicking to yield sticky-ends in target DNA sequences flanked by a 5′ PAM [98].
Recently, several studies have shown that, although CRISPR-Cas systems generally function in three distinct stages, involving peculiar molecular processes and various Cas molecular machines, the adaptation and interference steps can actually be coupled [48, 99–101], which is consistent with the priming hypothesis [48, 102–104]. Specifically, differential binding determines whether cognate target DNA should be destroyed as part of the interference pathway, or whether partially complementary sequences should be directed towards the adaptation path [48]. The coupling of the adaptation and interference stages also reflects their co-dependence on Cas9 and PAM sequences in type II systems [100, 101, 105], and implicates a ‘cut-and-paste’ model rather than ‘copy and paste’ [100].
Overall, a broad genetic and functional diversity of CRISPR-Cas immune systems occurs in the genomes of many bacteria and most archaea. Common denominators include DNA-encoded immunity within CRISPR arrays that yield small guide RNAs, which define sequence-specific targets for Cas nucleases and subsequent nucleic acid cleavage. The universal cas1 and cas2 genes, implicated in polarized, sequence- and structure-specific integrase-mediated spacer acquisition during the adaptation stage [106–108], are present in all characterized types and subtypes in the two main classes. By contrast, there is substantial variability between classes, types and subtypes concerning the nature, sequence and structure of the CRISPR RNAs and Cas proteins involved, the reliance on and location of PAM sequences, and the nature of the target nucleic acid. Altogether, this illustrates the extensive multi-dimensional diversity of CRISPR-Cas systems, their native biological functions, and the relative potential for various biotechnological and industrial applications.
The diversity of CRISPR-Cas systems reflects their various functional roles. Although the primary established function of CRISPR-Cas systems is adaptive immunity against invasive genetic elements such as plasmids and viruses, several studies have independently implicated them in other functions, including endogenous transcriptional control, as well as resistance to stress, pathogenicity and regulation of biofilm formation [63, 109–114].
Future studies are anticipated to determine the rationale for the distribution biases in various phylogenetic groups, for the absence of CRISPR-Cas systems in so many bacteria, and to unravel the functional links between immunity and other key biological processes such as DNA homeostasis and repair. One intriguing conundrum about CRISPR-Cas systems is their absence in approximately half of the bacterial genomes sequenced to date, despite their intuitive evolutionary value. Another important consideration is whether the observed biases in proto-spacer sampling during adaptation correlate with efficiency biases for the interference stage. Specifically, spacer adaptation biases have been repeatedly observed in type I systems [115, 116] and in type II systems [105, 117], implicating replication-dependent DNA breaks at replication forks, Chi sites and interplay with the RecBCD DNA repair machinery, and so it will be important to determine whether these also explain spacer efficiency variability during interference.
Applications of native and engineered CRISPR-Cas systems in bacteria
Although the large majority of the CRISPR literature focuses on genome editing applications in eukaryotes, CRISPR-Cas systems arguably afford the most applications in both native and engineered forms in bacteria [118, 119]. Actually, most of the alleged CRISPR literature does not employ bona fide clustered regularly interspaced short palindromic repeats, but instead features crRNA-guided Cas9 proteins. Given the aforementioned CRISPR-Cas system diversity, and the available molecular biology tools for bacteria, we are thus on the cusp of full exploitation in microbes. There are three primary ways to harness CRISPR-Cas systems, depending on the CRISPR immunity stage, Cas machinery and outcome being exploited (Fig. 3).
Applications and targets of CRISPR-Cas systems. CRISPR-Cas systems can target various types of nucleic acids, including invasive and mobile DNA (green box), or endogenous sequences (blue box). In their native environment, CRISPR-Cas systems naturally target mobile and exogenous DNA elements. Conversely, engineered systems are typically designed to target self-DNA to trigger endogenous modifications. Targeting can be directed at bacteriophage DNA to provide anti-viral defense (upper left). Likewise, Cas nucleases can be directed at plasmid DNA in order to prevent uptake and dissemination of undesirable sequences or to cure the host of plasmid sequences (center left). Targeting can also be directed at mobile DNA elements such as transposons so as to maintain DNA integrity and ensure homeostasis (lower left). When aiming the CRISPR-Cas machinery towards the cell’s own chromosomal content, the purpose is typically to induce endogenous DNA repair pathways to drive editing of the DNA sequence (upper center). Catalytically deactivated variants of Cas nucleases can be used as DNA-binding proteins to block transcription (CRISPRi, upper right), or can be fused to transcriptional activators to activate transcription (CRISPRa, center right). Alternatively, Cas nucleases can be reprogrammed to trigger a lethal auto-immune response, leading to cell death (bottom right). CRISPR sequences themselves can be used for genotyping, by using the series of vaccination events as a genetic historical record (lower center). Cas CRISPR associated, CRISPR clustered regularly interspaced short palindromic repeat, CRISPRa CRISPR activation, CRISPRi CRISPR interference
First, the outcomes of native vaccination events can be used to genotype bacteria by comparing and contrasting the spacer acquisition events to unravel the evolutionary path of a strain isolate, or to delve into assessing the genetic composition and diversity of a population (Fig. 3). This approach has proven valuable for the typing of bacterial pathogens in which CRISPR array diversity reflects functional acquisitions over time, such as in Escherichia coli, Yersinia pestis, Mycobacterium tuberculosis, Salmonella enterica and Clostridium difficile [120]. This method is also instrumental for the detection and monitoring of industrially valuable workhorses associated with bioprocessing and food manufacturing, such as probiotics and starter cultures [17, 121–124]. Similar approaches, in combination with deep sequencing technology, have shown tremendous potential for the analysis of complex microbial populations, for the determination of clonal population diversity, and for the analysis of co-evolutionary dynamics and arms-races between bacteria and phages [105, 117, 125–127]. Studies can specifically investigate the evolutionary dynamics of hosts and phage populations and unravel selection events and counter-selective mutational patterns that allow bacteria to thrive in inhospitable conditions [128, 129]. In fact, early metagenomic work on CRISPR diversity and interplay with phage sequences from the Banfield laboratory was crucial in establishing the CRISPR field [130, 131]. More recently, studies of CRISPR-based interplay between bacteria and viruses have expanded to human-associated microbial populations, including in the oral cavity and gastrointestinal tract [132–135].
Second, CRISPR-Cas immune systems can be used to vaccinate against invasive genetic elements [13]. Either naturally or by engineering, CRISPR-Cas systems can be exploited to provide resistance against phages [136] or preclude uptake and dissemination of undesirable genetic material such as antibiotic resistance genes [22] or possibly target mobile genetic elements such as transposons to ensure genome homeostasis. In addition to blocking bacteriophages, CRISPR interference is a potent barrier to natural DNA transformation that can be harnessed to prevent the acquisition of virulence traits [137]. Perhaps surprisingly, the industrial exploitation of CRISPR-Cas systems for the genesis of robust and sustainable starter cultures used for the manufacturing of fermented dairy products has been implemented commercially in consumer products for several years [120, 136, 138]. Actually, exploiting first-generation CRISPR patent applications that are over a decade old, and building off early scientific discoveries about adaptive spacer acquisition in Streptococcus thermophilus, naturally generated bacteria that have been screened for vaccination events against phage isolated from commercial settings have been exploited on a global scale since 2011. Of course, natural CRISPR-immunized strains might have been used for a long time, unbeknownst to us. Practically, the breadth and depth of phage resistance can be built up iteratively through multiple rounds of selection of natural vaccination events that eventually yield a sustainable starter culture with increased life span in the food industry. Similar approaches hold much potential for the improvement of industrial workhorses valuable to the bio-manufacturing industry.
Third, endogenous or engineered Cas machinery can be repurposed for self-DNA targeting in a wide range of applications that encompass genome editing and targeted killing (Fig. 3). Many studies have documented the nimble potential of the sgRNA–Cas9 technology for ‘traditional’ genome editing, to knock out, insert or delete genes [57–59]. Furthermore, deactivated versions of Cas9 (dCas9) have been generated by inactivation of the RuvC and HNH nickase domains to turn the nuclease into a DNA binding protein able to control transcription, either by blocking RNA polymerases (CRISPR interference, CRISPRi) or by promoting transcription when tethered to transcriptional activators (CRISPR activation, CRISPRa). The use of both endogenous and engineered CRISPR-Cas systems for transcriptional control in bacteria has already been documented [139–141]. More recently, functional variants of Cas9 associated with fluorophores or methylase domains have been used for imaging and epigenome modification [142, 143], respectively. These applications have redefined genome editing beyond the alteration of the DNA sequence per se, and now enable the editing of any sequence in any cell in many ways. Despite the Cas9-based genome editing bias in eukaryotes, their implementation in bacteria is on the rise [118, 144–147]. In bacteria, a promising recent application of self-targeting is programmable killing [148], opening new avenues for the genesis of next-generation smart antimicrobials based on various CRISPR-Cas systems [148–152]. Specifically, engineered Cas9 systems, as well as native Cas9 and Cascade machines, have been successfully re-programmed for sequence-specific targeted killing of a bacterial population, which allows the manipulation of mixed consortia, and the select eradication of defined genotypes of interest [148]. This has successfully been implemented to target E. coli, Streptococcus and Staphylococcus aureus, both in in vitro and in vivo models [148–151]. This is an opportunity to properly select and leverage particular CRISPR-Cas systems that might be better suited for efficient killing, such as type I systems that rely on the Cas3 endo- and exo-nuclease, which digests the target DNA following initial cleavage (Fig. 2), and thus affords the cell fewer opportunities to repair cleaved DNA. Moving forward, there is much potential for this technology to develop narrow-range antibiotics that can be customized for the alteration of microbiomes. This also opens intriguing prospects for programmable eradication of select cell populations in eukaryotes.
Altogether, these various applications illustrate the functional diversity of CRISPR-Cas systems (Fig. 3) and set the stage for the customized selection and development of various molecular machines to expand the molecular biology toolbox. In some ways, type I systems can be construed as a powerful ‘hammer’, which heavily hits and destroys target DNA with the Cas3 exonuclease. Type II systems could be used as nimble ‘screwdrivers’, which precisely target DNA with the Cas9 endonuclease. Similarly, the recently characterized type V systems [64] can be perceived as screwdrivers with a different propensity (flat-head versus phillips) for precise targeting of DNA with the Cpf1 endonuclease [98]. Type III systems can be employed as ‘box cutters’ that can cleave either DNA or RNA with the signature Cas10 nuclease. Given how much our understanding of system diversity has increased in the past 15 years, the diversity of CRISPR-Cas systems will predictably further increase as we deepen our knowledge of microbial genomics, and valuable Cas molecular machines might be unearthed in the future. Altogether, these native and engineered systems hold tremendous potential for a broad range of bacterial applications (Fig. 4).
Exploitation of endogenous and engineered CRISPR-Cas systems in bacteria. Exogenous DNA sequences can be targeted by CRISPR-Cas systems to build up phage resistance in food starter cultures (to vaccinate yoghurt strains against bacteriophage), to prevent the uptake and dissemination of plasmids that encode undesirable traits such as antibiotic resistance genes (to immunize probiotic strains used in dietary supplements), or to ensure the genetic integrity and genomic homeostasis of valuable cultures (to fend off mobile genetic elements such as transposons and prophages) (upper panels). Unique records of iterative vaccination events captured as a series of spacers in CRISPR arrays can be used as sequencing targets for the detection, monitoring and typing of strains of interest, which include food cultures, spoilage organisms or pathogens (center panels). By contrast, self-targeting and engineered applications can be used in industrial settings to improve industrial workhorses by genome editing (indicated by ‘scissors’ symbol), or by re-directing the metabolic flux of various pathways for synthetic and yield purposes (lower panels). Lethal self-targeting can also be harnessed for the select eradication of pathogens or contaminants of interest. CRISPRa CRISPR activation, CRISPRi CRISPR interference
Keep calm and CRISPR on
Although the advent of the sgRNA–Cas9 technology for eukaryotic genome editing is merely two years old, the success of this disruptive technology is undeniable [1, 38]. It is noteworthy to point out that the scientific community was primed for the use and rapid implementation of this technology, given the historical use of the powerful RNAi technology on the one hand, and the rise of TALEN-, meganuclease- and zinc-finger nuclease (ZFN)-based genome editing applications on the other. Indeed, many were ready, and well-positioned, if not eager, to unleash the potential of this powerful technology. In hindsight, there are many attributes of CRISPR-Cas systems that render them valuable, including programmability, transferability, efficiency, specificity, affordability, rapidity of implementation, precision, ease of use, and ability to multiplex both guides and systems. Nevertheless, this is still a nascent technology, which needs improvements, especially as it relates to size (Cas9 is arguably cumbersome), targeting flexibility (broadening the PAM space) and efficiency (ability to recognize and cleave targets with specificity and efficiency). Perhaps a longer-term improvement consists of being able to select the most efficient spacer sequences since not all CRISPR spacers or RNA guides provide equal targeting of phage or target sequences, respectively, and adequate prediction of common outcomes (ability of viruses to mutate targeted sequences, or propensities of various DNA repair pathways to alter the cleaved sites). Already, biochemical and structural insights [43, 153–157] are fuelling efforts under way to engineer guides and Cas nucleases for improved functionalities, including smaller variants and PAM-targeting flexibility. In parallel, analysis of Cas nuclease diversity and orthogonality [156, 158–162] will accelerate the rational design of next-generation engineered nucleases. Likewise, lessons from RNAi are instrumental in optimizing the composition and structure of functional CRISPR guides for improved activity and specificity. Finally, the characterization of additional CRISPR-Cas systems in general, and more Cas effector proteins in particular, will broaden the set of forthcoming molecular tools available for various applications.
Already, there are a few valuable lessons regarding Cas effector proteins that have been gathered from CRISPR applications in bacteria that could prove useful to the broad scientific community. In particular, it is noteworthy to point out that, per se, immune systems must afford both specificity and efficiency, so as to prevent auto-immunity and ensure survival, respectively. This is particularly crucial in antiviral CRISPR defence given the speed with which phage co-opt the host cellular machinery, and the ease with which they can mutate to escape sequence-specific targeting. Indeed, stealth and specific targeting of viral DNA occurs through Cas effector protein recognition of bona fide sequences, and their specific nucleolytic destruction. CRISPR-based eradication of phages and toxic DNA thus occurs on the scale of minutes following infection, ensuring efficiency. Likewise, targeting hinging on protospacer recognition ensures that lethal self-targeting events are avoided, providing specificity.
Using recent history and the current momentum to foretell the short-term future of the CRISPR craze, it appears that: first, the pace at which the field is moving forwards is not abating, as indicated by literature output, citation rates and funding trends; second, the coverage has extended feverishly beyond the scientific press, into the mass media; and finally some of the most enthralling level of interest lies in the business commitment and commercial potential of that technology, illustrated by financial investment levels spanning a broad range of business segments, such as medicine, food, agriculture and biotechnology. As the fascinating CRISPR story continues to unfold, and the IP, ethical and awards debates consume attention, it will be crucial to ensure that we keep calm and CRISPR on to ensure we do not hinder but, instead, unleash and further advance this powerful technology.
Abbreviations
- Cas:
-
CRISPR-associated sequences
- Cascade:
-
CRISPR-associated complex for antiviral defense
- CRISPR:
-
clustered, regularly interspaced short palindromic repeats
- crRNA:
-
CRISPR RNA
- PAM:
-
protospacer adjacent motif
- sgRNA:
-
single guide RNA
- tracrRNA:
-
trans-activating CRISPR RNA
References
Pennisi E. The CRISPR craze. Science. 2013;341:833–6.
Barrangou R, Marraffini LA. CRISPR-Cas systems: Prokaryotes upgrade to adaptive immunity. Mol Cell. 2014;54:234–44.
Sorek R, Lawrence CM, Wiedenheft B. CRISPR-mediated adaptive immune systems in bacteria and archaea. Annu Rev Biochem. 2013;82:237–66.
Barrangou R. CRISPR–Cas systems and RNA-guided interference. Wiley Interdiscip Rev RNA. 2013;4:267–78.
Westra ER, Swarts DC, Staals RH, Jore MM, Brouns SJ, van der Oost J. The CRISPRs, they are a-changin’: how prokaryotes generate adaptive immunity. Annu Rev Genet. 2012;46:311–39.
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol. 1987;169:5429–33.
Jansen R, Embden JD, Gaastra W, Schouls LM. Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol. 2002;43:1565–75.
Mojica FJ, Diez-Villasenor C, Soria E, Juez G. Biological significance of a family of regularly spaced repeats in the genomes of Archaea. Bacteria and mitochondria. Mol Microbiol. 2000;36:244–6.
Pourcel C, Salvignol G, Vergnaud G. CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology. 2005;151:653–63.
Mojica FJ, Diez-Villasenor C, Garcia-Martinez J, Soria E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol. 2005;60:174–82.
Bolotin A, Quinquis B, Sorokin A, Ehrlich SD. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology. 2005;151:2551–61.
Makarova KS, Grishin NV, Shabalina SA, Wolf YI, Koonin EV. A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol Direct. 2006;1:7.
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315:1709–12.
Brouns SJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJ, Snijders AP, et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science. 2008;321:960–4.
Marraffini LA, Sontheimer EJ. CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science. 2008;322:1843–5.
Hale CR, Zhao P, Olson S, Duff MO, Graveley BR, Wells L, et al. RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell. 2009;139:945–56.
Horvath P, Romero DA, Coute-Monvoisin AC, Richards M, Deveau H, Moineau S, et al. Diversity, activity, and evolution of CRISPR loci in Streptococcus thermophilus. J Bacteriol. 2008;190:1401–12.
Deveau H, Barrangou R, Garneau JE, Labonte J, Fremaux C, Boyaval P, et al. Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J Bacteriol. 2008;190:1390–400.
Mojica FJ, Diez-Villasenor C, Garcia-Martinez J, Almendros C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology. 2009;155:733–40.
Wiedenheft B, van Duijn E, Bultema JB, Waghmare SP, Zhou K, Barendregt A, et al. RNA-guided complex from a bacterial immune system enhances target recognition through seed sequence interactions. Proc Natl Acad Sci U S A. 2011;108:10092–7.
Semenova E, Jore MM, Datsenko KA, Semenova A, Westra ER, Wanner B, et al. Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence. Proc Natl Acad Sci U S A. 2011;108:10098–103.
Garneau JE, Dupuis ME, Villion M, Romero DA, Barrangou R, Boyaval P, et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature. 2010;468:67–71.
Reeks J, Naismith JH, White MF. CRISPR interference: a structural perspective. Biochem J. 2013;453:155–66.
Jackson RN, Wiedenheft B. A Conserved structural chassis for mounting versatile CRISPR RNA-guided immune responses. Mol Cell. 2015;58:722–8.
Gong B, Shin M, Sun J, Jung CH, Bolt EL, van der Oost J, et al. Molecular insights into DNA interference by CRISPR-associated nuclease-helicase Cas3. Proc Natl Acad Sci U S A. 2014;111:16359–64.
Hochstrasser ML, Taylor DW, Bhat P, Guegler CK, Sternberg SH, Nogales E, et al. CasA mediates Cas3-catalyzed target degradation during CRISPR RNA-guided interference. Proc Natl Acad Sci U S A. 2014;111:6618–23.
Huo Y, Nam KH, Ding F, Lee H, Wu L, Xiao Y, et al. Structures of CRISPR Cas3 offer mechanistic insights into Cascade-activated DNA unwinding and degradation. Nat Struct Mol Biol. 2014;21:771–7.
Mulepati S, Bailey S. In vitro reconstitution of an Escherichia coli RNA-guided immune system reveals unidirectional, ATP-dependent degradation of DNA target. J Biol Chem. 2013;288:22184–92.
Rutkauskas M, Sinkunas T, Songailiene I, Tikhomirova MS, Siksnys V, Seidel R. Directional R-loop formation by the CRISPR-Cas surveillance complex cascade provides efficient off-target site rejection. Cell Rep. 2015. doi:10.1016/j.celrep.2015.01.067.
Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y, Pirzada ZA, et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature. 2011;471:602–7.
Sapranauskas R, Gasiunas G, Fremaux C, Barrangou R, Horvath P, Siksnys V. The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res. 2011;39:9275–82.
Gasiunas G, Barrangou R, Horvath P, Siksnys V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci U S A. 2012;109:E2579–86.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337:816–21.
Magadan AH, Dupuis ME, Villion M, Moineau S. Cleavage of phage DNA by the Streptococcus thermophilus CRISPR3-Cas system. PLoS One. 2012;7:e40913.
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, et al. RNA-guided human genome engineering via Cas9. Science. 2013;339:823–6.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339:819–23.
Jiang W, Bikard D, Cox D, Zhang F, Marraffini LA. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat Biotechnol. 2013;31:233–9.
Ledford H. CRISPR, the disruptor. Nature. 2015;522:20–4.
Barrangou R. RNA-mediated programmable DNA cleavage. Nat Biotechnol. 2012;30:836–8.
Jiang F, Zhou K, Ma L, Gressel S, Doudna JA. A Cas9-guide RNA complex preorganized for target DNA recognition. Science. 2015;348:1477–81.
Sternberg SH, Redding S, Jinek M, Greene EC, Doudna JA. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature. 2014;507:62–7.
Jiang F, Doudna JA. The structural biology of CRISPR-Cas systems. Curr Opin Struct Biol. 2015;30:100–11.
Jinek M, Jiang F, Taylor DW, Sternberg SH, Kaya E, Ma E, et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science. 2014;343:1247997.
Sashital DG, Jinek M, Doudna JA. An RNA-induced conformational change required for CRISPR RNA cleavage by the endoribonuclease Cse3. Nat Struct Mol Biol. 2011;18:680–7.
Sashital DG, Wiedenheft B, Doudna JA. Mechanism of foreign DNA selection in a bacterial adaptive immune system. Mol Cell. 2012;46:606–15.
Jore MM, Lundgren M, van Duijn E, Bultema JB, Westra ER, Waghmare SP, et al. Structural basis for CRISPR RNA-guided DNA recognition by Cascade. Nat Struct Mol Biol. 2011;18:529–36.
Szczelkun MD, Tikhomirova MS, Sinkunas T, Gasiunas G, Karvelis T, Pschera P, et al. Direct observation of R-loop formation by single RNA-guided Cas9 and Cascade effector complexes. Proc Natl Acad Sci U S A. 2014;111:9798–803.
Blosser TR, Loeff L, Westra ER, Vlot M, Kunne T, Sobota M, et al. Two distinct DNA binding modes guide dual roles of a CRISPR-Cas protein complex. Mol Cell. 2015;58:60–70.
Horvath P, Barrangou R. CRISPR/Cas, the immune system of bacteria and archaea. Science. 2010;327:167–70.
Bhaya D, Davison M, Barrangou R. CRISPR-Cas systems in bacteria and archaea: versatile small RNAs for adaptive defense and regulation. Annu Rev Genet. 2011;45:273–97.
Sontheimer EJ, Barrangou R. The bacterial origins of the CRISPR genome-editing revolution. Hum Gene Ther. 2015;26:413–24.
van Erp PB, Bloomer G, Wilkinson R, Wiedenheft B. The history and market impact of CRISPR RNA-guided nucleases. Curr Opin Virol. 2015;12:85–90.
Fineran PC, Charpentier E. Memory of viral infections by CRISPR-Cas adaptive immune systems: acquisition of new information. Virology. 2012;434:202–9.
Barrangou R, Birmingham A, Wiemann S, Beijersbergen RL, Hornung V, Smith A. Advances in CRISPR-Cas9 genome engineering: lessons learned from RNA interference. Nucleic Acids Res. 2015;43:3407–19.
Sternberg SH, Doudna JA. Expanding the biologist's toolkit with CRISPR-Cas9. Mol Cell. 2015;58:568–74.
Charpentier E, Richter H, van der Oost J, White MF. Biogenesis pathways of RNA guides in archaeal and bacterial CRISPR-Cas adaptive immunity. FEMS Microbiol Rev. 2015;39:428–41.
Sander JD, Joung JK. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol. 2014;32:347–55.
Doudna JA, Charpentier E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science. 2014;346:1258096.
Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell. 2014;157:1262–78.
Doudna JA. Genomic engineering and the future of medicine. JAMA. 2015;313:791–2.
Charpentier E, Marraffini LA. Harnessing CRISPR-Cas9 immunity for genetic engineering. Curr Opin Microbiol. 2014;19:114–9.
Barrangou R, May AP. Unraveling the potential of CRISPR-Cas9 for gene therapy. Expert Opin Biol Ther. 2015;15:311–4.
Barrangou R. The roles of CRISPR-Cas systems in adaptive immunity and beyond. Curr Opin Immunol. 2015;32:36–41.
Makarova KS, Wolf YI, Alkhnbashi OS, Costa F, Shah SA, Saunders SJ, et al. An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol. 2015. doi:10.1038/nrmicro3569.
Makarova KS, Aravind L, Wolf YI, Koonin EV. Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR-Cas systems. Biol Direct. 2011;6:38.
Makarova KS, Koonin EV. Annotation and classification of CRISPR-Cas systems. Methods Mol Biol. 2015;1311:47–75.
Sinkunas T, Gasiunas G, Fremaux C, Barrangou R, Horvath P, Siksnys V. Cas3 is a single-stranded DNA nuclease and ATP-dependent helicase in the CRISPR/Cas immune system. EMBO J. 2011;30:1335–42.
Sinkunas T, Gasiunas G, Waghmare SP, Dickman MJ, Barrangou R, Horvath P, et al. In vitro reconstitution of Cascade-mediated CRISPR immunity in Streptococcus thermophilus. EMBO J. 2013;32:385–94.
Mulepati S, Bailey S. Structural and biochemical analysis of nuclease domain of clustered regularly interspaced short palindromic repeat (CRISPR)-associated protein 3 (Cas3). J Biol Chem. 2011;286:31896–903.
Westra ER, van Erp PB, Kunne T, Wong SP, Staals RH, Seegers CL, et al. CRISPR immunity relies on the consecutive binding and degradation of negatively supercoiled invader DNA by Cascade and Cas3. Mol Cell. 2012;46:595–605.
van Erp PB, Jackson RN, Carter J, Golden SM, Bailey S, Wiedenheft B. Mechanism of CRISPR-RNA guided recognition of DNA targets in Escherichia coli. Nucleic Acids Res. 2015;43:8381–91.
Jackson RN, Golden SM, van Erp PB, Carter J, Westra ER, Brouns SJ, et al. Structural biology. Crystal structure of the CRISPR RNA-guided surveillance complex from Escherichia coli. Science. 2014;345:1473–9.
Jackson RN, Lavin M, Carter J, Wiedenheft B. Fitting CRISPR-associated Cas3 into the helicase family tree. Curr Opin Struct Biol. 2014;24:106–14.
Mulepati S, Heroux A, Bailey S. Crystal structure of a CRISPR RNA-guided surveillance complex bound to a ssDNA target. Science. 2014;345:1479–84.
Mulepati S, Orr A, Bailey S. Crystal structure of the largest subunit of a bacterial RNA-guided immune complex and its role in DNA target binding. J Biol Chem. 2012;287:22445–9.
Haurwitz RE, Jinek M, Wiedenheft B, Zhou K, Doudna JA. Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science. 2010;329:1355–8.
Haurwitz RE, Sternberg SH, Doudna JA. Csy4 relies on an unusual catalytic dyad to position and cleave CRISPR RNA. EMBO J. 2012;31:2824–32.
Hatoum-Aslan A, Maniv I, Samai P, Marraffini LA. Genetic characterization of antiplasmid immunity through a type III-A CRISPR-Cas system. J Bacteriol. 2014;196:310–7.
Rouillon C, Zhou M, Zhang J, Politis A, Beilsten-Edmands V, Cannone G, et al. Structure of the CRISPR interference complex CSM reveals key similarities with cascade. Mol Cell. 2013;52:124–34.
Samai P, Pyenson N, Jiang W, Goldberg GW, Hatoum-Aslan A, Marraffini LA. Co-transcriptional DNA and RNA cleavage during type III CRISPR-Cas immunity. Cell. 2015;161:1164–74.
Staals RH, Zhu Y, Taylor DW, Kornfeld JE, Sharma K, Barendregt A, et al. RNA targeting by the type III-A CRISPR-Cas Csm complex of Thermus thermophilus. Mol Cell. 2014;56:518–30.
Tamulaitis G, Kazlauskiene M, Manakova E, Venclovas C, Nwokeoji AO, Dickman MJ, et al. Programmable RNA shredding by the type III-A CRISPR-Cas system of Streptococcus thermophilus. Mol Cell. 2014;56:506–17.
Bailey S. The Cmr complex: an RNA-guided endoribonuclease. Biochem Soc Trans. 2013;41:1464–7.
Cocozaki AI, Ramia NF, Shao Y, Hale CR, Terns RM, Terns MP, et al. Structure of the Cmr2 subunit of the CRISPR-Cas RNA silencing complex. Structure. 2012;20:545–53.
Hale CR, Cocozaki A, Li H, Terns RM, Terns MP. Target RNA capture and cleavage by the Cmr type III-B CRISPR-Cas effector complex. Genes Dev. 2014;28:2432–43.
Ramia NF, Spilman M, Tang L, Shao Y, Elmore J, Hale C, et al. Essential structural and functional roles of the Cmr4 subunit in RNA cleavage by the Cmr CRISPR-Cas complex. Cell Rep. 2014;9:1610–7.
Shao Y, Cocozaki AI, Ramia NF, Terns RM, Terns MP, Li H. Structure of the Cmr2-Cmr3 subcomplex of the Cmr RNA silencing complex. Structure. 2013;21:376–84.
Spilman M, Cocozaki A, Hale C, Shao Y, Ramia N, Terns R, et al. Structure of an RNA silencing complex of the CRISPR-Cas immune system. Mol Cell. 2013;52:146–52.
Staals RH, Agari Y, Maki-Yonekura S, Zhu Y, Taylor DW, van Duijn E, et al. Structure and activity of the RNA-targeting type III-B CRISPR-Cas complex of Thermus thermophilus. Mol Cell. 2013;52:135–45.
Taylor DW, Zhu Y, Staals RH, Kornfeld JE, Shinkai A, van der Oost J, et al. Structural biology. Structures of the CRISPR-Cmr complex reveal mode of RNA target positioning. Science. 2015;348:581–5.
Terns RM, Terns MP. The RNA- and DNA-targeting CRISPR-Cas immune systems of Pyrococcus furiosus. Biochem Soc Trans. 2013;41:1416–21.
Zhang J, Rouillon C, Kerou M, Reeks J, Brugger K, Graham S, et al. Structure and mechanism of the CMR complex for CRISPR-mediated antiviral immunity. Mol Cell. 2012;45:303–13.
Zhang J, White MF. Expression and purification of the CMR (type III-B) complex in Sulfolobus solfataricus. Methods Mol Biol. 2015;1311:185–94.
Zhu X, Ye K. Crystal structure of Cmr2 suggests a nucleotide cyclase-related enzyme in type III CRISPR-Cas systems. FEBS Lett. 2012;586:939–45.
Zhu X, Ye K. Cmr4 is the slicer in the RNA-targeting Cmr CRISPR complex. Nucleic Acids Res. 2015;43:1257–67.
Karvelis T, Gasiunas G, Miksys A, Barrangou R, Horvath P, Siksnys V. crRNA and tracrRNA guide Cas9-mediated DNA interference in Streptococcus thermophilus. RNA Biol. 2013;10:841–51.
Barrangou R. Cas9 targeting and the CRISPR revolution. Science. 2014;34:707–8.
Zetsche B, Gootenberg JS, Abudayyeh OO, Slaymaker IM, Makarova KS, Essletzbichler P, et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell. 2015. doi:10.1016/j.cell.2015.09.038.
Heler R, Marraffini LA, Bikard D. Adapting to new threats: the generation of memory by CRISPR-Cas immune systems. Mol Microbiol. 2014;93:1–9.
Heler R, Samai P, Modell JW, Weiner C, Goldberg GW, Bikard D, et al. Cas9 specifies functional viral targets during CRISPR-Cas adaptation. Nature. 2015;519:199–202.
Wei Y, Terns RM, Terns MP. Cas9 function and host genome sampling in type II-A CRISPR-Cas adaptation. Genes Dev. 2015;29:356–61.
Datsenko KA, Pougach K, Tikhonov A, Wanner BL, Severinov K, Semenova E. Molecular memory of prior infections activates the CRISPR/Cas adaptive bacterial immunity system. Nat Commun. 2012;3:945.
Swarts DC, Mosterd C, van Passel MW, Brouns SJ. CRISPR interference directs strand specific spacer acquisition. PLoS One. 2012;7:e35888.
Fineran PC, Gerritzen MJ, Suarez-Diez M, Kunne T, Boekhorst J, van Hijum SA, et al. Degenerate target sites mediate rapid primed CRISPR adaptation. Proc Natl Acad Sci U S A. 2014;111:E1629–38.
Paez-Espino D, Morovic W, Sun CL, Thomas BC, Ueda K, Stahl B, et al. Strong bias in the bacterial CRISPR elements that confer immunity to phage. Nat Commun. 2013;4:1430.
Nunez JK, Kranzusch PJ, Noeske J, Wright AV, Davies CW, Doudna JA. Cas1-Cas2 complex formation mediates spacer acquisition during CRISPR-Cas adaptive immunity. Nat Struct Mol Biol. 2014;21:528–34.
Nunez JK, Lee AS, Engelman A, Doudna JA. Integrase-mediated spacer acquisition during CRISPR-Cas adaptive immunity. Nature. 2015;519:193–8.
Rollie C, Schneider S, Brinkmann AS, Bolt EL, White MF. Intrinsic sequence specificity of the Cas1 integrase directs new spacer acquisition. Elife. 2015. doi:10.7554/eLife.08716.
Sampson TR, Napier BA, Schroeder MR, Louwen R, Zhao J, Chin CY, et al. A CRISPR-Cas system enhances envelope integrity mediating antibiotic resistance and inflammasome evasion. Proc Natl Acad Sci U S A. 2014;111:11163–8.
Sampson TR, Saroj SD, Llewellyn AC, Tzeng YL, Weiss DS. A CRISPR/Cas system mediates bacterial innate immune evasion and virulence. Nature. 2013;497:254–7.
Sampson TR, Weiss DS. Cas9-dependent endogenous gene regulation is required for bacterial virulence. Biochem Soc Trans. 2013;41:1407–11.
Louwen R, Horst-Kreft D, de Boer AG, van der Graaf L, de Knegt G, Hamersma M, et al. A novel link between Campylobacter jejuni bacteriophage defence, virulence and Guillain-Barre syndrome. Eur J Clin Microbiol Infect Dis. 2013;32:207–26.
Louwen R, Staals RH, Endtz HP, van Baarlen P, van der Oost J. The role of CRISPR-Cas systems in virulence of pathogenic bacteria. Microbiol Mol Biol Rev. 2014;78:74–88.
Westra ER, Buckling A, Fineran PC. CRISPR-Cas systems: beyond adaptive immunity. Nat Rev Microbiol. 2014;12:317–26.
Levy A, Goren MG, Yosef I, Auster O, Manor M, Amitai G, et al. CRISPR adaptation biases explain preference for acquisition of foreign DNA. Nature. 2015;520:505–10.
Yosef I, Goren MG, Qimron U. Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli. Nucleic Acids Res. 2012;40:5569–76.
Paez-Espino D, Sharon I, Morovic W, Stahl B, Thomas BC, Barrangou R, et al. CRISPR immunity drives rapid phage genome evolution in Streptococcus thermophilus. MBio. 2015;6:e00262–15.
Jiang W, Marraffini LA. CRISPR-Cas: New tools for genetic manipulations from bacterial immunity systems. Annu Rev Microbiol. 2015. doi:10.1146/annurev-micro-091014-104441.
Selle K, Barrangou R. Harnessing CRISPR-Cas systems for bacterial genome editing. Trends Microbiol. 2015;23:225–32.
Barrangou R, Horvath P. CRISPR: new horizons in phage resistance and strain identification. Annu Rev Food Sci Technol. 2012;3:143–62.
Briner AE, Barrangou R. Lactobacillus buchneri genotyping on the basis of clustered regularly interspaced short palindromic repeat (CRISPR) locus diversity. Appl Environ Microbiol. 2014;80:994–1001.
Briner AE, Lugli GA, Milani C, Duranti S, Turroni F, Gueimonde M, et al. Occurrence and diversity of CRISPR-Cas systems in the genus Bifidobacterium. PLoS One. 2015;10:e0133661.
Barrangou R, Briczinski EP, Traeger LL, Loquasto JR, Richards M, Horvath P, et al. Comparison of the complete genome sequences of Bifidobacterium animalis subsp. lactis DSM 10140 and Bl-04. J Bacteriol. 2009;191:4144–51.
Broadbent JR, Neeno-Eckwall EC, Stahl B, Tandee K, Cai H, Morovic W, et al. Analysis of the Lactobacillus casei supragenome and its influence in species evolution and lifestyle adaptation. BMC Genomics. 2012;13:533.
Sun CL, Barrangou R, Thomas BC, Horvath P, Fremaux C, Banfield JF. Phage mutations in response to CRISPR diversification in a bacterial population. Environ Microbiol. 2013;15:463–70.
Sun CL, Thomas BC, Barrangou R, Banfield JF. Metagenomic reconstructions of bacterial CRISPR loci constrain population histories. ISME J. 2015. doi:10.1038/ismej.2015.162.
Weinberger AD, Sun CL, Plucinski MM, Denef VJ, Thomas BC, Horvath P, et al. Persisting viral sequences shape microbial CRISPR-based immunity. PLoS Comput Biol. 2012;8:e1002475.
Emerson JB, Andrade K, Thomas BC, Norman A, Allen EE, Heidelberg KB, et al. Virus-host and CRISPR dynamics in Archaea-dominated hypersaline Lake Tyrrell, Victoria, Australia. Archaea. 2013;2013:370871.
Heidelberg JF, Nelson WC, Schoenfeld T, Bhaya D. Germ warfare in a microbial mat community: CRISPRs provide insights into the co-evolution of host and viral genomes. PLoS One. 2009;4:e4169.
Andersson AF, Banfield JF. Virus population dynamics and acquired virus resistance in natural microbial communities. Science. 2008;320:1047–50.
Tyson GW, Banfield JF. Rapidly evolving CRISPRs implicated in acquired resistance of microorganisms to viruses. Environ Microbiol. 2008;10:200–7.
Stern A, Mick E, Tirosh I, Sagy O, Sorek R. CRISPR targeting reveals a reservoir of common phages associated with the human gut microbiome. Genome Res. 2012;22:1985–94.
Rho M, Wu YW, Tang H, Doak TG, Ye Y. Diverse CRISPRs evolving in human microbiomes. PLoS Genet. 2012;8:e1002441.
Naidu M, Robles-Sikisaka R, Abeles SR, Boehm TK, Pride DT. Characterization of bacteriophage communities and CRISPR profiles from dental plaque. BMC Microbiol. 2014;14:175.
Pride DT, Sun CL, Salzman J, Rao N, Loomer P, Armitage GC, et al. Analysis of streptococcal CRISPRs from human saliva reveals substantial sequence diversity within and between subjects over time. Genome Res. 2011;21:126–36.
Barrangou R, Coute-Monvoisin AC, Stahl B, Chavichvily I, Damange F, Romero DA, et al. Genomic impact of CRISPR immunization against bacteriophages. Biochem Soc Trans. 2013;41:1383–91.
Bikard D, Hatoum-Aslan A, Mucida D, Marraffini LA. CRISPR interference can prevent natural transformation and virulence acquisition during in vivo bacterial infection. Cell Host Microbe. 2012;12:177–86.
Barrangou R, Klaenhammer TR. Microbiology: Bacteria get vaccinated. Nature. 2014;513:175–6.
Bikard D, Jiang W, Samai P, Hochschild A, Zhang F, Marraffini LA. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucleic Acids Res. 2013;41:7429–37.
Luo ML, Mullis AS, Leenay RT, Beisel CL. Repurposing endogenous type I CRISPR-Cas systems for programmable gene repression. Nucleic Acids Res. 2015;43:674–81.
Rath D, Amlinger L, Hoekzema M, Devulapally PR, Lundgren M. Efficient programmable gene silencing by Cascade. Nucleic Acids Res. 2015;43:237–46.
Hilton IB, D'Ippolito AM, Vockley CM, Thakore PI, Crawford GE, Reddy TE, et al. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat Biotechnol. 2015;33:510–7.
Hilton IB, Gersbach CA. Enabling functional genomics with genome engineering. Genome Res. 2015;25:1442–55.
Oh JH, van Pijkeren JP. CRISPR-Cas9-assisted recombineering in Lactobacillus reuteri. Nucleic Acids Res. 2014;42:e131.
van Pijkeren JP, Britton RA. Precision genome engineering in lactic acid bacteria. Microb Cell Fact. 2014;13 Suppl 1:S10.
Peters JM, Silvis MR, Zhao D, Hawkins JS, Gross CA, Qi LS. Bacterial CRISPR: accomplishments and prospects. Curr Opin Microbiol. 2015;27:121–6.
Selle K, Klaenhammer TR, Barrangou R. CRISPR-based screening of genomic island excision events in bacteria. Proc Natl Acad Sci U S A. 2015;112:8076–81.
Gomaa AA, Klumpe HE, Luo ML, Selle K, Barrangou R, Beisel CL. Programmable removal of bacterial strains by use of genome-targeting CRISPR-Cas systems. MBio. 2014;5:e00928–00913.
Bikard D, Euler CW, Jiang W, Nussenzweig PM, Goldberg GW, Duportet X, et al. Exploiting CRISPR-Cas nucleases to produce sequence-specific antimicrobials. Nat Biotechnol. 2014;32:1146–50.
Citorik RJ, Mimee M, Lu TK. Sequence-specific antimicrobials using efficiently delivered RNA-guided nucleases. Nat Biotechnol. 2014;32:1141–5.
Beisel CL, Gomaa AA, Barrangou R. A CRISPR design for next-generation antimicrobials. Genome Biol. 2014;15:516.
Yosef I, Manor M, Kiro R, Qimron U. Temperate and lytic bacteriophages programmed to sensitize and kill antibiotic-resistant bacteria. Proc Natl Acad Sci U S A. 2015;112:7267–72.
Anders C, Jinek M. In vitro enzymology of Cas9. Methods Enzymol. 2014;546:1–20.
Nishimasu H, Ran FA, Hsu PD, Konermann S, Shehata SI, Dohmae N, et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell. 2014;156:935–49.
Nishimasu H, Cong L, Yan WX, Ran FA, Zetsche B, Li Y, et al. Crystal structure of Staphylococcus aureus Cas9. Cell. 2015;162:1113–26.
Briner AE, Donohoue PD, Gomaa AA, Selle K, Slorach EM, Nye CH, et al. Guide RNA functional modules direct Cas9 activity and orthogonality. Mol Cell. 2014;56:333–9.
van der Oost J, Westra ER, Jackson RN, Wiedenheft B. Unravelling the structural and mechanistic basis of CRISPR-Cas systems. Nat Rev Microbiol. 2014;12:479–92.
Chylinski K, Le Rhun A, Charpentier E. The tracrRNA and Cas9 families of type II CRISPR-Cas immunity systems. RNA Biol. 2013;10:726–37.
Chylinski K, Makarova KS, Charpentier E, Koonin EV. Classification and evolution of type II CRISPR-Cas systems. Nucleic Acids Res. 2014;42:6091–105.
Fonfara I, Le Rhun A, Chylinski K, Makarova KS, Lecrivain AL, Bzdrenga J, et al. Phylogeny of Cas9 determines functional exchangeability of dual-RNA and Cas9 among orthologous type II CRISPR-Cas systems. Nucleic Acids Res. 2014;42:2577–90.
Esvelt KM, Mali P, Braff JL, Moosburner M, Yaung SJ, Church GM. Orthogonal Cas9 proteins for RNA-guided gene regulation and editing. Nat Methods. 2013;10:1116–21.
Kleinstiver BP, Prew MS, Tsai SQ, Topkar VV, Nguyen NT, Zheng Z, et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature. 2015;523:481–5.
Acknowledgments
The author is supported by start-up funds from North Carolina State University, and the North Carolina Biotechnology Center. The author acknowledges his many colleagues and collaborators in the CRISPR field for their contributions, colourful conversations and insights into these fascinating systems.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
RB is a cofounder, shareholder and advisor of Intellia Therapeutics, and is a director, shareholder and advisor of Caribou Biosciences. RB is also an inventor on several patents related to CRISPR-Cas systems and their various uses.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Barrangou, R. Diversity of CRISPR-Cas immune systems and molecular machines. Genome Biol 16, 247 (2015). https://doi.org/10.1186/s13059-015-0816-9
Published:
DOI: https://doi.org/10.1186/s13059-015-0816-9