Abstract
Background
We report a rare case of Mammalian orthoreovirus (MRV) infection in a child with a primary immunodeficiency (PID). Infections with Mammalian orthoreovirus are very rare and probably of zoonotic origin. Only a few cases have been described so far, including one with similar pathogenesis as in our case.
Case presentation
The patient, age 11, presented with flu-like symptoms and persistent severe diarrhea. Enterovirus has been detected over several months, however, exact typing of a positive cell culture remained inconclusive. Unbiased metagenomic sequencing then detected MRV in stool samples from several time points. The sequencing approach further revealed co-infection with a recombinant Coxsackievirus and Adenovirus. MRV-specific antibodies detected by immunofluorescence proved that the patient seroconverted.
Conclusion
This case highlights the potential of unbiased metagenomic sequencing in supplementing routine diagnostic methods, especially in situations of chronic infection with multiple viruses as seen here in an immunocompromised host. The origin, transmission routes and implications of MRV infection in humans merit further investigation.
Similar content being viewed by others
Background
Individuals suffering from primary immunodeficiencies (PIDs) are prone to a variety of infections, and some types of PIDs can predispose the affected individuals to particular pathogens. Infections that are usually controlled and asymptomatic in immunocompetent individuals often cause chronic, active disease in immunocompromised individuals [1].
Rapid diagnosis of viral infections is crucial in immunocompromised patients. While a range of established molecular tests can detect specific viruses, high-throughput metagenomic sequencing is based on virus-sequence independent amplification of nucleic acids isolated directly from clinical samples; as such, it has the potential to identify any virus in an “open diagnostics” approach [2, 3]. Hence, this approach can detect rare viruses that are not included in routine diagnostic panels and viruses with sequence variations that would otherwise be missed [4]. Here we used metagenomic sequencing to complement routine diagnostic methods in a child with a PID suffering from persistent diarrhea.
Case presentation
We report on a female child living in Switzerland with a combined B- and T-cell immunodeficiency, hypogammaglobulinaemia and autoimmunity (diabetes mellitus) under immunoglobulin replacement therapy. In early February 2014, the patient, aged 11 at that time, presented with flu-like symptoms with cough, headache, fever and severe diarrhea. Although the other symptoms resolved, the diarrhea persisted. In view of low initial calprotectin levels (100–400 μg/g), a fecal marker for intestinal inflammation, the diarrhea was considered as caused by the underlying PID and not by an infectious agent. However, a rise in calprotectin level to 4000 μg/g in March 2015 prompted us to initiate virological investigations. Enterovirus had been detected over several months using a specific RT-PCR (Additional file 1: Table S1) and was thus considered as causative agent although other viruses were not tested for during this time period.
As symptoms continued, additional testing was performed. A stool sample (November 3rd 2015) was positive in Caco-2 cell culture showing a cytopathic effect (CPE) after 9 days of incubation. The culture sample stained positive with a pool of anti-enterovirus monoclonal antibodies, but we did not succeed to further subtype the suspected enterovirus using routine immunostaining methods. In order to identify the virus amplified in the Caco-2 cell culture, we applied unbiased metagenomic sequencing as previously described [4]. For three sampling time points in September and November 2015, we sequenced both the Caco-2 culture supernatant and the original stool suspension. We detected many reads of Mammalian orthoreovirus 3 (MRV-3) in the cell culture supernatant and reads of Coxsackievirus A (CV-A) in the original stool suspension (Table 1). No virus reads were detected in the supernatant of a non-infected Caco-2 cell culture used as a negative control.
To trace the timing of these two virus infections we conducted a retrospective analysis of the available stool samples by both cell culture and metagenomic sequencing (Table 1). In cell cultures, a CPE was visible after about 14 days of culturing. By sequencing the cell culture supernatants, we found numerous reads for Human adenovirus C (HAdV-C) in these earlier time points, but none for MRV-3. By sequencing original stool suspensions, we identified reads corresponding to the previously identified CV-A.
We verified all the metagenomics analyses by specific PCRs designed for the isolates in this study (Table 1). In addition to confirming MRV-3 in the three cell cultures supernatants with MRV-3 reads, we also detected MRV-3 in two corresponding stool suspensions by specific PCR. CV-A was confirmed in all stool suspensions and in one of two colon biopsies; HAdV-C was confirmed in all tested cell cultures and stool suspensions (Table 1).
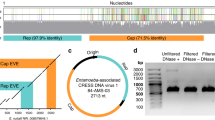
To define the MRV-3 infection in more detail, we combined sequencing reads from all time points and reconstructed a consensus sequence that covered the full coding sequences in all 10 segments of the MRV-3 isolate (GenBank KX932029 - KX932038). Phylogenetic analyses showed that the MRV-3 sequence isolated in this study was most related in all 10 segments to an isolate from a child in Slovenia [5] and further clustered with isolates from bats in Germany [6] and pigs in Italy [7] (Fig. 1a and Additional file 1: Figures S1-S9).
Mammalian orthoreovirus infection confirmed by phylogenetic analysis and immunofluorescence staining. a Phylogenetic analysis of Mammalian orthoreovirus segment S1 isolated in this study (circle, mew716_S1_type_3_human) reveals a close relationship with previously described isolates identified in a child in Slovenia (triangle, KF154730), bats in Germany (JQ412761), and pigs in Italy (KX343206). Phylogenetic trees were constructed in MEGA7 using the Maximum Likelihood method based on the Kimura 2-parameter model. Bootstrap values from 1000 tries are shown. MRV-type and host species are depicted if available. b Anti-MRV-3 immunofluorescence of patient plasma before and after seroconversion on MRV-3-infected and uninfected Caco-2 cells
In order to proof active replication and diagnosis of MRV infection, we performed immunofluorescence for the detection of MRV-specific antibodies of the patient. Patient plasma from time points after MRV detection were positive for anti-MRV IgG antibodies showing that the patient seroconverted (Fig. 1b).
We were able to reconstruct the full-length coxsackievirus genome present in this patient using additional sequence information from sequencing with CV-A22 serotype consensus primers. Genotyping and bootscan analysis revealed that the isolate (GenBank KX932039) probably resulted from recombination between CV-A19 and CV-A22 (Additional file 1: Figures S10 and S11). The detected HAdV-C was type 2, strain human/ARG/A15932/2002/2[P2H2F2] (JX173079.1).
Discussion and conclusions
Using a metagenomic sequencing approach, we detected multiple virus infections in a child with PID and persistant diarrhea. Although routine PCR methods detected Enterovirus in stool samples for a prolonged period of time, an attempt to subtype the virus after cell culturing was not successful. In order to resolve this, we performed metagenomic sequencing of cell culture supernatants and stool suspensions.
The most unexpected finding was the infection with MRV-3, proven by metagenomic sequencing and serology of samples from several time points. MRVs belong to the Reoviridae family, a group of non-enveloped dsRNA viruses with 10 genome segments and have been isolated from a wide range of mammalian hosts and in a variety of clinical contexts [5,6,7]. To date, the diseases associated with MRV infections in humans include respiratory disease [8], meningitis [9,10,11], acute necrotizing encephalopathy [12] and (in a similar case involving a child living in Slovenia) acute gastroenteritis [5]. Zoonotic transmission is often suspected [6, 13, 14]. Our patient lives close to a farm and might have come in contact with pets, farm animals and animal feces making a zoonotic infection conceivable, however, due to the lack of data on MRV distribution and appropriate samples we can only hypothesize on the transmission route [7]. Stool samples from 16 children with suspicion of gastrointestinal infection treated at the same hospital during 2015 all tested negative with qPCR (data not shown).
The virus growing in the initial Caco-2 cell culture was therefore MRV-3 and not an enterovirus as suggested by the serotyping assay. In fact, the manufacturer’s datasheet for the reagent used states that there is potential for cross-reaction with Hepatitis A, Reovirus 3, and some Rhinovirus and Astrovirus strains. The latter highlights the genuine difficulty in accurate detection of highly diverse virus families with numerous genotypes such as Enteroviruses where typing by specific PCR can be challenging due to the high susceptibility to recombination and the emergence of novel strains [15]. Notably, CV-A was detected in stool suspensions but not in cell culture supernatants. While cell culturing was critical for MRV-3 detection, CV-A, in line with a general difficulty to culture coxsackieviruses [16], did not infect Caco-2 cells. The reason that MRV-3 was not detected by metagenomic sequencing in the original stool suspension, but only after amplification in cell culture, is likely because it was at levels too low to be detected with the applied depth of sequencing.
In summary, this case highlights the complexity of infections in immunocompromised hosts and reveals limitations of routine diagnostic methods. A combination of traditional cell culture, metagenomic sequencing, verification by specific PCRs and serology proved key to detect a persistent co-infection with three clinically relevant viruses. While the presence of these viruses in stool specimen suggests a link with the observed pathogenesis, it cannot be defined to what extent each of the viruses contributed to the initial flu-like symptoms and prolonged diarrhea. Based on the timing of virus detection, the earlier detected CV-A and HAdV-C are more likely to be involved in the child’s condition than the later emerging MRV-3.
Abbreviations
- CPE:
-
Cytopathic effect
- CV-A:
-
Coxsackievirus A
- HAdV-C:
-
Human adenovirus C
- MRV:
-
Mammalian orthoreovirus
- PCR:
-
Polymerase chain reaction
- PID:
-
Primary immunodeficiency
References
Dropulic L, Cohen J. Severe viral infections and primary immunodeficiencies. Clin Infect Dis. 2011;53(9):897–909.
Delwart EL. Viral metagenomics. Rev Med Virol. 2007;17(2):115–31.
Mokili JL, Rohwer F, Dutilh BE. Metagenomics and future perspectives in virus discovery. Current Opinion in Virology. 2012;2(1):63–77.
Lewandowska DW, Zagordi O, Zbinden A, Schuurmans MM, Schreiber P, Geissberger FD, Huder JB, Böni J, Benden C, Mueller NJ, et al. Unbiased metagenomic sequencing complements specific routine diagnostic methods and increases chances to detect rare viral strains. Diagn Microbiol Infect Dis. 2015;83(2):133–8.
Steyer A, Gutierrez-Aguire I, Kolenc M, Koren S, Kutnjak D, Pokorn M, Poljsak-Prijatelj M, Racki N, Ravnikar M, Sagadin M, et al. High similarity of novel orthoreovirus detected in a child hospitalized with acute gastroenteritis to mammalian orthoreoviruses found in bats in Europe. J Clin Microbiol. 2013;51(11):3818–25.
Kohl C, Lesnik R, Brinkmann A, Ebinger A, Radonic A, Nitsche A, Muhldorfer K, Wibbelt G, Kurth A. Isolation and characterization of three mammalian orthoreoviruses from European bats. PLoS One. 2012;7(8):e43106.
Lelli D, Beato MS, Cavicchio L, Lavazza A, Chiapponi C, Leopardi S, Baioni L, De Benedictis P, Moreno A. First identification of mammalian orthoreovirus type 3 in diarrheic pigs in Europe. Virol J. 2016;13(1):139.
Chua KB, Crameri G, Hyatt A, Yu M, Tompang MR, Rosli J, McEachern J, Crameri S, Kumarasamy V, Eaton BT, et al. A previously unknown reovirus of bat origin is associated with an acute respiratory disease in humans. Proc Natl Acad Sci U S A. 2007;104(27):11424–9.
Johansson PJ, Sveger T, Ahlfors K, Ekstrand J, Svensson L. Reovirus type 1 associated with meningitis. Scand J Infect Dis. 1996;28(2):117–20.
Hermann L, Embree J, Hazelton P, Wells B, Coombs RTK. Reovirus type 2 isolated from cerebrospinal fluid. Pediatr Infect Dis J. 2004;23(4):373–5.
Tyler KL, Barton ES, Ibach ML, Robinson C, Campbell JA, O’Donnell SM, Valyi-Nagy T, Clarke P, Wetzel JD, Dermody TS. Isolation and molecular characterization of a novel type 3 reovirus from a child with meningitis. J Infect Dis 2004;189(9):1664-1675.
Ouattara LA, Barin F, Barthez MA, Bonnaud B, Roingeard P, Goudeau A, Castelnau P, Vernet G, Paranhos-Baccala G, Komurian-Pradel F. Novel human reovirus isolated from children with acute necrotizing encephalopathy. Emerg Infect Dis. 2011;17(8):1436–44.
Lelli D, Moreno A, Lavazza A, Bresaola M, Canelli E, Boniotti MB, Cordioli P. Identification of Mammalian orthoreovirus type 3 in Italian bats. Zoonoses Public Health. 2013;60(1):84–92.
Chua KB, Voon K, Yu M, Keniscope C, Abdul Rasid K, Wang L-F. Investigation of a potential zoonotic transmission of orthoreovirus associated with acute influenza-like illness in an adult patient. PLoS One. 2011;6(10):e25434.
Lukashev AN. Role of recombination in evolution of enteroviruses. Rev Med Virol. 2005;15(3):157–67.
Wenner HA, Lenahan MF. Propagation of group a Coxsackie viruses in tissue cultures. II. Some interactions between virus and mammalian cells. Yale J Biol Med. 1961;34:421–38.
Acknowledgements
Not applicable
Funding
Funding was provided by the Clinical Research Priority Program “Viral Infectious Diseases” of the University of Zurich. The funding body did not have any role in the design of the study, in the collection, analysis, and interpretation of data and in writing the manuscript.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Author information
Authors and Affiliations
Contributions
SP, CB, TG, JR and JPS cared for the patient and provided clinical data and materials. RC, JB and AZ analyzed and interpreted routine diagnostic results. DWL, FDG and CS performed sequencing experiments and PCR assays. DWL, OZ and MH analyzed the sequencing data. MK and MK performed cell culture and immunostaining. DWL, AT and MH wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Non-invasive samples were obtained from patient mew716 in the frame of the Viral Metagenome Study of the Clinical Research Priority Program ‘Viral Infectious Diseases’ of the University of Zurich. The ethics committee of the canton of Zurich approved the study and written informed consent was obtained.
Consent for publication
Written informed consent for the publication of this case report was obtained from both parents of the child.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional file
Additional file 1:
Portable Document Format. Table S1. Routine screening results for Enterovirus. Figures S1 to S9. Phylogenetic analyses of all Mammalian orthoreovirus segments isolated in this study. Figure S10. Coxsackievirus BLAST genotyping. Figure S11. Coxsackievirus Bootscan Analysis. Supplementary Methods. Supplementary References. (PDF 456 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Lewandowska, D.W., Capaul, R., Prader, S. et al. Persistent mammalian orthoreovirus, coxsackievirus and adenovirus co-infection in a child with a primary immunodeficiency detected by metagenomic sequencing: a case report. BMC Infect Dis 18, 33 (2018). https://doi.org/10.1186/s12879-018-2946-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12879-018-2946-7