Abstract
Background
Extended spectrum β-lactamases (ESBLs) are a group of beta-lactamase enzymes that confer resistance to the oxyimino-cephalosporins and monobactams. The emergence of ESBL - producing genes possesses a serious threat for treating infections since it is associated with multi-drug resistance. This study was focused to identify the ESBLs producing genes from Escherichia coli isolates from clinical samples from a referral-level tertiary care hospital in Lalitpur.
Methods
This was a cross-sectional study conducted from September 2018 to April 2020 at the Microbiology Laboratory of Nepal Mediciti Hospital. Clinical samples were processed, and culture isolates were identified and characterized following standard microbiological techniques. An antibiotic susceptibility test was performed by a modified Kirby-Bauer disc diffusion method as recommended by Clinical and Laboratory Standard Institute guidelines.Extended -spectrum beta-lactamases were phenotypically confirmed by the combined disc method. The ESBL-producing genes blaTEM, blaCTX−M and blaSHV were confirmed by PCR.
Results
Of the 1449 total E. coli isolates, 22.29% (323/1449) isolates were multi-drug resistant (MDR). Among the total MDR E. coli isolates, 66.56% (215/323) were ESBL producers. The maximum number of ESBL E. coli was isolated from urine 90.23% (194) followed by sputum 5.58% (12), swab 2.32% (5), pus 0.93% (2), and blood 0.93% (2). The antibiotic susceptibility pattern of ESBL E. coli producers showed the highest sensitivity toward tigecycline (100%) followed by polymyxin b, colistin and meropenem. Out of 215 phenotypically confirmed ESBL E. coli, only 86.51% (186) isolates were found to be positive by PCR for either blaTEM or blaCTX−M genes. Among the ESBL genotypes, the most common were blaTEM 63.4% (118) followed by blaCTX−M 36.6% (68).
Conclusion
The emergence of MDR and ESBL – producing E. coli isolates with high antibiotic – resistant rates to commonly used antibiotics and increased predominance of major gene types blaTEM is a serious concern to the clinicians and microbiologists. Periodic monitoring of antibiotic susceptibility and associated genes would help guide the rationale use of antibiotics for treating the predominant pathogen E. coli in the hospitals and healthcare facilities of the communities.
Similar content being viewed by others
Introduction
Extended-spectrum beta-lactamases (ESBLs) are the group of beta-lactamase enzymes, that hydrolyze and cause resistance to the oxyimino-cephalosporins (cefotaxime, ceftazidime, ceftriaxone, cefuroxime and cefepime) and monobactams (aztreonam), but not the cephamycins (cefoxitin and cefotetan) or carbapenems (imipenem, meropenem, and ertapenem), produced by Escherichia coli and Klebsiella pneumoniae [1].
The emergence of resistant bacteria worldwide is a threat to favorable outcomes of treatment of common infections in community and hospital settings. E. coli is one of the commonest pathogens to exhibit multi-drug resistance. Important risk factors for infection with MDR and ESBL E. coli are prolonged antibiotic exposure, overstay in hospital, increased use of third-generation cephalosporins, severe illness, increased use of intravenous devices or catheters [2].
ESBL was first detected during 1983–1990 in different countries [3]. Distinct epidemic clones with TEM and SHV enzymes have been found in Europe, including SHV-12, CTX-M-9, CTX-M-3 andCTX-M-15 [4].
The prevalence of ESBL - producing organisms is more than 20% in Asia and South Africa [5].
In Nepal, also, due to the increasing incidence of ESBL - producing E. coli, the cost associated with the treatment has increased. The detection of major genes such as blaTEM, blaCTX−M and blaSHV in ESBL producing E. coli by molecular methods and their antibiotic resistance pattern can provide valuable information about their epidemiology and help in the formulation of rational antimicrobial therapy [6]. Therefore, this study was conducted with the objectives of determining the spectrum of MDR and ESBL E. coli producing strains and molecular characterization of these resistant genes. Characterization of ESBL E. coli at the molecular level would be useful for developing better treatment strategy and prevention of the disease.
Materials and methods
Sample Processing and identification of Bacteria
A cross-sectional study was conducted in the Microbiology Laboratory of Nepal Mediciti Hospital, Bhaisepati, Nepal from September 2018 to April 2020. Ethical approval was obtained from the Ethical Review Board of Nepal Health Research Council (NHRC), Kathmandu, Nepal. A total of 16,542 clinical samples sent to the microbiology laboratory were processed and cultured following standard microbiological techniques. Bacterial isolates were identified by cultural and morphological characteristics, Gram stain and biochemical tests (triple sugar iron, indole, citrate, urease and motility).
Antibiotic susceptibility tests
Antibiotic susceptibility testing was performed by a modified Kirby-Bauer disc diffusion method as recommended by the Clinical and Laboratory Standard Institute guidelines. The antibiotics used were amikacin (30 µg), gentamicin (10 µg), ciprofloxacin (30 µg), ceftriaxone (30 µg), cefotaxime (30 µg), ceftazidime (30 µg), nitrofurantoin (300 µg), norfloxacin (10 µg), nalidixic acid (30 µg) ofloxacin (5 µg), cotrimoxazole (25 µg),cefixime (5 µg), cefepime (30 µg), tigecycline (15 µg), imipenem (10 µg), meropenem (10 µg), polymyxin b (300 µg), and colistin (10 µg). Plates were incubated aerobically at 37 °C for 24 h. Zone diameter in millimeters was measured and organisms were identified as sensitive, resistant, and intermediate as per CLSI 2022 guidelines [7]. E. coli strain ATCC 25,922 was used as a control strain.
Screening of ESBL
The screening for ESBL was done by the disk diffusion technique using 3rd generation cephalosporins (ceftazidime, cefotaxime and ceftriaxone). Isolates resistant to more than one of these agents were identified as possible ESBL producers [7].
Confirmation of ESBL
For confirmation of ESBL, a combined disc test was performed using ceftazidime (30 µg) alone and ceftazidime with clavulanic acid (30 µg/10µg) and cefotaxime (30 µg) and cefotaxime with clavulanic acid (30 µg/10µg). A difference in the zone of inhibition by ≥ 5 mm of either of ceftazidime clavulanic acid with ceftazidime alone and cefotaxime clavulanic acid with cefotaxime alone was interpreted as confirmed ESBL [7].
Gene detection
From confirmed ESBL E. coli, plasmid DNA was extracted using an alkaline hydrolysis method. These plasmid DNAs were used as a template for PCR amplification using blaTEM, blaCTX−M and blaSHV specific primers (Marcogen, Korea). For PCR amplification, 1.5 µl plasmid DNA was added to 25 µl mixture containing 13 µl master mixture (Solis Biodyne, Estonia), 10.5 µl nuclease-free water, and 0.5 µl each reverse and forward primers. PCR was performed in 5 Prime/02 thermal cycler using optimized conditions (Bibby Scientific, U.K.). For blaTEM gene identification, initial denaturation at 94oC for 5 min followed by 30 cycles of each of denaturation (95oC for 45 s), annealing (50oC for 45 s), and extension (72oC for 30 s), and final extension (72oC for 10 min). For blaSHV and blaCTX−M genes, initial denaturation at 94oC for 5 min followed by 30 cycles of each denaturation at 95oC for 45 s, annealing at 56oC for 45 s and 62oC for 45 s respectively, and extension at 72oC for 30 s, and final extension at 72oC for 10 min. The amplified product was subjected to gel electrophoresis (2% gel stained with ethidium bromide) at 70 V for 45 min. A DNA ladder of 100 bp was used to estimate the molecular weight of the amplified products.
Control of the phenotypic tests and PCR
For the ESBL test, E. coli (ATCC 25,922) and Klebsiella pneumoniae (ATCC 700,603) were taken as negative control and positive control respectively. Confirmed E. coli strains harboring blaTEM, blaSHV, and blaCTX−M were taken as positive control and nuclease-free water as negative control.
Statistical analysis
Data were entered and percentage calculations were analyzed using the Statistical Package for Social Science (SPSS) version 21.
Results
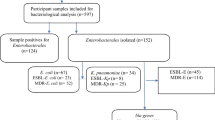
1449 E. coli isolates were recovered from various clinical samples. The highest number of E. coli was isolated from urine followed by sputum, swab, pus, blood, fluid, foley’s tip,vaginal swab, catheter tip, BAL, biopsy, bile suction tube, CVP tip, and ET tube. Of the 1449 total E. coli isolates, 22.29% (323/1449) isolates were multi-drug resistant. Among the total MDR E. coli isolates, 66.56% (215/323) isolates were ESBL producers. The maximum number of ESBL E. coli was isolated from urine 90.23% (194), followed by sputum 5.58% (12), swab 2.32% (5), pus 0.93% (2), and blood 0.93% (2) (Table 1).
The antibiotic susceptibility pattern of ESBL producing E. coli showed the highest sensitivity toward tigecycline (100%) followed by polymyxin b, colistin, nitrofurantoin, amikacin, meropenem,gentamicin, imipenem, ciprofloxacin, norfloxacin, and ofloxacin (Table 2).
Out of 215 phenotypically confirmed ESBL E. coli, only 86.51% (186) isolates were found to be positive by PCR using blaTEM, blaCTX−M and blaSHV specific primers (Table 3). 13.49% (29) were negative for any resistant gene tested. Among the ESBL genotypes, the most common were blaTEM 63.4% (118) followed by blaCTX−M 36.6% (68) (Figs. 1 and 2). The co-existence of blaTEM and blaCTX−M in ESBL -producing E. coli was 20.96% (39). No ESBL E. coli isolates co-harbored blaSHV and blaTEM, blaCTX−M and blaSHV genes.
Discussion
Despite the discovery of antibiotics, the emergence of MDR and ESBLs producing bacteria due to the extensive use of extended spectrum cephalosporins (ESCs) since the early 1980s is a significant evolution in antimicrobial resistance [8]. Several other factors, including misuse of drugs, inappropriate antibiotic treatment, and extensive use of antimicrobials have also contributed to the emergence of drug-resistant bacteria. This study was conducted in the department of microbiology laboratory, Nepal Mediciti Hospital during a period of September 2018 to April 2020 with the aim of understanding the antibiotic susceptibility profile of MDR and ESBL -producing E. coli.
It was found that the highest number of E. coli isolates was recovered from urine .With regard to urinary tract infection, E. coli showed great extent of resistance to nalidixic acid, co-trimoxazole and third generation cephalosporins. A similar pattern of resistance in urinary isolates of E. coli was shown in Nepal [9, 10]. In contrast to our results, Fanta et al. reported 73% E. coli isolates were ceftriaxone-resistant [11]. This may be due to the irrational use of third-generation cephalosporins [12]. However, a significant degree of susceptibility was found to nitrofurantoin followed by amikacin and gentamicin. Similar findings have been reported in various studies [13,14,15]. This may be due to the rational use of these drugs in urinary tract infections (UTIs) cases since nitrofurantoin is reserved drug for UTIs.
In this study, analysis of the antibiotic susceptibility of E. coli isolated from sputum, blood, swab, and pus demonstrated a significant degree of susceptibility toward tigecycline (100%) followed by colistin (98–100%), polymyxin b (97–100%), meropenem (91–96%), and imipenem (79–90%). Similar results are shown in other studies [16, 17]. It was found, a higher resistant pattern toward cephalosporins (22%to 93%), fluoroquinolones (26–85%), and aminoglycosides (8–59%) compared to urine isolates. Several studies conducted in Nepal have showed similar results [14, 18]. In contrast to our study, Kubone et al. noted a higher susceptibility pattern toward cephalosporins, fluoroquinolones, and aminoglycosides [19]. The increased level of drug resistance is a major concern worldwide since these are the first-line drugs recommended internationally [20] and are irrationally used in the public and private sectors [21].
This study noted 22.29% (323/1449) MDR E. coli isolates that were suspected of being ESBL producers were confirmed by the combined disc method. The prevalence of ESBL E. coli was 66.56% (215/323), which was alarmingly high. Several studies reported high prevalence i.e.40–70% of ESBL E. coli among MDR E. coli [10, 15, 17, 22, 23]. However, the study by Onyedibe et al. in 2018 observed only 18.6% ESBL E. coli [24], which is an analogous result to other study [25]. This is not similar to our study due to the variation in geography, study design, and selection of the type of antimicrobial agents. The indiscriminate use of beta-lactam antibiotics leads to the generation of selective pressures, which have led to the selection of various mutated forms of beta lactamase [26]. The antibiotic profiles of ESBL E. coli was found to be higher sensitivity toward tigecycline (100%), polymyxin b (100%), colistin (100%) followed by amikacin (91.6%), meropenem (90.2%), and imipenem (68.8%). The susceptibility to nitrofurantoin was 93.8% against ESBL E. coli isolated from urine. So, it could be the drug of choice for treating infection caused by ESBL producing E. coli similar to the previous studies conducted in India [27, 28].
In this study, out of 323 MDR E. coli isolates, ESBL E. coli phenotypes were found to be positive in 66.56% (215) isolates. Similar findings were reported by Ozcakar et al. (2011) and Dalela et al. (2012) [29, 30]. 13.49% (29) phenotypic ESBL - positive E. coli isolates lacked blaTEM, blaCTX−M and blaSHV genes. That could be false - positive results by phenotypic methods or can be the possible presence of other ESBL encoding genes such as SFO, BES, BEL, TLA, GES, PER and VEB types and structural changes in penicillin-binding proteins that result in resistance to β-lactam antibiotics [29, 31,32,33].
In this study, the overall prevalence of ESBL genes was 86.51% (186), which is similar to other findings reported by Dirar et al., 2020 in Sudan and Ahmad et al., 2019 in Iraq [34, 35]. PCR revealed blaTEM, blaCTX−M and blaSHV genes in ESBL - producing E. coli were 63.4% (118), 36.6% (68), and 0.0 (0) respectively. In this study, blaTEM was the most predominant genotype of ESBL among E. coli isolates. Similar findings were reported by Dirar et al., 2020 in Sudan, Ahmad et al.,2019 in Iraq, Noha et al., 2020 in upper Egypt, Pandit et al., 2020 in Nepal, Michael et al., 2018 in Iraq, Sahoo et al. 2019 in India, and Jena et al., 2017 in India [34,35,36,37,38,39,40].
In this study, the prevalence of blaCTX−M genes was found to be 36.6% (68), which concurs with various reports demonstrating the extensive worldwide dissemination of blaCTX−M genes in ESBL - producing E. coli isolates [41]. However, another study from Nepal has reported the high prevalence of blaCTX−M genes (100%) by Lohani et al., 2019 and (91.4%) by Parajuli et al., 2016 [42, 43].
The differences in frequencies of the prevalence of these genes may be because of differences in time by which isolates were collected and differences in volume and type of antibiotic consumption [44].
Furthermore, multiple harboring of genes in a single ESBL - producing E. coli were also noted. The most common combination gene was blaTEM + blaCTX−M type 20.96% (39). Our finding is in agreement with the study by Lohani et al., where (21.2%) of blaTEM and blaCTX−M genes were reported [42]. The presence of multiple genotypes in a single isolate might be the result of a complex antibiotic resistance pattern [45].
In this study, blaSHV type E. coli was not detected, similar to the study in Nigeria [46]. However, several findings in Nepal reported the prevalence of blaSHV gene at low frequency [42, 43].
Conclusion
This study highlights the emergence of MDR and ESBL -producing E. coli isolates with high antibiotic -resistant rates to commonly used antibiotics and increased predominance of major gene types blaTEM. No resistance was documented to tigecycline, polymyxin b, and colistin suggesting the suitable drug of choice for treating ESBL -producing E. coli infections. Periodic molecular detection and identification of ESBL -producing bacterial isolates could inform the rationale use of antibiotics to preserve antibiotics for the future.
Data Availability
All data generated in the study have been included in the manuscript.
Abbreviations
- ESBL:
-
Extended Spectrum Beta Lactamase
- MDR:
-
Multi Drug Resistant
- bla:
-
β-lactamase-coding gene
- ATCC:
-
American Type Culture Collection
- CLSI:
-
Clinical Laboratory Standard Institute
- CTX-M:
-
Cefotaximase, Munich
- TEM:
-
Temoniera gene
- SHV:
-
Sulfhydryl variable
- UTIs:
-
Urinary Tract Infections
References
Ghafourian S, Sadeghifard N, Soheili S, Sekawi Z. Extended spectrum beta- lactamases: definition, classification and Epidemiology.Curr. Issues Mol Biol. 2015;17(1):11–22. https://doi.org/10.21775/cimb.017.011.
Chaudhary U, Aggarwal R. Extended spectrum β-lactamases (ESBL): emerging threat to clinical therapeutics. Indian J Med Microbiol. 2004;22(2):7. https://doi.org/10.1016/S0255-0857(21)02884-X.
Shaikh S, Fatima J, Shakil S, Mohd S, Rizvi D, Mohammad AK. Antibiotic resistance and extended spectrum beta-lactamases: types, epidemiology and treatment. Saudi J Biol Sci. 2015;22(1):90–101. https://doi.org/10.1016/j.sjbs.2014.08.002.
Perilli M, Segatore B, Mugnaioli C, Celenza G, Rossolini GM, Stefani S, Luzzaro F, Pini B, Amicosante G. Persistence of TEM-52/TEM-92 and SHV-12 extended-spectrum beta-lactamases in clinical isolates of Enterobacteriaceae in Italy. Microb Drug Resist. 2011;17(4):521–4. https://doi.org/10.1089/mdr.2011.0059.
Mshana SE, Kamugisha E, Mirambo M, Chakraborty T, Lyamuya EF. Prevalence of multiresistant gram-negative organisms in a tertiary hospital in Mwanza,Tanzania. BMC Res Notes. 2009;2:49. https://doi.org/10.1186/1756-0500-2-49.
Kaur M, Aggarwal A. Occurrence of the CTX-M, SHV and the TEM genes among the Extended Spectrum Beta-Lactamase Producing Isolates of Enterobacteriaceae in a Tertiary Care Hospital of North India. J Clin Diagn Res. 2013;7:642–5. https://doi.org/10.7860/JCDR/2013/5081.2872.
Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial susceptibility testing; 32nded.CLSI supplement M100.Clinical and Laboratory Standards Institute,USA,2022.
Rawat D, Nair D. Extended-spectrum β-lactamases in gram negative bacteria. J Glob Infect Dis.2010; 2(3): 263–74. https://doi.org/10.4103/0974-777X.68531.
Guragain N, Pradhan A, Dhungel B, Banjara MR, Rijal KR, Ghimire P. Extended spectrum beta-lactamase producing gram negative bacterial isolates from urine of patients visiting Everest Hospital, Kathmandu, Nepal. Tribhuvan Univ J Microbiol. 2019;6:26–31. https://doi.org/10.3126/tujm.v6i0.26575.
Khanal LK, Amatya R, Sah AK, Adhikari RP, Khadka S, Sapkota J, Rai SK. Prevalence of extended spectrum beta lactamase producing Escherichia coli and Klebsiella spp. from urinary specimen in a tertiary care hospital. Nepal Med Coll J. 2022;24(1):75–80. https://doi.org/10.3126/nmcj.v24i1.44145.
Fanta G, Eshetu M, Mekidim M, Gemechu Z. Antimicrobial resistance profile of different clinical isolates against third-generation cephalosporins. J Pharm. 2018;7:5070742. https://doi.org/10.1155/2018/5070742.
Mitman SL, Amato HK, Saraiva-Garcia C, Loayza F, Salinas L, Kurowski K, et al. Risk factors for third-generation cephalosporin-resistant and extended-spectrum β- lactamase-producing Escherichia coli carriage in domestic animals of semirural parishes east of Quito,Ecuador. PLOS Glob Public Health. 2022;2(3):e0000206. https://doi.org/10.1371/journal.pgph.0000206.
Salem MA, Ahmed FA. Bacterial profile of urinary tract infection and antimicrobial susceptibility pattern among patients attending at Bushra Medical Laboratory, Tripoli, Libya. J Gastroenterol Hepatol Res. 2018;7(4):2671–5. http://www.ghrnet.org/index.php/joghr/article/view/2380.
Subedi S, Chaudhary M, Shrestha B, High. MDR AND ESBL producing Escherichia coli and Klebsiella pneumoniae from urine, Pus and Sputum samples. Br J Med &Medical Res. 2016;13(10):1–10. https://doi.org/10.9734/BJMMR/2016/23350.
Rimal U, Thapa S, Maharjan R. Prevalence of extended spectrum beta-lactamase producing Escherichia coli and Klebsiella species from urinary specimens of children attending Friendship International Children’s hospital. Nepal J Biotechnol. 2017;5(1):32–8. https://doi.org/10.3126/njb.v5i1.18868.
Naqid IA, Balatay AA, Hussein NR, Saeed KA, Ahmed HA, et al. Antibiotic susceptibility pattern of Escherichia coli isolated from various clinical samples in Duhok City, Kurdistan Region of Iraq. Int J Infect. 2020;7(3):e103740. https://doi.org/10.5812/iji.103740.
Shilpakar A, Ansari M, Rai KR, Rai G, Rai SK. Prevalence of multidrug-resistant and extended-spectrum beta-lactamase producing Gram-negative isolates from clinical samples in a tertiary care hospital of Nepal. Trop Med Health. 2021;49(1):23. https://doi.org/10.1186/s41182-021-00313-3.
Yadav K, Prakash S. Screening of ESBL producing Multidrug resistant E.coli from urinary tract infection suspected cases in Southern Terai of Nepal. J Infect Dis Diagn. 2017;2:2. https://doi.org/10.4172/2576-389x.1000116.
Kubone PZ, Mlisana KP, Govinden U, Abia ALK, Essack SY. Antibiotic susceptibility and molecular characterization of uropathogenic Escherichia coli associated with community-acquired urinary tract infections in urban and rural settings in South Africa. Trop Med Infect Disease. 2020;5(4):176. https://doi.org/10.3390/tropicalmed5040176.
Iftekhar A, Md.Bodiuzzaman R, Sakina S. Antibiotic resistance in Bangladesh: Asystematic review. Int J Infect Dis. 2019;80:54–61. https://doi.org/10.1016/j.ijid.2018.12.017.
Rijal KR, Banjara MR, Dhungel B, et al. Use of antimicrobials and antimicrobial resistance in Nepal: a nationwide survey. Sci Rep. 2021. https://doi.org/10.1038/s41598-021-90812-4.11,11554.
Manandhar S, Zellweger RM, Maharjan N, et al. A high prevalence of multi-drug resistant gram-negative bacilli in a Nepali tertiary care hospital and associated widespread distribution of extended-spectrum beta-lactamase (ESBL) and carbapenemase-encoding genes. Ann Clin Microbiol Antimicrob. 2020;19:48. https://doi.org/10.1186/s12941-020-00390-y.
Sadeghi M, Sedigh Ebrahim-Saraie H, Mojtahedi A. Prevalence of ESBL and AmpC genes in E. coli isolates from urinary tract infections in the north of Iran. New Microbes New Infect. 2022;45:100947. https://doi.org/10.1016/j.nmni.2021.100947.
Onyedibe K, Shobowale E, Okolo M, Iroezindu M, Afolaranmi T, Nwaokorie F, Opajobi S, Isa S, Egah D. Low prevalence of carbapenem resistance in clinical isolates of extended spectrum beta lactamase (ESBL) producing Escherichia coli in north central, Nigeria. Advances in Infectious Diseases.2018; 8:109–120. https://doi.org/10.4236/aid.2018.83011
Vachvanichsanong P, McNeil EB,Dissaneewate P. Extended-spectrum beta- lactamase Escherichia coli and Klebsiella pneumoniae urinary tract infections.Epidemiology and Infection.2021;149,e12,1–7. https://doi.org/10.1017/S0950268820003015
Chanu TR, Shah PK, Soni S, Ghosh A. Phenotypic detection of extended spectrum, AmpC, Metallo beta-lactamases and their coexistence in clinical isolates of commonly isolated gram negative bacteria in GKGH hospital. Bhuj IP Int J Med Microbiol Trop Dis. 2019;5(1):52–6. https://doi.org/10.18231/2581-4761.2019.0012.
Gharavi MJ, Zarei J, Roshani-Asl P, Yazdanyar Z, Sharif M, Rashidi N. Comprehensive study of antimicrobial susceptibility pattern and extended spectrum beta-lactamase (ESBL) prevalence in bacteria isolated from urine samples. Sci Rep. 2021;11(1):578. https://doi.org/10.1038/s41598-020-79791-0.
Kumar MS, Arunagirinathan N, Ravikumar M. Antibiotic susceptibility profile of extended spectrum β-lactamase producing Escherichia coli, Klebsiella pneumoniae and Klebsiella oxytoca from urinary tract infections. Research Journal of Pharmacy and Technology. 2021;14(8):4425–4428. https://doi.org/10.52711/0974-360X.2021.00768
Ozcakar ZB, Yalcinkaya F, Kavaz A, Kadioglu G, Elhan AH, Aysev D, Guriz H, Ekim M. Urinary tract infections owing to ESBL-producing bacteria:microorganisms change- clinical pattern does not. Acta Paediatr. 2011;100(8):61–4. https://doi.org/10.1111/j.1651-2227.2011.02262.x.
Dalela G, Gupta S, Jain DK, Mehta P. Antibiotic resistance pattern in uropathogens at a tertiary care hospital at Jhalawar with special reference to ESBL, AmpC β- lactamase and MRSA Production. J Clin D R. 2012;6(4):645–51.
Nass T, Poirel L, Nordmann P. Minor extended spectrum β-lactamases.Clin MicrobiolInfect.2008;14 (Suppl.1):42–52. https://doi.org/10.1111/j.1469-0691.2007.01861.x
Marthie ME, Chrisna V, Eddy PM, Michael GD, Anwar AH, Marleen MK. Detection of blaSHV,blaTEM and blaCTX–M antibiotic resistance genes in randomly selected bacterial pathogens from the Steve Biko Academic Hospital. Immunol Med Microbiol. 2009;56:191–6. https://doi.org/10.1111/j.1574-695x.2009.00564.x.
Majid K, Majid B, Fateh R. Detection of TEM,SHV and CTX-M antibiotic resistance genes in Escherichia coli isolates from infected wounds. Med Lab J. 2017;11(2):30–5. https://doi.org/10.18869/acadpub.mlj.11.2.30.
Dirar MH, Bilal NE, Ibrahim ME, Hamid ME. Prevalence of extended-spectrum β- lactamase (ESBL) and molecular detection of blaTEM, blaSHV and blaCTX–M genotypes among Enterobacteriaceae isolates from patients in Khartoum. Sudan Pan Afr Med J. 2020;37:213. https://doi.org/10.11604/pamj.2020.37.213.24988.
Ahmad Hamad P, Khadija KM. Prevalence of blaTEM, blaSHV,and blaCTX–M genes among ESBL-producing Klebsiella pneumoniae and Escherichia coli isolated from thalassemia in Erbil,Iraq. Mediterranean J Hematol Infect Dis. 2019;11(1):e2019041. https://doi.org/10.4084/mjhid.2019.041.
Noha AH, Ahmed SK, Eman MF, Adel MH, Medhat AF. Molecular characterization of extended-spectrum β lactamase-producing E.colirecovered from community- acquired urinary tract infections in Upper Egypt. Sci Rep. 2020;10:2772. https://doi.org/10.1038/s41598-020-59772-z.
Pandit R, Awal B, Shrestha SS, Joshi G, Rijal BP, Parajuli NP. Extended-spectrum β- lactamase(esbl) genotypes among multidrug-resistant uropathogenic Escherichia coli clinical isolates from a teaching hospital of Nepal. Interdisciplinary Perspectives on Infectious Diseases.2020;8. https://doi.org/10.1155/2020/6525826
Michael NS, Saadi AT. Detection of blaCTX–M, blaTEM01 and blaSHV genes in Multidrug Resistant Uropathogenic E. coli isolated from patients with recurrent urinary tract infections. Int J Med Res Health Sci. 2018;7(9):81–9.
Sahoo S, Otta S, Swain B, Kar SK. Detection and genetic characterization of extended- spectrum beta-lactamases producers in a tertiary care hospital. J Lab Physicians. 2019;11(3):253–8. https://doi.org/10.4103/jlp.jlp_31_19.
Jena J, Sahoo RK, Debata NK, Subudhi E. Prevalence of TEM, SHV, and CTX-M genes of extended-spectrum beta-lactamases-producing Escherichia coli strains isolated from urinary tract infections in adults. Biotech. 2017;7(4):244. https://doi.org/10.1007/s13205-017-0879-2.
Canton R, Gonzalez-Alba JM, Galan JC. CTX-M enzymes:origin and diffusion. Front Microbiol. 2012;3:110. https://doi.org/10.3389/fmicb.2012.00110.
Lohani B, Thapa M, Sharma L, Adhikari H, Sah AK, Khanal AB, Basnet RB, Aryal M. Predominance of CTX-M type extended Spectrum β lactamase (ESBL) producers among clinical Isolates of Enterobacteriaceae in a Tertiary Care Hospital, Kathmandu, Nepal. The OpenMicrobiology. Journal. 2019;13:28–33. https://doi.org/10.2174/1874285801913010028.
Parajuli NP, Maharjan P, Joshi G, Khanal PR. Emerging perils of extended spectrum β-lactamase Producing Enterobacteriaceae clinical isolates in a teaching hospital of nepal. Biomed Research International. 2016;7. https://doi.org/10.1155/2016/1782835
Al-Agamy HM, Shibl AM, Tawfik AF. Prevalence and molecular characterization of extended-spectrum β-lactamase-producing Klebsiella pneumoniae. Riyadh Saudi Arabia Annals of Saudi Medicine. 2009;29(4):253. https://doi.org/10.4103/0256-4947.55306.
Polse RF, Yousif SY, Assafi MS. Prevalence and molecular characterization of extended spectrum β-lactamases producing uropathogenic Escherichia coli isolated in Zakho, Iraq. J Microbiol Infect Dis. 2016;6(4):163–7. https://doi.org/10.5799/jmid.328863.
Adekanmbi AO, Akinpelu MO, Olaposi AV, Oyelade AA. Extended spectrum beta-lactamase encoding gene-fingerprints in multidrug resistant Escherichia coli isolated from wastewater and sludge of a hospital treatment plant in Nigeria. Int J Environ Stud. 2021;78(1):140–50. https://doi.org/10.1080/00207233.2020.1778271.
Acknowledgements
We would like to acknowledge the Nepal Mediciti Hospital, Nepal for providing ethical approval and research work.We also thank to the microbiology laboratory staff for their valuable support to complete this work.
Funding
There was no funding support to complete this research.
Author information
Authors and Affiliations
Contributions
A. Mahesh Chaudhary: Laboratory work, analysis and interpretation of data, drafting manuscript and finalization, and final approval to be submitted.B. Prof. Dr. Indrani Jadhav: Analysis and interpretation of data, supervision, revision of the manuscript, and final approval to be submitted.C. Dr. Megha Raj Banjara: Supervision, revision of the manuscript, and final approval to be submitted.
Corresponding author
Ethics declarations
Ethical approval
Ethical approval was obtained from Nepal Health Research Council (NHRC).
Consent form
Consent form was completed before collection of data and samples.
Competing interests
The authors declare no conflicts of interest.
Consent for publication
Not Applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.


Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Chaudhary, M.K., Jadhav, I. & Banjara, M.R. Molecular detection of plasmid mediated blaTEM, blaCTX−M,and blaSHV genes in Extended Spectrum β-Lactamase (ESBL) Escherichia coli from clinical samples. Ann Clin Microbiol Antimicrob 22, 33 (2023). https://doi.org/10.1186/s12941-023-00584-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12941-023-00584-0