Abstract
Background
This study aimed to elucidate the roles of miR-1254 in cervical cancer progression and to explore the underlying mechanisms.
Methods
The expression levels of miR-1254 in normal-cancer cervical tissues and cells were measured using quantitive real-time polymerase chain reaction (qRT-PCR). The invasive and proliferative abilities of cervical cancer cell lines transfected with negative control (NC) mimic or miR-1254 mimic were measured using transwell, CCK-8, and colony formation assays. The binding sites between CD36 and miR-1254 were determined using luciferase reporter assays. The correlation of CD36 and miR-1254 with cervical cancer development was re-confirmed by co-transfection of miR-1254 mimic and CD36 overexpression using CCK-8, colony formation, transwell and western blot assays.
Results
MiR-1254 was expressed at significantly lower levels in the cervical cancer cell lines and tissues than in the controls. The functional assays revealed that upregulation of miR-1254 inhibited the invasion and proliferation of cervical cancer cells. The luciferase reporter assays demonstrated that CD36 messenger RNA and miR-1254 bound to one another. CD36 overexpression reversed the inhibitory effects of upregulated miR-1254 in the cervical cancer cells, suggesting that miR-1254 regulates cervical cancer progression by modulating CD36.
Conclusion
miR-1254 attenuated the invasion and proliferation of cervical cancer cells by modulating the expression levels of CD36.
Similar content being viewed by others
Background
Cervical cancer is the fourth most commonly diagnosed cancer and gynaecological cause of cancer-related deaths worldwide [1]. Several studies have found that HPV infection exerts important effects on the development of cervical cancer, which causes abnormal expression of numerous tumour-associated genes and microRNAs (miRNAs) [2, 3]. Integrating surgery and adjuvant chemotherapy are the preferred methods for treating cervical cancer; however, high rates of occurrence and metastasis lead to poor prognosis [4]. These findings suggest that it is necessary to further explore the molecular mechanisms of cervical cancer progression to discover novel therapeutic targets to block the development of cervical cancer.
Over the past few decades, increasing attention has been paid to the role of miRNAs in the onset and development of cervical cancer [5]. MiRNAs are a type of non-coding RNAs that contain fewer than 22 nucleotides and many RNA-binding sites for various target genes. They can regulate the expression levels of target messenger RNAs (mRNAs) to exert modulatory effects on the development of many tumours, including cervical cancer [6]. For example, upregulation of miR-1258 suppresses the invasion, migration and proliferation of cervical cancer cells by binding to the 3′UTR of E2F1 mRNA to promote apoptosis [7]. MiR-489 plays an anti-tumour role in cervical cancer by inactivating PI3K and AKT and upregulating the expression levels of P53 [8]. Similarly, miR-1254 is also well understood, and several studies have found that it acts as an inhibitor in some malignant tumours. MiR-1254 overexpression has been reported to inhibit the invasion, migration, proliferation, and epithelial–mesenchymal transition (EMT) of gastric cancer cells by increasing the expression of Smurf1 and inactivating the PI3K/AKT signalling pathway [9]. In addition, upregulation of miR-1254 also attenuated the migration ability of colon cancer cells by targeting PSMD10 [10]. The study findings suggest that miR-1254 is an important regulator of cancer development. In cervical cancer, Zhou et al. found that miR-1254 was significantly downregulated in cervical cancer after comparing its expression levels among 181 paired cancer-normal cervical tissues using quantitative real-time polymerase chain reaction (qRT-PCR) [11]. Nevertheless, studies on the effects and specific mechanisms of miR-1254 in cervical cancer development are scarce. These results stimulated our interest in exploring the specific effects of miR-1254 in cervical cancer. Thus, we performed this study to elucidate the effects of miR-1254 on cervical cancer progression, to analyse the underlying regulatory mechanisms and to explore new targets to block cervical cancer development.
Methods
Clinical specimen collection and processing
A total of 30 paired cancer-normal cervical tissues, obtained during surgery, were collected from patients, and the diagnosis was confirmed via postoperative pathology. No patients had been treated with radiotherapy or chemotherapy prior to surgery. The patients were divided into two groups according to the WHO criteria for the classification of cervical cancer, including 15 patients classified under stage I–II and 15 patients classified under stage III–IV. After surgical resection, all the tissues were stored in liquid nitrogen. Before collection of these samples, study approve was obtained by the Ethics Review Board of Renmin Hospital of Wuhan University, and written informed consent from all patients enrolled in this study.
Cell culture
The normal cervical cell line (Ect1/E6E7), cancerous cell lines (HeLa, C33a, SiHa and CaSki) and 293T cells were obtained from the American Type Culture Collection. The cells were then incubated in DMEM containing 10% foetal bovine serum (FBS, Invitrogen, Carlsbad, CA, USA) and 1% penicillin/streptomycin (Invitrogen), and cultured at 37 °C with 5% CO2.
RNA extraction and qRT-PCR
Total RNA was extracted from the cells and tissues using a TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. cDNA was synthesised using a reverse transcription kit (Takara, Dalian, China). Subsequently, the expression levels of miRNAs and CD36 were measured using qRT-PCR with SYBR II Premix Taq (Takara). GAPDH was used as an internal control and the fold-expression levels of miR-1254 and CD36 were calculated using the 2-ΔΔCT method. All experiments were carried out in triplicates. The miR-1254 primer was obtained from Sangon Biotech (Shanghai, China). The primers of relevant genes used were as follows: miR‐1254 forward primer: 5′‐AGCCTGGAAGCTGGAGCCTGCAGT‐3′; miR-1254 reverse primer: 5′‐GCGAGCACAGAATTAATACGAC‐3′; CD36 forward primer: 5′-TGTGCAAAATCCACAGGAAG-3′; CD36 reverse primer: 5′-GCCACAGCCA GATTGAGAAC-3′. GAPDH forward: 5′-CTGGGCTACACTGAGCACC-3′, GAPDH reverse: 5′-AAGTGGTCGTTGAGGGCAATG-3′.
Plasmid construction and transfection
MiR-1254 mimics (GenePharma, Hangzhou, China) were used to upregulate the expression levels of miR-1254, while miR-1254 inhibitors (GenePharma) were transfected into the cells to reduce miR-1254 expression. Upregulation of CD36 in the target cervical cancer cells was performed using pIRES2-ZsGreen1-CD36 (Hanbio, Shanghai, China) and labelled CD36-ov. Empty pIRES2-ZsGreen1 plasmid was also used as a control (CD36-ctrl). SiHa and CaSki cervical cancer cells were transferred into 6-well plates at a density of 1 × 105 cells per well. Then, the target cells were transfected with miR-1254 or miR-NC mimic, miR-1254 inhibitor, NC inhibitor, CD36-ov, or CD36-ctrl using Lipofectamine 3000 (Invitrogen) according to the manufacturer’s instructions.
Cell proliferation assay
The SiHa and CaSki cervical cancer cells transfected with miR-1254 mimic or NC mimic and CD36-ov or CD36-ctrl were transferred into 96-well plates at 1 × 103 cells per well. After incubation for 4, 24, 48, 72, and 96 h, the OD value of the transfected cells at 450 nm was detected by CCK-8 (Dojindo, Kumamoto, Japan) using a microplate reader. The experiments were performed in triplicates.
Colony-formation assay
The transfected SiHa and CaSki cervical cancer cells were transferred into 6-well plates at 1 × 103 cells per well and then cultured in an incubator at 37 °C for 7 days. After the formation of visible colonies, the cells were stained using a Giemsa stain kit (BA-4017, BASO, China) according to the manufacturer’s protocol. Subsequently, the number of colonies was counted using an optical microscope (Olympus DX51, Olympus, Denmark).
Luciferase reporter assay
The mutant and wild-type vectors of the pMIR-CD36-3′-UTR were purchased from GenePharma (Hangzhou, China). The SiHa and CaSki cervical cancer cells were co-transfected with miR-1254 mimic or NC mimic and pMIR-CD36-3′-UTR mutant-type vector or wild-type vector. Subsequently, the cells were cultured at 37 °C for 48 h. The luciferase activity of the target cells was measured using the dual-luciferase assay kit (Promega, Madison, WI, USA), according to the manufacturer’s instructions.
Cell invasion and migration assays
The effects of miR-1254 on the invasion of the SiHa and CaSki cervical cancer cells were measured using transwell chambers with an 8-µm pore size (Corning, NY, USA). Briefly, the transfected SiHa and CaSki cells incubated with serum-free DMEM were seeded into the upper chamber, which had been coated with 50 µl Matrigel (BD, Beijing, China). We added 600 µl DMEM containing 10% FBS (Invitrogen) to the lower chamber. After 24 h of culture, the non-invading SiHa and CaSki cells were removed, and the invading cells were fixed with 4% paraformaldehyde and stained with haematoxylin–eosin. The number of invaded cells was counted using an optical microscope (Olympus DX51).
Western blot
Protein samples of the SiHa and CaSki cells co-transfected with miR-1254 and CD36-ov or CD36-ctrl were extracted in radio immunoprecipitation assay lysis buffer (KeyGen BioTech, Nanjing, China). Subsequently, they were quantified using a bicinchoninic acid kit (KeyGen BioTech, Nanjing, China) according to the manufacturer’s protocol. The equal masses of proteins were separated using 12% SDS-PAGE and then transferred to PVDF membranes (Millipore, Billerica, MA, USA). The membranes were blocked using 5% non-fat milk in Tris-buffered saline with Tween-20 (TBS-T). The blots were incubated with GAPDH and CD36 antibodies overnight at 4 °C. After washing of the membranes with TBS-T for three times (10 min/wash), the membranes were incubated with secondary antibodies at 37 °C for 1 h. We repeated the washing step with TBS-T, and the protein signals were analysed using an enhanced chemiluminescence detection kit (Thermo Fisher, USA).
Statistical analysis
Data were processed using SPSS 19.0 and expressed as means ± standard deviation. All experiments were performed at least in triplicates. Differences between the two groups were analysed using the paired Student’s t-test, and statistical significance was set at P values of < 0.05.
Results
Reduced miR-1254 expression was observed in the human cervical cancer tissues and cell lines
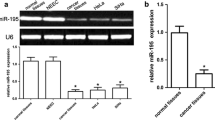
The expression levels of miR-1254 in the 30 paired normal-cancer cervical tissues were measured using qRT-PCR. We found that the miR-1254 expression was significantly lower in the cancerous tissues than in the normal tissues (Fig. 1A). Furthermore, the expression levels of miR-1254 were lower in the patients with advanced pathological stages (Fig. 1B). These results suggest that miR-1254 is related to the progression of cervical cancer and may indicate a poor prognosis. In addition, the expression levels of miR-1254 in the normal cervical cell lines (Ect1/E6E7) and cancerous cell lines (HeLa, SiHa, CaSki and C33a) were also examined, and a significant low expression was found in the latter (Fig. 1C).
MiR-1254 levels in the cervical cancer tissues and cell lines. A Low miR-1254 levels were measured in the cervical cancer samples compared to those in the adjuvant tissues. B The miR-1254 levels were lower in the patients with cervical cancer with late pathological stages. C MiR-1254 levels in the cervical normal and cancer cell lines. *P < 0.05
Upregulation of miR-1254 attenuated the proliferative and invasive abilities of the cervical cancer cells
We transfected miR-1254 mimic or NC mimic into the SiHa and CaSki cells to construct miR-1254 overexpression cell models, because of the lower miR-1254 expression in the SiHa and CaSki cells. After transfection, an elevated miR-1254 expression level was confirmed on qRT-PCR. The CCK-8 and colony formation assays revealed that miR-1254 overexpression inhibited the proliferation of the SiHa and CaSki cervical cancer cells (Fig. 2A–D). The transwell assay demonstrated that elevated expression levels of miR-1254 suppressed the invasion of the SiHa and CaSki cervical cancer cells (Fig. 2E, F).
Upregulation of miR-1254 inhibited the progression of cervical cancer. A, B The CCK-8 assay showed that transfection with miR-1254 mimic inhibited the proliferation in the CaSki and SiHa cervical cancer cells. C, D The colony formation assay showed that miR-1254 overexpression significantly reduced the number of colonies. E, F Transfection with miR-1254 mimic suppressed the invasion in the CaSki and SiHa cervical cancer cells examined using a transwell assay (magnification, ×100). *P < 0.05
MiR-1254 targeted CD36
The binding sites of miR-1254 and CD36 were reported in OSCC in a recent study [12]. Herein, we found the same relationship using the TargetScan database (http://www.targetscan.org) and confirmed this using a luciferase reporter assay (Fig. 3A, B). The luciferase activity was significantly lower in the miR-1254 mimic and wild-type CD36-3′-UTR co-transfected groups than in the mutant-type CD36-3′-UTR group (Fig. 3B). To further verify this, we measured the expression levels of CD36 in the SiHa and CaSki cells that had been transfected with miR-1254 mimic or NC mimic, miR-1254 inhibitor or NC inhibitor. We found that upregulated miR-1254 reduced CD36 expression, while downregulated miR-1254 increased CD36 expression (Fig. 3C, D). Taken together, these results suggest that CD36 is a target gene of miR-1254.
MiR-1254 targeted CD36. A Binding sites for miR-1254 and CD36. B The association between miR-1254 and CD36 was confirmed using luciferase activity assays. C, D The expression levels of CD36 mRNA were measured using quantitative real-time polymerase chain reaction after downregulation or upregulation of miR-1254 to further confirm the correlation between CD36 and miR-1254. *P < 0.05
Overexpression of CD36 rescued the effects of miR-1254 on cervical cancer cell proliferation and invasion
We constructed miR-1254 and CD36 co-overexpressing SiHa and CaSki cervical cancer cells using miR-1254 mimic and pIRES2-ZsGreen1-CD36. A control group was constructed using miR-1254 mimic and empty plasmids. After that, the expression of CD36 at the protein level was examined via western blot, and a high CD36 expression was detected in the pIRES2-ZsGreen1-CD36 group (Fig. 4A, B). Subsequently, the CCK-8 and colony formation assays showed that the proliferative ability of the cervical cancer cells was significantly enhanced after the expression of CD36 increased (Figs. 4C, D, 5A, B). The transwell assays showed that upregulation of CD36 rescued the promotive effects of miR-1254 on cervical cancer cell invasion (Fig. 5C, D). These results suggest that miR-1254 suppresses cervical cancer progression by targeting CD36, and increased CD36 expression reverses the inhibitory effects of miR-1254.
CD36 overexpression rescued the effects of miR-1254 on the cervical cancer cells. A, B Western blot and quantitative real-time polymerase chain reaction revealed that transfection of pIRES2-ZsGreen1-CD36 significantly increased the CD36 expression levels in the CaSki and SiHa cells transfected with miR-1254 mimic. C, D The CCK-8 assay revealed that upregulation of CD36 promoted the proliferative ability of the CaSki and SiHa cells transfected with miR-1254 mimic. *P < 0.05
CD36 overexpression rescued the suppressive effects of miR-1254 on cervical cancer cell proliferation and invasion. A, B The colony formation assays revealed that upregulation of CD36 promoted the proliferative ability of the SiHa and CaSki cells transfected withmiR-1254 mimic. C, D The transwell assays showed that CD36 overexpression enhanced the invasive ability of the SiHa and CaSki cells transfected with miR-1254 mimic (magnification, ×100). *P < 0.05
Discussion
In recent years, substantial dysregulated miRNAs with important regulatory effects have been found in cervical cancer cell lines and tissues [13,14,15]. MiRNAs exert effects on all biological behaviours of cervical cancer onset and development, including apoptosis, proliferation, invasion, migration, and drug resistance [16,17,18]. For example, miR-20b enhances the invasive ability of cervical cancer cells by upregulating EMT [19]. Wang et al. found that the proliferative and invasive abilities of cervical cancer could be enhanced by downregulation of miR-214, which was completed by modulating the PI3K/AKT/mTOR signalling pathway or targeting ARL2, EZH2, and FOXM1 [20,21,22,23]. MiR-145 suppresses the proliferation and metastasis of cervical cancer cells by inhibiting the Wnt/β-catenin pathway [24].
Among these miRNAs, miR-1254 is well known. Dysregulation of miR-1254 has crucial effects on cancer onset and development. Some studies have reported that miR-1254 plays an important anti-tumour role in various cancers [9, 10]. For example, its downregulation was examined in OSCC cells, and functional experiments showed that miR-1254 overexpression inhibited the metastasis of OSCC by modulating the MAPK signalling pathway [25]. However, some studies have identified the oncogenic effects of miR-1254 [12, 26]. However, to date, no reports have revealed the specific role and mechanism of miR-1254 during the biological progression of cervical cancer up to date. In this present study, we explored the specific role of miR-1254 in cervical cancer development and analysed its underlying mechanisms. Significant downregulation of miR-1254 was observed in both cervical cancer cell lines and tissues, suggesting a negative association between miR-1254 expression and cervical cancer progression. Further functional experiments suggested that increasing miR-1254 levels suppressed the invasion and proliferation of cervical cancer cells. The role of miR-1254 in regulating cervical cancer and its potential molecular mechanism might provide a novel direction for the study of cervical cancer occurrence and development.
Several lines of evidence suggest that CD36, a receptor protein, is related to the onset and progression of tumours and normally acts as a cancer promoter. For example, Pascual et al. found that blocking CD36 inhibited the metastasis of human oral cancer in mouse models [27]. CD36 also plays a scavenger receptor role and regulates cancer stem characteristics in glioblastoma [28]. Zhao et al. found that CD36 regulates cell cycle arrest, apoptosis and migration of breast cancer cells by acting as a fatty acid translocase (FAT) [29]. By acting synergistically with TGF-β, CD36 enhances EMT and promotes metastasis in cervical cancer cells [30]. These lines of evidence suggest that CD36 may be a potential therapeutic target for the treatment of various cancers, including cervical cancer. Additionally, data from the TargetScan database (http://www.targetscan.org) showed that CD36 is a direct target of miR-1254. Thus, we wondered whether miR-1254 could affect CD36 expression in cervical cancer cells. The luciferase reporter assays confirmed the binding sites between CD36 and miR-1254. We constructed miR-1254 and CD36 co-overexpressing cervical cancer cells and performed CCK-8, colony formation, and transwell experiments. The analyses showed that increased CD36 overexpression reversed the suppressive effects of miR-1254 mimic on cervical cancer cell proliferation and invasion compared with miR-1254 mimic alone. Therefore, miR-1254 exerts anti-tumour effects in cervical cancer by modulating CD36 expression.
This study has several limitations. Firstly, the number of cervical cancer and normal tissues was insufficient. More clinical tissues should be used to confirm the expression levels of miR-1254 in cervical cancer tissues. Secondly, the effects of decreased miR-1254 levels on cervical cancer progression should be investigated subsequently. Thirdly, the expression levels of miR-1254 in blood samples should be detected to explore its potential as a biological marker for diagnosis and treatment.
Conclusions
Taken together, the miR-1254 levels were lower in the cervical cancer tissues and cell lines, and miR-1254 exerted anti-tumour effects on cervical cancer progression. Upregulation of miR-1254 inhibited the invasion and proliferation of the SiHa and CaSki cervical cancer cells. Increased expression of CD36 rescued the effects of miR-1254 upregulation on cervical cancer progression, suggesting that miR-1254 suppresses cervical cancer development by modulating CD36.
Availability of data and materials
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Abbreviations
- qRT-PCR:
-
Quantitive real-time polymerase chain reaction
- CCK-8:
-
Cell counting kit-8
- NC:
-
Negative control
- miRNAs:
-
MicroRNAs
- mRNAs:
-
Messenger RNAs
- EMT:
-
Epithelial–mesenchymal transition
- FBS:
-
Foetal bovine serum
- FAT:
-
Fatty acid translocase
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Granados Lopez AJ, Lopez JA. Multistep model of cervical cancer: participation of miRNAs and coding genes. Int J Mol Sci. 2014;15:15700–33.
Berman TA, Schiller JT. Human papillomavirus in cervical cancer and oropharyngeal cancer: one cause, two diseases. Cancer. 2017;123:2219–29.
Liu Y, Yang Y, Li L, Liu Y, Geng P, Li G, Song H. LncRNA SNHG1 enhances cell proliferation, migration, and invasion in cervical cancer. Biochem Cell Biol. 2018;96:38–43.
Miao J, Regenstein JM, Xu D, Zhou D, Li H, Zhang H, Li C, Qiu J, Chen X. The roles of microRNA in human cervical cancer. Arch Biochem Biophys. 2020;690: 108480.
Pedroza-Torres A, Campos-Parra AD, Millan-Catalan O, Loissell-Baltazar YA, Zamudio-Meza H, Cantu de Leon D, Montalvo-Esquivel G, Isla-Ortiz D, Herrera LA, Angeles-Zaragoza O, et al. MicroRNA-125 modulates radioresistance through targeting p21 in cervical cancer. Oncol Rep. 2018;39:1532–40.
Peng X, Zhang Y, Gao J, Cai C. MiR-1258 promotes the apoptosis of cervical cancer cells by regulating the E2F1/P53 signaling pathway. Exp Mol Pathol. 2020;114: 104368.
Juan C, Hua Q, Ruping Z, Tingting W. miRNA-489 as a biomarker in diagnosis and treatment of cervical cancer. Bratisl Lek Listy. 2018;119:278–83.
Jiang M, Shi L, Yang C, Ge Y, Lin L, Fan H, He Y, Zhang D, Miao Y, Yang L. miR-1254 inhibits cell proliferation, migration, and invasion by down-regulating Smurf1 in gastric cancer. Cell Death Dis. 2019;10:32.
Chu YM, Peng HX, Xu Y, Yang DM, Zhou FL, Li J, Kuai R, Lin Y. MicroRNA-1254 inhibits the migration of colon adenocarcinoma cells by targeting PSMD10. J Dig Dis. 2017;18:169–78.
Zhou J, Liu X, Wang CH, Wang D, Du JJ. Decreased expression of miR-1254 is associated with cancer aggressiveness and predicts poor outcome in cervical cancer. Eur Rev Med Pharmacol Sci. 2018;22:2997–3001.
Chen R, Zhang Y, Zhang X. MiR-1254 functions as a tumor suppressor in oral squamous cell carcinoma by targeting CD36. Technol Cancer Res Treat. 2019;18:1533033819859447.
Hua FF, Liu SS, Zhu LH, Wang YH, Liang X, Ma N, Shi HR. MiRNA-338-3p regulates cervical cancer cells proliferation by targeting MACC1 through MAPK signaling pathway. Eur Rev Med Pharmacol Sci. 2017;21:5342–52.
Zhu L, Tu H, Liang Y, Tang D. MiR-218 produces anti-tumor effects on cervical cancer cells in vitro. World J Surg Oncol. 2018;16:204.
Gao C, Zhou C, Zhuang J, Liu L, Liu C, Li H, Liu G, Wei J, Sun C. MicroRNA expression in cervical cancer: novel diagnostic and prognostic biomarkers. J Cell Biochem. 2018;119:7080–90.
Yao R, Zheng H, Wu L, Cai P. miRNA-641 inhibits the proliferation, migration, and invasion and induces apoptosis of cervical cancer cells by directly targeting ZEB1. Onco Targets Ther. 2018;11:8965–76.
Sanches JGP, Xu Y, Yabasin IB, Li M, Lu Y, Xiu X, Wang L, Mao L, Shen J, Wang B, et al. miR-501 is upregulated in cervical cancer and promotes cell proliferation, migration and invasion by targeting CYLD. Chem Biol Interact. 2018;285:85–95.
Liu S, Wang H, Mu J, Wang H, Peng Y, Li Q, Mao D, Guo L. MiRNA-211 triggers an autophagy-dependent apoptosis in cervical cancer cells: regulation of Bcl-2. Naunyn Schmiedebergs Arch Pharmacol. 2019;393(3):359–70.
Cheng Y, Geng L, Zhao L, Zuo P, Wang J. Human papillomavirus E6-regulated microRNA-20b promotes invasion in cervical cancer by targeting tissue inhibitor of metalloproteinase 2. Mol Med Rep. 2017;16:5464–70.
Wang JM, Ju BH, Pan CJ, Gu Y, Li MQ, Sun L, Xu YY, Yin LR. MiR-214 inhibits cell migration, invasion and promotes the drug sensitivity in human cervical cancer by targeting FOXM1. Am J Transl Res. 2017;9:3541–57.
Wang F, Tan WH, Liu W, Jin YX, Dong DD, Zhao XJ, Liu Q. Effects of miR-214 on cervical cancer cell proliferation, apoptosis and invasion via modulating PI3K/AKT/mTOR signal pathway. Eur Rev Med Pharmacol Sci. 2018;22:1891–8.
Yang Y, Liu Y, Li G, Li L, Geng P, Song H. microRNA-214 suppresses the growth of cervical cancer cells by targeting EZH2. Oncol Lett. 2018;16:5679–86.
Peng R, Men J, Ma R, Wang Q, Wang Y, Sun Y, Ren J. miR-214 down-regulates ARL2 and suppresses growth and invasion of cervical cancer cells. Biochem Biophys Res Commun. 2017;484:623–30.
Li Q, Yu X, Yang L. MiR-145 inhibits cervical cancer progression and metastasis by targeting WNT2B by Wnt/beta-catenin pathway. Int J Clin Exp Pathol. 2019;12:3740–51.
Lu M, Chen WH, Wang CY, Mao CQ, Wang J. Reciprocal regulation of miR-1254 and c-Myc in oral squamous cell carcinoma suppresses EMT-mediated metastasis and tumor-initiating properties through MAPK signaling. Biochem Biophys Res Commun. 2017;484:801–7.
Li H, Yang T, Shang D, Sun Z. miR-1254 promotes lung cancer cell proliferation by targeting SFRP1. Biomed Pharmacother. 2017;92:913–8.
Pascual G, Avgustinova A, Mejetta S, Martin M, Castellanos A, Attolini CS, Berenguer A, Prats N, Toll A, Hueto JA, et al. Targeting metastasis-initiating cells through the fatty acid receptor CD36. Nature. 2017;541:41–5.
Hale JS, Otvos B, Sinyuk M, Alvarado AG, Hitomi M, Stoltz K, Wu Q, Flavahan W, Levison B, Johansen ML, et al. Cancer stem cell-specific scavenger receptor CD36 drives glioblastoma progression. Stem Cells. 2014;32:1746–58.
Zhao J, Zhi Z, Wang C, Xing H, Song G, Yu X, Zhu Y, Wang X, Zhang X, Di Y. Exogenous lipids promote the growth of breast cancer cells via CD36. Oncol Rep. 2017;38:2105–15.
Deng M, Cai X, Long L, Xie L, Ma H, Zhou Y, Liu S, Zeng C. CD36 promotes the epithelial–mesenchymal transition and metastasis in cervical cancer by interacting with TGF-beta. J Transl Med. 2019;17:352.
Acknowledgements
Not applicable.
About this supplement
This article has been published as part of Journal of Translational Medicine Volume 19 Supplement 2 2021: Cancer Immunotherapy. The full contents of the supplement are available at https://translational-medicine.biomedcentral.com/articles/supplements/volume-19-supplement-2.
Funding
The publication costs were covered by the corresponding authors.
Author information
Authors and Affiliations
Contributions
ZY and ZJ provided the detection of the manuscript. ZY and LX wrote and edited the manuscript. LX, YJ and ZY collected associated data. LX and ZJ drew the figures and tables. ZY and YJ revised the manuscript. ZY, LX and ZJ provided the guidance throughout the preparation of this manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The study was approved by Ethics Review Board of Renmin Hospital of Wuhan University.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Zhang, J., Li, X., Yang, J. et al. MiR-1254 suppresses the proliferation and invasion of cervical cancer cells by modulating CD36. J Transl Med 19 (Suppl 2), 531 (2022). https://doi.org/10.1186/s12967-022-03582-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12967-022-03582-6