Abstract
Background
We implemented a longitudinal study to determine the incidence of Brucella infection in cattle, camels, sheep and goats that were being raised in a pastoral area in Isiolo County, Kenya. An initial cross-sectional survey was implemented to identify unexposed animals for follow up; that survey used 141 camels, 216 cattle, 208 sheep and 161 goats. Sera from these animals were screened for Brucella spp. using the Rose Bengal Plate test (RBPT), a modified RBPT, and an indirect multispecies Enzyme Linked Immunosorbent Assay (iELISA). Results of RBPT and iELISA were interpreted in parallel to determine seroprevalence. A total of 30 camels, 31 cattle, 22 sheep and 32 goats that were seronegative by all the above tests were recruited in a subsequent longitudinal study for follow up. These animals were followed for 12 months and tested for anti-Brucella antibodies using iELISA. Seroconversion among these animals was defined by a positive iELISA test following a negative iELISA result in the previous sampling period. All seropositive samples were further tested using real-time PCR-based assays to identify Brucella species. These analyses targeted the alkB and BMEI1162 genes for B. abortus, and B. melitensis, respectively. Data from the longitudinal study were analysed using Cox proportional hazards model that accounted for within-herds clustering of Brucella infections.
Results
The overall incidence rate of Brucella infection was 0.024 (95% confidence interval [CI]: 0.014–0.037) cases per animal-months at risk. Brucella infection incidence in camels, cattle, goats and sheep were 0.053 (0.022–0.104), 0.028 (0.010–0.061), 0.013 (0.003–0.036) and 0.006 (0.0002–0.034) cases per animal-months at risk, respectively. The incidence rate of Brucella infection among females and males were 0.020 (0.009–0.036) and 0.016 (0.004–0.091), respectively. Real-time PCR analyses showed that B. abortus was more prevalent than B. melitensis in the area. Results of multivariable Cox regression analysis identified species (camels and cattle) as an important predictor of Brucella spp. exposure in animals.
Conclusions
This study estimated an overall brucellosis incidence of 0.024 cases per animal-months at risk with camels and cattle having higher incidence than sheep and goats. These results will inform surveillance studies in the area.
Similar content being viewed by others
Background
Brucellosis is an important zoonotic disease that affects a great variety of hosts such as livestock (cattle, sheep, goats and camels), humans and wildlife [1]. Whereas this disease has been successfully controlled or eradicated in livestock populations in many developed countries including New Zealand, Japan and Australia [2], it remains a major problem affecting both livestock production and humans in Kenya [3], and also other parts of Africa [4]. Brucellosis causes direct production losses resulting from abortions, stillbirths, infertility, the mortality of calves/kids/lambs, longer calving intervals, reduced draught power, poor weight gain, and reduced milk production [5]. The etiological agent of this disease is an intracellular gram-negative coccobacillus of the genus Brucella. The main Brucella spp. that affect livestock species include B. abortus (cattle, camels), B. melitensis (sheep, goats), B. suis (pigs), and B. ovis (sheep) [1]. Humans serve as incidental hosts for Brucella spp. with B. melitensis, B. abortus, B. suis and B. canis being the main pathogenic species [6]. While Brucella spp. may show host preference, inter-species transmission of this pathogen may occur in areas with intense interactions between livestock and wildlife [7], or in mixed livestock production systems [8]. For example, cattle are often infected by B. suis and B. melitensis [9]. B. abortus has also been detected in pigs [10] and small ruminants (sheep and goats) [11].
There are limited studies that have been carried out to understand the epidemiology of Brucella spp. in Kenya [3] even though this pathogen is known to be endemic in pastoral areas [12]. Many seroprevalence studies have been done in the country involving livestock and humans. In pastoral areas, seroprevalences in humans often range between < 1–46.5% [3], while in livestock, they range between 3 and 40% [3, 8, 13, 14]. Seroprevalance in livestock is often associated with advanced age, large herd sizes, communal herding, and pastoralism [8, 12, 14], while in humans, advanced age, consumption of raw meat or unpasteurised milk and poor access to health services are known risk factors [2, 15]. Although seroprevalence estimates provide useful insights on the distribution of burden, they may be confusing for some diseases like brucellosis whose antibodies persist in circulation for months following recovery of the infection. In such cases, measures of incidence would provide more realistic indicators of burden.
There are also major challenges with screening of Brucella in humans and animals. This is a major limitation in remote areas due to limited veterinary and animal health personnel, poor laboratory infrastructure, and lack of biocontainment facilities required for culturing the agent [16, 17]. Due to these limitations, the diagnosis of brucellosis in animals is mainly performed using serological tests such as Rose Bengal Plate Test (RBPT), Milk Ring Test (MRT), Serum Agglutination Test (SAT), Complement Fixation Test (CFT), Enzyme Linked Immunosorbent Assay (iELISA), Competitive Enzyme Immunoassays and Fluorescence Polarization Assay (FPA) which can be performed in laboratories with simple equipment [17]. Among these tests, RBPT is more convenient for low-income countries since it is less technologically demanding, less expensive and yields good results if standardized correctly under local conditions with proven bacteriological samples [18]. These serological tests are often used in series or parallel depending on the overall objective of the study. Whereas the parallel testing of sera increases the overall diagnostic sensitivity, this testing strategy also reduces specificity unlike in series testing.
This study was implemented to determine the seroprevalence and incidence of Brucella infection in cattle, camels, sheep, and goats raised in a common (pastoral) area in Isiolo County, northern Kenya. Our study further identified the potential risk factors associated with Brucella spp. seroprevalence and incidence in animals as well as the main Brucella species circulating among livestock animals in the area. Our findings form the basis for further One Health surveillance studies.
Results
Descriptive results
A total of 841 animals consisting 382 cattle, 185 sheep, 174 goats and 100 camels were sampled in the cross-sectional survey. The total number of cattle and camel herds sampled were 10 and 3 respectively, while sheep and goat flocks were 8 and 10, respectively. The overall median herd size for all animals was 25 (range; 8–110) while those of cattle and camels were 26 (range; 8–110) and 40 (range; 10–50), respectively. For sheep and goats, the median flock sizes were respectively 17.5 (range; 2–51) and 11 (range; 3–64). A considerable proportion of the sampled animals, 309 (36.7%) were not included in the analysis due to either missing epidemiological data or they were not tested using RBPT and mRBPT due to logistical constraints. The overall apparent seroprevalence of Brucella spp. at animal-level was 11.3% (95% CI; 8.6–14.0, n = 532) based on the parallel interpreted results of the conventional RBPT and iELISA tests. In decreasing order, the true seroprevalences of Brucella spp. in camels, cattle, goats and sheep were 23.7% (95% CI; 3.36–50), 14.4% (95% CI; 10.3–18.1), 13.5% (95% CI; 8.2–21.4) and 2.7% (95% CI; 0.9–7.8), respectively. The apparent seroprevalences of Brucella spp. by the animal-level independent factors used in the study are presented in Table 1. These varied significantly between the livestock species (Fisher’s exact 2-tailed p = 0.002) and sex (p = 0.027). More female animals (12.6%; 95% CI; 9.7–16.0) tested positive for Brucella spp. antibodies compared to males (4.5% 95% CI; 1.1–8.4) (Table 1). We also found a statistically significant difference between Brucella spp. seroprevalence and the age of animals (P = 0.002); adult animals had higher seroprevalence (13.4%; 95% CI; 10.3–17.0) compared to weaners (6.7%; 95% CI; 1.8–16.2) and young animals (0.0%) (Table 1). Results on cross-tabulation of Brucella spp. seroprevalence by species, sex, age and pregnancy status are summarized in Table 2.
There was a substantial level of agreement between the three serological tests (Cohen’s Kappa statistic k = 0.78). Nevertheless, the proportion of seropositive animals detected by the three tests differed significantly (Cochran’s Q test = 18.5, df = 2, p < 0.001). Further post-hoc pair-wise analysis using McNemar’s χ2 showed significant differences between RBPT and mRBPT (p < 0.001), RBPT and iELISA (p = 0.001), but not between mRBPT and iELISA (p = 0.302). Overall, more seropositive animals were detected by the iELISA (10.3%; 95% CI; 7.7–13.0), followed in order by mRBPT (9.4%; 95% CI; 7.1–11.8) and RBPT (7.0%; 95% CI; 5.1–9.1).
A total of 60 seropositive samples were further tested using real-time PCR-based assays. The real-time PCR assay targeting the genus-specific (bcsp31) gene detected genus Brucella DNA in 49 (81.7%) samples; all of which tested positive for B. abortus. There was no B. melitensis DNA detected in any of these samples.
Risk factor analysis
Table 1 shows the results of the independent variables assessed for their association with Brucella spp. seroprevalence using univariable mixed-effects logistic regression models. The results of the final multivariable logistic regression model showed that Brucella spp. seropositivity was significantly lower among male animals (p = 0.012) compared to females (Table 3). Among the livestock species sampled, sheep had statistically significant lower odds of Brucella spp. seropositivity than cattle (Table 3). The ICC for within-herd/flock clustering of animals was estimated to be 0.10 (95% CI; 0.02–0.13).
Brucella infection incidence results
Table 4 shows the number of observed Brucella infection cases and the estimated animal-months at risk stratified by livestock species, sex and age. The estimated overall incidence rate of Brucella infection in all animals was 0.024 (95% CI; 0.014–0.037) cases per animal-months at risk. The incidence of Brucella infection in camel, cattle, goats and sheep were 0.053 (95% CI; 0.022–0.104), 0.028 (95% CI; 0.010–0.061), 0.013 (95% CI; 0.003–0.036) and 0.006 (95% CI; 0.0002–0.034) cases per animal-months at risk, respectively (Table 4). Considering animal sex, the incidence rate of Brucella infection were respectively 0.020 (95% CI; 0.009–0.036) and 0.016 (95% CI; 0.004–0.091) among females and males, while based on age, young animals had a slightly higher incidence rate compared to adults (Table 4).
The results of the univariable Cox regression analysis are shown in supplementary file 1. Among the investigated risk factors, only species (camels and cattle) was identified as a significant predictor of Brucella spp. exposure in animals by multivariable Cox regression analysis (Table 5). The results of the global test used to assess the proportional hazard assumption indicated that this assumption was satisfied (χ2 = 7.4, df = 5, p = 0.190). Also, all the investigated covariates had p-values of > 0.05.
Discussion
To the best of our knowledge, this is the first study to determine the incidence of Brucella infection in livestock in a pastoral area in Kenya. In both cross-sectional and longitudinal studies, the seropositivity of Brucella spp. in animals was due to natural exposure as vaccination of animals against brucellosis is not done in the area, except in a few commercial farms in other parts of the country. The proportion of Brucella spp. seropositive animals detected by mRBPT (9.4%) and iELISA (10.3%) did not differ significantly. Both tests also detected a significantly higher number of seropositive animals than RBPT (7.0%). This finding confirms that mRBPT provides comparable results as iELISA, which is known to have higher sensitivity and specificity, and therefore this test (mRBPT) can be used for more surveillance activities in pastoral areas.
The overall animal-level seroprevalence and incidence rate of Brucella infection found in this study were 11.3% (95% CI; 8.6–14.0) and 0.024 (95% CI; 0.014–0.037) cases per animal-months at risk, respectively. While estimates of Brucella infection incidence in livestock remain largely unknown in many developing countries including Kenya, partly due to weak surveillance systems and under-reporting, the overall seroprevalence of Brucella spp. found in this study was within the ranges previously reported in other pastoral areas (e.g., 7.5 to 40%) in Africa [8]. Both findings confirm that brucellosis is prevalent in the area. Furthermore, animal infections were also clustered within herds/flocks (ICC = 0.10), in agreement with other studies [14, 19]. Infections of animals by Brucella spp. could cause high livestock production losses since this disease is contagious and many animals within a herd could become infected [2]. For example, it is estimated that about 20% of cattle in herds with high exposure levels (> 30%) could abort, while milk yields could reduce by 20–25% among aborting animals [5]. Livestock infections by Brucella spp. also poses a continuous risk for humans [20], but this study could not confirm this linkage because there was no human component. However, earlier studies conducted in resource-limited areas have found livestock infections to be positively correlated with humans’ exposure [13, 21]. Human infections could occur through direct contact with sick animals or their fluids while relieving dystocia, disposing aborted material, but also by eating undercooked meat, raw/contaminated milk, or dairy products [22]. Indeed, a previous study conducted in the area (Isiolo and Marsabit Counties) detected B. abortus and B. melitensis in milk samples from camels, cattle, sheep and goats [23]. In addition, human brucellosis has also been reported in Isiolo county, among veterinarians, laboratory personnel, and individuals with febrile illness [24].
This study found a significantly higher incidence of Brucella infection in camels and cattle compared to sheep and goats. Brucella spp. seroprevalence by livestock species also followed a similar pattern as that of incidence. If seroprevalence results were to be interpreted without the incidence data, it could have been concluded that cattle and camels have higher seroprevalences than sheep and goats because they live longer and hence are more likely to manifest cumulative exposures over time. However, the observed similarities in the patterns of Brucella seroprevalence and incidence suggests that camels and cattle have a higher force of infection which manifests as a significantly higher incidence. These infection patterns could be attributed to relative susceptibility of the livestock species sampled in the area to the prevalent Brucella species reported in our study. Indeed, a recent study implemented in other pastoral areas of Kenya found that cattle and camels were readily infected with B. abortus compared to goats and sheep [11]. This observation could also be connected to the livestock grazing lifestyles used by farmers in the area. For example, it was observed during sampling that cattle and camels were normally raised together in pastoral systems unlike sheep and goats which were grazed within farms. Given these production systems, the effective contact rates between susceptible and Brucella-infected animals were therefore likely to be higher among cattle and camel herds compared with sheep and goats. This is due to sharing of pasture and watering sources between several herds and/or the uncontrolled movement of livestock that are typical of pastoral production systems [12].
Our real-time PCR results showed that B. abortus which primarily infects cattle and camels was more prevalent in the area compared to B. melitensis which naturally infects sheep and goats. A total of 11 (18.3%) samples that were positive by serological tests did not amplify with genus-specific primers for Brucella species, and also with species-specific primers for both B. abortus and B. melitensis. This finding could be due to low yields of Brucella DNA in serum samples [25]. The World Organisation for Animal Health (OIE) recommends the use of sequential ELISA tests, as employed in this study, to confirm exposure of animals to Brucella. PCR test, though conclusive compared to ELISA, may also not be sensitive enough to pick some of the infections that could become sequestered in tissues [26]. Nonetheless, the detection of B. abortus DNA in sheep and goats indicated cross-species transmission (spill-over) from cattle or camels to these hosts which is commonly reported in mixed livestock production systems [11, 27, 28]. However, more studies need to be conducted to determine the relative transmission rates of the various Brucella spp. between multiple host species. The higher prevalence of B. abortus in livestock in the area compared to other Brucella spp. also suggest the presence of underlying biological or ecological mechanisms that influences Brucella infection. For example, B. abortus pathogen has also been shown to survive in the environment (soil, vegetation) for a long period of time (e.g., 21–81 days) depending on soil moisture, temperature and sunlight [29]. The environmental persistence of B. abortus could also indirectly increase the transmission levels of this pathogen if contaminated pastures or watering sources are shared between animals. This would be one entry point for studies on Brucella ecology to be conducted to generate more knowledge on how environment influences Brucella epidemiology.
The results from the cross-sectional survey showed that Brucella spp. seropositivity was significantly associated with animals’ age and sex; adults and female animals had higher levels of exposure compared to young animals and males, respectively. Older animals probably had longer exposure time compared with young ones which could have increased their chances of infection, while females could have had repeated exposure to Brucella spp. as they are more likely to stay longer in a herd than males since they are kept for breeding purposes [13]. Besides, the results obtained from multivariable Cox regression analysis did not show significant associations between Brucella spp. exposure and animals’ sex or age given that the procedure used to estimate incidence through the calibration of animal-time at risk provided a reliable comparison of the risk of exposure between animals.
The main limitation of this study is that the seroprevalences of Brucella spp. in sheep, goats, and camels were estimated using smaller sample sizes than required. This could have led to a low statistical power [30]. Also, a fairly large number of animals were sampled per herd in a few herds which could also have lowered the precision of seroprevalence estimate. Furthermore, the use of a parallel approach to determine the seropositivity of Brucella spp. in animals, and to select seronegative animals for follow-up in the longitudinal phase of the study, could also have led to the overestimation of seroprevalence due to low specificity.
Conclusion
The Brucella infection incidence and seroprevalance estimates obtained in this study demonstrated that brucellosis is prevalent in the area. Brucella infection incidence was significantly higher among camels and cattle compared to sheep and goats. Brucella abortus was more prevalent than B. melitensis. Given that livestock infections by Brucella spp. poses a public health risk for the livestock keepers in Isiolo County, further One Health surveillance studies are required to determine exposure and incidence of this pathogen in humans and to inform control interventions. Vaccination of livestock which is rarely implemented in the area is also recommended because infected livestock animals are the key sources of infections in humans.
Methods
Study area
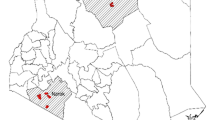
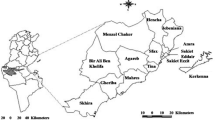
This study was conducted in Kinna ward in Isiolo County, northern Kenya (Fig. 1). The area was selected purposively due to good accessibility and reliable security. In addition, a previous survey that involved the screening of milk for Brucella spp. using milk ring test and real time PCR indicated that Kinna had a higher prevalence of Brucella spp. compared to other areas that were surveyed in Isiolo and the neighbouring Marsabit counties [23]. Pastoral livestock production system is the main cultural and economic activity for the local people because the area is semi-arid [31]. The average annual rainfall is 580 mm [31], and ranges between 350 and 600 mm [32]. Rainfall in the area has a bimodal distribution; long rains occur from March to May while the short rains occur in November to December [32]. The mean annual temperature in the county range between 24 °C and 30 °C [33].
Study design and sampling procedure
This study used both cross sectional and longitudinal study designs. The cross-sectional survey was done in December 2017 as a preliminary step to select animals for the longitudinal phase of the study which was conducted between December 2017 and December 2018. The first longitudinal follow-up of negative animals was initiated immediately after the cross-sectional study. The sample size required for the cross-sectional survey was estimated using the formula; n = (1.96)2p(1-p)/d2 [34]. Based on previous seroprevalence surveys, the expected seroprevalences (p) of Brucella spp. in camels, cattle, sheep and goats were 10.3, 16.9, 16.1 and 11.9% [3], while the precision (d) of the test was set at 0.05. The initial sample sizes estimated for each livestock species were adjusted for design effects (DE) to account for the within-household clustering of observations. We derived the DE using the formula DE = 1 + ρ(m-1)), where ρ is the estimated intra-cluster correlation coefficient (ICC) at the household level and assumed that three animals (m) would be sampled in each cluster (household) [35]. An ICC of 0.1 was used to estimate the design effect; this value was adopted from previous similar studies [36, 37]. In general, the ICC values for infectious diseases range from 0.05 to 0.2, except for highly infectious pathogens that could exceed 0.5 [38]. A sample size of 871 animals, including 170 camels, 259 cattle, 249 sheep and 193 goats, was required after adjustment of the initial sample sizes for design effect. However, given that this study used archived sera from a previous study, all the 841 sera samples comprising 382 cattle, 100 camels, 185 sheep and 174 goats that had been collected from a few households (herds) in that study, were all included. Households with at least cattle, sheep and goats were included in the sampling frame since these are the common livestock species found in the area. All the animals sampled during the cross-sectional survey were ear-tagged for easy identification.
For the longitudinal study, animals from each species that were seronegative for Brucella spp. by all the three serological tests, from the samples used in the cross-sectional survey, were randomly selected. In total, these included 31 cattle, 22 sheep, 32 goats and 30 camels. These animals were sampled at monthly intervals for a period of 1 year.
Sample collection
In both cross-sectional and longitudinal studies, about 10 ml venous blood samples were collected from all the animals recruited in plain vacutainers through jugular venepuncture. For the longitudinal study, sampling was done at monthly intervals. In each event, data on animals’ sex, age (young, weaner or adult), pregnancy status (yes/no) and species of the animals kept in the source herd were also obtained using a questionnaire. Serum was extracted from the blood samples after centrifugation at 5000 rpm for 10 min. The samples were transported at − 20 °C to the International Livestock Research Institute (ILRI), Nairobi for laboratory analysis.
Serological testing
Serum samples collected in the cross-sectional survey were tested for antibodies against Brucella spp. using three serological tests – conventional RBPT, modified RBPT and indirect ELISA (iELISA). Those collected in the longitudinal study were tested using iELISA only.
The conventional RBPT followed the procedure described by Nielsen [39]. In brief, 25 μl of the serum sample and an equal volume of the Rose Bengal reagent (antigen) (IDvet Innovative Diagnostics, France) were dispensed onto a white tile next to each other using micropipettes and sterile disposable tips. A sterile applicator stick was then used to mix the test serum sample and the reagent, followed by gentle agitation of the tile for 4 min. Samples showing any visible agglutination to the antigen within the 4 min were classified as positive while those with no agglutination were classified as negative. Serum samples were also re-tested using a modified RBPT (mRBPT) [40]. The testing procedure used for the mRBPT was the same as that of the conventional RBPT described above, except that 75 μl of the serum sample was mixed with 25 μl of the Rose Bengal reagent in each test.
The iELISA technique tested samples for anti-Brucella spp. antibodies (IgG1); this used multispecies IDvet kit (IDvet Innovative Diagnostics, France) which could detect infections with either B. abortus, B. melitensis or B. suis. In brief, we analysed the test and reference sera (positive and negative controls) in duplicates for each test plate and measured the optical densities (ODs) of all the wells at 450 nm. The ratio of the OD of test serum (S) to that of positive control (P) expressed as a percentage was calculated using the formula below:
In the case of the cross-sectional survey, we classified animals as negative if \({S}\left/{P}\right.\) was ≤110%, inconclusive (borderline) if between 110 and 120% and positive if ≥120% as recommended by the manufacturer. We re-tested animals with inconclusive iELISA results; those that returned borderline results after re-testing were included as negatives in the analysis. For the longitudinal survey, a new infection (seroconversion) among recruited animals was determined by a positive iELISA test following a negative iELISA result in the previous sampling period.
The diagnostic sensitivity and specificity estimates of these tests were, 87%/97.8%, 98.9%/100%, 92%/100% and 97.8%/100% in camels, cattle, sheep and goats, respectively, for conventional RBPT and 97.2%/99.8%, 96.6%/100%, 100%/100% and 100%/100% in camels, cattle, sheep and goats, respectively, for iELISA [18, 41]. For mRBPT, there is very limited data on the diagnostic sensitivity and specificity of this test among livestock species.
Molecular detection of Brucella DNA using real-time PCR
Samples that tested positive by any of the above three serological tests were further subjected to real-time PCR-based assays to detect genus Brucella DNA and for species identification. Genomic DNA was extracted from these samples using the QIAamp blood DNA extraction kit (Qiagen, USA), following the manufacturer’s instructions. Briefly, 200 μl of each serum sample was mixed with 20 μl proteinase K and 200 μl of lysis buffer. The lysate was then taken through the stages of digestion, deactivation, and elusion, according to the manufacturer’s guidelines. The quality and quantity of the extracted DNA was first determined using Nano-Drop spectrophotometer (ThermoFisher Scientific, USA) before DNA samples were stored at − 20 °C until they could be tested.
We performed real-time PCR on all the extracted DNA samples using an ABI 7500 thermocycler machine (Applied Biosystems, Life Technologies, Singapore). The sequences of the oligonucleotide primers and probes used in this study are presented in Table 6. The DNA samples were first amplified using genus-specific primers that targeted the bcsp31 gene to detect Brucella DNA at the genus-level. All the DNA samples that tested positive for genus Brucella were further amplified using species-specific primers that targeted the alkB and BMEI1162 genes for B. abortus, and B. melitensis, respectively [42] (Table 6). The PCR reactions for both the genus and species-specific assays were performed in duplicate, using 20 μl reaction volume containing; 10 μl of 2X PerfeCTa qPCR masterMix, 0.5 μl of each of the pairs of primers (10 nM), 0.25 μl of each of the three probes (10 nM), 2.25 μl of nuclease free water and 4 μl of the (extracted) DNA template. The amplification conditions were as follows; one cycle at 95 °C for 10 min as initial denaturation followed by 40 cycles at 95 °C for 15 s for denaturation, and 1 min for both annealing and extension at 60 °C. Reference strains of B. abortus 544 and B. melitensis 16 M (from Friedrich-Loeffler-Institute, Germany) were included in all the PCR runs as positive controls, alongside the non-template negative controls. Samples that showed clear amplification plot, accompanied with a cycle threshold (ct) value of < 39 were considered as positive.
Statistical analysis
Data entered into Microsoft Excel 2016 was first cleaned before being imported into R statistical software, version 3.6.0 [43] for analysis. All descriptive analyses including the calculation of apparent seroprevalence of Brucella spp. was done using the CrossTable function in gmodels package [44], while the 95% confidence intervals (CI) of the respective estimates were adjusted for within-household clustering using the epi.conf function in epiR package [45]. The outcome variable (apparent animal-level seroprevalence of Brucella spp.) used in our analysis was based on paralleled interpreted results of both conventional RBPT and iELISA. Animals (camels, cattle, sheep and goats) were classified as positive if they had anti-Brucella spp. antibodies by either RBPT, or iELISA tests, and negative if no anti-Brucella spp. antibodies were detected by both tests. We calculated the true seroprevalance (TP) of Brucella spp. for each livestock species from the apparent animal-level seroprevalence (AP) using the formula below;
where AP is the apparent seroprevalence; Se1 and Sp1 denoted sensitivity and specificity estimates of the conventional RBPT, respectively, while Se2 and Sp2 were the respective sensitivity and specificity estimates of the iELISA test for each livestock species [46]. The level of agreement between the three serological tests was estimated using Cohen’s Kappa statistic. We also used the Cochran’s Q test to compare the proportion of Brucella spp. seropositive animals detected by the three tests followed by post-hoc pairwise comparisons of the tests using McNemar’s χ2. Fisher’s exact test was also used to determine the association between categorical predictors (animal’s sex, age, pregnancy status, and species) and the outcome variable. The aggregated data from all the animals was also further subset by the livestock species and the above categorical predictors assessed for their association with the outcome variable.
For the cross sectional data, risk factor analysis was done at the animal-level. We did not perform analysis at herd-level. The categorical variables listed above were first tested for their independent association with Brucella spp. seropositivity using univariable mixed-effects logistic regression models. Variables with p-value ≤0.15 in the univariable models [34], were further analysed using a multivariable mixed-effects logistic regression model. Data were fitted in both univariable and multivariable models using the glmer function in the lme4 package [47], with the household ID (representing herds/flocks) being entered as a random variable (random effect) to account for the within-herd/flock clustering of observations. The final multivariable model selected comprised only significant covariates (p ≤ 0.05) and was used to estimate the intra-cluster correlation coefficient (ICC) due to the clustering of animals within herds/flocks. The variance components of this model were extracted using the re_var function in the lme4 package [47], and the ICC estimated through bootstrap simulation.
For the analysis of the longitudinal data, we first removed cases that were classified as being positive during the cross-sectional study to remain with uninfected animals. Animals were followed on monthly basis until they got exposed. If seroconversion took place between two sampling dates, we used the second sampling date as the time at which the animal was exposed to Brucella spp. Animal-time (in months) at risk for each animal was obtained and aggregated to obtain an exact denominator for the overall Brucella infection incidence. The numerator was the total number of new Brucella spp. cases recorded during the follow-up period. The estimation of the incidence rates with their respective 95% CI (overall incidence as well as by livestock species, sex and age) were performed using the epi.conf function in epiR package [45].
This study also determined the hazard rate ratio for the above categorical variables using univariable and multivariable Cox proportional hazards models. In these analyses, Brucella spp. exposure in animals and time at which the exposure was detected, were both included in the Cox regression models as the outcome of interest. We fitted data to these models using the coxph function in the survival package [48] and accounted for within-herd/flock clustering of animals using the household ID as a random effect. The proportional hazard assumption was evaluated statistically for each covariate and globally using the cox.zph function in survival package [48].
Availability of data and materials
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Change history
20 May 2022
A Correction to this paper has been published: https://doi.org/10.1186/s12917-022-03228-1
Abbreviations
- RBPT:
-
Rose Bengal Plate Test
- mRBPT:
-
Modified Rose Bengal Plate Test
- iELISA:
-
Indirect Enzyme Linked Immunosorbent Assay
- ICC:
-
Intra-cluster correlation coefficient
- ILRI:
-
International Livestock Research Institute
- PCR:
-
Polymerase Chain Reaction
References
Olsen S, Palmer M. Advancement of knowledge of Brucella over the past 50 years. Vet Pathol. 2014;51(6):1076–89.
Franc K, Krecek R, Häsler B, Arenas-Gamboa A. Brucellosis remains a neglected disease in the developing world: a call for interdisciplinary action. BMC Public Health. 2018;18(1):125.
Njeru J, Wareth G, Melzer F, Henning K, Pletz M, Heller R, et al. Systematic review of brucellosis in Kenya: disease frequency in humans and animals and risk factors for human infection. BMC Public Health. 2016;16(1):853.
Ducrotoy M, Bertu W, Matope G, Cadmus S, Conde-Álvarez R, Gusi A, et al. Brucellosis in sub-Saharan Africa: current challenges for management, diagnosis and control. Acta Trop. 2017;165:179-193. Available from: https://doi.org/10.1016/j.actatropica.2015.10.023.
McDermott J, Grace D, Zinsstag J. Economics of brucellosis impact and control in low-income countries. Rev Sci Tech. 2013;32(1):249–61. Available from:. https://doi.org/10.20506/rst.32.1.2197.
Franco MP, Mulder M, Gilman RH, Smits HL. Human brucellosis. Lancet Infect Dis. 2007;7(12):775–86.
Godfroid J, Scholz H, Barbier T, Nicolas C, Wattiau P, Fretin D, et al. Brucellosis at the animal/ecosystem/human interface at the beginning of the 21st century. Prev Vet Med. 2011;102(2):118–31.
McDermott JJ, Arimi S. Brucellosis in sub-Saharan Africa: epidemiology, control and impact. Vet Microbiol. 2002;90(1-4):111-34. Available from: https://doi.org/10.1016/s0378-1135(02)00249-3.
Aparicio ED. Epidemiology of brucellosis in domestic animals caused by Brucella melitensis, Brucella suis and Brucella abortus. Rev Sci Tech. 2013;32(1):53–60.
Akoko J, Pelle R, Kivali V, Schelling E, Shirima G, Machuka EM, et al. Serological and molecular evidence of Brucella species in the rapidly growing pig sector in Kenya. BMC vet res. 2020;16:1-7. Available from: https://doi.org/10.1186/s12917-020-02346-y.
Akoko J, R P, Lukambagire A, Machuka E, Nthiwa D, Mathew C, et al. Molecular epidemiology of Brucella species in mixed livestock-human ecosystems in Kenya. Sci. Rep. 2021. Available from: https://doi.org/10.1038/s41598-021-88327-z.
Racloz V, Schelling E, Chitnis N, Roth F, Zinsstag J. Persistence of brucellosis in pastoral systems. Rev Sci Tech. 2013;32(1):61–70.
Kairu-Wanyoike S, Nyamwaya D, Wainaina M, Lindahl J, Ontiri E, Bukachi S, et al. Positive association between Brucella spp. seroprevalences in livestock and humans from a cross-sectional study in Garissa and Tana River counties, Kenya. PLoS Negl trop dis. 2019;13(10). Available from: https://doi.org/10.1371/journal.pntd.0007506.
Nthiwa D, Alonso S, Odongo D, Kenya E, Bett B. Zoonotic pathogen Seroprevalence in cattle in a wildlife–livestock Interface, Kenya. Ecohealth. 2019;16(4):712-25. Available from: https://doi.org/10.1007/s10393-019-01453-z.
Seleem MN, Boyle SM, Sriranganathan N. Brucellosis: a re-emerging zoonosis. Vet Microbio. 2010;140(3):392–8.
Pappas G, Panagopoulou P, Christou L, Akritidis N. Biological weapons. Cell Mol Life Sci. 2006;63(19–20):2229–36.
Godfroid J, Nielsen K, Saegerman C. Diagnosis of brucellosis in livestock and wildlife. CMJ. 2010;51(4):296-305. Available from: https://doi.org/10.3325/cmj.2010.51.296.
Gusi AM, Bertu WJ, Jesús de Miguel M, Dieste-Pérez L, Smits HL, Ocholi RA, et al. comparative performance of lateral flow immunochromatography, iELISA and rose Bengal tests for the diagnosis of cattle, sheep, goat and swine brucellosis. PLoS Negl trop dis. 2019;13(6):e0007509. Available from: https://doi.org/10.1371/journal.pntd.0007509.
Boukary AR, Saegerman C, Abatih E, Fretin D, Bada RA, De Deken R, et al. Seroprevalence and potential risk factors for Brucella spp. infection in traditional cattle, sheep and goats reared in urban, periurban and rural areas of Niger. PLoS one. 2013;8(12):e83175. Available from: https://doi.org/10.1371/journal.pone.0083175.
Kunda J, Fitzpatrick J, French N, Kazwala R, Kambarage D, Mfinanga GS, et al. Quantifying risk factors for human brucellosis in rural northern Tanzania. PLoS one. 2010;5(4). Available from: https://doi.org/10.1371/journal.pone.0009968.
Osoro EM, Munyua P, Omulo S, Ogola E, Ade F, Mbatha P, et al. Strong association between human and animal Brucella seropositivity in a linked study in Kenya, 2012–2013. Am J Trop Med Hyg. 2015;93(2):224–31.
Dadar M, Shahali Y, Whatmore AM. Human brucellosis caused by raw dairy products: a review on the occurrence, major risk factors and prevention. Int J Food Microbiol. 2019;292(2):39–47.
Wainaina M, Aboge GO, Omwenga I, Ngaywa C, Ngwili N, Kiara H, et al. Detection of Brucella spp. in raw milk from various livestock species raised under pastoral production systems in Isiolo and Marsabit counties, northern Kenya. Trop. Anim. Health prod. 2020;52(6):3537-3544. Available from: https://doi.org/10.1007/s11250-020-02389-1.
Teng Y-H, Teng J-J, Chao S, Chao H, Waghela SD. Comparison of the rose bengal plate and the complement fixation tests with the tube agglutination test for diagnosis of human brucellosis. Open J Clin Diagn. 2017;7(3):73. Available from: https://doi.org/10.4236/ojcd.2017.73008.
Tomaso H, Kattar M, Eickhoff M, Wernery U, Al Dahouk S, Straube E, et al. Comparison of commercial DNA preparation kits for the detection of Brucellae in tissue using quantitative real-time PCR. BMC infect dis. 2010;10(1):100. Available from: https://doi.org/10.1186/1471-2334-10-100.
de Figueiredo P, Ficht TA, Rice-Ficht A, Rossetti CA, Adams LG. Pathogenesis and immunobiology of brucellosis: review of Brucella–host interactions. Am J Pathol. 2015;185(6):1505-17. Available from: https://doi.org/10.1016/j.ajpath.2015.03.003.
Wareth G, Melzer F, Tomaso H, Roesler U, Neubauer H. Detection of Brucella abortus DNA in aborted goats and sheep in Egypt by real-time PCR. BMC res notes. 2015;8(1):212. Available from: https://doi.org/10.1186/s13104-015-1173-1.
Selim A, Attia K, Ramadan E, Hafez YM, Salman A. Seroprevalence and molecular characterization of Brucella species in naturally infected cattle and sheep. Prev vet med. 2019;171:104756. Available from: https://doi.org/10.1016/j.prevetmed.2019.104756.
Aune K, Rhyan JC, Russell R, Roffe TJ, Corso B. Environmental persistence of Brucella abortus in the greater Yellowstone area. J Wildl Manag. 2012;76(2):253-61. Available from: https://doi.org/10.1002/jwmg.274.
McNeish DM, Harring JR. Clustered data with small sample sizes: comparing the performance of model-based and design-based approaches. Commun stat-Simul C. 2017;46(2):855-69. Available from: https://doi.org/10.1080/03610918.2014.983648.
King-Okumu C, Wasonga OV, Jarso I, Salah YMS. Direct use values of climate-dependent ecosystem services in Isiolo County, Kenya. International Institute for Environment and Development (IIED): London, UK. 2016. Available from: https://pubs.iied.org/pdfs/10142IIED.pdf?
Lamuka PO, Njeruh FM, Gitao GC, Abey KA. Camel health management and pastoralists’ knowledge and information on zoonoses and food safety risks in Isiolo County, Kenya. Pastoralism. 2017;7(1):20. Available from: https://doi.org/10.1186/s13570-017-0095-z.
Noor IM, Bebe BO, Guliye AY. Analysis of an emerging peri-urban camel production in Isiolo County, northern Kenya. J Camelid Sci. 2012;5(1):41–61 Available from: http://www.isocard.net/images/journal/FILEf22f5ef375fdb37.pdf.
Dohoo IR, Martin SW, Stryhn H. Methods in epidemiologic research. Charlottetown: Prince Edward Island, Canada, VER Inc.; 2012.
Fosgate GT. Practical sample size calculations for surveillance and diagnostic investigations. Journal of veterinary diagnostic investigation. 2009;21(1):3-14. PMID: J Vet Diagn Invest. 2009;21:3–14.
Kanouté YB, Gragnon BG, Schindler C, Bonfoh B, Schelling E. Epidemiology of brucellosis, Q fever and Rift Valley fever at the human and livestock interface in northern Côte d’Ivoire. Acta Trop. 2017;165:66–75 https://doi.org/10.1016/j.actatropica.2016.1002.1012.
Segura-Correa J, Domínguez-Díaz D, Avalos-Ramírez R, Argaez-Sosa J. Intraherd correlation coefficients and design effects for bovine viral diarrhoea, infectious bovine rhinotracheitis, leptospirosis and neosporosis in cow–calf system herds in North-eastern Mexico. Prev. Vet. Med. 2010;96(3–4):272–275.Available from: https://doi.org/10.1016/j.prevetmed.2010.07.006.
Otte M, Gumm I. Intra-cluster correlation coefficients of 20 infections calculated from the results of cluster-sample surveys. Prev Vet Med. 1997;31(1–2):147–50.
Nielsen K. Diagnosis of brucellosis by serology. Vet Microbiol. 2002;90(1–4):447–59 https://doi.org/10.1016/S0378-1135(02)00229-8.
Ferreira A, Cardoso R, Dias I, Mariano I, Belo A, Preto I, et al. Evaluation of a modified rose Bengal test and an indirect enzyme-linked Immunosorbent assay for the diagnosis of Brucella melitensis infection in sheep. Vet Res. 2003;34(3):297-305. Available from: https://doi.org/10.1051/vetres:2003005.
Wernery U. Camelid brucellosis: a review. Revue scientifique et technique (International Office of Epizootics) 2014;33(3):839–857.
Probert WS, Schrader KN, Khuong NY, Bystrom SL, Graves MH. Real-time multiplex PCR assay for detection of Brucella spp., B. abortus, and B. melitensis. J Clin Microbiol. 2004;42(3):1290–3.
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. 2019. Available from: https://cran.r-project.org/bin/windows/base/old/3.6.0/
Warnes GR, Bolker B, Lumley T, Johnson RC. gmodels, various R programming tools for model fitting. R package version 2.15. 0. 2009.
Stevenson M, Stevenson MM, BiasedUrn I. Package ‘epiR’; 2019.
Noordhuizen JPTM, Frankena K, Thrusfield MV, Graat E. Application of quantitative methods in veterinary epidemiology: Wageningen Pers; 2001.
Bates D, Mächler M, Bolker B, Walker S. Fitting linear mixed-effects models using lme4. arXiv preprint arXiv:14065823. 2014.
Therneau TM, Lumley T. Package ‘survival’. Survival analysis Published on CRAN. 2014;2.
Acknowledgements
We appreciate Mr. Fredrick Otieno (ILRI) for creating the map used in this study. The serum samples used in this study were obtained from a previous project titled ‘Developing Optimum Vaccination Strategies for Rift Valley Fever (RVF) in East Africa’ led by Dr. Bernard Bett, ILRI. The positive controls used in this study were provided by Prof. Heinrich Neubauer and Dr. Falk Melzer from Friedrich-Loeffler-Institut, Germany. We also acknowledge support from the project: Co-infection with Rift Valley fever virus, Brucella spp. and Coxiella burnetii in humans and animals in Kenya: Disease burden and ecological factors funded by the Defense Threat Reduction Agency for covering the time used by the co-authors to develop the manuscript.
Funding
The analysis of the samples was implemented under the project: Co-infection with Rift Valley fever virus, Brucella spp. and Coxiella burnetii in humans and animals in Kenya: Disease burden and ecological factors (Co-infection project) funded by the Defense Threat Reduction Agency, contract number HDTRA 11910031. Additional support for staff time was supported by the Agriculture for Nutrition and Health CGIAR program led by IFPRI. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Concept development: JN, DN, JA, HO, BB; Formal analysis: DN, BB; Funding acquisition: BB; Investigation: JN, DN, JA, HO, BB; Methodology: JN, DN, JA, BB; Project administration: BB; Supervision: BB; Writing - original draft: JN, DN; Writing –review & editing: all authors. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The ethical and animal use approvals for this study were respectively provided by the Institutional Review Committee (reference number ILRI-IREC 2017–19) and the animal care and use committee (reference number IACUC-RC2018–03) at the International Livestock Research Institute (ILRI), Nairobi. Furthermore, written informed consent for livestock sampling and the questionnaire survey was also obtained from all the livestock owners.
Consent for publication
Not applicable.
Competing interests
The authors have declared that no competing interests exist.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: the authors pointed out several corrections to be made in the article.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Njeru, J., Nthiwa, D., Akoko, J. et al. Incidence of Brucella infection in various livestock species raised under the pastoral production system in Isiolo County, Kenya. BMC Vet Res 17, 342 (2021). https://doi.org/10.1186/s12917-021-03036-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12917-021-03036-z