Abstract
Background
Although vertebrates are bilaterally symmetric organisms, their internal organs are distributed asymmetrically along a left-right axis. Disruption of left-right axis asymmetric patterning often occurs in human genetic disorders. In zebrafish embryos, Kupffer’s vesicle, like the mouse node, breaks symmetry by inducing asymmetric expression of the Nodal-related gene, spaw, in the left lateral plate mesoderm (LPM). Spaw then stimulates transcription of itself and downstream genes, including lft1, lft2, and pitx2, specifically in the left side of the diencephalon, heart and LPM. This developmental step is essential to establish subsequent asymmetric organ positioning. In this study, we evaluated the role of krüppel-like factor 8 (klf8) in regulating left-right asymmetric patterning in zebrafish embryos.
Methods
Zebrafish klf8 expression was disrupted by both morpholino antisense oligomer-mediated knockdown and a CRISPR-Cas9 system. Whole-mount in situ hybridization was conducted to evaluate gene expression patterns of Nodal signalling components and the positions of heart and visceral organs. Dorsal forerunner cell number was evaluated in Tg(sox17:gfp) embryos and the length and number of cilia in Kupffer’s vesicle were analyzed by immunocytochemistry using an acetylated tubulin antibody.
Results
Heart jogging, looping and visceral organ positioning were all defective in zebrafish klf8 morphants. At the 18–22 s stages, klf8 morphants showed reduced expression of genes encoding Nodal signalling components (spaw, lft1, lft2, and pitx2) in the left LPM, diencephalon, and heart. Co-injection of klf8 mRNA with klf8 morpholino partially rescued spaw expression. Furthermore, klf8 but not klf8△zf overexpressing embryos showed dysregulated bilateral expression of Nodal signalling components at late somite stages. At the 10s stage, klf8 morphants exhibited reductions in length and number of cilia in Kupffer’s vesicle, while at 75% epiboly, fewer dorsal forerunner cells were observed. Interestingly, klf8 mutant embryos, generated by a CRISPR-Cas9 system, showed bilateral spaw expression in the LPM at late somite stages. This observation may be partly attributed to compensatory upregulation of klf12b, because klf12b knockdown reduced the percentage of klf8 mutants exhibiting bilateral spaw expression.
Conclusions
Our results demonstrate that zebrafish Klf8 regulates left-right asymmetric patterning by modulating both Kupffer’s vesicle morphogenesis and spaw expression in the left LPM.
Similar content being viewed by others
Background
Despite the outward appearance of bilateral symmetry in vertebrates, internal organs exhibit substantial left-right asymmetry. In humans, genetic disorders that affect left-right asymmetric patterning may result in organ heterotaxy [1], complex congenital heart disease, and asplenia/polysplenia [2]. In order to study the various processes that establish left-right asymmetry in a laboratory setting, several vertebrates, including mice and zebrafish, have been utilized. Largely based on these animal studies, the major developmental processes which establish asymmetry are known to include: symmetry-breaking in the node, the transfer of asymmetric Nodal expression from the node to the left lateral plate mesoderm (LPM), asymmetric expression of Nodal and downstream genes in the left LPM, and the completion of left-right asymmetric organ morphogenesis [3, 4].
Clockwise rotation of nodal cilia creates a directional nodal flow, which is responsible for the preferential activation of Nodal expression on the left side of the embryo [5]. Not surprisingly, mutations in genes involved in ciliogenesis [6] or its regulation [7, 8] have been found to disrupt normal left-right patterning. Leftward nodal flow generates an initial accumulation of NODAL protein on the left side of the embryo. Subsequently, self-enhancement and lateral-inhibition systems involving NODAL, LEFTY1 and LEFTY2 reinforce the asymmetric distribution and restrict Nodal gene expression to the left side of the organism [9].
Nodal signalling is initiated by the binding of NODAL to the ACTIVIN receptor and EGF-CFC co-receptor, which results in the formation of an intracellular regulatory complex. This complex consists of phosphorylated SMAD2, SMAD4 and FoxH1, and directly activates target gene transcription [10]. Left-side specific enhancers (ASEs) with FoxH1 binding motifs are present in the murine Nodal and Lefty2 genes [11]. Thus, NODAL amplifies its own expression in the left LPM via SMADs/FoxH1 interaction with the ASE. Simultaneously, NODAL induces Lefty2 expression, which inhibits low-level NODAL signalling, and thereby restricts Nodal expression to the left LPM. This asymmetric NODAL activation induces expression of Pitx2 in the left LPM, via its ASE. PITX2 is a homeodomain transcription factor implicated in left-right asymmetric organ morphogenesis [12], and loss of Pitx2 expression has been shown to affect the asymmetric distribution of internal organs in several vertebrates [13].
In zebrafish embryos, Kupffer’s vesicle (KV) performs a similar role to the mouse node in initiating left-right asymmetric patterning [14]. KV is derived from dorsal forerunner cells (DFCs), which are formed via a Nodal signalling-dependent ingression of surface enveloping layer cells from the dorsal blastoderm margin. This ingression occurs at the blastula stage, when embryonic epiboly initiates [15]. DFCs then migrate toward the vegetal pole and organize into multiple rosette-like, epithelial structures at the end of gastrulation. These epithelial rosettes then merge into a single epithelial rosette and differentiate into the ciliated KV, with the vesicle lumen arising from apical membrane expansion during early somite stages. Tilted cilia are positioned with the basal body at the posterior pole of DFCs, and these motile cilia are asymmetrically distributed along the anterior-posterior axis of KV. Furthermore, cilia asymmetry is established by the Rho kinase, Rock2b, and is essential to generate an anti-clockwise swirling flow that commences asymmetric Nodal signalling [16,17,18]. In addition to rock2b, deficiencies in several genes involved in the movement, formation or positioning of cilia, such as dnah9, ift88, and vangl2, have been shown to disrupt normal left-right patterning [14, 19, 20].
Three nodal-related genes, namely ndr2 (cyc), ndr1 (sqt), and southpaw (spaw), have been identified in zebrafish [10]. Among these genes, spaw exhibits the earliest expression in the left-side of the LPM, and stimulation of its own transcription during somitogenesis shifts its expression domain from the posterior to the anterior left LPM [21, 22]. Furthermore, morpholino knockdown of spaw decreases expression of genes encoding Nodal signalling components, including spaw, lefty1 (lft1), lefty2 (lft2), and pitx2, in the left LPM [22], affecting the left-right asymmetric distribution of heart, pancreas, and diencephalon. Together, these studies demonstrate the essential role of Spaw, and underscore its relevance as a NODAL homolog in establishing left-right asymmetry of teleosts [22, 23].
Similar to mouse embryos, different repressors of the Nodal signalling pathway have been reported to modulate the induction or maintenance of asymmetric Nodal signalling in teleosts. At the 4–6 s stages, spaw is expressed bilaterally in KV, while charon is expressed in a region adjacent to KV, where it antagonizes Spaw activity and contributes to biased spaw expression in the left LPM [24]. Furthermore, repression of Spaw activity in the right LPM or cardiac field by Lft1 or Lft2 is also essential in the establishment of left-right patterning. Notably, lft1 expression in the notochord is induced via binding of BMP4 with BMP receptor 1aa at the early segmentation stage, while lft2 expression in the left cardiac field is activated by Spaw in the anterior LPM [25,26,27]. Despite this detailed knowledge about Spaw repressor proteins, it is still unknown whether asymmetric spaw expression in the left LPM can be regulated by transcription factors.
Krüppel-like factor 8 (KLF8) is a member of the KLF family of transcription factors [28, 29], and participates in a broad range of developmental processes. KLF proteins contain C-terminal zinc finger DNA binding motifs, and distinct N-terminal regulatory elements. KLF8, like KLF3 and KLF12, possesses a regulatory domain that interacts with C-terminal binding protein (CtBP) [30]. Interaction of KLF8 with the co-repressor CtBP inhibits embryonic Gamma-Globin gene expression [31, 32], a role confirmed in klf8-deficient mice [33]. KLF8 also functions as a mediator of focal adhesion kinase to activate cyclin D1 expression, modulating cell cycle progression [34]. Recently, we used morpholino knockdown and rescue experiments to show that zebrafish Klf8 has a novel function in cerebellar development. In this context, Klf8 modulates the expression of p53 and met to maintain ptf1a-expressing neuronal progenitors, which are required for proper development of cerebellar Purkinje and granule cells [35]. In addition, we noted that klf8 morphants often exhibited a no-loop heart at 48 h post fertilization (hpf).
In this study, we demonstrate that zebrafish Klf8 plays an additional role in regulating left-right asymmetric patterning. Heart jogging, looping and visceral organ positioning were defective in klf8 morphants. At 18–22 s stages, expression levels of spaw, lft1, lft2, and pitx2 were decreased or eliminated in the left LPM, diencephalon, and heart of the majority of klf8 morphants. In contrast, klf8 overexpression resulted in bilateral expression of spaw and its downstream target genes in these tissues. Both dorsal forerunner cell number, and the length and number of cilia in KV were also affected in klf8 morphants. However, klf8 CRISPR-Cas mutant embryos showed bilateral spaw expression in the LPM, which may have been partly due to compensatory upregulation of klf12b.
Methods
Ethics approval
All animal procedures were approved by the Institutional Animal Care and Use Committee of Academia Sinica (Protocol ID: 15–12-918).
Zebrafish maintenance and staging
Adult AB zebrafish, Tg(sox17:gfp) s870/+ and klf8 mutants (klf8 d25, klf8 i17), generated by a CRISPR-Cas9 system were maintained in high density, self-circulation systems (Aqua Blue), or 20 L aquaria supplied with filtered fresh water and aeration under a photo period of 14 h light and 10 h dark. Embryos were maintained at 28.5 °C, and morphological criteria were defined as described [36].
Plasmid construction, morpholino and mRNA injection
The full-length klf8 coding sequence or klf8 lacking zinc finger domain (klf8△zf) was cloned into the T7TS vector, and used as template to synthesize capped mRNA with the mMESSAGE mMACHINE T7 Kit (Ambion). Previously published morpholinos (MOs) were used [35], including two MOs that prevent Klf8 protein translation: klf8-MO1atg (2.2 ng) and klf8-MO2atg (1.9 ng), two MOs that prevent klf8 mRNA splicing: MODO (0.73 ng) and MOAC (0.73 ng), and one control MO: klf8-4 mm MO1 (2.2 ng). An additional MO to prevent klf8 mRNA splicing: MODO2 (0.73 ng; 5′- TGGGTCACATCCATCTCACCTGATC -3′; targets the donor site of exon 3) was used. A 1.5-fold greater dosage of P53 MOsp [37] was injected, as compared to the co-injected klf8-MOs. To verify upregulation of klf12b in homozygous klf8 d25 mutant F6 embryos, klf12b MO (5 or 10 ng) was used. klf12b MO (5′-ATTCCGTCTAGCATTAACATCCTGT-3′), which is complementary to 20 bases of the coding region including the ATG start codon (underlined) and five bases of the 5′ untranslated region, was used. The indicated MO or mRNA was microinjected into one- or two-cell zygotes using a Nanoject II automatic injector (Drummond).
Whole mount in situ hybridization, whole mount immunohistochemistry, and photography
Whole mount in situ hybridization was performed on embryos treated with 0.003% phenylthiocarbamide, using digoxigenin-antisense RNA probes and alkaline phosphatase-conjugated anti-digoxigenin antibody. Various templates derived from pGEMT or pGEMT-Easy vectors were linearized, and the following antisense RNA probes were generated (restriction site and promoter in parentheses): bmp2b (BamHI/T7), charon (BamHI /Sp6), myl7 (NcoI /SP6), gata5 (SacII/SP6), gata 6 (SalI/T7), lft1 (SalI/T7), lft2 (HindIII/SP6), ntl (XhoI/T7), oep (NcoI/SP6), pitx2c (EcoRI/T7), and spaw (SpeI/T7). To produce the dvr1 antisense RNA probe, PCR product that was generated using M13 forward and M13 reverse primers was used as a template and transcribed by T7 RNA polymerase.
To detect changes in KV cilia, one- or two-cell zygotes of Tg(sox17:gfp) were microinjected with different klf8-MOs. The 10s stage embryos were fixed in 4% paraformaldehyde for 2 h at room temperature (RT) and dehydrated in methanol at −20 °C. After dehydration, the embryos were permeabilized using acetone at −20 °C and washed by Phosphate-buffered saline with tween 20 (PBST) followed by blocking in 5% serum. The embryos were incubated with anti-acetylated tubulin antibody (1:250, Sigma-Aldrich) for 2 h, at RT, followed by mouse Alexa Fluor 568 for 1 h, at RT (1:250, Invitrogen).
To investigate DFC number alteration, one- or two-cell zygotes of Tg(sox17:gfp) were microinjected with different klf8-MOs. 75% embryos were fixed in 4% paraformaldehyde at 4 °C overnight. After dehydration, the embryos were permeabilized using acetone at −20 °C for 7 min and treated with 0.15 M Tris-HCl, pH 9.0 at 70 °C for 15 min. The embryos were washed with PBST followed by blocking in 1% blocking solution (Roche) at RT for 2 h. The embryos were incubated with anti-GFP antibody (1: 200) at 4 °C overnight, followed by rabbit Alexa Fluor 488 (1:200, Invitrogen) incubation for 1 h, at RT. Nuclei were then stained with Hoechst 33,341 (1:1000 in PBST, Invitrogen).
Brightfield embryo images were taken using an AxioCam HRC camera mounted on a Zeiss Imager M1 microscope. Fluorescent images were taken using a Leica TCS-SP5-MP confocal microscope.
Generation of klf8 mutants using CRISPR-Cas9 system
klf8 mutants were generated using a CRISPR-Cas9 system targeting exon 2. sgRNA was designed by ZGENEBIO BIOTECH INC. (Taipei, Taiwan) and DNA template was amplified from pZGB plasmids containing klf8 sgRNA. PCR was conducted using forward (5′-ACACAGGAAACAGCTATGACCATG-3′) and reverse (5′-GATCCG CACCGACTCGGTGCCACTTT-3′) primers, and klf8 sgRNA were synthesized using the MEGAshortscript T7 Transcription Kit (Ambion, Austin, TX, USA). klf8 sgRNA (86.3 pg) and capped nls-zCas9-nls mRNA (34.5 pg, Addgene) [38] were co-injected into one-cell zygotes. Genomic DNA was isolated from pools of 10 embryos at 24 hpf. PCR was conducted using forward (5′- TCTTTCTACTCCTCCCCCAACTAA-3′) and reverse (5′- CCACACCCCTTTCCAATAACTCTA-3′) primers and amplified DNA was then digested with T7 endonuclease I to evaluate insertion and deletion efficiency. The rest of the embryos were reared to adulthood. Injected fish were designated as the F0 generation. To detect the DNA sequence alterations induced by klf8 sgRNA, genomic DNA was isolated from clipped tail fin of adult F1 fish, and high resolution melt analysis was performed. PCR was conducted in a 20 μL reaction comprising 8–12 ng genomic DNA, 3.5 mM MgCl2, 1× Master Mix containing Taq DNA polymerase, dNTP mix and LightCycler 480 ResoLight dye, and 5 pmol each of forward (5′- ATCTCAGAACTCGGGTCACTTTTT-3′) and reverse (5′- CCACCATACACTCCACCTCCTC-3′) primers. The PCR conditions were 95 °C for 300 s, 1 cycle for pre-incubation, 95 °C for 10 s, 65 °C to 53 °C with a 0.5 °C gradient decline for 10 s, and 72 °C for 10 s, for 70 cycles of amplification, and 95 °C for 60 s, 40 °C for 60 s, 65 °C to 95 °C for 1 s melting interval for high resolution melting. DNA sequencing was conducted to confirm F1 adult fish with induced DNA sequence alterations. Two klf8 F1 mutants including klf8 d25 with a 25 bp deletion and klf8 i17 with a 17 bp insertion in the exon 2 klf8 sgRNA target site were crossed with wild type fish to produce the F2 generation. Subsequently, homozygous F4 klf8 d25 and klf8 i17 mutants were generated by intercrossing of respective heterozygous F3 mutants and maintained to the F5 generation.
Reverse transcription PCR (RT-PCR) and reverse transcription quantitative real-time PCR (RT-qPCR)
RT-PCR was used to evaluate the efficacy of sp.-MOs. cDNA from 24 hpf, forward primer (5′- ATCAAGCCGGAGCCAGAGGAGGTG-3′) and reverse primer (5′-GCCGTCGGTGAAGTGCCAGGTG-3′) were used. RT-qPCR [39] was used to compare expression levels of klf3, klf12a or klf12b in wild type and homozygous F5 klf8 d25 and klf8 i17 mutant embryos. cDNA was generated by a GoScript Reverse transcription system (Promega) using total RNA isolated from 10 to 12 s stage wild type or two homozygous klf8 d25 and klf8 i17 mutant embryos. RT-qPCR was conducted in a 10 μL reaction containing 1× LightCycler 480 SYBR Green I Mix (Roche), respective primer pairs (5 μg) and 1/10 cDNA from wild type or mutant embryos. PCR conditions were 95 °C for 10 min, then 55 cycles of, 95 °C for 10 s, 60 °C for 10 s, and 72 °C for 10 s, followed by a 4 °C pause. Primer pairs for klf3 were forward (5′-TATCCAAGTGGACATTACTGTGGG-3′) and reverse (5′-CAGTGGGCAACACAGAACGGCAG-3′). Primer pairs for klf12a were forward (5′-GAGCGGGTCTCTTTCTGCCAGTG-3′) and reverse (5′-CAATAAACCGTATGAGGGAAAGGC-3′). Primer pairs for klf12b were forward (5′-GGCAATCCCTGCTCCTCAGAAAC-3′) and reverse (5′-CCACATCGTAGACTCCAAAATGCG-3′).
Quantification of cilia number and length and lumen of Kupffer’s vesicles as well as DFC number
The cilia length and number were quantified using LAS AF and MetaMorph software according to the following steps: (i) merge images with LAS AF for MetaMorph analysis; (ii) “Threshold Image” was set to demarcate cilia and KV cell locations, and the image was converted to grayscale; (iii) from the Arithmetic menu, “Logical AND” was selected, and KV cilia regions were defined; (iv) “Calibrate Distances” was set to define units of length (μm); and (v) the length and number of cilia were quantified by selecting “Integrated Morphometry Analysis” in the Measure menu. The area of KV lumen was quantified using ImageJ software as follows: (i) The merged grayscale images from MetaMorph were loaded in ImageJ; (ii) “Elliptical selections” was selected to demarcate cilia area; (iii) “Set scale” was selected to define units of area (μm2); and (iv) the area was determined by selecting “Measure” in the Analyze menu.
In order to evaluate the DFC number, immunofluorescence confocal images of Tg(sox17:gfp) 75% epiboly embryos were merged using ImageJ software using the following steps: (i) Images were loaded into ImageJ; (ii) “Images to stack” was selected from the “Stacks” item in the Image menu; (iii) “Stacks Focuser” was selected from the “Stacks” item in the Plugins menu. DFC number was then manually counted from images with merged GFP and nuclei.
Statistics
Two-tailed Student’s t-tests with unequal variance were conducted to compare number of cilia, cilia length, lumen area and DFC. To compare the klf8 mRNA rescue effect, Fisher’s Exact Test was used. Statistical tests were performed with Excel software. Differences with p < 0.05 were considered to be statistically significant.
Results
klf8 morphants display abnormal heart jogging, looping and visceral organ positions
During a previous study investigating the role of Klf8 in cerebellar development [35], we noted that klf8 morphants often exhibited a no-loop heart at 48 hpf. Because cardiac development is asymmetric, this observation suggested that Klf8 may regulate the general process of left-right patterning. Thus we performed klf8 knockdown experiments with previously validated klf8-MOs and systematically evaluated heart morphogenesis by whole-mount in situ hybridization, using a myl7 antisense RNA probe. A phenotypically normal, L-jog heart tube was readily detected in wild type and klf8-4 mm MO1-injected embryos, while embryos injected with klf8-MO1atg or klf8-MO2atg often showed no-jog (32% for MO1atg, 33% for MO2atg) or right-jog (4% for MO1atg, 3% for MO2atg) heart tubes at 24 hpf (Fig. 1a). Consequently, embryos injected with klf8-MO1atg or klf8-MO2atg frequently developed a no-loop heart (53% for MO1atg, 40% for MO2atg) as compared to the vast majority of wild type or klf8-4 mm MO1-injected embryos showing a D-loop heart at 48 hpf (Fig. 1b). At 72 hpf, a similar percentage of klf8-MO1atg- or klf8-MO2atg-injected embryos displayed a no-loop heart (40% for MO1atg, 38% for MO2atg) as compared to wild type or klf8-4 mm MO1-injected embryos, which almost invariably had a D-loop heart (Fig. 1c). Embryos showing a delayed heart cone phenotype were also identified in the klf8-MO1atg (13%) or klf8-MO2atg (27%) groups at 24 hpf (Fig. 1a). However, this delayed phenotype was not observed in the 48 hpf or 72 hpf time points. By these stages, the delayed morphants caught up developmentally and displayed either D-loop, no-loop or L-loop hearts (Fig. 1b&c). We also examined the position of digestive organs in klf8 morphants by whole-mount in situ hybridization, using a gata6 antisense RNA probe. In klf8-MO1atg- or klf8-MO2atg-injected embryos, frequent occurrence of organ dysmorphologies were observed. Reversal of liver and pancreas position (20% for MO1atg, 11% for MO2atg), only intestine development (15% for MO1atg, 3% for MO2atg), and bilateral liver and pancreas (1% for MO1atg, 1% for MO2atg) were all frequently detected in klf8 morphants, but were rare events in wild type or control embryos at 54 hpf (Fig. 1d). These results confirmed that klf8 loss-of-function affects left-right patterning in zebrafish embryos.
Knockdown of zebrafish klf8 caused defects in heart jogging and looping, and visceral organ positions. a klf8-MO1atg or klf8-MO2atg -injected embryos stained with myl7 exhibited left (L)-jog, no-jog or right (R)-jog and were compared to stained wild-type or 4 mm-MO1-injected control embryos at 24 hpf. b myl7 stained embryos injected with klf8-MO1atg or klf8-MO2atg displayed D-loop, no-loop or L-loop heart and were compared to wild type and control embryos at 48 hpf. c myl7 stained embryos injected with klf8-MO1atg or klf8-MO2atg displayed D-loop, no-loop or L-loop heart and were compared to wild type and control embryos at 72 hpf. d At 54 hpf, gata6 stained wild type or embryos injected with klf8-4 mm MO1, klf8-MO1atg or klf8-MO2atg exhibited organ positions that were classified as: (Normal) normal positions of liver-left, pancreas-right and intestine with left looping, (Intestine only) only intestine without left looping, (Reverse) reversed position of liver-right, pancreas-left and no looping intestine, or (Bilateral) bilateral extension of liver and pancreas. A, atrium; I, intestine; L, liver; P, pancreas; V, ventricle
klf8 deficiency affects the level and pattern of expression for genes in the Nodal signalling pathway
Genes encoding Nodal signalling components, including spaw, lft1, lft2, and pitx2, were asymmetrically expressed in the left side of the diencephalon, heart, or lateral plate mesoderm (LPM) during the 18–22 s stages in zebrafish embryos (Fig. 2a–m). Disruption in the expression of left-side specific Nodal signalling genes results in organ heterotaxy. We observed that spaw expression in the left LPM was either decreased (no expression in the anterior LPM and low expression in the posterior LPM, 27%) or absent (46%) in many klf8-MO1atg-injected embryos (Fig. 2d–f). A similar effect (32% decreased, 25% absent, 8% bilateral, 5% right) was observed following klf8-MO2atg injection (Fig. 2b-f). Consistent with these results, the expression of genes downstream of spaw (lft1, lft2, and pitx2) was also absent or decreased in the left diencephalon, heart, and LPM of most klf8-MO1atg or klf8-MO2atg -injected embryos (Fig. 2h–o). Additionally, we injected three splicing MOs (MODO2, MOAC, and MODo) to block splicing of klf8 mRNA (Additional file 1: Figure S1, A). RT-PCR indicated that splicing of klf8 mRNA was effectively disrupted in MOsp-MOs-injected embryos at 24 hpf (Additional file 1: Figure S1, B). Expression of spaw in the left LPM was either decreased (8%) or eliminated (63%) in most 18 s stage embryos injected with MOsp-MOs (Additional file 1: Figure S1, C).
Genes encoding Nodal signalling components exhibited decreased or abolished expression in klf8 morphants. The majority of klf8-MO1atg or klf8-MO2atg -injected embryos showed decreased or abolished spaw (arrow) expression in the left lateral plate mesoderm (LPM) at the 18 s stage (a-f). The majority of klf8 morphants failed to express lft1 in the left diencephalon (arrowhead) or heart (asterisk) (g- i), lft2 in the left heart (asterisk) (j-l), and pitx2 in the left LPM (white arrow) (m-o) at the 22 s stage. Dorsal views of embryos are shown. A, absent; B, bilateral; D, decreased; L, left; R, right
Klf8 was previously shown to repress p53 expression and induce met expression to modulate the development of Purkinje cells and proliferation of granule cells [35]. To confirm that defective left-right patterning did not arise from induction of p53 due to klf8 deficiency, we analysed heart looping and gene expression of Nodal signalling components in embryos that were co-injected with p53-MOsp, alongside klf8-MO1atg or klf8-MO2atg.
Of the embryos co-injected with p53-MOsp, together with klf8-MO1atg or klf8-MO2atg, 35–39% exhibited a no-loop heart at 72 hpf, similar to embryos injected with klf8 MOs alone (Additional file 2: Figure S2, A). Likewise, expression levels of spaw, lft1, lft2, and pitx2 were reduced or eliminated in the left LPM, diencephalon, and heart of high percentages of co-injected embryos during the 18–22 s stages (Additional file 2: Figure S2, B-E). We also found that the expression levels of gata5 and oep (which are known to be expressed in the LPM at the 22 s stage) were unaffected by klf8 knockdown (Additional file 2: Figure S2, F-M). Together these data clearly indicate that increased p53 expression and apoptosis are not responsible for the decreased expression of genes involved in Nodal signalling.
To further confirm that the decrease in spaw expression is a consequence of klf8 loss-of-function, we performed rescue experiments by co-injecting embryos with klf8-MO1atg and klf8 mRNA. Approximately 69% of klf8-MO1atg-injected embryos exhibited eliminated or decreased spaw expression in the left LPM at the 18 s stage (Fig. 3a, e). However, this proportion showed a statistically significant reduction to 44% for embryos co-injected with klf8-MO1atg and klf8 mRNA (Fig. 3b-e). Taken together, these results demonstrate that klf8 loss-of-function causes downregulation of Nodal signalling component genes.
Reduced spaw expression in klf8 morphants was partially rescued by co-injection of klf8 mRNA. Representative embryos showing spaw expression in the left LPM (spaw+) or absent spaw expression (spaw-) are shown (a-d). Percentages of embryos with asymmetric spaw expression or no spaw expression are shown with indicated treatments (e). Statistical significance was determined by Fisher’s Exact Test. *p < 0.05
Overexpression of klf8 mRNA causes bilateral expression of genes involved in Nodal signalling
Since klf8 knockdown reduced expression of genes involved in Nodal signalling, we hypothesized that klf8 overexpression may have the opposite effect. While the majority of embryos injected with 100 pg of LacZ mRNA expressed spaw exclusively in the left LPM at the 18 s stage, 24% of embryos injected with 50 pg and 51% of embryos injected with 100 pg of klf8 mRNA expressed spaw bilaterally in the LPM (Fig. 4c, e). Thus, a dose-dependent effect of klf8 expression was revealed. Moreover, embryos overexpressing klf8 also frequently exhibited bilateral expression patterns of lft1, lft2, and pitx2 in the diencephalon, heart, and LPM at 19–22 s stages (Fig. 4h–t). On the other hand, ntl expression in the notochord was not altered in 22 s stage embryos overexpressing klf8 as compared to LacZ-overexpressing embryos (Fig. 4u-w). In order to investigate whether the zinc finger DNA binding domain of Klf8 is involved in regulating the expression pattern of spaw or its downstream genes, we injected mRNA for klf8 lacking the zinc finger DNA binding domain (klf8△zf). We found that injection of 100 pg klf8△zf only induced a low percentage of embryos to exhibit bilateral expression of spaw (6.3%), lft1 (3.6%), lft2 (1.6%) or pitx2 (11.9%), compared to higher rates of bilateral spaw (48.3%), lft1 (31.6%), lft2 (37.5%) or pitx2 (44.3%) expression in klf8 injected embryos at 18 s or 19–22 s stages (Additional file 3: Figure S3). These results demonstrate that overexpression of klf8 does not affect the midline structure, but induces ectopic expression of spaw and its downstream genes when the Klf8 zinc finger DNA binding domain is intact.
Overexpression of klf8 mRNA caused bilateral expression of Nodal signalling component genes. Injection of klf8 mRNA induced bilateral expression of spaw in the LPM (arrow) at the 18 s stage, in a dose-dependent manner (a-e). Embryos injected with klf8, but not LacZ, exhibited bilateral expression of lft1 in the diencephalon (arrowhead) and heart (asterisk; f-j), lft2 in the heart (asterisk; k-o), and pitx2 in the LPM (white arrow) (p-t) at the 19–22 s stage. Expression of ntl in the notochord is similar in wild type and embryos injected with LacZ or klf8 mRNA (u-w). A, absent; B, bilateral; L, left; R, right
klf8 deficiency affects morphogenesis of Kupffer’s vesicle and asymmetric charon expression
Since asymmetric flow, generated by rotation of cilia within KV, is essential to initiate left-right asymmetric patterning, and KV is derived from dorsal forerunner cells (DFCs), we then investigated whether klf8 knockdown affected cilia or DFC number during KV morphogenesis. Individual klf8-MOs or control MO were microinjected into one- or two-cell Tg(sox17:gfp) zygotes, and immunofluorescence was conducted using anti-GFP antibody. We found that the number of DFCs at the dorsal margin was significantly reduced in klf8-MO1atg- (average of 26.6 DFCs) or klf8-MO2atg- (average of 27.6 DFCs) injected Tg(sox17:gfp) embryos as compared to wild type (average of 36.2 DFCs) or klf8-4 mm MO1- (average of 35.7 DFCs) injected embryos at 75% epiboly (Fig. 5a-d, o). Cilia were then detected by immunofluorescence staining of 10s stage embryos using an anti-acetylated tubulin antibody. KV lumen size was smaller, but not significantly so, in Tg(sox17:gfp) embryos injected with klf8-MO1atg, klf8-MO2atg, or MOsp-MOs as compared to wild type or klf8-4 mm MO1-injected control embryos at 10s stage (Fig. 5e-n, p). Significantly reduced number and length of KV cilia were detected in embryos injected with different klf8-MOs as compared to wild type and control embryos (Fig. 5q, r). Since asymmetric charon expression on the right side of the KV was influenced by strength and direction of KV flow [40], we also examined whether klf8 knockdown affected asymmetric charon expression around KV. The majority (61% for MO1atg, 57% for MO2atg) of embryos injected with different klf8-MOs revealed symmetric charon expression with reduced expression area around KV as compared to wild type and control embryos at the 10s stage (Additional file 4: Figure S4). These results indicate that KV morphogenesis, cilia length, cilia number and asymmetric charon expression were affected in klf8 knockdown embryos.
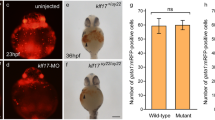
Knockdown of klf8 affected dorsal forerunner cell number, KV cilia number and length. Cell number of dorsal forerunner cells (DFCs) was affected in klf8-MO1atg- or klf8-MO2atg -injected embryos as compared to klf8-4 mm MO1-injected Tg(sox17:gfp) control embryos (4 mm) at 75% epiboly (a-d). Images of KV cilia stained with acetylated tubulin antibody in klf8 morphants (g-i) and control embryos (e, f) at 10s stage (e-i). Images of acetylated tubulin stained KV cilia and GFP stained DFCs in Tg(sox17:gfp) embryos injected with different klf8 MOs (l-n), klf8-4 mm MO1 (k) or wild type (j) embryos at 10s stage (j-n). DFC number (o), KV cilia number (q), length (r) but not lumen area (p) were affected in klf8 morphants as compared to control embryos. Statistical significance was determined by Student’s t-test. *p < 0.05, **p < 0.01, ***p < 0.001. Error bars indicate standard deviation
Generation of klf8 mutant by a CRISPR-Cas9 system
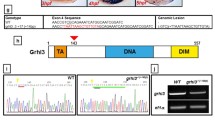
In order to confirm our morphant results, we generated klf8 mutants using a CRISPR-Cas9 system. Although we designed three klf8 sgRNAs targeting to exon 2 or exon 3, only klf8 sgRNA1, which targets to exon 2, was successful in producing mutants. Administration of sgRNA1 induced efficient deletion or insertion of bases in exon 2 and resulted in two klf8 mutant alleles (klf8 d25 and klf8 i17) (Fig. 6a). The klf8 d25 mutant had a 25 bp deletion, which produced a 35 amino acid-long misframed Klf8 protein, while the klf8 i17 mutant had a 17 bp insertion that generated a 49 amino acid-long misframed Klf8 protein (Fig. 6b).
Generation of klf8 mutants by CRISPR-Cas9 gene editing and the effect on spaw expression. a klf8 genomic structure with klf8 sgRNA (blue lettering) targeted to exon 2. Protospacer adjacent motif (PAM) sequence is shown in red. b Nucleotide and predicted amino acid sequences of klf8 in wild type, klf8 I17 and klf8d25 mutants are shown. Deleted nucleotides are shown by a red dashed line, while inserted nucleotides are shown in green lettering. Representative images of embryos with different spaw expression patterns in the LPM at 18 s stage (arrow; c-f). g Percentage of embryos displayed left (L), right (R), decreased (D) or bilateral (B) expression of spaw in the LPM from intercross of respective klf8 d25 or klf8 i17 F2 heterozygous mutants. Deduced percentage of wild type (+/+), heterozygote (+/−) or homozygote (−/−) genotype of embryos from intercross of respective klf8 d25 or klf8 i17 F2 heterozygous mutants exhibited bilateral spaw expression pattern. Expression levels of klf3 (h, k), klf12a (i, l) or klf12b (j, m) were compared between wild type and respective klf8 d25 and klf8 i17 F5 homozygous mutant embryos at 10–12 s stages (h-m). Knockdown of klf12b reduced the percentage of embryos with bilateral spaw expression in the LPM of klf8 d25 F6 homozygous mutant embryos, but the reduction did not reach significance (p = 0.45 for the comparison between klf8 d25 and klf8 d25+ 5 ng klf812b MO, p = 0.05 for the comparison between klf8 d25 and klf8 d25+ 10 ng klf812b MO) (n). Statistical significance was determined by Student’s t-test. * p < 0.05. Error bars indicate standard deviation
Of note, in 24 hpf homozygous F3 embryos of klf8 d25 and klf8 i17 mutants, we did not observe smaller eyes and abnormal cerebellar morphology that were detected in klf8 morphant embryos [35]. Next, we investigated whether spaw expression was affected in klf8 mutant embryos. Bilateral spaw expression was more frequently observed in embryos from the intercross of respective klf8 d25 (27 out of 69, 39.1%) or klf8 i17 (9 out of 35, 25.7%) F2 heterozygous mutants as compared to wild type embryos (7%) at 18 s stage (Fig. 6c-g). In order to evaluate the genotype of F3 embryos from klf8 i17 and klf8 d25 mutants with bilateral spaw expression, we sequenced seven klf8 i17 and 18 klf8 d25embryos with the phenotype. From sequencing data, we obtained two (28.6%) wild type, three (42.9%) heterozygotes and two (28.5%) homozygotes from a total of seven klf8 i17 F3 embryos, as well as three (16.7%) wild type, seven (38.9%) heterozygotes and eight (44.4%) homozygotes from a total of 18 klf8 d25 embryos. We then deduced that 7.3% of klf8 i17 or 17.4% of klf8 d25 F3 embryos were homozygous mutant embryos that also had bilateral spaw expression. This observation was based on the following calculation [0.285 (% of sequenced embryos that were homozygous) × 9 (total number with bilateral spaw expression) / 35 (total number of embryos) = 7.3% for klf8 i17; 0.444 × 27 / 69 = 17.4% for klf8 d25]. The other F3 embryos that exhibited bilateral spaw expression were also deduced to be either heterozygous mutant embryos (11% in klf8 i17, 15.2% in klf8 d25) or sibling wild type (7.4% in klf8 i17, 6.5% in klf8 d25) based on similar calculations.
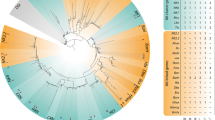
Because human KLF8, KLF3 and KLF12 form a subgroup in phylogenetic tree analysis due to the presence of CtBP-binding sites [41], we wondered whether expression of zebrafish klf3, klf12a, or klf12b may be upregulated to compensate for klf8 deficiency. We discovered substantial upregulation of klf12b in klf8 d25 F5 homozygous mutant embryos and downregulation of klf12a in klf8 i17 F5 homozygous mutant embryos at 10–12 s stages, while no alteration of klf3 expression was observed in either klf8 d25 or klf8 i17 mutant embryos (Fig. 6h-m). Subsequently, we knocked down klf12b in klf8 d25 F6 homozygous mutant embryos and evaluated the spaw expression pattern at 18 s stage (Fig. 6n). In klf8 d25 F6 homozygous mutant embryos, bilateral (50.3%) and decreased (6.3%) spaw expression patterns were detected. In homozygous mutant embryos injected with 5 or 10 ng klf12b MO, we found a dose-dependent reduction that did not reach significance in the percentage (42.1% for 5 ng, 29.3% for 10 ng) of embryos with bilateral spaw expression, which was accompanied by the increased occurrence of right (1.6% for 5 ng, 1.7% for 10 ng), decreased (15.9% for 5 ng, 16.4% for 10 ng) or absent (4.0% for 5 ng, 3.4% for 10 ng) spaw expression patterns.
These results indicate that although klf8 mutant embryos display different spaw expression patterns than morphant embryos, this effect may be partly attributed to a compensatory induction of klf12b expression.
Discussion
Establishing asymmetric spaw expression in the LPM is essential for left-right patterning in zebrafish embryos. Expression of spaw is first apparent in bilateral cells flanking KV between the 4–6 s stages, while asymmetric spaw expression emerges in the posterior LPM during the 10–12 s stages, and extends to the anterior LPM by the 18 s stage [22]. Knockdown of spaw abolishes spaw expression in the LPM but not in peri-KV domains [21], suggesting that autoregulation of spaw occurs only in the left LPM. We used morpholino antisense oligomers to knockdown klf8, and found that spaw expression in the left LPM was reduced or eliminated in the majority of 18–22 s stage morphants (Fig. 2, Additional file 2: Figure S2). Spaw activates expression of itself, as well as lft1, lft2, and pitx2 in the left LPM during the segmentation stage [22]. As such, the observed reduction or elimination of lft1, lft2, and pitx2 in the left diencephalon, heart or LPM at 18–22 s stages in the majority of klf8 morphants was not unexpected (Fig. 2, Additional file 2: Figure S2). Overall, defects in the expression of spaw and its downstream genes, and the subsequent defects in internal organ patterning observed in klf8 morphants are consistent with those detected in spaw morphants [22]. Although spaw/sfw mutant embryos more frequently displayed a D-loop heart (68%), compared to klf8 morphants, the difference may be attributed to heart specific actomyosin activity. Furthermore, in sfw mutant embryos, expression of laterality genes including lft1, lft2, and pitx2 were lost, and spaw expression did not propagate toward the anterior of the left LPM [42]. In our study, we saw that overexpression of klf8 but not klf8△zf resulted in bilateral expression of spaw, lft1, lft2, and pitx2 at 18 s or 19–22 s stages (Fig. 4, Additional file 3: Figure S3), demonstrating a requirement for the Klf8 zinc finger DNA binding domain. Expression of ntl in the notochord was found to be unaltered in klf8-overexpressing embryos, suggesting that the midline structure is in intact. Overall, our results demonstrate that Klf8 may regulate asymmetric spaw expression in the left LPM, which in turn affects left-side specific expression of lft1, lft2, and pitx2 in zebrafish embryos.
Proper morphogenesis of KV is also important for initial asymmetric spaw expression in the posterior LPM. The development of KV involves the formation of DFCs from surface epithelial cells, ingression at the dorsal germ ring margin, DFC migration, formation of rosette-like epithelial structures, coalescence of epithelial rosettes, and differentiation of ciliated KV with interior lumen [15, 43]. Both Tbx16 and Ntl were shown to regulate a mesenchymal to epithelial transition that participates in the formation of rosette-like epithelia [44]. Wnt11- and Prickle1a-mediated planar cell polarity signalling, as well as Cnpy1-mediated FGF signalling, were shown to regulate cell adhesion between adjacent dorsal forerunner cells to maintain cluster formation [45, 46]. Defects in these signalling events resulted in small KV lumen, with shortened and decreased number of KV cilia. In klf8 morphants, a similar smaller KV lumen, with decreased number and length of KV cilia was frequently observed at the 10s stage. These abnormal structures may result from lower number of DFCs that was observed in 75% epiboly morphants (Fig. 5). Whether Klf8 may participate in Wnt11- and Prickle1a-mediated planar cell polarity signalling, or Cnpy1-mediated FGF signalling to modulate DFC cluster formation, remains to be determined.
Zebrafish KV architecture is asymmetric along the anterior-posterior axis, with more ciliated cells in the anterior region. Furthermore, the positioning of the basal body of motile cilia at the posterior end of the epithelial cells may result in cilia tilting [17, 47]. These motile cilia then use a vortical motion to generate swirling fluid flow consisting of a relatively stronger leftward flow across the anterior pole of KV and a weaker rightward flow at the posterior end [16, 18]. Based on experimental tracking of native particles within the KV of wild type, did −/− mutant and dnah7 morphants, and simulated flow by mathematically modelling, it was determined that a threshold of 30 cilia, with dorsal anterior clustering, is essential to generate proper swirling flow in the anti-clockwise direction [40]. In control embryos, with strong left-sided flow across the anterior pole of KV, asymmetric expression is established for charon in the right side of the KV, and spaw in the left LPM. In embryos with non-directional flow, symmetric charon expression and a lack of spaw expression may be found. Embryos without motile cilia, and therefore no KV flow, may exhibit symmetric and slightly weaker charon expression and bilateral spaw expression in the posterior LPM [40]. In klf8 morphant embryos, injected with different klf8 MOs, a significantly reduced number of cilia (< 30), with random distribution was detected (Fig. 5). KV with such a cellular architecture may exhibit a weak and homogenous fluid flow, resulting in the symmetric charon expression around KV that was detected in the majority of klf8 morphants, and leading to drastically reduced spaw expression at late somite stage (Additional file 4: Figure S4, Fig. 2). Overall, our results clearly indicate that Klf8 is required for normal KV morphogenesis, which is known to be critical for initiating asymmetric spaw expression in the left LPM.
In mouse and chick, BMP signalling plays either a positive or negative role in regulating asymmetric Nodal expression. Moreover, the presence of BMP antagonists, such as Noggin, Chordin, or Caronte, can relieve BMP-inhibition to promote asymmetric Nodal expression in the left LPM [48,49,50,51]. In zebrafish embryos, heat–activated BMP2b expression inhibits spaw expression, while heat-activated noggin3 induced bilateral spaw expression, indicating that BMP signalling is required to repress spaw expression in the right LPM of early segmentation stage embryos, [25]. In addition, expression of Lft1 in the midline, and Lft2 in the left cardiac field, serve to generate a posterior or anterior barrier to restrict Spaw activity to the left LPM during segmentation stages in zebrafish embryos [27]. Dvr1, a zebrafish Vg1 ortholog, was also shown to facilitate the transfer of spaw expression from the peri-KV region to the left LPM. Thus, reduced or absent expression of spaw and downstream lft1 and lft2 in the LPM, diencephalon, notochord, or heart were detected in dvr1 morphants [52]. In order to investigate whether Klf8 may regulate asymmetric spaw expression via modulation of expressions of bmp2b or dvr1, we then compared expression of these two genes between klf8 morphants and control embryos (Additional file 5: Figure S5). Similar bmp2b expression level around tailbud region was identified in 3 s stage wild type and embryos injected with klf8-MO1atg, klf8-MO2atg or klf8- 4 mm MO1. Likewise, no alteration of dvr1 expression around the tailbud region was detected in wild type and embryos injected with klf8-MO1atg, klf8-MO2atg or klf8-4 mm MO1 at bud stage. Thus, Klf8 does not act via BMP2b or Dvr1 signalling pathway to regulate asymmetric spaw expression, and the underlying mechanism remains to be determined.
In addition to our studies with klf8 morphants, we generated klf8 mutants by a CRISPR-Cas9 system (Fig. 6). Intriguingly, obvious phenotypic differences were found between morphants and mutants. In 24 hpf homozygous F3 embryos of klf8 d25 and klf8 i17 mutants, we did not observe smaller eyes and abnormal cerebellar morphology that were detected in klf8 morphant embryos [35]. In addition, bilateral spaw expression was detected in the LPM of klf8 d25 and klf8 i17 mutants at the 18 s stage (Fig. 6). Discrepant phenotypes between mutants created by TALENs or CRISPR-Cas genome editing systems and antisense morpholino mediated-morphants have been frequently encountered. Previously, differences have been attributed to off-target effects of morpholinos [53], or compensatory effects, which have been described in vasculature development [54], reproduction [55], or neurogenesis [56]. With regard to the two klf8 mutant alleles from our study, more klf8 d25 mutant embryos showed bilateral spaw expression in the LPM, compared to klf8 i17 mutants (Fig. 6g, n). This difference in outcome may correlate with aberrant upregulation of klf12b in klf8 d25 that was further confirmed by klf12b knockdown, but not in klf8 i17, mutants (Fig. 6n). Similar compensatory inconsistency was found in stmn4 mutants, which had a low (< 10%) portion of embryos showing similar phenotype to stmn4 morphants. Interestingly, the authors found that stmn1b was upregulated to compensate in stmn4△5 but not stmn4△4 mutants [56]. In our study, we observed that in response to klf8 deficiency, klf12b, a member of a subgroup of KLF family with a CtBP interaction site, was induced to compensate for the loss of klf8. However aberrant upregulation of klf12b further resulted in bilateral spaw expression. On the contrary, downregulation of klf12a was detected in klf8 i17 mutants (Fig. 6). In these mutants, bilateral spaw expression was observed to a lesser degree, suggesting that klf12a may have undergone functional divergence with klf12b, and as such, klf12a may play a role in restricting spaw expression to the left side of embryos. Overall, klf8 mutant embryos showed bilateral spaw expression, which was quite different from klf8 morphants that exhibited reduced or eliminated spaw expression in the LPM. This dissimilar phenotype may have been partly related to the compensatory induction of klf12b expression in the mutant embryos.
Conclusions
In this report, we have demonstrated a novel role for zebrafish Klf8 in left-right asymmetric patterning. During gastrulation, Klf8 may regulate DFC cell number to control proper KV morphogenesis, which is essential to initiate asymmetric spaw expression in the left LPM. During somitogenesis, Klf8 may further modulate asymmetric spaw expression in the left LPM to ensue asymmetric organ positioning.
Abbreviations
- ASE:
-
Left-side specific enhancers
- CRISPR-Cas:
-
Clustered regularly interspaced short palindromic repeats- CRISPR-associated system
- CtBP:
-
C-terminal binding protein
- DFCs:
-
Dorsal forerunner cells
- Hpf:
-
Hours post fertilization
- klf8:
-
krüppel-like factor 8
- KV:
-
Kupffer’s vesicle
- lft1:
-
lefty1
- lft2:
-
lefty2
- LPM:
-
Lateral plate mesoderm
- MO:
-
Morpholino oligomer
- Ndr:
-
Nodal-related genes
- RT:
-
Room temperature
- RT-PCR:
-
Reverse transcription PCR
- RT-qPCR:
-
Reverse transcription quantitative real-time PCR
- S:
-
Somite
References
Bisgrove BW, Morelli SH, Yost HJ. Genetics of human laterality disorders: insights from vertebrate model systems. Annu Rev Genomics Hum Genet. 2003;4:1–32.
Kosaki K, Casey B. Genetics of human left-right axis malformations. Semin Cell Dev Biol. 1998;9:89–99.
Hamada H, Meno C, Watanabe D, Saijoh Y. Establishment of vertebrate left-right asymmetry. Nat Rev Genet. 2002;3:103–13.
Shiratori H, Hamada H. The left-right axis in the mouse: from origin to morphology. Development. 2006;133:2095–104.
Hirokawa N, Tanaka Y, Okada Y. Left-right determination: involvement of molecular motor KIF3, cilia, and nodal flow. Cold Spring Harb Perspect Biol. 2009;1:a000802.
Nonaka S, Tanaka Y, Okada Y, Takeda S, Harada A, Kanai Y, Kido M, Hirokawa N. Randomization of left-right asymmetry due to loss of nodal cilia generating leftward flow of extraembryonic fluid in mice lacking KIF3B motor protein. Cell. 1998;95:829–37.
Neugebauer JM, Amack JD, Peterson AG, Bisgrove BW, Yost HJ. FGF signalling during embryo development regulates cilia length in diverse epithelia. Nature. 2009;458:651–4.
Thomas J, Morle L, Soulavie F, Laurencon A, Sagnol S, Durand B. Transcriptional control of genes involved in ciliogenesis: a first step in making cilia. Biol Cell. 2010;102:499–513.
Nakamura T, Mine N, Nakaguchi E, Mochizuki A, Yamamoto M, Yashiro K, Meno C, Hamada H. Generation of robust left-right asymmetry in the mouse embryo requires a self-enhancement and lateral-inhibition system. Dev Cell. 2006;11:495–504.
Schier AF. Nodal signaling in vertebrate development. Annu Rev Cell Dev Biol. 2003;19:589–621.
Saijoh Y, Adachi H, Sakuma R, Yeo CY, Yashiro K, Watanabe M, Hashiguchi H, Mochida K, Ohishi S, Kawabata M, Miyazono K, Whitman M, Hamada H. Left-right asymmetric expression of lefty2 and nodal is induced by a signaling pathway that includes the transcription factor FAST2. Mol Cell. 2000;5:35–47.
Shiratori H, Sakuma R, Watanabe M, Hashiguchi H, Mochida K, Sakai Y, Nishino J, Saijoh Y, Whitman M, Hamada H. Two-step regulation of left-right asymmetric expression of Pitx2: initiation by nodal signaling and maintenance by Nkx2. Mol Cell. 2001;7:137–49.
Ryan AK, Blumberg B, Rodriguez-Esteban C, Yonei-Tamura S, Tamura K, Tsukui T, de la Pena J, Sabbagh W, Greenwald J, Choe S, Norris DP, Robertson EJ, Evans RM, Rosenfeld MG, Izpisua Belmonte JC. Pitx2 determines left-right asymmetry of internal organs in vertebrates. Nature. 1998;394:545–51.
Essner JJ, Amack JD, Nyholm MK, Harris EB, Yost HJ. Kupffer’s vesicle is a ciliated organ of asymmetry in the zebrafish embryo that initiates left-right development of the brain, heart and gut. Development. 2005;132:1247–60.
Oteiza P, Koppen M, Concha ML, Heisenberg CP. Origin and shaping of the laterality organ in zebrafish. Development. 2008;135:2807–13.
Amack JD. Salient features of the ciliated organ of asymmetry. BioArchitecture. 2014;4:6–15.
Wang G, Cadwallader AB, Jang DS, Tsang M, Yost HJ, Amack JD. The Rho kinase Rock2b establishes anteroposterior asymmetry of the ciliated Kupffer’s vesicle in zebrafish. Development. 2011;138:45–54.
Smith DJ, Montenegro-Johnson TD, Lopes SS. Organized chaos in Kupffer’s vesicle: how a heterogeneous structure achieves consistent left-right patterning. BioArchitecture. 2014;4:119–25.
Bisgrove BW, Snarr BS, Emrazian A, Yost HJ. Polaris and Polycystin-2 in dorsal forerunner cells and Kupffer’s vesicle are required for specification of the zebrafish left-right axis. Dev Biol. 2005;287:274–88.
Borovina A, Superina S, Voskas D, Ciruna B. Vangl2 directs the posterior tilting and asymmetric localization of motile primary cilia. Nat Cell Biol. 2010;12:407–12.
Wang X, Yost HJ. Initiation and propagation of posterior to anterior (PA) waves in zebrafish left-right development. Dev Dyn. 2008;237:3640–7.
Long S, Ahmad N, Rebagliati M. The zebrafish nodal-related gene southpaw is required for visceral and diencephalic left-right asymmetry. Development. 2003;130:2303–16.
Ahmad N, Long S, Rebagliati M. A southpaw joins the roster: the role of the zebrafish nodal-related gene southpaw in cardiac LR asymmetry. Trends Cardiovasc Med. 2004;14:43–9.
Hashimoto H, Rebagliati M, Ahmad N, Muraoka O, Kurokawa T, Hibi M, Suzuki T. The Cerberus/Dan-family protein Charon is a negative regulator of Nodal signaling during left-right patterning in zebrafish. Development. 2004;131:1741–53.
Chocron S, Verhoeven MC, Rentzsch F, Hammerschmidt M, Bakkers J. Zebrafish Bmp4 regulates left-right asymmetry at two distinct developmental time points. Dev Biol. 2007;305:577–88.
Smith KA, Noel E, Thurlings I, Rehmann H, Chocron S, Bakkers J. Bmp and nodal independently regulate lefty1 expression to maintain unilateral nodal activity during left-right axis specification in zebrafish. PLoS Genet. 2011;7:e1002289.
Lenhart KF, Lin SY, Titus TA, Postlethwait JH, Burdine RD. Two additional midline barriers function with midline lefty1 expression to maintain asymmetric Nodal signaling during left-right axis specification in zebrafish. Development. 2011;138:4405–10.
Kaczynski J, Cook T, Urrutia R. Sp1- and Kruppel-like transcription factors. Genome Biol. 2003;4:206.
Pearson R, Fleetwood J, Eaton S, Crossley M, Bao S. Kruppel-like transcription factors: a functional family. Int J Biochem Cell Biol. 2008;40:1996–2001.
Lahiri SK, Zhao J. Kruppel-like factor 8 emerges as an important regulator of cancer. Am J Transl Res. 2012;4:357–63.
van Vliet J, Turner J, Crossley M. Human Kruppel-like factor 8: a CACCC-box binding protein that associates with CtBP and represses transcription. Nucleic Acids Res. 2000;28:1955–62.
Zhang P, Basu P, Redmond LC, Morris PE, Rupon JW, Ginder GD, Lloyd JA. A functional screen for Kruppel-like factors that regulate the human gamma-globin gene through the CACCC promoter element. Blood Cells Mol Dis. 2005;35:227–35.
Funnell AP, Mak KS, Twine NA, Pelka GJ, Norton LJ, Radziewic T, Power M, Wilkins MR, Bell-Anderson KS, Fraser ST, Perkins AC, Tam PP, Pearson RC, Crossley M. Generation of mice deficient in both KLF3/BKLF and KLF8 reveals a genetic interaction and a role for these factors in embryonic globin gene silencing. Mol Cell Biol. 2013;33:2976–87.
Zhao J, Bian ZC, Yee K, Chen BPC, Chien S, Guan J-L. Identification of transcription factor KLF8 as a downstream target of focal adhesion kinase in its regulation of cyclin D1 and cell cycle progression. Mol Cell. 2003;11:1503–15.
Tsai MY, Lu YF, Liu YH, Lien HW, Huang CJ, Wu JL, Hwang SP. Modulation of p53 and met expression by Kruppel-like factor 8 regulates zebrafish cerebellar development. Dev Neurobiol. 2015;75:908–26.
Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF. Stages of embryonic development of the zebrafish. Dev Dyn. 1995;203:253–310.
Chen J, Ruan H, Ng SM, Gao C, Soo HM, Wu W, Zhang Z, Wen Z, Lane DP, Peng J. Loss of function of def selectively up-regulates Delta113p53 expression to arrest expansion growth of digestive organs in zebrafish. Genes Dev. 2005;19:2900–11.
Jao LE, Wente SR, Chen W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc Natl Acad Sci U S A. 2013;110:13904–9.
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem. 2009;55:611–22.
Sampaio P, Ferreira RR, Guerrero A, Pintado P, Tavares B, Amaro J, Smith AA, Montenegro-Johnson T, Smith DJ, Lopes SS. Left-right organizer flow dynamics: how much cilia activity reliably yields laterality? Dev Cell. 2014;29:716–28.
McConnell BB, Yang VW. Mammalian Kruppel-like factors in health and diseases. Physiol Rev. 2010;90:1337–81.
Noel ES, Verhoeven M, Lagendijk AK, Tessadori F, Smith K, Choorapoikayil S, den Hertog J, Bakkers J. A Nodal-independent and tissue-intrinsic mechanism controls heart-looping chirality. Nat Commun. 2013;4:2754.
Matsui T, Ishikawa H, Bessho Y. Cell collectivity regulation within migrating cell cluster during Kupffer’s vesicle formation in zebrafish. Front Cell Dev Biol. 2015;3:27.
Amack JD, Wang X, Yost HJ. Two T-box genes play independent and cooperative roles to regulate morphogenesis of ciliated Kupffer’s vesicle in zebrafish. Dev Biol. 2007;310:196–210.
Oteiza P, Koppen M, Krieg M, Pulgar E, Farias C, Melo C, Preibisch S, Muller D, Tada M, Hartel S, Heisenberg CP, Concha ML. Planar cell polarity signalling regulates cell adhesion properties in progenitors of the zebrafish laterality organ. Development. 2010;137:3459–68.
Matsui T, Thitamadee S, Murata T, Kakinuma H, Nabetani T, Hirabayashi Y, Hirate Y, Okamoto H, Bessho Y. Canopy1, a positive feedback regulator of FGF signaling, controls progenitor cell clustering during Kupffer’s vesicle organogenesis. Proc Natl Acad Sci U S A. 2011;108:9881–6.
Wang G, Manning ML, Amack JD. Regional cell shape changes control form and function of Kupffer’s vesicle in the zebrafish embryo. Dev Biol. 2012;370:52–62.
Schlange T, Arnold HH, Brand T. BMP2 is a positive regulator of Nodal signaling during left-right axis formation in the chicken embryo. Development. 2002;129:3421–9.
Mine N, Anderson RM, Klingensmith J. BMP antagonism is required in both the node and lateral plate mesoderm for mammalian left-right axis establishment. Development. 2008;135:2425–34.
Fujiwara T, Dehart DB, Sulik KK, Hogan BL. Distinct requirements for extra-embryonic and embryonic bone morphogenetic protein 4 in the formation of the node and primitive streak and coordination of left-right asymmetry in the mouse. Development. 2002;129:4685–96.
Raya A, Izpisua Belmonte JC. Unveiling the establishment of left-right asymmetry in the chick embryo. Mech Dev. 2004;121:1043–54.
Peterson AG, Wang X, Yost HJ. Dvr1 transfers left-right asymmetric signals from Kupffer's vesicle to lateral plate mesoderm in zebrafish. Dev Biol. 2013;382:198–208.
Kok FO, Shin M, Ni CW, Gupta A, Grosse AS, van Impel A, Kirchmaier BC, Peterson-Maduro J, Kourkoulis G, Male I, DeSantis DF, Sheppard-Tindell S, Ebarasi L, Betsholtz C, Schulte-Merker S, Wolfe SA, Lawson ND. Reverse genetic screening reveals poor correlation between morpholino-induced and mutant phenotypes in zebrafish. Dev Cell. 2015;32:97–108.
Rossi A, Kontarakis Z, Gerri C, Nolte H, Holper S, Kruger M, Stainier DY. Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature. 2015;524:230–3.
Spicer OS, Wong TT, Zmora N, Zohar Y. Targeted mutagenesis of the hypophysiotropic Gnrh3 in Zebrafish (Danio rerio) reveals no effects on reproductive performance. PLoS One. 2016;11:e0158141.
Lin MJ, Lee SJ. Stathmin-like 4 is critical for the maintenance of neural progenitor cells in dorsal midbrain of zebrafish larvae. Sci Rep. 2016;6:36188.
Acknowledgments
We thank the members of the Core Facility of the Institute of Cellular and Organismic Biology, Academia Sinica, for their assistance with DNA sequencing and confocal imaging. We thank the Taiwan Zebrafish Core Facility at Academia Sinica (TZCAS) for providing the ASAB wild type strain. We thank ZIRC and TZCAS for providing Tg(sox17:gfp) s870/+ fish line. We thank Dr. M. Rebagliati for providing full-length spaw cDNA. We thank Ms. Mei-Chen Chen for fish maintenance.
Funding
This work was supported by Academia Sinica Innovative Translational Agricultural Research Program [542300 to S.P.L.H.] and the Ministry of Science and Technology, Taiwan [MOST 105-2313-B-001-005-MY3 to S.P.L.H.].
Availability of data and materials
All materials are available by the corresponding author.
Author information
Authors and Affiliations
Contributions
CYL, MYT, YCC, YRL, YHL, YFL, HCL, HWL, CHY performed experiments; CYL, MYT, CJH and SPLH analysed data; SPLH conceived the project and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
All animal procedures were approved by the Institutional Animal Care and Use Committee of Academia Sinica (Protocol ID: 15-12-918).
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional files
Additional file 1: Figure S1.
Knockdown of klf8 expression by splicing morpholino oligomers resulted in embryos with reduced or absent spaw expression in the left LPM. A klf8 genomic structure showing position of translational morpholino oligomers (klf8-MO1atg, klf8-MO2atg) and splicing morpholino oligomers (klf8 DO2, klf8 AC, klf8 DO). Arrows indicate the positions of forward and reverse primers. B RT-PCR showing the efficacy of klf8 splicing morpholino oligomers. C The majority of embryos injected with klf8 splicing MOs had decreased or absent spaw expression in the left LPM. (TIFF 153 kb)
Additional file 2: Figure S2.
Heart looping and downregulated expression of spaw and its downstream genes were not caused by induction of p53 expression in klf8 morphants. Embryos co-injected with p53-MOsp and klf8-MO1atg or klf8-MO2atg displayed no-loop or L-loop heart defects at 72 hpf (A). The majority of embryos co-injected with p53-MOsp and klf8-MO1atg or klf8-MO2atg exhibited decreased or absent expression of spaw (B), lft1 (C), lft2 (D), or pitx2 (E) in the left LPM, diencephalon or heart at the 18–22 s stages. Expression levels of gata5 and oep which are known to be expressed in the LPM at the 22 s stage were unaffected by klf8 knockdown (F-M). (TIFF 797 kb)
Additional file 3: Figure S3.
Overexpression of klf8 but not klf8△zf mRNA induced bilateral expression of Nodal signalling component genes. Percentages of embryos injected with either 100 pg of klf8 or klf8△zf mRNA that exhibit left (L), right (R), decreased (D) or bilateral (B) expression of spaw in the LPM, lft1 in the diencephalon and heart, lft2 in the heart, and pitx2 in the LPM at 18 s or 19–22 s stages. (TIFF 74 kb)
Additional file 4: Figure S4.
Symmetric charon expression around KV was observed in the majority of klf8 morphants. Representative images of embryos showing stronger charon expression on the right side (A) or left side (B) and symmetric charon expression on both sides (C) of KV are shown. Quantification of different charon expression patterns in embryos injected with different klf8 MOs, control MO or wild type embryo is shown (D). Statistical significance was determined by Student’s t-test. * p < 0.05. Error bars indicate standard deviation. (TIFF 397 kb)
Additional file 5: Figure S5.
Expression level of bmp2b or dvr1 around tailbud region was not affected by klf8 knockdown. Representative images show similar expression level of bmp2b (A-D) or dvr1 (E-H) around the tailbud region in the wild type or embryos injected with klf8-MO1atg, klf8-MO2atg or klf8-4 mm MO1 at 3 s or bud stages. (TIFF 382 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Lin, CY., Tsai, MY., Liu, YH. et al. Klf8 regulates left-right asymmetric patterning through modulation of Kupffer’s vesicle morphogenesis and spaw expression. J Biomed Sci 24, 45 (2017). https://doi.org/10.1186/s12929-017-0351-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12929-017-0351-y