Abstract
Background
Among the cytochrome P450 enzymes (CYP), families 1-3 constitute almost half of total CYPs in mammals and play a central role in metabolism of a wide range of pharmaceuticals. This study investigated gene and protein expression and cellular localisation of CYP1A, CYP2A, CYP2C, CYP2D and CYP2E in equine intestine and liver. Real-time polymerase chain reaction (RT-PCR) was used to analyse gene expression, western blot to examine protein expression and immunohistochemical analyses to investigate cellular localisation.
Results
CYP1A and CYP2C were the CYPs with the highest gene expression in the intestine and also showed considerable gene expression in the liver. CYP2E and CYP2A showed the highest gene expression in the liver. CYP2E showed moderate intestinal gene expression, whereas that of CYP2A was very low or undetectable. For CYP2D, rather low gene expression levels were found in both intestine and the liver. In the intestine, CYP gene expression levels, except for CYP2E, exhibited patterns resembling those of the proteins, indicating that intestinal protein expression of these CYPs is regulated at the transcriptional level. For CYP2E, the results showed that the intestinal gene expression did not correlate to any visible protein expression, indicating that intestinal protein expression of this CYP is regulated at the post-transcriptional level. Immunostaining of intestine tissue samples showed preferential CYP staining in enterocytes at the tips of intestinal villi in the small intestine. In the liver, all CYPs showed preferential localisation in the centrilobular hepatocytes.
Conclusions
Overall, different gene expression profiles were displayed by the CYPs examined in equine intestine and liver. The CYPs present in the intestine may act in concert with those in the liver to affect the oral bioavailability and therapeutic efficiency of substrate drugs. In addition, they may play a role in first-pass metabolism of feed constituents and of herbal supplements used in equine practice.
Similar content being viewed by others
Background
Among the cytochrome P450 enzymes (CYP), families 1-3 constitute almost half of total CYPs in mammals. These are generally designated xenobiotic-metabolising enzymes and play a central role in metabolism of a wide range of xenobiotics [1]. The xenobiotic-metabolising CYPs are mainly expressed in the liver, but several are also expressed in extrahepatic tissues, particularly in organs in direct contact with xenobiotics, such as the intestines and the respiratory tissues [2],[3]. Studies in different species have shown that each tissue may have its unique CYP expression pattern [2],[3].
To date, three equine CYPs within families 1-3 have been cloned and functionally characterised. These are CYP2D50 [4], CYP2C92 [5] and CYP3A96 [6]. A substantial difference has been reported in the metabolic capacity of these equine CYPs compared with the human orthologs. For example, the metabolic capacity of CYP2D50, CYP2C92 and CYP3A96 is about 20- to 180-fold lower with regard to dextromethon-O-demethylation, diclofenac-oxidation and testosterone-6β-hydroxylation, respectively, than that of the human orthologs, CYP2D6, CYP2C9 and CYP3A4 [4]-[6].
There has been a continual increase in the volume of sales of drugs used in equine practice [7] and many drugs used in equine therapy are substrates for CYPs. There is also an increasing retail market for herbal supplements for horses [8]. However, little is known about drug-drug interactions and drug-herb interactions in the horse. Due to differences in expression level and substrate specificity of the CYP enzymes, it is difficult to extrapolate findings on the pharmacokinetics and pharmacodynamics of drugs between species [9],[10]. Thus there is clearly a need for increased knowledge of the expression pattern and metabolic capacity of xenobiotic-metabolising CYP enzymes in the horse.
Previous studies by our group have examined expression of isoenzymes belonging to the CYP3A subfamily in the equine intestine and liver [11],[12]. In the intestinal mucosa, CYP3A is considered to play an important role for the oral availability of substrate drugs and other xenobiotics, by sequential CYP3A-mediated metabolism in the intestine and liver [12],[13]. However, studies in man and rodents have shown that, in addition to the CYP3A subfamily, other CYP subfamilies are also present in the intestinal mucosa and may play a role in modulating drug and xenobiotic bioavailability [13],[14].
In the present study, we examined gene and protein expression and cellular localisation of CYPs of the 1A, 2A, 2C, 2D and 2E subfamilies in the intestine and liver of the horse. Gene expression was examined using quantitative real-time reverse transcriptase polymerase chain rraction (RT-qPCR). For the subfamilies CYP2C and CYP2D primers were designed based on conserved gene regions, thus enabling amplification of all isoenzymes belonging to these subfamilies. For the subfamilies CYP1A, CYP2A and CYP2E, primers were designed based on a single sequence. The reason for this was that these were the only sequences identified for these equine subfamilies at the time when this study was performed. Protein expression was examined by western blot. Immunohistochemical analysis was used to examine the cellular localisation of the CYPs in the intestine and liver of the horse.
Methods
Tissue sample collection
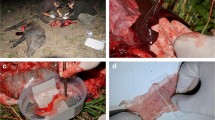
Tissues from healthy Swedish standard-bred trotters (n =8) were collected at a local slaughterhouse. The tissues were from 5- to 18-year-old mares and geldings. In accordance with Swedish National Food Administration regulations, the horses were examined by a veterinarian before slaughter. In addition, the passports of the horses were checked to ensure that all horses in the study were free of medication during the stipulated period prior to slaughter. Samples were collected from the liver and various parts of the intestines (Table 1), and tissue preparation was performed as described in [15]. In brief, for RNA analysis the mucosal surface was scraped smoothly with a scalpel, and samples were collected in Rnase-free plastic tubes and snap-frozen in liquid nitrogen. Liver tissue was treated in a similar way. Samples for protein expression were collected as described for RNA samples and transported to the laboratory in 0.9% NaCl at 4°C. Samples for immunohistochemical analysis (0.5 cm × 0.5 cm) were transported to the laboratory in 0.9% NaCl at 4°C and thereafter fixed in 4% paraformaldehyde.
Isolation of RNA
Total RNA was prepared using the NucleoSpin RNA II kit containing deoxyribonuclease I (DNase I) (Macherey-Nagel, Düren, Germany) according to the manufacturer’s instructions. The purity and integrity of the RNA were checked as described previously [16]. In brief, prior to real-time RT-PCR analysis the integrity of RNA was verified by examining ribosomal RNA 28 S and 18 S on 1% agarose gel containing 18% formaldehyde. Only RNA samples with a 260/280 nm ratio exceeding 1.8 were selected for real-time PCR. The exact amount of RNA was quantified using the RNA-specific Quant-iT RiboGreen protocol (Molecular Probes, Eugene, OR, USA) and a microplate reader (Wallac 1420 VICTOR2TM, software version 2.0, Turku, Finland).
CYP gene expression
The gene sequences of the CYP2D subfamily were predicted by NCBI Map Viewer according to Yasukochi et al. [17]. The genes of the CYP2C subfamily were derived from drnelson homepage [18]. All other CYP genes were derived from the NCBI genome database (http://www.ncbi.nlm.nih.gov/). Primers for the genes of the CYP2C and CYP2D subfamilies were designed based on conserved regions, thus enabling amplification of all isoenzymes belonging to the same subfamily (Table 2). Primers for CYP1A, CYP2A and CYP2E were designed based on one sequence (Table 2). The primers were designed in Primer 3 (v. 0.4.0) [19] and synthesised by CyberGene AB, Stockholm, Sweden. Gene expression was examined by quantitative reverse transcriptase-polymerase chain reaction (RT-qPCR) using Rotor-Gene 3000 (Corbett Research, Mortlake, Australia) by applying a one-step QuantiTect®-SYBR®green RT-PCR kit (Qiagen, Inc., Valencia, CA, USA) according to the manufacturer’s recommendations. The primer concentration was 0.4 μM and 300 ng total RNA were used. Copy number was determined using a standard curve derived from serial dilutions of known concentrations of the purified PCR products in the customary way. Copy number was calculated using the following formula: molecules μL-1 = (A × 6.022 × 1023) (660 × B)-1, where A is the concentration of the PCR product (g μL-1), B is the plasmid length, 6.022 × 1023 is Avogadro’s number, and 660 is the average molecular weight of one base pair. The amplification efficiency was calculated as (10(-1/slope) - 1) × 100. The terminology used was in accordance with Bustin et al. [20]. The standard curve was created from purified PCR product stock solution that was serially diluted to give: 2×107, 2×106, 2×105, 2×104, 2×103 copies μL-1. This was repeated several times to test the efficacy and reproducibility of the qPCR. Standard curves were created automatically and accepted when the slope was on average -3.69 ± 0.05 (97% efficacy) and the correlation coefficient (r2) was 1.00. The specificity of the PCR products obtained was checked in connection with each real-time RT-PCR run, using melt curve analysis. In addition, the PCR products were validated by checking the specificity and size on 1% agarose gel. The PCR products generated were sequenced at the Centre for Genomic Research, Karolinska Institute, Stockholm, Sweden.
CYP protein expression
Preparation of microsomes and determination of protein concentration were performed as described previously [15]. In brief, tissues were transported from the slaughterhouse in 0.9% NaCl at 4°C and within 40 min the samples (30 g) were homogenised in 10 mL cold 0.01 m KH2PO4/0.32 m KCl buffer, pH 7.4, containing 20% glycerol (vol/vol) and Complete EDTA-free Protease Inhibitor Cocktail Tablets (Roche Diagnostics GmbH, Mannheim, Germany). This was followed by three centrifugation steps; one at 10 000 × g for 20 min and two at 105 000 × g for 60 min. Aliquots of 50 μg of microsomal protein were separated on 10% Tris-HCl polyacrylamide gels under reducing conditions, as described previously [15]. Separated proteins were electroblotted to nitrocellulose membranes and transfer of proteins was confirmed by staining with Ponceau S (Sigma-Aldrich, St. Louis, MO, USA). Membranes were blocked in 10% dry milk and incubated with primary antibodies (Table 3), diluted in phosphate-buffered saline (PBS) containing 0.1% Tween-20 (PBST). For CYP1A, CYP2A, CYP2D and CYP2E, a secondary goat-anti-rabbit antibody (AQ132P, Chemicon International, Temecula, CA, USA) conjugated with horseradish peroxidase (HRP) was used, diluted 1:50 000 in 2% advanced blocking buffer (GE Healthcare, Uppsala, Sweden). HRP was detected by Enhanced Chemiluminescence (ECL) western blot Detection reagents (GE Healthcare, Uppsala, Sweden). For CYP2C, a secondary mouse-anti-sheep antibody (213-032-177, Jackson ImmunoResearch Laboratories, West Grove, PA, USA) was used and diluted 1:1000 as described above. The ECL reagents were used according to the manufacturer’s instructions. The Chemi-Doc Gel Quantification System (Bio-Rad, Hercules, CA, USA) and Quantity-One software were applied to detect the intensity of the bands. The blotting membranes were stripped using 65 mM Tris–HCl, 100 mM 2-mercaptoethanol, 2% SDS for 5 min at 55-60°C, rinsed three times with PBST and thereafter incubated with the next primary and secondary antibodies.
Immunohistochemical analysis
Pieces of intestine and liver tissue were fixed in formaldehyde, dehydrated and embedded in paraffin. From these, 5 μm tissue sections were taken and deparaffinised, hydrated and rinsed in phosphate-buffered saline (PBS). The rinsing procedure, involving two rinsing intervals in PBS for 10 min, was repeated after each of the steps described below. Endogenous peroxidase activity was blocked with 1.6% H2O2 in PBS, followed by incubation for 1 hour in PBS with 10% normal goat serum (for CYP1A, CYP 2A, CYP 2D and CYP 2E) or normal rabbit serum (for CYP2C). Endogenous avidin and biotin were blocked using the Avidin/Biotin Blocking kit (Vector Laboratories Inc., Burlingame, CA, USA). Thereafter, the tissue sections were incubated overnight at 4°C with the primary antibodies diluted in PBS as shown in Table 3. The secondary antibodies (goat-anti-rabbit or rabbit-anti-sheep; Vector Laboratories Inc., Burlingame, CA, USA) were used at 1:200 dilution and applied for 1 h at room temperature. The antigen-antibody complex was conjugated with avidin-biotin peroxidase using the ABC Vectastatin kit and then visualised with DAB staining, according to the supplier’s recommendations (Dako, Glostrup, Denmark). Finally, the sections were counterstained with haematoxylin (VWR International AB, Stockholm, Sweden). Negative controls were run in parallel with non-immune rabbit (CYP1A, 2A, 2E, 2D) or sheep (CYP2C) IgG (Abcam, Cambridge, UK) in amounts equivalent to those used for the antibodies against the CYP enzymes.
Statistical analysis
All statistical analyses were performed using Minitab® software, version 15 (Minitab Inc., State College, PA, USA). Levene’s test was used to test whether the data followed a normal distribution. A paired t-test or a one-way analysis of variance (ANOVA) was performed to test for differences between groups. Dunnett’s test was used as a post hoc test in combination with ANOVA. Differences were considered significant at P < 0.05.
Results
CYP gene expression
For CYP2A, the gene expression level was very low or even undetectable in all parts of the intestine, whereas gene expression in the liver was high for this CYP (Table 4). For the other CYPs, the CYP expression levels were higher in all parts of the small intestine (duodenum, jejunum and ileum) than in the large intestine (caecum and colon). CYP1A and CYP2C showed the highest intestinal gene expression. Both these CYPs also showed considerable hepatic gene expression levels. CYP2E showed moderate intestinal gene expression and high hepatic gene expression. For CYP2D, the gene expression levels were rather low in both intestine and liver.
Statistical analysis of the differences between the hepatic and small intestinal gene expression levels revealed that CYP2A and CYP2E had significantly higher hepatic than intestinal gene expression (P < 0.05). For CYP1A, CYP2C and CYP2D, there were no statistical differences between gene expression levels in the small intestine and liver.
CYP protein expression
Representative western blots of microsomal proteins in the liver resulted in detection of immunopositive bands at 56-58 kDa for all CYPs (Figure 1). For CYP1A, CYP2C and CYP2D, immunopositive bands of this size were seen in the small intestine, whereas in the large intestine such bands were either absent or barely detectable (Figure 1). For CYP2E, it was not possible to clearly detect immunopositive bands at 56-58 kDa in any part of the intestine. For CYP2A, the western blots showed a complete lack of intestinal immunostaining.
Immunohistochemistry
For CYP1A, CYP2C and CYP2D, the immunohistochemical analysis showed marked immunostaining of the enterocytes of the small intestine (duodenum, jejunum and ileum) and weak immunostaining of the enterocytes of the large intestine (caecum and colon). There was no detectable intestinal immunostaining for CYP2A and CYP2E. The immunostaining observed for CYP1A, CYP2C and CYP2D was strongest in enterocytes at the tips of intestinal villi in the small intestine, with preferential localisation in the outer parts of the cytoplasm of the enterocytes, towards the intestinal lumen (Figure 2A-C). Goblet cells did not stain and there was no immunostaining in the lamina propria or muscularis mucosae. In the liver, immunostaining was seen for all CYPs, with the strongest staining in hepatocytes in central parts of the hepatic lobuli (Figure 2D).
Cellular CYP-localisation in the intestines and the liver. Immunohistochemical staining of CYP1A (A), CYP2C (B), CYP2D (C) and CYP2E (D) enzymes in the duodenum (C), jejunum (A,B), and liver (D) in horse. The immunoreactivity of the CYP antibodies, indicated by arrows, are shown in brown colour. The nuclei of the cells blue colored with Heamatoxylin are easily seen. Legends: cv, central vein; e, enterocyte; g, goblet cell; lp, lamina propia; mv, microvilli of enterocytes. Magnifications are indicated by bars.
Discussion
This study showed that gene expression levels in equine intestine and liver usually display particular patterns for the different CYP isoforms. In the intestine, gene expression was high for CYP2C and CYP1A, low for CYP2E and CYP2D and almost undetectable for CYP2A. In the liver, gene expression was high for CYP2A and CYP2E, moderate for CYP2C and CYP1A and low for CYP2D. The intestinal gene expression of the CYPs examined in the present study was higher in the small intestine than in the large intestine. This was also observed for CYP3A in our previous studies in the horse [11],[12]. Similar distribution patterns in gene expression of CYPs along the gastrointesinal tract have been observed in other species [13],[21],[22].
The present study demonstrated high levels of gene expression for CYP1A and CYP2C in the equine small intestine. As mentioned, our previous studies have shown that CYP3A is also highly expressed in equine intestine [11],[12]. CYP3A and CYP2C represent the major CYPs expressed in human intestinal mucosa, but this is not generally the case for CYP1A [13],[23]. It has been shown that CYP1A1 expression in the human intestine is highly inducible, whereas there is consitutive expression of CYP3A and CYP2C [23]. The high expression of CYP1A observed here in the equine intestine can be related to the presence of CYP1A-inducing components in the diet of the horse. These differences in CYP isoform expression in the intestine must be considered when extrapolating data between species.
The CYP2D gene and protein expression was found to be low in both the liver and intestine of the horse. Yasukoch & Satta [17] examined the evolution of the CYP2D gene cluster and found that the number of members within the CYP2D subfamily varies between species. For example, primates have two to three CYP2D genes, whereas rodents, rabbits and horses have seven, five and six CYP2D genes, respectively. It has been suggested that the expansion of members within the CYP2D subfamily in herbivores might be related to the fact that several plant toxins are substrates for the CYP2D enzyme [17],[24]. It is interesting that the expansion of the CYP2D subfamily in the equine genome was not accompanied by prominent gene or protein expression in the equine liver and intestine in the present study. The human CYP2D6 is known to be a polymorphic CYP isoenzyme with highly variable gene expression [25].
CYP2A and CYP2E were highly expressed in the equine liver, but both had very low gene expression levels in the intestine. This is consistent with reported levels in humans, where CYP2E1 is highly expressed in the liver, but only weakly expressed in the intestine for review see [26]. Bièche et al.[27] have shown high hepatic gene expression and very low intestinal gene expression for the three members of the human CYP2A subfamily.
Our present and prevous studies have shown that the gene expression levels of CYP1A, CYP2C and CYP3A in the equine small intestine were comparable to those in the liver. These results differ from observations in humans and dogs, in which the CYP expression levels in the liver are generally much higher compared than those in the small intestine [27],[28]. It is possible that the high levels of CYPs in the equine intestine relate to the fact that the horse is a herbivorous species, which means that the diet may contain various CYP-inducing substrates, including phytonutrients and phytotoxicants. Consequently, during their evolution horses may have developed a more effective intestinal CYP system than omnivores or carnivores such as humans and dogs.
In the equine intestine and the liver, the CYP gene expression levels, except for CYP2E, exhibited expression patterns resembling those of the proteins, as shown by western blot analysis (Figure 1). This confirms findings in other species indicating that CYPs in general are regulated at the transcriptional level [29],[30]. As regards CYP2E, our results showed that the intestinal gene expression detected in the PCRs did not correlate to any clearly detectable CYP2E protein expression in the western blots. This indicates that the protein expression of CYP2E is regulated at the post-transcriptional level. Similarly, studies with human liver biopsies have shown that the mRNA levels for CYP2E1 do not correlate to the CYP2E1 protein levels [31]. In addition, studies by Rodriguez-Antona et al.[32] have shown that there is no significant correlation between CYP2E mRNA expression and CYP2E-related metabolic activity in human liver samples.
Our immunohistochemical analyses showed that for the CYPs for which intestinal immunostaining was observed (CYP1A, CYP2C and CYP2D), there was preferential localisation of the staining in the enterocytes at the tips of the villi in the small intestine. We have previously shown that this staining pattern also applies for CYP3A in the equine intestine [11]. Similar findings have been made in other species [33],[34]. In the liver, marked immunostaining was seen for all CYPs, with the strongest staining in hepatocytes in central parts of the hepatic lobuli. These results also corroborate those in other species (for review see [35]).
Many CYPs have been shown to be metabolically active in horses and, on the whole, oxidative drug metabolism appears more extensive in horses than in man [36]. Many drugs used in equine therapy, such as quinolones [37], dexamethasone [38], ivermectin [39],[40], benzimidazoles [41],[42], ketamine [43], meloxicam [44], omeprazole [45], phenylbutazone [46], praziquantel [47],[48] and pyrantel [47], are substrates for the CYP enzymes. Several herbal supplements used in equine practice have also been reported to be CYP substrates. Examples are quercetin, the active component in devil’s claw root [49]; ginsengoides, the active components in ginseng [49]; and silymarin, the active component in meadowsweet [50]. It is also known that CYP-inducible components, such as flavonoids [51], are present in the normal diet of the horse, which may indicate that the equine CYPs have been strongly subjected to positive selection. It is apparent that there is a need for further studies on the expression patterns, metabolic capacities and inducibility of CYP enzymes in the horse.
Conclusions
This study demonstrated differing gene and protein expression profiles of the five CYPs studied in equine intestine and liver. The CYPs present in the intestine may act in concert with those in the liver to affect the oral bioavailability and therapeutic efficiency of substrate drugs. In addition, they may play a role in first-pass metabolism of equine feed constituents and of herbal supplements used in equine practice.
Authors’ contributions
ET, PL, HT participated in the design of the study, collected the materials. ET performed the laboratory analyses. PL performed the immunohistochemistry. All authors participated in interpreting the data and drafting the manuscript. All authors read and approved the final manuscript.
References
Nebert DW, Russell DW: Clinical importance of the cytochromes P450. Lancet. 2002, 360: 1155-1162. 10.1016/S0140-6736(02)11203-7.
Ding X, Kaminsky LS: Human extrahepatic cytochromes P450: functiona in xenobiotic metabolism and tissue-selective chemical toxicity in the respiratory and gastrointestinal tracts. Annu Rev Pharmacol Toxicol. 2003, 43: 149-173. 10.1146/annurev.pharmtox.43.100901.140251.
Tydén E, Löfgren M, Hakhverdyan M, Tjälve H, Larsson P: The genes of all seven CYP3A isoenzymes identified in the equine genome are expressed in the airways of horses. J Vet Pharmacol Ther. 2013, 36: 370-375. 10.1111/jvp.12012.
Knych HK, Stanley SD: Complementary DNA cloning, functional expression and characterization of a novel cytochrome P450, CYP2D50, from equine liver. Biochem Pharmacol. 2008, 76: 904-911. 10.1016/j.bcp.2008.07.016.
Knych HK, DeStefano SC, Buckpitt AB, Stanley SD: Equine cytochrome P450 2C92: cDNA cloning, expression and initial characterization. Arch Biochem Biophys. 2009, 485: 49-55. 10.1016/j.abb.2009.02.009.
Knych HK, McKemie S, Stanley D: Molecular cloning, expression, and initial characterization of members of the CYP3A family in horses. Drug Metab Disp. 2010, 38: 1820-1827. 10.1124/dmd.110.032953.
Girma K: Försäljningsutveckling av djurläkemedel 2011. [Sales development of veterinary drugs 2011].Board of Agriculture 2012, 7-10.,
Williams CA, Lamprecht ED: Some commonly fed herbs and other functional foods in equine nutrition: a review. Vet J. 2008, 178: 21-31. 10.1016/j.tvjl.2007.06.004.
Nebbia C, Dacasto M, Rossetto Giaccherino A, Giuliano A, Carletti M: Comparative expression of liver cytochrome P450-dependent monooxygenases in the horse and in other agricultural and laboratory species. Vet J. 2003, 165: 53-64. 10.1016/S1090-0233(02)00174-0.
Fink-Gremmels J: Implications of hepatic cytochrome P450-related biotransformation processes in veterinary sciences. Eur J Clin Pharm. 2008, 585: 502-509. 10.1016/j.ejphar.2008.03.013.
Tydén E, Olsen L, Tallkvist J, Larsson P, Tjälve H: CYP3A in horse intestines. Toxicol Appl Pharm. 2004, 201: 112-119. 10.1016/j.taap.2004.05.015.
Tydén E, Löfgren M, Pegolo S, Capolongo F, Tjälve H, Larsson P: Differential gene expression of CYP3A isoforms in equine liver and intestines. J Vet Pharmacol Ther. 2012, 35: 588-595. 10.1111/j.1365-2885.2012.01379.x.
Paine MF, Hart HL, Ludington SS, Haining RL, Rettie AE, Zeldin DC: The human intestinal cytochrome P450 "pie". Drug Metab Dispos. 2006, 34: 880-886. 10.1124/dmd.105.008672.
Mitschke D, Reichel A, Fricker G, Moenning U: Characterization of cytochrome P450 protein expression along the entire length of the intestine of male and female rats. Drug Metab Dispos. 2008, 36: 1039-1045. 10.1124/dmd.107.019687.
Tydén E, Olsen L, Tallkvist J, Tjälve H, Larsson P: Cytochrome P450 3A, NADPH cytochrome P450 reductase and cytochrome b5 in the upper airways in horse. Res Vet Sci. 2008, 85: 80-85. 10.1016/j.rvsc.2007.09.012.
Tydén E, Tallkvist J, Tjälve H, Larsson P: P-glycoprotein in intestines, liver, kidney and lymphocytes in horse. J Vet Pharmacol Ther. 2009, 32: 167-176. 10.1111/j.1365-2885.2008.01017.x.
Yasukochi Y, Satta S: Evolution of the CYP2D gene cluster in humans and four non-human primates. Genes Genet Syst. 2011, 86: 109-116. 10.1266/ggs.86.109.
Nelson DR: The cytochrome P450 homepage. Hum Genom. 2009, 4: 59-65.
Rozen S, Skaletsky HJ: Primer3 on the WWW for general users and for biologist programmers. Bioinformatics Methods and Protocols. Methods in Molecular Biology Humana Press Totowa 2000. Edited by: Krawetz S, Misener S. 2000, 365-386. software version 4.0.http://primer3.ut.ee/, [http://primer3.ut.ee/]
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT: The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem. 2009, 55: 611-622. 10.1373/clinchem.2008.112797.
Zhang Q-Y, Dunbar D, Ostrowska A, Zeisloft S, Yang J, Kaminsky LS: Characterization of human small intestinal cytochromes P-450. Drug Metab Disp. 1999, 27: 804-809.
Shin HC, Kim HR, Cho HJ, Yi H, Cho SM, Lee DG, El-Aty AM, Kim JS, Sun D, Amidon GL: Comparative gene expression of intestinal metabolizing enzymes. Biopharm Drug Disp. 2009, 30: 411-421. 10.1002/bdd.675.
Buchthal J, Grund KE, Buchman A, Schrenk D, Beaume P, Bock KW: Induction of cytochrome P4501A by smoking or omeprazole in comparison with UDP-glucuronsyltansferase in biopsies of human duodenal mucosa. Eur J Clin Pharmacol. 1995, 47: 431-435. 10.1007/BF00196857.
Fonne-Pfister R, Meyer UA: Xenobiotic and endobiotic inhibitors of cytochrome P-450dbl function, the target of the debrisoquine/sparteine type polymorphism. Biochem Pharmacol. 1988, 37: 3829-3835. 10.1016/0006-2952(88)90063-9.
Ingelman-Sundberg M: Genetic polymophism of cytochrome P450 D6 (CYP2D6): clinical consequences, evolutionary aspects and functional diveristy. Pharmacogenomics. 2005, 5: 6-13. 10.1038/sj.tpj.6500285.
Zanger UM, Schwab M: Cytochrome P450 enzymes in drug metabolism: Regulation of gene expression, enzyme activities, and impact of genetic varation. Pharmacol Therapeut. 2013, 138: 103-141. 10.1016/j.pharmthera.2012.12.007.
Bièche I, Narjoz C, Asselah T, Vacher S, Marcellin P, Lidereau R, Beaune P, de Waziers I: Reverse transcriptase-PCR quantification of mRNA levels from cytochrome (CYP)1, CYP2 and CYP3 families in 22 different human tissues. Pharmacogenet Genom. 2007, 17: 731-742. 10.1097/FPC.0b013e32810f2e58.
Mealey KL, Jabbes M, Spencer E, Akey JM: Differential expression of CYP3A12 and CYP3A26 mRNAs in canine liver and intestine. Xenobiotica. 2008, 38: 1305-1312. 10.1080/00498250802446146.
Gonzales F, Lin S, Yano M: Regulation of cytochrome P450 genes: molecular mechanisms. Pharmacogenet. 1993, 3: 51-57. 10.1097/00008571-199302000-00006.
Dogra S, Whitelaw M, May B: Transcriptional activation of cytochrome p450 genes by different classes of chemical inducers. Clin Exp Pharmacol Physiol. 1998, 25: 1-9. 10.1111/j.1440-1681.1998.tb02135.x.
Sumida A, Kinoshita K, Fukada T, Matsuda H, Yamamoto I, Inaba T, Azuma J: Relationship between mRNA levels quantified by reversed transcription - competitive PCR and metabolic activity of CYP3A4 and CYP2E1 in human liver. Bichem Biophys Res Commun. 1999, 262: 499-503. 10.1006/bbrc.1999.1233.
Rodrigues-Antona C, Donato M, Pureja E, Goméz-Lechón M-J, Castett J: Cytochrome P450 mRNA expression in human liver and its relationship with enzyme activity. Arch Biochem Biophys. 2001, 393: 308-315. 10.1006/abbi.2001.2499.
Enayetallah AE, French RA, Thibodeau MS, Grant DF: Distribution of soluble epoxide hydrolase and cytochrome P450 2C8, 2C9 and 2 J2 in human tissues. J Histochem Cytochem. 2006, 52: 447-454. 10.1177/002215540405200403.
Uno S, Dragin N, Miller ML, Dalton TP, Gonzalez FJ, Nebert DW: Basal and inducible CYP1 mRNA quantitation and protein localization throughout the mouse gastrointestinal tract. Free Radic Biol Med. 2008, 44: 570-583. 10.1016/j.freeradbiomed.2007.10.044.
Thelen K, Dressman JB: Cytochrome P450-mediated metabolism in the human gut wall. J Pharm Pharmacol. 2009, 61: 541-558. 10.1211/jpp.61.05.0002.
Scarth JP, Teale P, Kuuranne T: Drug metabolism in the horse: a review. Drug Test Anal. 2011, 31: 19-53. 10.1002/dta.174.
Toda T, Ohi K, Kudo T, Yoshida T, Ikarashi N, Ito K, Sugiyama K: Ciprofloxacin suppresses Cyp3a in mouse liver by reducing lithocholic acid-producing intestinal flora. Drug Metab Pharmacokinet. 2009, 24: 201-208. 10.2133/dmpk.24.201.
Pascussi JM, Drocourt L, Fabre JM, Maurel P, Vilarem MJ: Dexamethasone induces pregnane X receptor and retinoid X receptor-alpha expression in human hepatocytes: synergistic increase of CYP3A4 induction by pregnane X receptor activators. Mol Pharmacol. 2000, 58: 361-372.
Zeng Z, Andrew NW, Arison BH, Luffer-Atlas D, Wang RW: Identification of cytochrome P4503A4 as the major enzyme responsible for the metabolism of ivermectin by human liver microsomes. Xenobiotica. 1998, 28: 313-321. 10.1080/004982598239597.
Skálová L, Szotáková B, Machala M, Neca J, Soucek P, Havlasová J, Wsól V, Krídová L, Kvasnicková E, Lamka J: Effect of ivermectin on activities of cytochrome P450 isoenzymes in mouflon (Ovis musimon) and fallow deer (Dama dama). Chem Biol Interact. 2001, 137: 155-167. 10.1016/S0009-2797(01)00227-7.
Baliharova V, Velik J, Savlik M, Szotakova B, Lamka J, Tahotna L, Skalova L: The effects of fenbendazole, flubendazole and mebendazole on activities of hepatic cytochromes P450 in pig. J Vet Pharmacol Ther. 2004, 27: 85-90. 10.1111/j.1365-2885.2004.00557.x.
Velík J, Baliharová V, Fink-Gremmels J, Bull S, Lamka J, Skálová L: Benzimidazole drugs and modulation of biotransformation enzymes. Res Vet Sci. 2004, 76: 95-108. 10.1016/j.rvsc.2003.08.005.
Peters LM, Demmel S, Pusch G, Buters JT, Thormann W, Zielinski J, Leeb T, Mevissen M, Schmitz A: Equine cytochrome P450 2B6 genomic identification, expression and functional characterization with ketamine. Toxicol Appl Pharmacol. 2013, 266: 101-108. 10.1016/j.taap.2012.10.028.
Chesne C, Guyomard C, Guillouzo A, Schmid J, Ludwig E, Sauter T: Metabolism of Meloxicam in human liver involves cytochromes P4502C9 and 3A4. Xenobiotica. 1998, 28: 1-13. 10.1080/004982598239704.
Yang JC, Wang HL, Chern HD, Shun CT, Lin BR, Lin CJ, Wang TH: Role of omeprazole dosage and cytochrome P450 2C19 genotype in patients receiving omeprazole-amoxicillin dual therapy for Helicobacter pylori eradication. Pharmacotherapy. 2011, 31: 227-238. 10.1592/phco.31.3.227.
Ogg MS, Williams JM, Tarbit M, Goldfarb PS, Gray TJ, Gibson GG: A reporter gene assay to assess the molecular mechanisms of xenobiotic-dependent induction of the human CYP3A4 gene in vitro. Xenobiotica. 1999, 29: 269-279. 10.1080/004982599238669.
Li X-Q, Björkman A, Andersson TB, Gustafsson LL, Masimirembwa C: Identification of human cytochrome P450s that metabolise anti-parasitic drugs and predictions of in vivo drug hepatic clearance from in vitro data. Eur J Clin Pharmacol. 2003, 59: 429-442. 10.1007/s00228-003-0636-9.
Godawska-Matysik A, Kieć-Kononowicz K: Biotransformation of praziquantel by human cytochrome p450 3A4 (CYP 3A4). Acta Pol Pharm. 2006, 63: 381-385.
Zhou S, Koh HL, Gao Y, Gong ZY, Lee EJ: Herbal bioactivation: the good, the bad and the ugly. Life Sci. 2004, 74: 935-968. 10.1016/j.lfs.2003.09.035.
Doehmer J, Tewes B, Klein KU, Gritzko K, Muschick H, Mengs U: Assessment of drug-drug interaction for silymarin. Toxicol In Vitro. 2008, 22: 610-617. 10.1016/j.tiv.2007.11.020.
Hodek P, Trefil P, Stiborová M: Flavonoids-potent and versatile biologically active compounds interacting with cytochromes P450. Chem Biol Interact. 2002, 139: 1-21. 10.1016/S0009-2797(01)00285-X.
Acknowledgements
This study was finanically supported by the Swedish Foundation for Equine Research. Maria Löfgren is thanked for performing some of the gene expression analyses.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Tydén, E., Tjälve, H. & Larsson, P. Gene and protein expression and cellular localisation of cytochrome P450 enzymes of the 1A, 2A, 2C, 2D and 2E subfamilies in equine intestine and liver. Acta Vet Scand 56, 69 (2014). https://doi.org/10.1186/s13028-014-0069-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13028-014-0069-8