Abstract
Background
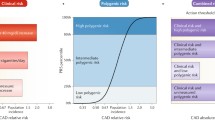
Polygenic risk scores (PRS) have been widely applied in research studies, showing how population groups can be stratified into risk categories for many common conditions. As healthcare systems consider applying PRS to keep their populations healthy, little work has been carried out demonstrating their implementation at an individual level.
Case presentation
We performed a systematic curation of PRS sources from established data repositories, selecting 15 phenotypes, comprising an excess of 37 million SNPs related to cancer, cardiovascular, metabolic and autoimmune diseases. We tested selected phenotypes using whole genome sequencing data for a family of four related individuals. Individual risk scores were given percentile values based upon reference distributions among 1000 Genomes Iberians, Europeans, or all samples. Over 96 billion allele effects were calculated in order to obtain the PRS for each of the individuals analysed here.
Conclusions
Our results highlight the need for further standardisation in the way PRS are developed and shared, the importance of individual risk assessment rather than the assumption of inherited averages, and the challenges currently posed when translating PRS into risk metrics.
Similar content being viewed by others
Background
Although genetics plays a substantial role in the development of common diseases, to date, optimising its contribution to disease prevention in individuals remains a challenge [1]. PRS are an emerging tool in genetics, the potential of which has been picked up by health systems, including in UK’s National Health Service [2], as a tool for improving the health of their populations. For some common diseases, such as Coronary Artery Disease, Type 2 Diabetes or Breast Cancer, PRS have been shown to help capture a sizable genetic contribution as part of the aetiology of high-risk individuals [3]. However, it remains to be demonstrated how PRS can be a useful tool for disease prevention at the level of the individual in many complex conditions [4].
There have already been attempts to implement PRS in a preventative healthcare setting. For instance, the MedSeq project [5] provided a benchmark study for application of cardiovascular disease PRS in a cohort of 100 individual whole genomes. A number of direct-to-consumer companies are also providing PRS in a preventative context, including testing of traits such as Breast Cancer and Type 2 Diabetes. Nonetheless, for many of these PRS tests, only a relatively small proportion of known variants are being tested (e.g., tens or dozens), compared to the total number included in some PRS, which for Type 2 Diabetes, for instance, is in the order of 7 million Single Nucleotide Polymorphisms (SNPs) [3]. The current provision of PRS for disease risk prevention is thus not yet at the same level as in PRS research, where there is a plethora of new PRS incorporated into centralised repositories. Repositories such as Cancer-PRSweb [6] displays 69 PRS for cancer alone, while the Polygenic Score Catalog [7] reports 751 (last accessed on 24 March 2021).
Here we propose a novel implementation for reuse and deployment of PRS collected from public repositories and supported by scientific literature. Due to the heterogeneity and overlap of available PRS, we perform a systematic curation of existing data sources following a set of purposely generated criteria for their selection. We include PRS from a wide range of common diseases related to cancer, cardiovascular, metabolic and autoimmune diseases. We apply selected PRS as proof-of-principle implementation to a family of four of Iberian Spanish origin, who underwent whole genome sequencing. We note that a population effect must be kept in mind when applying PRS to populations of differing ancestry. In a recent study comparing PRS trained with UK Biobank samples and applied to other European populations, highest performance of PRS was found for their corresponding population dataset, with performance drops if different European populations were tested [8]. While we use the dataset of a family as our test case so as to be able to compare results of several related family members, we believe our methodology could be applied to a single individual.
Since we sequence the whole genome of each family member, we do not impute alleles for any variant. Instead we extract the exact allele from processed sequencing data. By using 1000 Genomes Project (1000G) individual variant data [9] as PRS background distributions, we are able to assess the genetic risk of each family member by comparing the individual’s score against the scores of the 1000G cohort.
From a total 43 PRS initially selected as candidates, we apply 15, encompassing a total of 37,025,730 tested SNPs for each family member. For each individual PRS, risk percentiles are calculated using the PRS of participants within three 1000G cohorts: Iberian Spanish (IBS; n = 107), European (EUR; n = 503), and all 1000G individuals (ALL; n = 2,504). Over 98 billion allele effect calculations were performed in order to obtain the PRS for each of the participants used in this study. This allows us to identify if an individual is at the higher risk end tail of the PRS 1000G background distribution and estimate their relative risk for developing a disease.
Sequencing and data processing
Saliva samples were collected using Oragene OG-600 and sent for DNA extraction and sequencing. The DNA samples were randomly fragmented by Covaris technology and fragments of 350 bps were obtained. Fragment DNA ends were repaired and an ‘A’ base added at the 3’ end of each strand. Adapters were then ligated to both strands of the end repaired/dA tailed DNA fragment. Amplification by ligation-mediated PCR was performed and then single strand separation and cyclisation. DNA nanoballs were created and loaded into the patterned nanoarrays and pair-end reads read through on the BGISEQ-500 platform for each library to maximise the chances of a target of 30 × coverage. Software for base calling with default parameters and the sequence data of each individual were generated as paired-end reads, identified as ‘raw data’ and provided as fastq format.
Once fastq files were obtained, we used the Sentieon DNASeq pipeline [10] for all four samples. Sentieon is a toolkit analogous to GATK [11] but built on a highly optimised backend. It takes raw fastq files and maps them to the human reference genome using BWA-MEM [12]. As all the PRS we were analysing used GRCh37, so we mapped to that reference. For variant calling, Sentieon uses the recommended best practices for variant analysis with GATK, with local realignment around indels and base recalibration using GATK and duplicate reads removed by Picard tools. Poor calls were removed as part of the Sentieon DNASeq pipeline.
Family dataset
We selected this particular family dataset because it has been well studied in the past [13,14,15,16], which affords us a deep knowledge of the family’s phenotypes and disease history. Figure 1 shows the family pedigree. In it we have individuals PT00007A (Father), PT00008A (Mother) and two children (PT00009A and PT00002A; Daughter and Son). From here onwards, and for simplicity, we refer to family members as (Father, Mother, Daughter, Son).
When we analysed the variant output of all samples, we benchmarked against Fabric Genomics Clinical Grade Scoring Rules (http://help.fabricgenomics.com/hc/en-us/articles/206433937-Appendix-4-Clinical-Grade-Scoring-Rules; accessed 7/January/2020), where Clinical Grade is a measure of a variant file’s overall quality and fitness for clinical interpretation (Table 1). Coverage in values with a star indicates that the median coverage of coding variants exceeds 40. Genotype quality with a starred value: more than 95% of the coding variants have a quality above 40. Starred homozygous / heterozygous ratio: the ratio for the coding variants is between 0.5 and 0.61. Starred transition / transversion ratio: The ratio for the coding variants is between 2.71 and 3.08. We performed a further analysis of quality of variants by counting those that pass the default standard filters of quality for interpretation given our analysis software.
Ethical framework
All participants underwent a consent process and signed a consent form accepting the terms and conditions of this work as well as the potential consequences of performing this analysis. We drew on the Personal Genome Project UK [17] as our approach to informed consent. The consent process we developed included the following elements: (a) participants underwent extensive training on the risks of genetic analysis including the risks of publishing personal genetic data; (b) participants completed an exam to demonstrate their comprehension of the risks and protocols associated with participating in genetic analysis which may be published and (c) participants were judged truly capable of giving informed consent. Consent forms were signed by all family members. This ethical framework has been independently assessed and approved by the Ethics Committee of Universidad Internacional de La Rioja (code PI:029/2020).
Curation of PRS
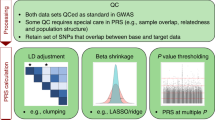
Underlying PRS data have been made available by the scientific community through the Polygenic Score Catalog [7] and Cancer-PRSweb [6]. Both resources provide centralised access to many PRS as well as data needed for their application, including SNP coordinates, effect alleles and their effect weights. We performed a curation process to identify PRS for application to our family use case of four. A dataset of 37,025,730 PRS SNPs was generated encompassing 15 common diseases (we call these common diseases ‘phenotypes’ from now onwards), together with risk alleles and weighted contributions for each SNP. Table 2 shows the two sets of criteria we followed to select PRS for our implementation model. The first set of criteria is based on study design and performance (Table 2a: Design Selection Criteria) and second on the requirements needed for their bioinformatics implementation (Table 2b: Bioinformatics Selection Criteria). In terms of design criteria, we chose PRS whose characteristics matched the following properties: (i) Underlying GWAS: the Genome Wide Association Study (GWAS) underlying the PRS could be traced to a recognisable consortium, and the phenotype in the GWAS was consistent with the phenotype in the resulting PRS. (ii) The PRS was trained in a second study using previously published PRS creation methods (e.g., LDpred [18] or clumping and thresholding [19]). (iii) The PRS was validated in a large independent cohort (we developed a preference for the UK Biobank for consistency reasons) [20]. (iv) Its area under the curve (AUC) or similar performance metric is above the 0.60 threshold except for Ischaemic stroke whose PRS performance (C-index 0.59) is comparable. (v) In case of more than one PRS being available for the same phenotype, we made a judgement of the study as a whole. (vi) The PRS should ideally have published risk metrics such as odds ratios, hazard ratios or fold increase.
Once we had filtered out PRS that did not fulfil the above design criteria, the remaining phenotypes underwent an extra filtering process according to a set of bioinformatics standards (Table 2b), required for us to run our pipelines successfully. These bioinformatics filtering criteria involved processing of the PRS raw data to establish that they fulfilled the following conditions: (i) Presence of at least 95% of SNPs in the 1000G Phase III distribution. (ii) Presence of risk alleles in either the reference or the alternative allele of the 1000G matching variant annotation. For this we check whether each SNP risk allele has an exact match to the reference or alternative allele in that coordinate position, discarding and labelling the SNP as ‘missing’ if otherwise (this enabled us to identify any reverse strand or similar bioinformatic inconsistencies) (iii) availability of effect weights and (iv) availability of coordinates in hg19. We used hg19 coordinates due to PRS source data being made available using this genome assembly. Any SNP that did not meet the above criteria was discarded but the phenotype was still used as long as it retains at least 95% of its source PRS SNPs.
How we calculate PRS for an individual
Our first step in calculating a PRS for a family member was to create background distributions so as to be able to put the score of a family member into context, and thus understand his or her relative risk. This is because source publications do not offer a translation of a raw PRS score directly into a risk measurement. Rather, they stratify different sections of a studied group into risk buckets (for example the top 5% of a distribution may be ascribed a particular odds ratio (OR)). Hence, when applying a PRS to an individual, it is necessary to know where that individual sits relative to others.
A PRS was calculated for each individual as the sum of the effect weights for all the risk alleles observed in the individual for a particular phenotype, divided by the total number of risk alleles reported for that phenotype. We calculated PRS following this method for each of the individuals in the final (Phase III) dataset of the 1000 Genomes Project (1000G), containing data for 2,504 participants. This required us to calculate the PRS of all 2,504 individuals in the 1000G project across all selected phenotypes.
Having generated raw scores for each of these 1000G individuals, we built distributions of raw scores according to different population groups within the 1000G cohort. We chose the subset of Iberians Spanish (IBS, n = 107), as all family members are of Spanish origin, the Europeans (EUR, n = 503), reflecting the ethnic background of the validation data sets for the PRS we selected, and the entire 1000G cohort (ALL, n = 2,504). ALL contains African, Admixed American, East Asian, South Asian and Europeans.
We then applied the same methodology for calculating a raw PRS score to the whole genome data of each of the four family members, and having determined the raw score for each family member for each phenotype, we placed that score inside the distribution of each population group already generated from the 1000G individuals (IBS, EUR and ALL).
Placing the individual in context this way allowed us to derive percentiles reflecting a family member’s position in a given population for a given phenotype. We could then readily compare these percentiles between individuals for each phenotype. We did this across the three different population groups in order to control for the impact of the ethnicity of the background population on the resulting percentile.
Scores from both 1000G participants and family members are thus calculated independently, producing a distribution of scores from which the percentile a family member occupies is generated.
PRS percentile inheritance patterns
We were interested to understand patterns of inheritance among family individuals for PRS percentiles. We set out to analyse how a high or low PRS percentile was explained in terms of risk being passed on from parents to children. This is a useful quality control and a way of adding credence to results, since it would be unusual for the same phenotype low risk observed in both parents engender high risk in offspring. In order to compare PRS percentiles between family individuals, we compare values relative to the 1000G EUR population distribution. We choose the EUR distribution percentile values for the remaining analyses because all selected PRS have as their validation dataset a European population such as those contained in the UK Biobank or FINRISK [21].
For the purposes of this analysis only, we have defined high risk as the individual falling above the 80th percentile of a given background distribution, as this is the threshold from which both Khera et al. [3] and Mars et al. [22] begin to quantify elevated risk. However, we acknowledge that the definition of a high-risk percentile is somewhat subjective, and we expand on this further below.
Translation of percentiles into risk metrics
For each family member we ascribe a relative risk. We note that when translating PRS percentiles into genetic risk metrics, each phenotype must be interpreted differently, as the risk metrics (e.g., odds ratios or hazard ratios), and risk thresholds vary from study to study. If a family member’s percentile is within a reported threshold of the PRS percentile source publication, we attribute a risk metric to that family member. We also pay attention to the Area Under the Curve (AUC) or other performance metrics described by each PRS source study.
Impact of population background distributions on risk percentiles
We considered the effect of background populations in risk calculation. There is a known risk that PRS predict less well in populations where the underlying GWAS and validation cohorts differ from the ancestry of the individual [23], as SNPs have different allele frequencies depending on ancestry. Therefore, an individual may be assigned different percentiles depending on background populations. This is important, given that odds ratios or hazard ratios are reported relative to intervals in PRS percentiles. If the choice of background population significantly changes an individual’s percentile PRS (i.e., > 20 percentile), their resulting odds or hazard ratio will then be different, affecting how we interpret risk. In order to determine whether or not the choice of background population made a difference to the results, we checked whether there are any noticeable differences in individual phenotype PRS percentiles depending on the choice of background distribution for each family member. For this, we compare whether tested individual percentiles for a phenotype change PRS quintiles depending on their background distribution. This choice of quintiles for binning risk distributions is a popular thresholding among the studies we curated [5, 22].
Case presentation
Our first set of criteria for selection of PRS considered the characteristics of the source study design, including recognisable GWAS consortia, performance metrics, presence of risk boundaries and independent cohort validation. We did not curate every single phenotype available, only those we judged promising candidates. Table 3 includes all phenotypes we researched after initial shortlisting. From an initial list of 43, we discarded 25 because a) there was not a clear consistency between the phenotype of the PRS and the phenotype in the underlying GWAS (for example All Cause Mortality, where the PRS is a composite of many separate GWAS); b) there was an alternative better performing candidate for the same phenotype (e.g., Coronary Artery Disease, Breast Cancer or Prostate Cancer); c) their performance metrics were below our acceptable threshold or were not available (e.g., Pancreatic Cancer, Multiple Myeloma, Uterine Cancer, Bladder Cancer, Squamous Cell Carcinoma, Epithelial Ovary Cancer, Lung Cancer, Non-Hodgkin’s Lymphoma, Cancer of other Lymphoid, Histiocytic Tissue, Cancer of Kidney, HDL Cholesterol, LDL Cholesterol, Triglycerides, Body Mass Index); d) their validation population was not the UK Biobank. We began with the PGS Catalog, and then complemented our selected set of PRS with Cancer-PRSweb phenotypes. From the Cancer-PRSweb we only considered their top 20 UK Biobank validated PRS, comparing them with phenotypes in the PGS Catalog where we found overlap. We tended to favour selection of standardised PRS such as those offered by the larger studies or the Cancer-PRSweb, as its blocks of performance metrics, risk boundaries, percentile thresholds and validation cohort metadata are well suited for benchmarking.
We had to reconcile conflicting criteria in the cases of Breast Cancer and Prostate Cancer PRS selection, and here we did deploy our judgement. For Breast Cancer, we selected the PRS from Khera et al. [3], although it has a lower performance than Mars et al. [22]. This was because overall the PRS for Khera et al. are high performing, and are all validated in the UK Biobank. For consistency therefore we retained the Breast Cancer phenotype from the Khera et al. study. For Prostate Cancer, despite being validated in the FINRISK consortium, we decided that the C-index of 0.86 in the Mars et al. study was sufficiently differentiated against that of Cancer-PRSweb (AUC of 0.71) that the Mars et al. PRS merited selection.
Concerning AUCs, we allowed any covariates that the source GWAS studies allowed. We recognise that this means that an AUC in one PRS is not exactly comparable to an AUC for another, as their design is not identical. Furthermore, we do not make a distinction between the type of method applied to calculation of the PRS (e.g., LDPred, Pruning and thresholding, etc.), accepting any method as long as it has been peer reviewed. Finally, we also note that some phenotypes are discrete while others are not, further affecting the choice of PRS calculation method.
Having made an initial selection of phenotypes whose study design met our eligibility criteria, we applied Table 2b’s bioinformatic filtering criteria scheme (Table 4). These bioinformatic requirements derived from the need to reliably replicate a PRS percentile as originally envisaged by the source publication. Because we use the 1000 Genomes (1000G) Project Phase III participants as our background PRS distributions, we required a high overlap (> 95%) of all PRS effect alleles and their weights between the SNPs identified by the study in question and the 1000G project.
A total of 15 phenotypes passed all our selection criteria for PRS implementation and testing. These phenotypes involved conditions related to cancer, cardiovascular, metabolic and autoimmune diseases. 8 of these phenotypes summed less than 10,000 (8,123) SNPs in total, whereas 6 phenotypes composed the vast majority of tested SNPs (37,017,607; 99.98%). We note that 11,954 SNPs were missing from our PRS calculation because they were not present as 1000G variants or their risk allele did not match the 1000G reference or alternative allele. However, the missing number of SNPs was never greater than 5% of the total for any of our selected phenotypes. The vast majority of PRS missed significantly fewer than 5% of the SNPs, and in fact more often than not, no SNPs were missed (9 out of 15 phenotypes missed none). The phenotype that proportionally misses the greatest number of SNPs is Basal Cell Carcinoma (missing 1 out of 24 SNPs; 4.17%), whereas Ischaemic Stroke missed the greatest absolute number: 11,103 SNPs (0.34%). We also note that all of the applied PRS were validated in the UK Biobank, with the exception of Prostate Cancer, which was validated on the FINRISK population.
Patterns of risk inheritance among family members
Weiner et al. [113] suggest that over a large group, the PRS of offspring is the average of the parents’ PRS and indeed we find that some averaging has taken place (Table 5). As an example, averaging plays a role in diminishing risk percentile in Daughter’s risk of Breast Cancer. Here, Daughter inherits a close to average parental percentile risk, diminishing her risk percentile for this condition when compared to her mother. We also observe some PRS where the offspring diverge from the average parental risk. For instance, we find that for Coronary Artery Disease, both children inherit a high percentile which carries over from Mother and has not been mitigated by Father. This departure from the averaging effect of PRS in offspring as observed in Coronary Artery Disease does not preclude, however, the overall pattern of averaging as suggested by Weiner et al. [113], and could be considered departures from the mean in a distribution.
Translation of percentiles into risk metrics
When consulting source publications to ascertain the risk of developing a disease phenotype given a particular percentile, we found that risks and their thresholds are differently described depending on the publication.
Khera et al. [3] for phenotypes Breast Cancer, Atrial Fibrillation, Coronary Artery Disease, Type 2 Diabetes and Inflammatory Bowel Disease offer odds ratios for patients in the top 20%, 10%, 5%, 1% and 0.5% of the distribution of risk versus the remaining part of the distribution (80%, 90%, 95%, 99%, 99.5%) as the reference group. 95% confidence intervals and P-values are also provided.
All Cancer-PRSweb phenotypes offer odds ratios for the top 25%, 10%, 5%, 2% and 1% of the PRS distribution versus the rest, together with 95% confidence intervals and P-values.
The source study led by Craig et al. [28], from which we take the Glaucoma phenotype, provides odds ratios for the top 50%, 20%, 10%, 5%, 2% and 1% versus the rest of the distribution. This study offers odds ratios for Father, whose risk percentile is 88.675, but also for individuals above 50%, i.e., Daughter and Son’s, whose percentile risks are 65.01 and 65.21, respectively.
From Mars et al. [22] we selected their Prostate Cancer PRS, extracting odds ratios and 95% confidence intervals per standard deviation increase, using the FINRISK (n = 21,813) population as the validation dataset. (We did not use other phenotypes from this publication as they are already covered by Khera et al. [3] and we decided to choose PRS from the latter).
Abraham et al. [109], which studies Ischaemic Stroke, does not provide odds ratios. Instead, they offer a hazard ratio per standard deviation by age 75, using the UKB as the validation dataset.
For each extracted odds/hazard ratio, each source publication must be considered independently when reporting for an individual. In most cases, the PRS percentile of the individual lies within a reported interval from the source thresholds, but there are exceptions.
Percentile thresholds vary from ≥ 50% to ≥ 99.5%, depending on the source publication. We note all lower end confidence intervals as being > 1 and P-values much lower than the significance threshold of 0.05 (risks very likely not to have occurred by chance). Table 6 summarises source publication extracted risks based on PRS percentiles for each family member.
Effect of background population in percentile calculation
It has been previously reported that PRS distributions are affected by population stratification [114]. In order to test whether for our selection of PRS distributions the choice of background population for percentile calculations are significantly different, we conducted the analysis of phenotype percentiles individually. We checked whether there are any significant differences at the level of individual phenotypes when comparing the effect of background PRS distributions. Table 7 highlights phenotypes for family members (Father, Mother, Daughter, Son) in the top (red) and bottom (green) PRS quintiles using three background 1000G distributions (IBS, EUR and ALL). Italicised red/green are shown for phenotype PRS in the top 5th or bottom 5th percentile, respectively. We observe that the pattern of red/green, although generally conserved across the three background distributions and between family members, also show differences. Differences within the same individual reflect how the PRS percentile changes when comparing it against a different 1000G population group. For example, we see differences in Basal Cell Carcinoma, Ischaemic Stroke and Type 2 Diabetes. Primarily, these differences follow two patterns: (a) lower percentiles for IBS/EUR than ALL; e.g., Basal Cell Carcinoma; (b) higher percentiles in IBS/EUR and lower for ALL; e.g., Ischaemic Stroke and Type 2 Diabetes, with all family individuals having much higher percentiles in the IBS and EUR background distribution than ALL.
We also observe similar percentiles when comparing across distributions. We note as examples Colorectal Cancer, Coronary Artery Disease and Testicular Cancer, where a family member’s PRS percentile is similar across the different background distributions.
Whether the percentile PRS is consistent among different populations does not depend on the source study. For instance, looking at results of PRS from Khera et al. [3], Type 2 Diabetes gives inconsistent results (i.e., quintiles differing by > 20 percentile points) across population groups, while Coronary Artery Disease gives greater consistency. Similarly, we find consistent percentiles in Cancer-PRSweb phenotypes (Testicular Cancer) and inconsistent ones (Basal Cell Carcinoma).
Discussion
Approaches combining the information from large numbers of genomic variants into PRS promise substantial improvement of risk prediction for common diseases and cancer [115]. Implementation of PRS at scale in health services, however, remains a challenge, particularly the translation of PRS into actionable benefits for individuals. Governments in various countries, including in the UK [2], have the ambition to use PRS in healthcare settings, which implies that existing PRS studies do need to be translated into actionable tools for use at the individual level. Such translation requires standardisation so that implementation can be scaled to large numbers of people. In this paper we developed a proof-of-principle implementation of publicly available PRS information, following a systematic curation, deployment and translation of PRS into personalised risk assessments, using a family of four as a test case. A selection of 15 common diseases and cancers (phenotypes) resulted from our curation process, encompassing 37 million SNPs. We applied PRS to 1000 Genomes Project (1000G) participants, using the effect weights of over 96 billion risk alleles to construct a background distribution of 15 PRS from which to infer risk percentiles for each of our four family members.
Our curated set of PRS from 15 diverse conditions span autoimmune, cancer, cardiovascular and metabolic diseases. In what follows we discuss our PRS curation, risk percentile generation and interpretation of disease risk assessments as well as opportunities and limitations that such a PRS implementation provides for disease prevention.
PRS deployment requires considerable curation
Among the many hundreds of PRS we found in online repositories, we note varying study designs, PRS performances, validation cohorts and risk metrics. For instance, Coronary Artery Disease (Polygenic Score Catalogue ID: PGS000013) is based on a model adjusting for covariates such as age, sex, ancestry Principal Component 1–4, genotyping chip. However, another PRS for Coronary Artery Disease such as ID: PGS000018, used different covariates, which make them more difficult to compare. While this makes it challenging to deploy existing PRS data into a coherent framework for testing of individuals, it also reflects the diverse study designs and analysis methods of the original studies. We developed a set of curation criteria, allowing us to shortlist candidate PRS. Our own judgement was needed in order to evaluate their final inclusion in our analysis. This meant that our selection criteria were not always strictly followed, reducing the potential for standardisation and scalability.
Percentile PRS calculation lies at the core of risk inference
The concept of putting the individual into the context of a wider population has allowed us to posit a template for turning PRS developed at the population level into a tool which can be applied to individuals. We believe in this approach, largely because the risk metric which results is one of relative risk, consistent with the methodology underpinning the PRS validation process.
By calculating the PRS for each individual within the 1000G, and then placing the family members within the context of that distribution, a robust method of translating population level PRS into relevant individually related scores was arrived at. This is further supported by using whole genome sequencing, and thus avoiding the need for imputation of alleles at any given PRS SNP, which we expect to provide more accurate results. Moreover, from a total of 37 million SNPs in 15 phenotypes, 99.98% passed our bioinformatics curation criteria, which allowed us to reliably implement the published PRS in both our tested individuals and background 1000G populations.
In addition, background populations were independent from the cohorts used for training and validation of the PRS. This allowed us to independently test the effect of the choice of background population (IBS, EUR and ALL) for percentile calculation.
Importance of considering risk inheritance patterns individually
Part of the objective of this study is to offer a method for applying PRS which have been trained on large cohorts to individuals. We do see the impact of averaging mitigating the individual parents’ risk in the offspring, however this does not apply to all phenotypes. For example, for Coronary Artery Disease high risk percentiles were observed in both Daughter and Son, despite a low risk percentile in Father.
This result highlights the importance of not assuming expected population-level averages when it comes to analysing individuals and families.
Role of background population in percentile calculation
When we look at the individual phenotypes, the quintile analysis revealed that the results for some phenotypes are consistent between ALL and EUR or IBS (e.g. Coronary Artery disease and Colorectal Cancer) and wholly different in other phenotypes (e.g. Type 2 diabetes and Basal Cell Carcinoma).
We note that studies with multiple PRS (e.g. Khera et al. (2018), Cancer-PRSweb, etc.) contain phenotypes differing by > 20 percentile points for the same individual across population groups, suggesting that the results we are seeing are not due to study design.
In order to explain this consistency between ALL and IBS or EUR for some phenotypes and inconsistency for others, we can hypothesise that for ‘consistent’ quintile PRS percentiles, the frequencies of variants of their PRS SNPs are conserved across the different populations. This may mean that such PRS are more portable than others across different ancestry groups, however we stress that this would require further work. Such work might seek to validate the more ‘consistent’ PRS in non-European population groups. In turn, this validation would require access to (and the existence of) large scale biobank data in such populations, which remains a challenge.
Translation of percentiles into risk metrics
Our method for translating PRS percentiles into risk metrics relied on their availability in source publications. In order for us to reuse source publication risk metrics they had to be associated to PRS percentile interval thresholds. We found risk metrics to be variably reported, with some studies reporting odds ratios, others hazard ratios and others still no risk metric at all. Furthermore, some studies reported risk over the 80th PRS percentile threshold while others over the 75th or even the top 50th percentile. Still others reported risk of one group relative to a reference group (for instance Mars et al. [22]), rather than relative to the rest (for instance Khera et al. [3]).
We translated genetic risk regardless of thresholds wherever available, but it was not possible to follow a uniform set of rules with which to report risk. For future developments, a standard set of thresholds and risk metrics with which to report genetic risk would be highly desirable for at scale implementation. This would allow for direct comparison between different PRS, creating a common basis for a discussion about disease risk for complex genetic disorders.
Additionally, when considering healthcare interventions, similar phenotype odds ratios with different AUC performances may lead to different levels of confidence. To illustrate this point, we can compare two different, high quality approaches, in Khera et al. [3] and Abraham et al. [109]. The Coronary Artery Disease PRS AUC in Khera et al. has an AUC of 0.81, suggesting that it is able to stratify individuals into different risk bins with a good level of accuracy. With such a robust PRS, certain preventative healthcare interventions for an individual in a high-risk bin might be justified by the PRS alone (for instance lifestyle adjustments). By contrast, Abraham et al.’s Ischaemic Stroke PRS [109] has a C-index of 0.58. With a significantly less robust PRS such as this, it is harder to justify intervening based on the PRS alone, even if the individual in question shows up in a high-risk part of the distribution, as there is less confidence that the risk is correctly attributed to that individual. However, that does not mean that such a PRS is without use. As Abraham et al. [109] point out, the Ischaemic Stroke PRS with a C-index of 0.58 is still comparable to other common predictors of Ischaemic Stroke, for instance, family history of stroke (C-index of 0.56) Systolic Blood Pressure (C-index of 0.57) or BMI (C-index of 0.57). Therefore, while this PRS is not robust enough to be used on a standalone basis, it nonetheless adds value to the overall assessment of risk of Ischaemic Stroke, and when combined with other risk factors including hypertension, raises the C-index to 0.635.
Limitations of this implementation
First and foremost, our analysis is limited by the size of our use case, the family of four. As a result, we seek only to offer a proof of concept, and some insights which we believe are generalisable to implementations at a larger scale. A greater number of subjects would be needed in order to validate the specific results presented here.
The second important limitation is population constraints. Our percentile risk calculation has only been performed for an Iberian family. A subsequent analysis could consider individuals from different ancestry backgrounds against different population cohorts. Further, our results may have been affected by the way that the 1000G population groups are constructed. The same Iberian Spanish participants of the 1000G are included in the European subset population, and in turn, all Europeans are included in the total population of the 1000G (ALL). We also note that the Iberian population is of small size (n = 107) in the 1000G, reducing the statistical significance of results using that population as background distribution. We are also conscious that all PRS used here were themselves derived from and validated in Northern European populations (White British or Finnish), which may also contribute inaccuracy to our risk analysis in the IBS subpopulation. Privé et al. [116] suggests portability of such scores to Southern European populations might reduce prediction performance to 86% of that observed in the source population.
There are also a number of limitations imposed by bioinformatics constraints. We require the overwhelming majority of PRS SNPs to be present in the 1000G population, which may rule out some high quality PRS. Furthermore, we were only able to select PRS whose number of SNPs not present in the 1000G dataset was smaller than 5% of the total. Missing SNPs in the PRS calculation will have a greater impact for phenotypes where the PRS had few SNPs (e.g., Basal Cell Carcinoma is the phenotype that proportionally misses the greatest proportion of SNPs, 1 out of 23; 4.35%), in contrast to those that included all SNPs genome wide (e.g., Type 2 Diabetes ~ 7 million SNPs). We nevertheless believe that the expected impact is not significant, since for the great majority of PRS we have used, considerably less than 5% of the SNPs were missed or none at all (see Table 4, ‘% Missing SNPs’). Another weakness of our current methodology is that we have excluded PRS that contained SNPs in the X and Y chromosomes, resulting in more missing SNPs in certain phenotypes, and so causing us to exclude them (e.g., Testosterone Levels). Finally, if the minor allele frequency (MAF) of the source PRS is different from that of the tested individuals, the PRS may have different performance. Given that there is no MAF information for IBS in public data resources, this has limited our ability to filter by MAF discordance between the tested and source PRS. However, as shown by [8] other populations within the European continent tested with UK Biobank PRS data still conserve AUC performance within meaningful levels.
Further work
Further work could include the application of this methodology to a greater number of individuals, which would allow the validation of results obtained here, at small scale. When considering phenotype selection, it would be useful to compare different PRS for the same phenotype by showing how concordant PRS values are across different 1000G populations. Finally, as suggested above, we would propose further research into understanding the potential of those PRS whose average percentiles in tested individuals do not significantly differ across background populations as this could be an indication that they are more portable across different ancestries.
Conclusion
We have presented a comprehensive set of 15 curated PRS encompassing autoimmune, metabolic, cancer and cardiovascular diseases. We offer a proof-of-principle approach for an implementation of individualised PRS analysis, with a test case of a family of four using background distributions from 1000G. These 1000G populations allow us to calculate PRS and extrapolate them into relative risk for individuals using as input whole genome variant data. Calculated risk percentiles from PRS allow us to infer relative risks for any of the diseases analysed here. We show how current lack of standards for risk reporting challenges our ability to implement PRS more straightforwardly. It is also noted that different disease risks cannot be uniformly interpreted as their differences in study design, performances and risk reporting are not standardised. We further explore the effect of background population on an individual PRS percentile by comparing how different 1000G populations affect resulting PRS percentile calculations. All in all, this work offers insight into how PRS can be translated into relative risks for individuals, and therefore showcases their potential for their deployment in a preventative healthcare setting.
Availability of data and materials
The sources of PRS used in this study are included in Table 3 and Table 4 and were downloaded from the PGS Catalog (https://www.pgscatalog.org) and Cancer-PRSweb (https://prsweb.sph.umich.edu:8443). Request to access the family genome variation data should be directed to Manuel Corpas (m.corpas@cpm.onl) and are available upon reasonable request.
Abbreviations
- 1000G:
-
1000 genomes project
- AUC:
-
Area under curve
- BMI:
-
Body mass index
- GWAS:
-
Genome wide association study
- hg19:
-
Human genome 19 reference assembly
- HR:
-
Hazard ratio
- IBS:
-
Iberian Spanish
- MAF:
-
Minor allele frequency
- OR:
-
Odds ratio
- PRS:
-
Polygenic risk score
- SNP:
-
Single nucleotide polymorphism
References
Lewis CM, Vassos E. Polygenic risk scores: from research tools to clinical instruments. Genome Med. 2020;12:44.
Department of Health and Social Care. Genome UK: the future of healthcare. 2020. https://www.gov.uk/government/publications/genome-uk-the-future-of-healthcare. Accessed 7 Apr 2021.
Khera AV, Chaffin M, Aragam KG, Haas ME, Roselli C, Choi SH, et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet. 2018;50:1219–24.
Fullerton JM, Nurnberger JI. Polygenic risk scores in psychiatry: Will they be useful for clinicians. F1000Res. 2019. https://doi.org/10.12688/f1000research.18491.1.
Machini K, Ceyhan-Birsoy O, Azzariti DR, Sharma H, Rossetti P, Mahanta L, et al. Analyzing and reanalyzing the genome: findings from the MedSeq project. Am J Hum Genet. 2019;105:177–88.
Fritsche LG, Patil S, Beesley LJ, VandeHaar P, Salvatore M, Ma Y, et al. Cancer PRSweb: an online repository with polygenic risk scores for major cancer traits and their evaluation in two independent biobanks. Am J Hum Genet. 2020;107:815–36.
Lambert SA, Gil L, Jupp S, Ritchie SC, Xu Y, Buniello A, et al. The polygenic score catalog as an open database for reproducibility and systematic evaluation. Nat Genet. 2021. https://doi.org/10.1038/s41588-021-00783-5.
Gola D, Erdmann J, Läll K, Mägi R, Müller-Myhsok B, Schunkert H, et al. Population bias in polygenic risk prediction models for coronary artery disease. Circ Genom Precis Med. 2020;13: e002932.
Genomes Project Consortium, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Kendig KI, Baheti S, Bockol MA, Drucker TM, Hart SN, Heldenbrand JR, et al. Sentieon DNASeq variant calling workflow demonstrates strong computational performance and accuracy. Front Genet. 2019;10:736.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491–8.
Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. 2013.
Glusman G, Cariaso M, Jimenez R, Swan D, Greshake B, Bhak J, et al. Low budget analysis of direct-to-consumer genomic testing familial data. F1000Res. 2012. https://doi.org/10.12688/f1000research.1-3.v1.
Corpas M. A family experience of personal genomics. J Genet Couns. 2012;21:386–91.
Corpas M, Valdivia-Granda W, Torres N, Greshake B, Coletta A, Knaus A, et al. Crowdsourced direct-to-consumer genomic analysis of a family quartet. BMC Genomics. 2015;16:910.
Corpas M, Megy K, Mistry V, Metastasio A, Lehmann E. Whole genome interpretation for a family of five. Front Genet. 2021;12: 535123.
PGP-UK Consortium. Personal Genome Project UK (PGP-UK): a research and citizen science hybrid project in support of personalized medicine. BMC Med Genomics. 2018;11:108.
Vilhjálmsson BJ, Yang J, Finucane HK, Gusev A, Lindström S, Ripke S, et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am J Hum Genet. 2015;97:576–92.
Privé F, Vilhjálmsson BJ, Aschard H, Blum MGB. Making the most of clumping and thresholding for polygenic scores. Am J Hum Genet. 2019;105:1213–21.
Bycroft C, Freeman C, Petkova D, Band G, Elliott LT, Sharp K, et al. The UK biobank resource with deep phenotyping and genomic data. Nature. 2018;562:203–9.
Borodulin K, Tolonen H, Jousilahti P, Jula A, Juolevi A, Koskinen S, et al. Cohort profile: the national FINRISK study. Int J Epidemiol. 2018;47:696–696i.
Mars N, Koskela JT, Ripatti P, Kiiskinen TTJ, Havulinna AS, Lindbohm JV, et al. Polygenic and clinical risk scores and their impact on age at onset and prediction of cardiometabolic diseases and common cancers. Nat Med. 2020;26:549–57.
Martin AR, Kanai M, Kamatani Y, Okada Y, Neale BM, Daly MJ. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet. 2019;51:584–91.
Meisner A, Kundu P, Zhang YD, Lan LV, Kim S, Ghandwani D, et al. Combined utility of 25 disease and risk factor polygenic risk scores for stratifying risk of all-cause mortality. Am J Hum Genet. 2020;107:418–31.
Reid S, Alexsson A, Frodlund M, Morris D, Sandling JK, Bolin K, et al. High genetic risk score is associated with early disease onset, damage accrual and decreased survival in systemic lupus erythematosus. Ann Rheum Dis. 2020;79:363–9.
Mavaddat N, Michailidou K, Dennis J, Lush M, Fachal L, Lee A, et al. Polygenic risk scores for prediction of breast cancer and breast cancer subtypes. Am J Hum Genet. 2019;104:21–34.
Schumacher FR, Al Olama AA, Berndt SI, Benlloch S, Ahmed M, Saunders EJ, et al. Association analyses of more than 140,000 men identify 63 new prostate cancer susceptibility loci. Nat Genet. 2018;50:928–36.
Craig JE, Han X, Qassim A, Hassall M, Cooke Bailey JN, Kinzy TG, et al. Multitrait analysis of glaucoma identifies new risk loci and enables polygenic prediction of disease susceptibility and progression. Nat Genet. 2020;52:160–6.
Wang Z, McGlynn KA, Rajpert-De Meyts E, Bishop DT, Chung CC, Dalgaard MD, et al. Meta-analysis of five genome-wide association studies identifies multiple new loci associated with testicular germ cell tumor. Nat Genet. 2017;49:1141–7.
Litchfield K, Levy M, Orlando G, Loveday C, Law PJ, Migliorini G, et al. Identification of 19 new risk loci and potential regulatory mechanisms influencing susceptibility to testicular germ cell tumor. Nat Genet. 2017;49:1133–40.
Litchfield K, Holroyd A, Lloyd A, Broderick P, Nsengimana J, Eeles R, et al. Identification of four new susceptibility loci for testicular germ cell tumour. Nat Commun. 2015;6:8690.
Kristiansen W, Karlsson R, Rounge TB, Whitington T, Andreassen BK, Magnusson PK, et al. Two new loci and gene sets related to sex determination and cancer progression are associated with susceptibility to testicular germ cell tumor. Hum Mol Genet. 2015;24:4138–46.
Ruark E, Seal S, McDonald H, Zhang F, Elliot A, Lau K, et al. Identification of nine new susceptibility loci for testicular cancer, including variants near DAZL and PRDM14. Nat Genet. 2013;45:686–9.
Chung CC, Kanetsky PA, Wang Z, Hildebrandt MAT, Koster R, Skotheim RI, et al. Meta-analysis identifies four new loci associated with testicular germ cell tumor. Nat Genet. 2013;45:680–5.
Turnbull C, Rapley EA, Seal S, Pernet D, Renwick A, Hughes D, et al. Variants near DMRT1, TERT and ATF7IP are associated with testicular germ cell cancer. Nat Genet. 2010;42:604–7.
Rapley EA, Turnbull C, Al Olama AA, Dermitzakis ET, Linger R, Huddart RA, et al. A genome-wide association study of testicular germ cell tumor. Nat Genet. 2009;41:807–10.
Law PJ, Berndt SI, Speedy HE, Camp NJ, Sava GP, Skibola CF, et al. Genome-wide association analysis implicates dysregulation of immunity genes in chronic lymphocytic leukaemia. Nat Commun. 2017;8:14175.
Berndt SI, Camp NJ, Skibola CF, Vijai J, Wang Z, Gu J, et al. Meta-analysis of genome-wide association studies discovers multiple loci for chronic lymphocytic leukemia. Nat Commun. 2016;7:10933.
Speedy HE, Di Bernardo MC, Sava GP, Dyer MJS, Holroyd A, Wang Y, et al. A genome-wide association study identifies multiple susceptibility loci for chronic lymphocytic leukemia. Nat Genet. 2014;46:56–60.
Berndt SI, Skibola CF, Joseph V, Camp NJ, Nieters A, Wang Z, et al. Genome-wide association study identifies multiple risk loci for chronic lymphocytic leukemia. Nat Genet. 2013;45:868–76.
Slager SL, Skibola CF, Di Bernardo MC, Conde L, Broderick P, McDonnell SK, et al. Common variation at 6p21.31 (BAK1) influences the risk of chronic lymphocytic leukemia. Blood. 2012;120:843–6.
Slager SL, Rabe KG, Achenbach SJ, Vachon CM, Goldin LR, Strom SS, et al. Genome-wide association study identifies a novel susceptibility locus at 6p21.3 among familial CLL. Blood. 2011;117:1911–6.
Di Bernardo MC, Crowther-Swanepoel D, Broderick P, Webb E, Sellick G, Wild R, et al. A genome-wide association study identifies six susceptibility loci for chronic lymphocytic leukemia. Nat Genet. 2008;40:1204–10.
Gudmundsson J, Thorleifsson G, Sigurdsson JK, Stefansdottir L, Jonasson JG, Gudjonsson SA, et al. A genome-wide association study yields five novel thyroid cancer risk loci. Nat Commun. 2017;8:14517.
Mancikova V, Cruz R, Inglada-Pérez L, Fernández-Rozadilla C, Landa I, Cameselle-Teijeiro J, et al. Thyroid cancer GWAS identifies 10q26.12 and 6q14.1 as novel susceptibility loci and reveals genetic heterogeneity among populations. Int J Cancer. 2015;137:1870–8.
Köhler A, Chen B, Gemignani F, Elisei R, Romei C, Figlioli G, et al. Genome-wide association study on differentiated thyroid cancer. J Clin Endocrinol Metab. 2013;98:E1674–81.
Gudmundsson J, Sulem P, Gudbjartsson DF, Jonasson JG, Masson G, He H, et al. Discovery of common variants associated with low TSH levels and thyroid cancer risk. Nat Genet. 2012;44:319–22.
Melin BS, Barnholtz-Sloan JS, Wrensch MR, Johansen C, Il’yasova D, Kinnersley B, et al. Genome-wide association study of glioma subtypes identifies specific differences in genetic susceptibility to glioblastoma and non-glioblastoma tumors. Nat Genet. 2017;49:789–94.
Kinnersley B, Labussière M, Holroyd A, Di Stefano A-L, Broderick P, Vijayakrishnan J, et al. Genome-wide association study identifies multiple susceptibility loci for glioma. Nat Commun. 2015;6:8559.
Walsh KM, Codd V, Smirnov IV, Rice T, Decker PA, Hansen HM, et al. Variants near TERT and TERC influencing telomere length are associated with high-grade glioma risk. Nat Genet. 2014;46:731–5.
Rajaraman P, Melin BS, Wang Z, McKean-Cowdin R, Michaud DS, Wang SS, et al. Genome-wide association study of glioma and meta-analysis. Hum Genet. 2012;131:1877–88.
Sanson M, Hosking FJ, Shete S, Zelenika D, Dobbins SE, Ma Y, et al. Chromosome 7p11.2 (EGFR) variation influences glioma risk. Hum Mol Genet. 2011;20:2897–904.
Shete S, Hosking FJ, Robertson LB, Dobbins SE, Sanson M, Malmer B, et al. Genome-wide association study identifies five susceptibility loci for glioma. Nat Genet. 2009;41:899–904.
Ransohoff KJ, Wu W, Cho HG, Chahal HC, Lin Y, Dai H-J, et al. Two-stage genome-wide association study identifies a novel susceptibility locus associated with melanoma. Oncotarget. 2017;8:17586–92.
Law MH, Bishop DT, Lee JE, Brossard M, Martin NG, Moses EK, et al. Genome-wide meta-analysis identifies five new susceptibility loci for cutaneous malignant melanoma. Nat Genet. 2015;47:987–95.
Iles MM, Law MH, Stacey SN, Han J, Fang S, Pfeiffer R, et al. A variant in FTO shows association with melanoma risk not due to BMI. Nat Genet. 2013;45:428–32.
Barrett JH, Iles MM, Harland M, Taylor JC, Aitken JF, Andresen PA, et al. Genome-wide association study identifies three new melanoma susceptibility loci. Nat Genet. 2011;43:1108–13.
Macgregor S, Montgomery GW, Liu JZ, Zhao ZZ, Henders AK, Stark M, et al. Genome-wide association study identifies a new melanoma susceptibility locus at 1q21.3. Nat Genet. 2011;43:1114–8.
Bishop DT, Demenais F, Iles MM, Harland M, Taylor JC, Corda E, et al. Genome-wide association study identifies three loci associated with melanoma risk. Nat Genet. 2009;41:920–5.
Brown KM, Macgregor S, Montgomery GW, Craig DW, Zhao ZZ, Iyadurai K, et al. Common sequence variants on 20q11.22 confer melanoma susceptibility. Nat Genet. 2008;40:838–40.
Huyghe JR, Bien SA, Harrison TA, Kang HM, Chen S, Schmit SL, et al. Discovery of common and rare genetic risk variants for colorectal cancer. Nat Genet. 2019;51:76–87.
Lin Y, Chahal HS, Wu W, Cho HG, Ransohoff KJ, Dai H, et al. Association between genetic variation within vitamin D receptor-DNA binding sites and risk of basal cell carcinoma. Int J Cancer. 2017;140:2085–91.
Chahal HS, Wu W, Ransohoff KJ, Yang L, Hedlin H, Desai M, et al. Genome-wide association study identifies 14 novel risk alleles associated with basal cell carcinoma. Nat Commun. 2016;7:12510.
Stacey SN, Helgason H, Gudjonsson SA, Thorleifsson G, Zink F, Sigurdsson A, et al. New basal cell carcinoma susceptibility loci. Nat Commun. 2015;6:6825.
Stacey SN, Sulem P, Gudbjartsson DF, Jonasdottir A, Thorleifsson G, Gudjonsson SA, et al. Germline sequence variants in TGM3 and RGS22 confer risk of basal cell carcinoma. Hum Mol Genet. 2014;23:3045–53.
Nan H, Xu M, Kraft P, Qureshi AA, Chen C, Guo Q, et al. Genome-wide association study identifies novel alleles associated with risk of cutaneous basal cell carcinoma and squamous cell carcinoma. Hum Mol Genet. 2011;20:3718–24.
Rafnar T, Sulem P, Stacey SN, Geller F, Gudmundsson J, Sigurdsson A, et al. Sequence variants at the TERT-CLPTM1L locus associate with many cancer types. Nat Genet. 2009;41:221–7.
Stacey SN, Gudbjartsson DF, Sulem P, Bergthorsson JT, Kumar R, Thorleifsson G, et al. Common variants on 1p36 and 1q42 are associated with cutaneous basal cell carcinoma but not with melanoma or pigmentation traits. Nat Genet. 2008;40:1313–8.
Klein AP, Wolpin BM, Risch HA, Stolzenberg-Solomon RZ, Mocci E, Zhang M, et al. Genome-wide meta-analysis identifies five new susceptibility loci for pancreatic cancer. Nat Commun. 2018;9:556.
Zhang M, Wang Z, Obazee O, Jia J, Childs EJ, Hoskins J, et al. Three new pancreatic cancer susceptibility signals identified on chromosomes 1q32.1, 5p15.33 and 8q24.21. Oncotarget. 2016;7:66328–43.
Wolpin BM, Rizzato C, Kraft P, Kooperberg C, Petersen GM, Wang Z, et al. Genome-wide association study identifies multiple susceptibility loci for pancreatic cancer. Nat Genet. 2014;46:994–1000.
Wu C, Kraft P, Stolzenberg-Solomon R, Steplowski E, Brotzman M, Xu M, et al. Genome-wide association study of survival in patients with pancreatic adenocarcinoma. Gut. 2014;63:152–60.
Amundadottir L, Kraft P, Stolzenberg-Solomon RZ, Fuchs CS, Petersen GM, Arslan AA, et al. Genome-wide association study identifies variants in the ABO locus associated with susceptibility to pancreatic cancer. Nat Genet. 2009;41:986–90.
Went M, Sud A, Försti A, Halvarsson B-M, Weinhold N, Kimber S, et al. Identification of multiple risk loci and regulatory mechanisms influencing susceptibility to multiple myeloma. Nat Commun. 2018;9:3707.
Mitchell JS, Li N, Weinhold N, Försti A, Ali M, van Duin M, et al. Genome-wide association study identifies multiple susceptibility loci for multiple myeloma. Nat Commun. 2016;7:12050.
Swaminathan B, Thorleifsson G, Jöud M, Ali M, Johnsson E, Ajore R, et al. Variants in ELL2 influencing immunoglobulin levels associate with multiple myeloma. Nat Commun. 2015;6:7213.
Chubb D, Weinhold N, Broderick P, Chen B, Johnson DC, Försti A, et al. Common variation at 3q26.2, 6p21.33, 17p11.2 and 22q13.1 influences multiple myeloma risk. Nat Genet. 2013;45:1221–5.
Weinhold N, Johnson DC, Chubb D, Chen B, Försti A, Hosking FJ, et al. The CCND1 c.870G>A polymorphism is a risk factor for t(11;14)(q13;q32) multiple myeloma. Nat Genet. 2013;45:522–5.
Broderick P, Chubb D, Johnson DC, Weinhold N, Försti A, Lloyd A, et al. Common variation at 3p22.1 and 7p15.3 influences multiple myeloma risk. Nat Genet. 2011;44:58–61.
O’Mara TA, Glubb DM, Amant F, Annibali D, Ashton K, Attia J, et al. Identification of nine new susceptibility loci for endometrial cancer. Nat Commun. 2018;9:3166.
Cheng TH, Thompson DJ, O’Mara TA, Painter JN, Glubb DM, Flach S, et al. Five endometrial cancer risk loci identified through genome-wide association analysis. Nat Genet. 2016;48:667–74.
Spurdle AB, Thompson DJ, Ahmed S, Ferguson K, Healey CS, O’Mara T, et al. Genome-wide association study identifies a common variant associated with risk of endometrial cancer. Nat Genet. 2011;43:451–4.
Rafnar T, Sulem P, Thorleifsson G, Vermeulen SH, Helgason H, Saemundsdottir J, et al. Genome-wide association study yields variants at 20p12.2 that associate with urinary bladder cancer. Hum Mol Genet. 2014;23:5545–57.
Figueroa JD, Ye Y, Siddiq A, Garcia-Closas M, Chatterjee N, Prokunina-Olsson L, et al. Genome-wide association study identifies multiple loci associated with bladder cancer risk. Hum Mol Genet. 2014;23:1387–98.
Rafnar T, Vermeulen SH, Sulem P, Thorleifsson G, Aben KK, Witjes JA, et al. European genome-wide association study identifies SLC14A1 as a new urinary bladder cancer susceptibility gene. Hum Mol Genet. 2011;20:4268–81.
Rothman N, Garcia-Closas M, Chatterjee N, Malats N, Wu X, Figueroa JD, et al. A multi-stage genome-wide association study of bladder cancer identifies multiple susceptibility loci. Nat Genet. 2010;42:978–84.
Kiemeney LA, Sulem P, Besenbacher S, Vermeulen SH, Sigurdsson A, Thorleifsson G, et al. A sequence variant at 4p16.3 confers susceptibility to urinary bladder cancer. Nat Genet. 2010;42:415–9.
Wu X, Ye Y, Kiemeney LA, Sulem P, Rafnar T, Matullo G, et al. Genetic variation in the prostate stem cell antigen gene PSCA confers susceptibility to urinary bladder cancer. Nat Genet. 2009;41:991–5.
Kiemeney LA, Thorlacius S, Sulem P, Geller F, Aben KKH, Stacey SN, et al. Sequence variant on 8q24 confers susceptibility to urinary bladder cancer. Nat Genet. 2008;40:1307–12.
Chahal HS, Lin Y, Ransohoff KJ, Hinds DA, Wu W, Dai H-J, et al. Genome-wide association study identifies novel susceptibility loci for cutaneous squamous cell carcinoma. Nat Commun. 2016;7:12048.
Phelan CM, Kuchenbaecker KB, Tyrer JP, Kar SP, Lawrenson K, Winham SJ, et al. Identification of 12 new susceptibility loci for different histotypes of epithelial ovarian cancer. Nat Genet. 2017;49:680–91.
Kuchenbaecker KB, Ramus SJ, Tyrer J, Lee A, Shen HC, Beesley J, et al. Identification of six new susceptibility loci for invasive epithelial ovarian cancer. Nat Genet. 2015;47:164–71.
Couch FJ, Wang X, McGuffog L, Lee A, Olswold C, Kuchenbaecker KB, et al. Genome-wide association study in BRCA1 mutation carriers identifies novel loci associated with breast and ovarian cancer risk. PLoS Genet. 2013;9: e1003212.
Rafnar T, Gudbjartsson DF, Sulem P, Jonasdottir A, Sigurdsson A, Jonasdottir A, et al. Mutations in BRIP1 confer high risk of ovarian cancer. Nat Genet. 2011;43:1104–7.
Song H, Ramus SJ, Tyrer J, Bolton KL, Gentry-Maharaj A, Wozniak E, et al. A genome-wide association study identifies a new ovarian cancer susceptibility locus on 9p22.2. Nat Genet. 2009;41:996–1000.
Byun J, Schwartz AG, Lusk C, Wenzlaff AS, de Andrade M, Mandal D, et al. Genome-wide association study of familial lung cancer. Carcinogenesis. 2018;39:1135–40.
McKay JD, Hung RJ, Han Y, Zong X, Carreras-Torres R, Christiani DC, et al. Large-scale association analysis identifies new lung cancer susceptibility loci and heterogeneity in genetic susceptibility across histological subtypes. Nat Genet. 2017;49:1126–32.
Wang Y, McKay JD, Rafnar T, Wang Z, Timofeeva MN, Broderick P, et al. Rare variants of large effect in BRCA2 and CHEK2 affect risk of lung cancer. Nat Genet. 2014;46:736–41.
Landi MT, Chatterjee N, Yu K, Goldin LR, Goldstein AM, Rotunno M, et al. A genome-wide association study of lung cancer identifies a region of chromosome 5p15 associated with risk for adenocarcinoma. Am J Hum Genet. 2009;85:679–91.
Skibola CF, Berndt SI, Vijai J, Conde L, Wang Z, Yeager M, et al. Genome-wide association study identifies five susceptibility loci for follicular lymphoma outside the HLA region. Am J Hum Genet. 2014;95:462–71.
Cerhan JR, Berndt SI, Vijai J, Ghesquières H, McKay J, Wang SS, et al. Genome-wide association study identifies multiple susceptibility loci for diffuse large B cell lymphoma. Nat Genet. 2014;46:1233–8.
Smedby KE, Foo JN, Skibola CF, Darabi H, Conde L, Hjalgrim H, et al. GWAS of follicular lymphoma reveals allelic heterogeneity at 6p21.32 and suggests shared genetic susceptibility with diffuse large B-cell lymphoma. PLoS Genet. 2011;7:e1001378.
Conde L, Halperin E, Akers NK, Brown KM, Smedby KE, Rothman N, et al. Genome-wide association study of follicular lymphoma identifies a risk locus at 6p21.32. Nat Genet. 2010;42:661–4.
Skibola CF, Bracci PM, Halperin E, Conde L, Craig DW, Agana L, et al. Genetic variants at 6p21.33 are associated with susceptibility to follicular lymphoma. Nat Genet. 2009;41:873–5.
Scelo G, Purdue MP, Brown KM, Johansson M, Wang Z, Eckel-Passow JE, et al. Genome-wide association study identifies multiple risk loci for renal cell carcinoma. Nat Commun. 2017;8:15724.
Turnbull C, Perdeaux ER, Pernet D, Naranjo A, Renwick A, Seal S, et al. A genome-wide association study identifies susceptibility loci for Wilms tumor. Nat Genet. 2012;44:681–4.
Inouye M, Abraham G, Nelson CP, Wood AM, Sweeting MJ, Dudbridge F, et al. Genomic risk prediction of coronary artery disease in 480,000 adults: implications for primary prevention. J Am Coll Cardiol. 2018;72:1883–93.
Wang M, Menon R, Mishra S, Patel AP, Chaffin M, Tanneeru D, et al. Validation of a genome-wide polygenic score for coronary artery disease in South Asians. J Am Coll Cardiol. 2020;76:703–14.
Abraham G, Malik R, Yonova-Doing E, Salim A, Wang T, Danesh J, et al. Genomic risk score offers predictive performance comparable to clinical risk factors for ischaemic stroke. Nat Commun. 2019;10:5819.
Klarin D, Busenkell E, Judy R, Lynch J, Levin M, Haessler J, et al. Genome-wide association analysis of venous thromboembolism identifies new risk loci and genetic overlap with arterial vascular disease. Nat Genet. 2019;51:1574–9.
Kuchenbaecker K, Telkar N, Reiker T, Walters RG, Lin K, Eriksson A, et al. The transferability of lipid loci across African, Asian and European Cohorts. Nat Commun. 2019;10:4330.
Flynn E, Tanigawa Y, Rodriguez F, Altman RB, Sinnott-Armstrong N, Rivas MA. Sex-specific genetic effects across biomarkers. Eur J Hum Genet. 2021;29:154–63.
Weiner DJ, Wigdor EM, Ripke S, Walters RK, Kosmicki JA, Grove J, et al. Polygenic transmission disequilibrium confirms that common and rare variation act additively to create risk for autism spectrum disorders. Nat Genet. 2017;49:978–85.
Khera AV, Chaffin M, Zekavat SM, Collins RL, Roselli C, Natarajan P, et al. Whole-genome sequencing to characterize monogenic and polygenic contributions in patients hospitalized with early-onset myocardial infarction. Circulation. 2019;139:1593–602.
Thomas M, Sakoda LC, Hoffmeister M, Rosenthal EA, Lee JK, van Duijnhoven FJB, et al. Genome-wide modeling of polygenic risk score in colorectal cancer risk. Am J Hum Genet. 2020;107:432–44.
Privé F, Aschard H, Carmi S, Folkersen L, Hoggart C, O’Reilly PF, et al. Portability of 245 polygenic scores when derived from the UK Biobank and applied to 9 ancestry groups from the same cohort. Am J Hum Genet. 2022;109:373.
Zheng X, Levine D, Shen J, Gogarten SM, Laurie C, Weir BS. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics. 2012;28(24):3326–8. https://doi.org/10.1093/bioinformatics/bts606 (Epub 2012 Oct 11. PMID: 23060615; PMCID: PMC3519454).
Acknowledgements
Not applicable.
Funding
Cambridge Precision Medicine Limited funded this project, including publication costs. Cambridge Precision Medicine Limited intellectual property was used in the design of the study and collection, analysis, and interpretation of data.
Author information
Authors and Affiliations
Contributions
MC and EL conceived the experiments and performed the analysis. MC wrote the paper with contributions from all authors. All authors read and approved the paper.
About this supplement
This article has been published as part of BMC Medical Genomics Volume 15 Supplement 3, 2022: Personal Genomes: Accessing, Sharing and Interpretation During Pandemic Times. The full contents of the supplement are available online at https://bmcmedgenomics.biomedcentral.com/articles/supplements/volume-15-supplement-3.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
All participants underwent a consent process and signed a consent form accepting the terms and conditions of this project as well as the potential consequences of performing this analysis. This project has been independently assessed and approved by the Ethics Committee of Universidad Internacional de La Rioja (code PI:029/2020).
Consent for publication
All participants of this project have given their informed consent to publish results as part of this publication.
Competing interests
At the time of this writing, MC, KM, AM and EL are associated with Cambridge Precision Medicine Limited.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix
Appendix
Principal Component Analysis (PCA) of Family Quartet in the context of 1000G. We used R packages (gdsfmt and SNPRelate [117] for PCA. We used in total ~ 22 K SNPs after linkage pruning. All the 1000G individuals appear in blue, while the family quartet in red. The left cluster corresponds to 1000G Asian individuals, the bottom-centre cluster to Europeans and the top right to Africans. The family quartet appears clearly in the European cluster.

Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Corpas, M., Megy, K., Metastasio, A. et al. Implementation of individualised polygenic risk score analysis: a test case of a family of four. BMC Med Genomics 15 (Suppl 3), 207 (2022). https://doi.org/10.1186/s12920-022-01331-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12920-022-01331-8