Abstract
Background
Cellulase can convert lignocellulosic feedstocks into fermentable sugars, which can be used for the industrial production of biofuels and chemicals. The high cost of cellulase production remains a challenge for lignocellulose breakdown. Trichoderma reesei RUT C30 serves as a well-known industrial workhorse for cellulase production. Therefore, the enhancement of cellulase production by T. reesei RUT C30 is of great importance.
Results
Two sets of novel minimal transcriptional activators (DBDace2-VP16 and DBDcre1-VP16) were designed and expressed in T. reesei RUT C30. Expression of DBDace2-VP16 and DBDcre1-VP16 improved cellulase production under induction (avicel or lactose) and repression (glucose) conditions, respectively. The strain TMTA66 under avicel and TMTA139 under glucose with the highest cellulase activities outperformed other transformants and the parental strain under the corresponding conditions. For TMTA66 strains, the highest FPase activity was approximately 1.3-fold greater than that of the parental strain RUT C30 at 120 h of cultivation in a shake flask using avicel as the sole carbon source. The FPase activity (U/mg biomass) in TMTA139 strains was approximately 26.5-fold higher than that of the parental strain RUT C30 at 72 h of cultivation in a shake flask using glucose as the sole carbon source. Furthermore, the crude enzymes produced in the 7-L fermenter from TMTA66 and TMTA139 supplemented with commercial β-glucosidase hydrolyzed pretreated corn stover effectively.
Conclusions
These results show that replacing natural transcription factors with minimal transcriptional activators is a powerful strategy to enhance cellulase production in T. reesei. Our current study also offers an alternative genetic engineering strategy for the enhanced production of industrial products by other fungi.
Similar content being viewed by others
Background
Lignocellulosic feedstocks are abundant and renewable resources in nature. They have been used to produce environment-friendly biofuels and chemicals, which have received increased attention for research [1]. The conversion of lignocellulosic feedstocks into fermentable sugars involving cellulase-based enzymatic saccharification is the key issue in large-scale production of biofuels and chemicals [2]. However, the industrial-scale production of biofuels and chemicals is limited in part by the high cost of cellulase enzymes [3,4,5]. Trichoderma reesei, which produces the enzymes necessary for the complete hydrolysis of lignocellulose, has been widely used for the production of commercial cellulase [6]. T. reesei RUT C30 has been proven to be a cellulase hyper-producer [7]. Therefore, improvement of cellulase production in T. reesei RUT C30 as a cellulosic biorefinery is of great importance.
The cellulase produced in T. reesei RUT C30 mainly comprise two cellobiohydrolases (CBHI and CBHII), two endoglucanases (EGI and EGII), and β-glucosidase I (BGLI) that synergistically hydrolyze lignocellulosic materials, together with related xylanases and auxiliary proteins [7,8,9]. Among the total proteins produced by T. reesei, CBHI and CBHII account for approximately 50–60 and 10–15%, respectively [10]. EGI and EGII are the two main endoglucanases and together account for 6–20% of the total produced proteins [11]. BGLI is the key enzyme involved in the complete conversion of cellobiose to glucose [12]. XYNI and XYNII are the main components of xylanase from T. reesei [13]. Additionally, swollen in, as a non-enzymatic cellulose attacking enzyme, and lytic polysaccharide monooxygenases (LPMOs) synergistically degrade lignocellulosic materials as auxiliary proteins [14, 15].
Expression of cellulase and xylanase in T. reesei is regulated by a combination of transcription factors such as XYRI, ACEII, and CREI. Among these factors, XYRI (xylanase regulator 1) is recognized as the primary transcriptional activator with a GAL4-like Zn(II)2Cys6 binuclear cluster domain [16]. Similar to XYRI, ACEII also acts as a transcriptional activator, consisting of a Zn(II)2Cys6 DNA-binding domain, which is responsible for the regulation of cellulase-encoding genes in T. reesei under induction conditions [17]. On the other hand, CREI is the primary negative regulator that mediates carbon catabolite repression (CCR). The expression of cellulase in T. reesei RUT C30 is inhibited by CREI when glucose is present [18].
In addition to protein engineering of cellulase chimeras to enhance specific activity [19], increasing cellulase production via the application of artificial transcription factors is an effective strategy [20,21,22]. An artificial transcription activator containing the two DNA-binding domains from CREI and ACEI and an effector domain from ACEII can regulate gene expression in T. reesei [20]. Zhang et al. [21] screened a mutant T. reesei strain U3 with enhanced cellulase production by transforming an artificial zinc finger protein library. Similarly, an artificial transcription activator linking the CREI-binding domain with the C-terminus of XYRI was shown to improve cellulase production in the recombinant strain T. reesei zxy-2 with glucose as the sole carbon source [22].
Here, we developed two sets of novel artificial transcriptional activators, which were designed as minimal transcriptional activators containing one DNA binding domain from either ACEII or CREI, and one strong transcriptional activation domain VP16 from the herpes simplex virus [23]. These minimal transcriptional activators DBDace2-VP16 and DBDcre1-VP16 were transformed into T. reesei RUT C30 to replace the native regulators ACEII and CREI, respectively, in the genome by homologous double exchange. We then investigated the effects of these minimal transcriptional activators (DBDace2-VP16 under induction conditions and DBDcre1-VP16 under repression conditions) on cellulase and xylanase generation. We suggest that the minimal transcriptional activators DBDace2-VP16 and DBDcre1-VP16 can enhance cellulase production.
Results
Construction of transformants with minimal transcriptional activators
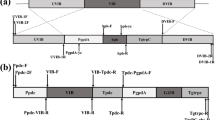
Each compact minimal transcription activator consists of one DNA binding domain (DBD) of the native transcription factor and the transcriptional activation domain VP16. The transcription factors ACEII and CREI were used as targets to be replaced. Three sets of ace2 DBDs with different lengths were selected to construct the DBDace2-VP16 (MTA58/66/81) minimal transcriptional activators. DBDace2–58 (amino acids 1–58) is the core DBD as previously demonstrated using DNA mobility shift assays [17]. DBDace2–66 (amino acids 1–66) includes a histidine-rich region (amino acids 53–66) with unknown function that is also found in other Zn(II)2Cys6-type regulatory proteins [17]. DBDace2–81 (amino acids 1–81) is the longest region responsible for DNA binding and is followed by the region corresponding to the activation domain. Similarly, three sets of cre1 DBDs were selected (cre1 96 , DBDcre1–109, and DBDcre1–139). However, cre1 96 does not represent the DNA sequence of amino acids 1–96 of CREI but is a chimera due to the loss of a 2477-bp fragment at position + 262 of cre1 in the T. reesei hyper-cellulolytic mutant RUT C30 [24]. The plasmids for minimal transcriptional activators DBDace2-VP16 (pMTA58/66/81) and DBDcre1-VP16 (pMTA96/109/139) (Fig. 1) were transformed into T. reesei RUT C30 to replace the original transcriptional factors (ace2 and cre1 96 , respectively) by homologous double exchange and eliminate the risk of any unpredictable mutagenesis caused by random insertion. We screened nine transformants each for DBDace2-VP16 (named TMTA58-1/-2/-3, TMTA66-1/-2/-3, and TMTA81-1/-2/-3) and for DBDcre1-VP16 (named TMTA96-1/-2/-3, TMTA109-1/-2/-3, and TMTA139-1/-2/-3). All transformants were identified as correct gene knockout strains harboring single-copy DNA integration (Additional file 1: Figure S1). The plasmids and transformants are listed in Table 1.
Construction of the minimal transcriptional activators. Plasmids for two sets of novel minimal transcriptional activators fusing the DNA-binding domains of ACEII or CREI with the VP16 activation domain and a short flexible linker (GGGGSGGGGS). Transformants TMTA58/66/81 and TMTA96/109/139 were obtained after xylose-induced marker rescue. The natural factors were replaced by the minimal transcriptional activators in transformants genomes
Growth of minimal transcriptional activator transformants
To determine whether the minimal transcriptional activators were involved in cellulase production under induction or repression conditions, we first investigated the growth of the transformants of the minimal transcriptional activators, along with the parental strain RUT C30, in glucose, lactose, and avicel. The growth of DBDace2-VP16 transformants TMTA58, TMTA66, and TMTA81 were examined in minimal medium (MM) with glucose, lactose, or avicel as the sole carbon source (Fig. 2a–c). The growth of DBDcre1-VP16 transformants TMTA96, TMTA109, and TMTA139 were detected in MM containing glucose as the sole carbon source (Fig. 2d). The growth of DBDace2-VP16 transformants exhibited no significant differences from the parental strain RUT C30 when cultured in glucose (Fig. 2a), lactose (Fig. 2b), or avicel (Fig. 2c). This suggests that DBDace2-VP16 is not involved in basic cellular metabolism. However, the growth of DBDcre1-VP16 transformants TMTA96, TMTA109, and TMTA139 showed delayed growth in glucose (Fig. 2d). Growth delays were more pronounced in TMTA139 strains than those in the TMTA96 and TMTA109 strains, indicating that DBDcre1-VP16 affected the primary metabolism of the cells in glucose.
Cell growth differences between transformants and RUT C30. Conidia (106/mL) of strains were incubated on MM medium supplemented with different carbon sources (2% w/v) for 72 h. RUT C30 and DBDace2-VP16 transformants were cultured on glucose (a), lactose (b), and avicel (c). RUT C30 and DBDcre1-VP16 transformants were cultured on glucose (d). The biomass dry weight per liter was calculated from the intracellular protein content for avicel based on 0.32 g of intracellular protein per g dry biomass. Error bars show the respective standard deviation of three biological replicates
Minimal transcriptional activator DBDace2-VP16 promotes cellulase production under induction conditions
To identify whether the minimal transcriptional activator DBDace2-VP16 can enhance cellulase production, cellulase activities, xylanase activities, and secreted protein concentrations from transformants and parental RUT C30 were examined in induction conditions (avicel and lactose) (Fig. 3). Distinctly, the pNPCase, FPase, and CMCase activities of TMTA58, TMTA66, and TMTA81 in both lactose- and avicel-containing media were superior compared to those of the parental strain RUT C30 (Fig. 3a–c). Moreover, the tested transformants exhibited better performance in avicel than in lactose. The FPase activity in TMTA66 strains cultured in avicel at 120 h was 5.2 U/mL, which was about 1.3-fold greater than that produced by RUT C30 (Fig. 3a). In addition, TMTA66 strains also showed the highest pNPCase (0.7 U/mL) and CMCase activities (28.6 U/mL), which were about 2.2- and 1.9-fold, respectively, when compared to those of RUT C30 in avicel at 120 h (Fig. 3b, c). On the other hand, the pNPGase and xylanase activities had no significant difference between TMTA58, TMTA66, TMTA81, and RUT C30 (Fig. 3d–f). In agreement with the noticeable increment of cellulase activities, 51% more secreted protein was detected in the culture supernatant of TMTA66 transformants compared to that of the parental strain RUT C30 (Fig. 3g).
Cellulase and xylanase production in DBDace2-VP16 transformants. DBDace2-VP16 transformants and RUT C30 were cultured on 2% avicel or lactose after a shift from glucose. FPase (a), pNPCase (b), CMCase (c), pNPGase (d), xylanase I (e), and xylanase II (f) activity (U/mL), and extracellular secreted protein concentration (g) were measured at 72, 96, and 120 h. Error bars show the respective standard deviation of three biological replicates; asterisks indicate significant differences (*p ≤ 0.05) between the transformants and the parental strain RUT C30, as assessed by Student’s t test
To further confirm the effects of DBDace2-VP16 on the synthesis of cellulase or total protein secretion, the transcript levels of cellulase-related genes including five main cellulase genes (cbh1, cbh2, egl1, egl2, and bgl1), two major xylanase genes (xyn1 and xyn2), one accessary protein gene (swo1), and two transcription factor genes (xyr1 and cre1 96 ) at 12 and 24 h were analyzed using RT-qPCR (Fig. 4). TMTA58, TMTA66, and TMTA81 exhibited increased expression of cellulase genes cbh1, cbh2, egl1, and egl2 compared to that of the parental strain in both lactose- and avicel-containing media (Fig. 4a, b), which is consistent with the result of enhanced cellulase activities. Moreover, avicel induced a higher ratio for cellulase gene expression than lactose. Notably, TMTA66 strains showed the strongest transcriptional activation among the strains. Using avicel as the sole carbon source, the transcript levels of cbh1, cbh2, egl1, and egl2 in the TMTA66 strain increased about 2.9-, 1.3-, 1.4-, and 1.7-fold, respectively, when compared to that in T. reesei RUT C30 at 24 h (Fig. 4a). While, the transcript levels of bgl1, xyn1, and xyn2 were not significantly upregulated in both avicel and lactose (Fig. 4a, b). We also detected an enhancement of transcript levels of swo1 compared to that in parental strain (Fig. 4a, 4b). In addition, the transcript levels of xyr1 and cre1 96 showed no significant differences in the TMTA58, TMTA66, and TMTA81 transformants in comparison to that in RUT C30. This indicates that the minimal transcriptional activator DBDace2-VP16 contributes to the expression of the cellulase genes due to the enhancement of its own transcriptional activation.
Comparison of transcript levels of main cellulase, xylanase, and transcription factors genes in DBDace2-VP16 transformants. Gene expression ratios (-fold) in DBDace2-VP16 transformants relative to RUT C30 on 2% avicel (a) and lactose (b) for 12 and 24 h after a shift from glucose. Gene expression ratios (-fold) were normalized to the corresponding gene expression at the same timepoint in the parental strain RUT C30. Values represent the mean of three biological replicates and error bars show the standard deviation; asterisks indicate significant differences (gene expression ratio greater than twofold or less than 0.5-fold between the transformants and the parental strain RUT C30)
Minimal transcriptional activator DBDcre1-VP16 promotes the cellulase and xylanase production under repression conditions
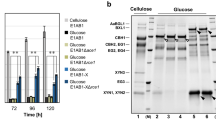
Given that CREI has been identified as a transcription factor responding to carbon source metabolite repression, glucose was used as the sole carbon source to analyze the effects of the minimal transcriptional activator DBDcre1-VP16 in the TMTA96, TMTA109, and TMTA139 transformants (Figs. 5, 6). Considering the growth delay of the TMTA96, TMTA109, and TMTA139 transformants compared to that of the parental strain RUT C30 in liquid culture (Fig. 2d), FPase, pNPCase, CMCase, and pNPGase activities were shown as U per mg T. reesei biomass to compare the differences in cellulase activities (Fig. 5). In TMTA96, TMTA109, and TMTA139, FPase, pNPCase, CMCase, and pNPGase activities were remarkably higher under glucose repression compared to those in the parental strain RUT C30 (Fig. 5a–d). The TMTA139 strain displayed the highest constitutive cellulase production using glucose as the sole carbon source compared to the other strains, with an FPase activity titer that was almost 26.5-fold higher than that obtained in T. reesei RUT C30 in shake flask culture at 72 h (Fig. 5a). The pNPCase (Fig. 5b), CMCase (Fig. 5c), and pNPGase activities (Fig. 5d) in TMTA139 were 0.07 U/mg biomass, 5.8 U/mg biomass and 0.05 U/mg biomass, which were about 31.5-, 22.4- and 24.4-fold, respectively, greater than that produced by T. reesei RUT C30 at 72 h. Moreover, the xylanase I (4.5 U/mg biomass) and xylanase II activities (5.5 U/mg biomass) in TMTA139 were approximately 11.8- and 11.5-fold, respectively, greater than that of RUT C30 at 72 h. Additionally, the extracellular protein concentration of TMTA96, TMTA109, and TMTA139 in glucose also increased 1.4-, 3.8-, and 7.7-fold compared to that of RUT C30 at 72 h (Fig. 5e), respectively, revealing that the protein production in TMTA96, TMTA109, and TMTA139 was improved in glucose. It could be confirmed by the SDS-PAGE (sodium dodecyl sulfate–polyacrylamide gel electrophoresis) assay (Additional file 1: Figure S2).
Cellulase and xylanase production in DBDcre1-VP16 transformants. DBDcre1-VP16 transformants and RUT C30 were cultured on 2% glucose. FPase (a), pNPCase (b), CMCase (c), pNPGase (d), xylanase I (e), and xylanase II (f) activity (U/g biomass), and extracellular secreted protein concentration (g) were measured at 48, 60, and 72 h. Error bars show the respective standard deviation of three biological replicates; asterisks indicate significant differences (*p ≤ 0.05) between the transformants and the parental strain RUT C30, as assessed by Student’s t test
Comparison of transcript levels of main cellulase, xylanase, transcription factor genes in DBDcre1-VP16 transformants. Gene expression ratios (-fold) in DBDcre1-VP16 transformants relative to RUT C30 on 2% glucose for 48 and 60 h. Gene expression ratios (-fold) were normalized to the corresponding gene expression at the same timepoint in the parental strain RUT C30. Values represent the mean of three biological replicates and error bars show the standard deviation; asterisks indicate significant differences (gene expression ratio greater than twofold or less than 0.5-fold between the transformants and the parental strain RUT C30)
To further investigate the effects of DBDcre1-VP16 on the synthesis of cellulase and xylanase, transcript levels of target genes at 48 and 60 h were analyzed using RT-qPCR (Fig. 6). The transcript levels of cbh1, cbh2, egl1, egl2, and bgl1 were markedly elevated in the TMTA96, TMTA109, and TMTA139 strains when cultivated in glucose, which is consistent with the observed increased cellulase activities. Among the DBDcre1-VP16 transformants, TMTA139 showed the best performance. The transcript levels of cbh1, cbh2, egl1, egl2, and bgl1 in the TMTA139 strain were upregulated 25.1-, 9.9-, 4.4-, 11.1-, and 4.0-fold, respectively at 60 h compared to those in the parental strain RUT C30 (Fig. 6). Moreover, the transcript levels of xyn1 and xyn2 in the TMTA139 strain were significantly upregulated 3.9- and 2.7-fold compared to that in RUT C30 at 60 h (Fig. 6). On the other hand, there was a notable increase in the transcription of xyr1 and ace2 in the DBDcre1-VP16 transformants (Fig. 6). In the TMTA139 strain, the transcript levels of xyr1 and ace2 were 6.8- and 4.8-fold, respectively, greater than that in RUT C30 at 60 h. The enhanced transcription of xyr1 and ace2 may account for the elevated transcript levels of cbh1, cbh2, egl1, egl2, and bgl1 [16, 17]. Additionally, the strong enhancement of the transcript levels of the swollenin gene swo1 (Fig. 6) was detected in the TMTA96, TMTA109, and TMTA139 strains.
Cellulase production in a jar fermentor and hydrolysis of corn stover by cellulase of TMTA66 and TMTA139 transformants
The strains TMTA66 in avicel and TMTA139 in glucose with the highest cellulase activities, consistent with the transcript levels, outperformed the other transformants and the parental strain RUT C30 in terms of cellulase production. Cellulase production by the TMTA66 and TMTA139 strains was further explored using a jar fermenter. The amount of secreted proteins and cellulase was significantly higher in these two strains than those in RUT C30 (Table 2). The TMTA66 strain showed maximum pNPCase (3.9 U/mL) and FPase (22.4 U/mL) activities after 5 days of cultivation. The TMTA139 strain reached an FPase activity of 4.5 U/mL after 3 days of cultivation (Table 2).
Saccharification by cellulase from TMTA66 and TMTA139 was determined via the hydrolysis of pretreated and biodetoxified corn stover [25]. The pretreated corn stover was first hydrolyzed to cellobiose during the catalysis of CBH and EG, and then cellobiose was hydrolyzed to glucose by ΒGL. Using the same FPase loading (15 U/g pretreated corn stover) without supplementation of commercial β-glucosidase, the glucose yields of the TMTA66 and TMTA139 strains showed similar low performances in saccharification of biomass to the glucose yield (66–71%) of the parental strain due to the lack of β-glucosidase produced by T. reesei (Fig. 7, Table 3, Additional file 1: Figure S3). With supplementation of commercial β-glucosidase, the glucose yields (98–99%) were almost the same as that obtained with the commercial enzyme CTec2 (Fig. 7, Table 3, Additional file 1: Figure S3). These results demonstrated that the enzymes from TMTA66 or TMTA139 supplemented with β-glucosidase were effective in hydrolyzing the pretreated corn stover.
Hydrolysis of pretreated corn stover by CTec2 and the crude enzyme from TMTA66 and TMTA139 using the same FPase loading. The crude enzymes from TMTA66 and TMTA139 were either supplemented with β-glucosidase (SUNSON®) (TMTA66+, TMTA139+) or without β-glucosidase (TMTA66, TMTA139) for a CBU/FPA ratio of 2. The commercial cellulase CTec2 (Novozymes) was used as the control. Enzyme was supplemented at 15 FPA/g biomass. Values represent the mean and standard deviation of triplicate measurements
Discussion
Cellulase and xylanase genes are majorly regulated at the transcript level in fungi [26]. In T. reesei, transcriptional activators play a significant role in the regulation of the production of cellulase and xylanase. Two important domains of transcription factors, the DNA-binding domain and the transcriptional domain, are essential for their function. Most of the transcription factors in T. reesei still retain other fungi-specific domains [16, 17, 27], which might have an on–off function for cellulase and xylanase regulation. The minimal transcriptional activators DBDace2-VP16 and DBDcre1-VP16 we constructed retained the core DNA binding domains of ACEII and CREI, respectively, did not contain redundant on–off regulation domains, and was fused with a strong transcriptional activation domain, VP16. VP16 was selected because of its compact structure (only 82 amino acids long) and stronger transcriptional activation than that of ACEII [28]. Retaining the core DNA binding domain of ACEII and CREI enabled our minimal transcriptional activators to competitively bind to promoters instead of the native transcription factors. The replacement of the VP16 strong transcriptional activation domain enhanced the transcription of downstream genes. Meanwhile, minimal transcriptional activators eliminated redundant on–off regulation domains to keep on an ON state. Thus, the minimal transcription activator DBDace2-VP16 increased cellulase expression under induction conditions (avicel and lactose) and DBDcre1-VP16 increased cellulase and xylanase expression under repression conditions (glucose).
Additionally, the minimal transcriptional activator DBDace2-VP16 enhanced cellulase production under induction conditions (Fig. 3). The transcript levels of related genes in DBDace2-VP16 transformants showed consistent results (Fig. 4). A previous study reported that the absence of ace2 decreased the transcript levels of the cellulase genes cbh1, cbh2, egl1, and egl2, and the total cellulase activity was reduced by 30–70% under cellulose induction conditions [17]. Therefore, the minimal transcription activator DBDace2-VP16 enhanced transcriptional activation compared to the original structure of ACEII, consequently increasing cellulase production. Interestingly, DBDace2-VP16 did not elevate the production of β-glucosidase and DBDace2-VP16 was ineffective in improving xylanase production. Deleting ace2 has no effect on xyn1 expression, according to a previous study [17]. It is feasible that xylanase production was not improved in DBDace2-VP16 transformants because of this. ACEII, however, can affect the expression of xyn2 [17, 29]. Phosphorylation and dimerization are required for ACEII to bind the xyn2 promoter [29]. The minimal transcriptional activator DBDace2-VP16 possibly impacted the phosphorylation and dimerization of the ACEII domain, inhibiting the enhancement of xyn2 expression in DBDace2-VP16 transformants. Moreover, the minimal transcriptional activator DBDace2-VP16 increased the transcript levels of swol gene, which contributed to the increased cellulase activity.
CREI acts as a carbon catabolite repressor of cellulase and xylanase [30]. However, minimal transcriptional activator DBDcre1-VP16 yielded a positive effect on cellulase and xylanase production (Figs. 5, 6). Mello-de-Sousa et al. [24] reported that the truncated CREI-96 protein of RUT C30 exhibits a positive regulation on the expression of target genes. Moreover, the CREI-96 protein can bind to the promoter of cbh1 and cbh2, resulting in increased gene expression with a simultaneous opening of chromatin structure in xyr1 [24]. XYRI is considered as the major transcriptional activator for cellulase and xylanase expression. The absence of xyr1 downregulates the expression of cellulase and xylanase in T. reesei [31]. Additionally, CREI binds to the promoter of xyn1 [32], and carbon catabolite repression (CCR) mediated by CREI can lead to the downregulated expression of xyn2 [33]. These explain our results regarding minimal transcriptional activator DBDcre1-VP16 acting as a positive regulator with enhanced xyr1 expression to improve cellulase and xylanase production in glucose (Fig. 6).
However, our minimal transcriptional activator transformants require further improvements. Though the high-yielding TMTA139 strain displayed constitutive cellulase production, with the production titer being 26.5-fold greater than that in T. reesei RUT C30 (Fig. 5a), this strain showed poor stability. After three generations, the TMTA139 strain began to retrogress, and the total FPase activity titer was substantially reduced; on the contrary, the growth rate significantly increased. Further optimization of TMTA139 strain stability is expected to be done in future studies.
Zhang et al. [22] reported the overexpression of an artificial transcription activator for constitutive cellulase production, which is the first report on cellulase production from glucose by T. reesei RUT C30 engineered with an artificial transcription factor. In related studies, artificial transcription activators were randomly inserted into the genome of T. reesei [20,21,22], which caused collateral mutations. Collateral mutations resulting in high cellulase production in T. reesei have been reported [34, 35]. Future studies will focus on overexpression, random insertion, and the combination of two sets of minimal transcriptional activators to further enhance cellulase production of superior cellulase-producing strains.
Transcription factors are switches that finely regulate gene expression and they enable organisms to better adapt to a particular environment. Transcription factors in T. reesei still retain their original evolutionary state [27]. To produce more cellulase and hemicellulase, minimal transcriptional activators offer an alternative genetic engineering strategy, and may enhance the production of industrial products in other fungi.
Conclusions
Minimal transcriptional activators were first constructed and transformed in T. reesei by fusing a DNA-binding domain of a native regulator with the VP16 activation domain to enhance cellulase production. Crude enzymes for both strains supplemented with commercial β-glucosidase were used to hydrolyze pretreated corn stover, and 98–99% glucose yields were obtained. These results demonstrate that replacing natural transcription factors with minimal transcription activators is a powerful strategy for enhancing cellulase production in T. reesei RUT C30. In addition, our current study also offers an alternative genetic engineering strategy for the enhanced production of industrial products in other fungi.
Methods
Strains
Trichoderma reesei RUT C30 (ATCC 56765), a hyper-cellulolytic strain, was used as the parental strain for genetic transformation and chromosomal DNA preparation. Escherichia coli DH5α and Agrobacterium tumefaciens AGL1 were used for plasmid construction and Agrobacterium-mediated transformation, respectively.
Construction of minimal transcriptional activators
The upstream and downstream sequences of ace2 and cre1 96 genes were amplified from T. reesei RUT C30 genomic DNA through PCR using the appropriate primers and used as the 5′ and 3′ homology arms of pMTA58/66/81 and pMTA96/109/139, respectively (Fig. 1). The different truncated versions of the DNA-binding domain sequence of ace2 (DBDace2–58, DBDace2–66, DBDace2–81) and cre 96 [24] were amplified from T. reesei RUT C30 genomic DNA via PCR, while the DBDcre1–109 and DBDcre1–139 (different truncated versions of the DNA-binding domain sequence of cre1) were cloned from T. reesei Qm9414 genomic DNA using the appropriate primers because the cre1 in RUT C30 was truncated and chimeric. All primers are listed in Additional file 1: Table S1. Linker and VP16 activation domain sequences were amplified from pG1V [28] through PCR. The resulting fragments were sequentially fused to linearized LML2.1 (digested by PacI/XbaI or SwaI) [36], by which resistance hygromycin B marker gene could be lost through xylose-induced Cre recombinase. All the vectors pMTA58, pMTA66, pMTA81, pMTA96, pMTA109, and pMTA139 (Fig. 1) were generated by the Seamless Cloning Kit (TransGen Biotech, Beijing, China). All plasmids were confirmed via DNA sequencing.
Transformation of T. reesei and characterization of the transformants
The generated plasmids were introduced into T. reesei RUT C30 via Agrobacterium-mediated transformation [37]. Clone verification and xylose-induced marker rescue [36] were performed to obtain the final transformants TMTA58/66/81 and TMTA96/109/139. The integration of the target gene-deleted constructs was analyzed using diagnostic PCR and sequencing. The single-copy DNA integration in transformed clones was verified by quantitative PCR (qPCR). The details are listed in Additional file 1: Table S1. For each minimal transcription activator, three final transformants were randomly selected and subcultured for subsequent enzyme production.
Biomass concentration assay
Conidia (106/mL) of final transformants and the parental strain RUT C30 were inoculated into 100 mL of minimal medium ((NH4)2SO4 5 g/L; Urea 0.3 g/L; KH2PO4 15 g/L; CaCl2 0.6 g/L; MgSO4 0.6 g/L; FeSO4·7H2O 5 mg/L; ZnSO4·7H2O 1.4 mg/L; CoCl2·6H2O 2 mg/L, pH 5.5) supplemented with 20 g/L glucose, lactose, or avicel in 500-mL Erlenmeyer flasks and incubated at 28 °C and 200 rpm for 72 h. Two milliliters of the culture liquid was collected every 12 h for biomass concentration assay. Biomass concentrations from glucose and lactose were measured gravimetrically according to the method of Corder et al. [38]. Biomass formation from avicel were indirectly determined by the amount of intracellular protein, as previously reported [39]. In brief, harvested mycelia were suspended in 1 mL 1 M NaOH in a reaction tube and the mixture was incubated for 2 h frequently being vortexed. The suspension was clarified via centrifugation at 14,000×g at 4 °C for 10 min. Total protein concentration of the suspension was determined by the Modified Lowry Protein Assay Kit (Sangon Biotech, Shanghai, China). The final protein content was corrected using a set of substrate controls where no inoculum was added to the avicel medium. The biomass dry weight was then calculated assuming an average content of 0.32 g intracellular protein per g of dry cell mass. Each experiment was performed in three biological replicates.
Cellulase production in a flask
To identify the cellulase production of TMTA58/66/81, 106/mL conidia of T. reesei strains were inoculated into 100 mL of Mandels-Andreotti medium [40] supplemented with 2% (w/v) glucose in a 500-mL shake flask incubated at 28 °C and 200 rpm for 36 h. Vegetative mycelia (0.4 g biomass) were collected by filtration, washed with distilled water, dried with sterile filter paper, and then subcultured into fresh 100 mL of Mandels-Andreotti medium supplemented with 2% (w/v) lactose or avicel in a 500-mL shake flask at 28 °C and 200 rpm. For TMTA96/109/139, 106/mL conidia of T. reesei strains were inoculated into 100 mL of Mandels-Andreotti medium supplemented with 2% (w/v) glucose in a 500-mL shake flask at 28 °C and 200 rpm for 72 h without medium replacement. Each experiment was performed in three biological replicates.
RNA extraction, reverse transcription, and real-time quantitative PCR (RT-qPCR) analysis
Total RNA was extracted using FastRNA Pro Red Kit (MP Biomedicals, Santa Ana, CA, USA) according to the manufacturer’s instructions. Synthesis of cDNA with 1000 ng total RNA was performed using TransScript All-in-One First-Strand cDNA Synthesis SuperMix for qPCR (TransGen, Beijing, China) following the manufacturer’s instructions. The transcript levels of target genes were assessed using real-time quantitative PCR (RT-qPCR) and normalized to that of the sar1 gene [41] using the 2−ΔΔCt method. The cycling conditions comprised 30 s initial denaturation and polymerase activation at 95 °C, followed by 40 cycles of 5 s at 95 °C and 60 s at 64 °C via an ABI StepOne Plus thermocycler (Applied Biosystems, Foster City, CA, USA). The primers are described in Additional file 1: Table S1. Threshold cycle (Ct) values and PCR efficiencies were used to calculate relative expression quantities by the ABI software. All samples were detected in three independent experiments with three replicates.
Enzyme assays, secreted protein concentration and SDS-PAGE assays
The supernatants collected via centrifugation (10,000×g for 10 min at 4 °C) were used for enzyme and secreted protein concentration assays. The FPase and CMCase activities were measured via the DNS method using glucose as a standard. One unit represents the amount of enzyme that formed 1 µmol of reducing sugar per minute during the hydrolysis reaction. The pNPCase and pNPGase activities were measured against p-nitrophenol-d-cellbioside (pNPC) and p-nitrophenyl β-d-glucopyranoside (pNPG) (Sigma-Aldrich, St. Louis, USA), respectively. One unit of pNPCase and pNPGase activity was defined as 1 μmol of p-nitrophenol released per minute during the hydrolysis reaction. Xylanase I and II activities were determined by xylan degradation at pH values of 3.7 and 5.0 [29], respectively. One unit of xylanase activity is defined as releasing 1 μmol of xylose reducing sugar equivalents per minute under the defined assay conditions. Protein concentration was determined using the Modified Lowry Protein Assay Kit (Sangon Biotech, Shanghai, China). All experiments were performed in three biological replicates. SDS-PAGE electrophoresis was carried out with 12% polyacrylamide separating gel.
Cellulase production in a jar
Selected dominant strains TMTA139 and TMTA66 were cultivated in a 7-L jar fermenter (BIOTECH-5BG-7000, Baoxing BIO-ENGINEERING EQUIPMENT, shanghai, China) with a final working volume of 3 L. The cultivations TMTA66 and parental strain RUT C30 were performed as follows. Conidia (106/mL) of strains were inoculated into 200 mL of Mandels-Andreotti medium supplemented with 1% (w/v) glucose and 1% (w/v) avicel in a 1-L shake flask and subsequently incubated with shaking (200 rpm) at 28 °C for 2 days. The culture was added to 2.8-L of fresh Mandels-Andreotti medium supplemented with 6% (w/v) avicel in a jar fermenter. Cultivation was carried out at 28 °C with 25% dissolved oxygen and 2 vvm (volumes of air per volume of liquid per minute) of aeration for 5 days. The pH was controlled within the range of 4.0–4.3 for the first 2 days and 5.0–5.2 thereafter.
The cultures of TMTA139 and the parental strain RUT C30 were performed as follows. Conidia (106/mL) of strains were inoculated into 200 mL of Mandels-Andreotti medium supplemented with 2% (w/v) glucose in a 1-L shake flask and incubated with shaking (200 rpm) at 28 °C for 2 days. The culture was poured into 2.8 L of Mandels-Andreotti medium fresh supplemented with 2% (w/v) glucose in the jar fermenter. Cultivation was carried out at 28 °C with 25% dissolved oxygen and 2 vvm of aeration for 3 days. The pH was controlled within the range of 5.0–5.2.
Enzymatic hydrolysis of corn stover by crude enzyme
Pretreated and biodetoxified corn stover was donated by Professor Jie Bao [25]. The pretreated corn stover was determined to contain 37.6% of cellulose and 4.4% hemicellulose in the dry mass. The crude enzymes produced by the T. reesei strains were supplemented with/without β-glucosidase (SUNSON®) with a CBU/FPA ratio of two to hydrolyze the corn stover. Hydrolysis experiments were performed in a flask containing 10% (w/v) pretreated corn stover as the substrate and FPase loading (15 U/g dry biomass) at 50 °C and pH 5.0 for 72 h. The methods for glucose analysis and calculation of glucose yield were based on the study by Li et al. [42].
Abbreviations
- MTA:
-
minimal transcriptional activators
- CBH:
-
cellobiohydrolases
- EG:
-
endoglucanases
- BGL1:
-
β-glucosidase I
- FPA/FPase:
-
the filter paper activity
- CMCase:
-
the CMC activity
- pNPCase:
-
the CBH activity
- pNPGase:
-
the β-glucosidase activity
- CBU:
-
cellobiase units
- U:
-
international units
- DNS:
-
3,5-dinitrosalicylic acid
- CCR:
-
carbon catabolite repression
- vvm:
-
volumes of air per volume of liquid per minute
- SDS-PAGE:
-
sodium dodecyl sulfate–polyacrylamide gel electrophoresis
- pNPC:
-
p-nitrophenol-d-cellobioside
- pNPG:
-
4-nitrophenyl-beta-d-galactopyranoside
- CMC:
-
sodium salt of caboxy methyl cellulose
- LPMOs:
-
lytic polysaccharide monooxygenases
References
Hahn-Hagerdal B, Galbe M, Gorwa-Grauslund MF, Liden G, Zacchi G. Bio-ethanol—the fuel of tomorrow from the residues of today. Trends Biotechnol. 2006;24:549–56.
Jäger G, Büchs J. Biocatalytic conversion of lignocellulose to platform chemicals. Biotechnol J. 2012;7:1122–36.
Biddy MJ, Davis R, Humbird D, Tao L, Dowe N, Guarnieri MT, Linger JG, Karp EM, Salvachúa D, Vardon DR, Beckham GT. The Techno-economic basis for coproduct manufacturing to enable hydrocarbon fuel production from lignocellulosic biomass. ACS Sust Chem Eng. 2016;4:3196–211.
Vicari KJ, Vicari KJ, Shatova T, Joo KK, Scarlata CJ, Humbird D, Wolfrum EJ, Beckham GT. Uncertainty in techno-economic estimates of cellulosic ethanol production due to experimental measurement uncertainty. Biotechnol Biofuels. 2012;5(1):1–12.
Kazi FK, Fortman J, Anex R, Kothandaraman G, Hsu D, Aden A, Dutta A. Techno-economic analysis of biochemical scenarios for production of cellulosic ethanol. Golden: National Renewable Energy Laboratory (NREL); 2010. https://doi.org/10.2172/982937.
Fang H, Xia L. High activity cellulase production by recombinant Trichoderma reesei ZU-02 with the enhanced cellobiohydrolase production. Bioresour Technol. 2013;144:693–7.
Peterson R, Nevalainen H. Trichoderma reesei RUT-C30—thirty years of strain improvement. Microbiology. 2012;158:58–68.
Bischof RH, Ramoni J, Seiboth B. Cellulases and beyond: the first 70 years of the enzyme producer Trichoderma reesei. Microb Cell Fact. 2016;15(1):106.
Kubicek CP, Mikus M, Schuster A, Schmoll M, Seiboth B. Metabolic engineering strategies for the improvement of cellulase production by Hypocrea jecorina. Biotechnol Biofuels. 2009;2:19.
Rosgaard L, Pedersen S, Langston J, Akerhielm D, Cherry JR, Meyer AS. Evaluation of minimal Trichoderma reesei cellulase mixtures on differently pretreated barley straw substrates. Biotechnol Prog. 2007;23(6):1270–6.
Miettinen-Oinonen A, Suominen P. Enhanced production of Trichoderma reesei endoglucanases and use of the new cellulase preparations in producing the stonewashed effect on denim fabric. Appl Environ Microbiol. 2002;68(8):3956–64.
Berlin A, Maximenko V, Gilkes N, Saddler J. Optimization of enzyme complexes for lignocellulose hydrolysis. Biotechnol Bioeng. 2007;97(2):287–96.
Zeilinger S, Mach RL, Schindler M, Herzog P, Kubicek CP. Different inducibility of expression of the two xylanase genes xyn1 and xyn2 in Trichoderma reesei. J Biol Chem. 1996;271:25624–9.
Saloheimo M, Paloheimo M, Hakola S, Pere J, Swanson B, Nyyssönen E, Bhatia A, Ward M, Penttilä M. Swollenin, a Trichoderma reesei protein with sequence similarity to the plant expansins, exhibits disruption activity on cellulosic materials. Eur J Biochem. 2002;269:4202–11.
Payne CM, Knott BC, Mayes HB, Hansson H, Himmel ME, Sandgren M, Stahlberg J, Beckham GT. Fungal cellulases. Chem Rev. 2015;115:1308–448.
Dos Santos Castro L, de Paula RG, Antonieto AC, Persinoti GF, Silva-Rocha R, Silva RN. Understanding the role of the master regulator XYR1 in Trichoderma reesei by global transcriptional analysis. Front Microbiol. 2016;7:175.
Aro N, Saloheimo A, Ilmen M, Penttila M. ACEII, a novel transcriptional activator involved in regulation of cellulase and xylanase genes of Trichoderma reesei. J Biol Chem. 2001;276:24309–14.
Lichius A, Seidl-Seiboth V, Seiboth B, Kubicek CP. Nucleo-cytoplasmic shuttling dynamics of the transcriptional regulators XYR1 and CREI under conditions of cellulase and xylanase gene expression in Trichoderma reesei. Mol Microbiol. 2014;94(5):1162–78.
Taylor LE 2nd, Knott BC, Baker JO, Alahuhta PM, Hobdey SE, Linger JG, Lunin VV, Amore A, Subramanian V, Podkaminer K, et al. Engineering enhanced cellobiohydrolase activity. Nat Commun. 2018;9:1186.
Su X, Chu X, Dong Z. Identification of elevated transcripts in a Trichoderma reesei strain expressing a chimeric transcription activator using suppression subtractive hybridization. World J Microb Biotechnol. 2009;25:1075–84.
Zhang F, Bai F, Zhao X. Enhanced cellulase production from Trichoderma reesei Rut-C30 by engineering with an artificial zinc finger protein library. Biotechnol J. 2016;11:1282–90.
Zhang X, Li Y, Zhao X, Bai F. Constitutive cellulase production from glucose using the recombinant Trichoderma reesei strain overexpressing an artificial transcription activator. Bioresour Technol. 2017;223:317–22.
Sadowski I, Ma J, Triezenberg S, Ptashne M. GAL4-VP16 is an unusually potent transcriptional activator. Nature. 1988;335:563.
Mello-de-Sousa TM, Gorsche R, Rassinger A, Poças-Fonseca MJ, Mach RL, Mach-Aigner AR. A truncated form of the Carbon catabolite repressor 1 increases cellulase production in Trichoderma reesei. Biotechnol Biofuels. 2014;7:129.
Qiu Z, Gao Q, Bao J. Constructing xylose-assimilating pathways in Pediococcus acidilactici for high titer d-lactic acid fermentation from corn stover feedstock. Bioresour Technol. 2017;245:1369–76.
Goncalves RD, Cupertino FB, Freitas FZ, Luchessi AD, Bertolini MC. A genome-wide screen for Neurospora crassa transcription factors regulating glycogen metabolism. Mol Cell Proteomics. 2011;10(11):M111.007963.
Lichius A, Bidard F, Buchholz F, Le Crom S, Martin J, Schackwitz W, Austerlitz T, Grigoriev IV, Baker SE, Margeot A, et al. Genome sequencing of the Trichoderma reesei QM9136 mutant identifies a truncation of the transcriptional regulator XYR1 as the cause for its cellulase-negative phenotype. BMC Genomics. 2015;16:326.
Wang W, Shi XY, Wei DZ. Light-mediated control of gene expression in filamentous fungus Trichoderma reesei. J Microbiol Methods. 2014;103:37–9.
Stricker AR, Trefflinger P, Aro N, Penttilä M, Mach RL. Role of Ace2 (Activator of Cellulases 2) within the xyn2 transcriptosome of Hypocrea jecorina. Fungal Genet Biol. 2008;45:436–45.
Ilmen M, Thrane C, Penttila M. The glucose repressor gene cre1 of Trichoderma: isolation and expression of a full-length and a truncated mutant form. Mol Gen Genet. 1996;251:451–60.
Akel E, Metz B, Seiboth B, Kubicek CP. Molecular regulation of arabinan and l-arabinose metabolism in Hypocrea jecorina (Trichoderma reesei). Eukaryot Cell. 2009;8:1837–44.
Mach RL, Strauss J, Zeilinger S, Schindler M, Kubicek CP. Carbon catabolite repression of xylanase I (xyn1) gene expression in Trichoderma reesei. Mol Microbiol. 1996;21(6):1273–81.
Rassinger A, Mello-de-Sousa TM, Regnat K, Derntl C, Mach RL, Mach-Aigner AR. Impact of xylanase expression-inducing compounds on DNA accessibility in Trichoderma reesei. Mycosphere. 2017;8(3):432–44.
Li C, Lin F, Li Y, Wei W, Wang H, Qin L, Zhou Z, Li B, Wu F, Chen Z. A β-glucosidase hyper-production Trichoderma reesei mutant reveals a potential role of cel3D in cellulase production. Microb Cell Fact. 2016;15:151.
Li C, Lin F, Zhou L, Qin L, Li B, Zhou Z, Jin M, Chen Z. Cellulase hyper-production by Trichoderma reesei mutant SEU-7 on lactose. Biotechnol Biofuels. 2017;10:228.
Zhang L, Zhao X, Zhang G, Zhang J, Wang X, Zhang S, Wang W, Wei D. Light-inducible genetic engineering and control of non-homologous end-joining in industrial eukaryotic microorganisms: LML 3.0 and OFN 1.0. Sci Rep. 2016;6:20761.
Michielse CB, Hooykaas PJ, Ca VDH, Ram AF. Agrobacterium-mediated transformation of the filamentous fungus Aspergillus awamori. Nat Protoc. 2008;3:1671–8.
Corder GW, Foreman DI. Nonparametric Statistics for Non-Statisticians: A Step-by-Step Approach. NY: Wiley; 2009.
Bischof R, Fourtis L, Limbeck A, Gamauf C, Seiboth B, Kubicek CP. Comparative analysis of the Trichoderma reesei transcriptome during growth on the cellulase inducing substrates wheat straw and lactose. Biotechnol Biofuels. 2013;6:127.
Mandels M, Andreotti RE. Problems and challenges in the cellulose to cellulase fermentation. Process Biochem. 1978;13:6–13.
Steiger MG, Mach RL, Mach-Aigner AR. An accurate normalization strategy for RT-qPCR in Hypocrea jecorina (Trichoderma reesei). J Biotechnol. 2010;145:30–7.
Li Y, Liu C, Bai F, Zhao X. Overproduction of cellulase by Trichoderma reesei RUT C30 through batch-feeding of synthesized low-cost sugar mixture. Bioresour Technol. 2016;216:503–10.
Authors’ contributions
WW1 initiated, designed, and coordinated the study and reviewed the manuscript. JZ planned and carried out experiments and measurements and interpreted experimental data. GZ analyzed the partial data. DW and WW2 supported the research funding. All authors read and approved the final manuscript.
Acknowledgements
We wish to thank Prof. Jie Bao (State key Lab of Bioreactor Engineering, East China University of Science and Technology, Shanghai, China.) for providing the pretreated and bio-detoxified corn stover.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
All data generated or analysed during this study are included in this published article and its additional file.
Consent for publication
All authors agreed to publish this article.
Ethics approval and consent to participate
Not applicable.
Funding
This research was supported by the National Natural Science Foundation of China (No. C010302-31500066) and the Fundamental Research Funds for the Central Universities (No. 222201714053).
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Author information
Authors and Affiliations
Corresponding author
Additional file
Additional file 1: Table S1.
All primers used for this study. Figure S1. The verification of the single-copy DNA integration in transformed clones by diagnostic PCR and qPCR. Figure S2. SDS-PAGE analysis of extracellular proteins secreted by T. reesei RUT C30 and TMTA96, TMTA109, TMTA139. Figure S3. Hydrolysis of pretreated corn stover by CTec2 and the crude enzyme from TMTA66, TMTA139 and RUT C30 using the same FPase loading.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Zhang, J., Zhang, G., Wang, W. et al. Enhanced cellulase production in Trichoderma reesei RUT C30 via constitution of minimal transcriptional activators. Microb Cell Fact 17, 75 (2018). https://doi.org/10.1186/s12934-018-0926-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12934-018-0926-7