Abstract
Background
Irritable bowel syndrome (IBS) is a common functional gastrointestinal disorder characterized by abdominal pain, discomfort, and changes in bowel habits. The mechanism underlying IBS remains unclear, and little evidence exists for clarifying the causal relationship between blood metabolites and IBS.
Methods
We conducted a Mendelian randomization (MR) study using two samples. Exposure data for 7824 Europeans were extracted from a genome-wide association study (GWAS) on metabolite levels. The IBS GWAS data from the GWAS database were used for the initial analysis. The primary analysis of causal relationships was conducted using inverse-variance weighting (IVW) with MR-Egger and weighted medians as supplementary analyses. Sensitivity analyses were performed using a combination of the Cochran’s Q test, MR-Egger intercept test, Mendelian randomization pleiotropy residual sum and outlier, and leave-one-out analysis. For significant associations, replication and meta-analyses were performed using additional independent IBS case GWAS data released by the FinnGen Consortium R9. To identify the metabolites, score regression, confounding analysis, and reverse MR were performed to further assess the causal relationships between the metabolites.
Results
After rigorous screening, we identified four known metabolites to be associated with IBS (stearate, odds ratio [OR]: 0.74, 95% confidence interval [CI]: 0.59–0.92; arginine, OR: 1.36, 95% CI: 1.07–1.74; 1-palmitoylglycerol, OR:1.49, 95% CI: 1.07–2.07; 1-palmitoylglycerophosphoinositol, OR: 0.84, 95% CI: 0.71–0.99).
Conclusions
MR analysis revealed a causal relationship between the four metabolites and IBS, providing preliminary evidence for the pathogenesis of IBS. Our results provide novel insights into the potential biomarkers of IBS.
Similar content being viewed by others
Background
Irritable bowel syndrome (IBS) is a common functional gastrointestinal illness characterized by abdominal pain, discomfort, and altered bowel patterns [1]. IBS is a symptom cluster brought on by a number of illnesses rather than a single condition. IBS development is influenced by altered metabolites, gut microbiota, intestinal immune function, motility, visceral sensation, brain-gut connections, and psychosocial state [2]. Changes in the gut microbiota are believed to play a role in the pathophysiology of IBS; however, the exact cause is unknown, and the illness is frequently accompanied by mental disorders such as anxiety and depression [3]. Various metabolites play pivotal roles in the pathophysiology of IBS [4]. Metabolites mediate numerous biological functions such as signal transmission and immune system control. Individuals with IBS have lower blood levels of short-chain fatty acids [5]. A more comprehensive and systematic investigation of the interactions between blood metabolites and IBS will enhance our understanding of the disease.
Recently, modern omics technologies, including metabolomics, have actively contributed to exploring the mechanisms underlying diseases [6]. Metabolomics involves the qualitative and quantitative assessment of metabolites (small molecules < 1.5 kDa) in bodily fluids. Owing to the intrinsic sensitivity of metabolomics, detecting subtle alterations in biological pathways is possible, which enhances the understanding of the mechanisms underlying various physiological states and abnormal processes, including diseases [7]. Over the past decade, metabolomics has been increasingly used to identify disease biomarkers, and it is regarded as an extremely powerful tool with immense potential for clinical translation [8]. The metabolome and associated pathways have enhanced our understanding of the pathophysiology and mechanisms underlying various diseases. However, studies on the risk relationship between blood metabolites and IBS are limited.
Mendelian randomization (MR), a causal inference method, has been extensively applied in genetic epidemiology [9]. Traditional observational studies are unable to eliminate the effects of environmental confounders. A definitive causal relationship cannot be established even when a strong statistical association is observed between exposure and outcome [10]. Compared with traditional observational studies, MR, employing genetic variants as instrumental variables (IVs), represents a broadly endorsed approach for mitigating potential confounders [11]. This method effectively circumvents the interference of reverse causation and confounders, thereby enabling a more accurate inference of the causal relationship between the exposure and outcome [12].
Stringent randomized controlled trials are highly recommended to determine causal associations; however, their implementation is often hindered by ethical considerations, temporal constraints, and spatial limitations. In the present study, MR was used to investigate the relationship between 486 blood metabolites and IBS. The primary objective of this study was to clarify the causal connections between blood metabolites and IBS and reveal the underlying mechanisms contributing to its onset.
Methods
Study design
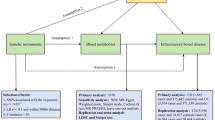
For MR analysis, three assumptions need to be met. These include the association assumption: a robust strong correlation exists between genetic variation (Z) and exposure factor (X); the independence assumption: genetic variation (Z) is independent of the confounding factors (U) influencing the “exposure factor (X) - outcome (Y)” relationship; the exclusion restriction assumption: genetic variation can only influence the outcome through the exposure factors, and cannot affect the outcome via other pathways [13]. This study utilized two sample MR and Mendelian randomization pleiotropy residual sum and outlier (MR-PRESSO) software packages, both implemented in the R software (4.2.3) (Fig. 1).
The study design of the correlation between blood metabolites and irritable bowel syndrome. Abbreviations: Assumption 1:the exposures of interest are significantly related to genetic instruments; Assumption 2: genetic tools are independent of confounding variables; Assumption 3: genetic tools are not linked to outcomes and only impact them through exposures
Data source
Genetic information on blood metabolites was obtained from the Metabolomics Genome-Wide Association Study (GWAS) server (https://metabolomics.helmholtz-muenchen.de/gwas/). Currently, this is the most thorough documentation available for the genetic loci of blood metabolites, made possible by a genome-wide association scan and advanced metabolic analysis performed by Shin et al. who identified ~ 2.1 million SNPs linked to 486 metabolites associated with human genetic variation [14]. The study included 7824 European citizens, including 6056 participants from the UK Twin Study and 1768 from the German KORA F4 Study. Of the 486 metabolites analyzed, 107 were categorized as unidentified because their chemical properties were ambiguous. Furthermore, 309 metabolites were chemically verified and were distributed across eight major metabolic categories: amino acids, carbohydrates, cofactors, vitamins, energy, lipids, nucleotides, peptides, and exogenous metabolism [15].
The GWAS summary data for IBS was sourced from the GWAS Catalog (https://www.ebi.ac.uk/gwas/) with accession number GCST90016564. This data originates from a large-scale IBS meta-analysis by Eijsbouts and colleagues published in Nature Genetics [16]. The sample includes 40,548 participants of European descent from the UK Biobank, meeting IBS diagnostic criteria based on DHQ Rome III symptom data, self-reported past IBS medical diagnosis, or electronic medical records [16].
Instrumental variable (IV) selection
Given the limited number of SNPs identified through stringent filtering, we relaxed the significance threshold to P < 1 × 10^-5, without altering other selection criteria. Subsequently, we designated linkage disequilibrium (LD) r2 < 0.01 within a 500 KB range, a standard commonly adopted in similar studies [16, 17]. In reverse causation, considering that the GWAS data for IBS possess a sufficient number of significant SNPs, to ensure that the instrumental variables in the model have adequate strength and thereby avoid potential weak instrument bias and violations of the exclusion restriction, we have chosen a genome-wide significance threshold (P < 5 × 10^− 8) as the criterion for including SNPs, and LD parameters set to r^2 = 0.001 and kb = 10,000 as conditions for removing LD. A rule of thumb is that the strength of the IVs, as measured by the F-statistic, should be at least 10. When the F-statistic is below 10, the estimates of causal effects can be severely biased [18]. However, a strict threshold may lead to an underestimation of genetic effects by excluding relevant IVs. Using an F-statistic threshold of F = 10 provides a balance, retaining robust instruments while minimizing the risk of weak instrument bias. Therefore, we calculated the R2 and F values for each SNP and excluded SNPs with an F value less than 10. Palindromic SNPs with intermediate-effect allele frequencies were excluded. Finally, the preserved SNPs were used for MR analysis.
Statistical power calculation
The statistical power was calculated using an online tool (https://shiny.cnsgenomic.org/mRnd/) [19, 20]. This tool calculates power using the asymptotic theory to detect causal effects inferred from IVs. The Type I error rate was set at 0.05, and power was computed using the R2 of IV, the proportion of cases with outcomes, and odds ratios (OR) obtained through IVW analysis.
Statistical analysis and sensitivity analysis
Following the above screening criteria, 486 blood metabolites were considered exposure variables. IVW random effects were used to analyze the causal relationship between blood metabolites and IBS. Assuming no horizontal pleiotropy for all SNPs, IVW estimates were derived from a summary study of Wald ratios for all genetic variations. Under this premise, IVW provides the most accurate assessment of causal effects [21]. We used the weighted median (WM) and MR-Egger methods for additional analyses to strengthen the dependability of the results and ensure their accuracy [22]. Four analytical methods were used for the sensitivity analysis: MR-PRESSO, MR-Egger intercept, Cochran-Q test, and leave-one-out analysis (LOO). MR-PRESSO is a recently developed MR method designed to detect and correct abnormal horizontal pleiotropy values, thereby providing accurate estimates [23]. The MR-Egger intercept was calculated to assess the horizontal pleiotropy better and reduce the risk of bias from faulty instrumental variables (IVs) and horizontal pleiotropy [24]. Cochran’s Q test was used to assess heterogeneity among SNPs [25]. LOO was performed to evaluate the influence of each variable on causal estimates by sequentially removing one SNP from the analysis to ascertain if a single SNP predominantly influenced the findings [26].
Employing the False Discovery Rate (FDR) method, the p-values of the IVW analysis results for 486 metabolites were corrected. Metabolites with an FDR less than 0.05 were considered to have a relatively convincing causal relationship [27]. In contrast, metabolites with an FDR greater than 0.05 but a p-value less than 0.05 were deemed to have a nominally significant causal relationship and were identified as potential risk factors.
Replication and meta-analysis
Despite setting strict screening criteria and conducting a sensitivity analysis to ensure data reliability, we repeated the IVW analysis in another IBS cohort using GWAS data of IBS cases released by the FinnGen consortium R9 (https://r9.finngen.fi/). Specifically, these data were obtained from a meta-analysis of GWAS conducted on 9323 Finnish patients with IBS and 301,931 controls. For this investigation, we utilized GWAS data with the accession number GCST012879, and for the replication, we used GWAS data from the FinnGen collaboration. Through a meta-analysis of the two MR analyses, we ultimately determined the blood metabolites that were responsible for the causative effects on IBS. The Review Manager (version 5.3) random-effects IVW model was used to perform the meta-analysis. This experiment was performed according to the protocol described by Cai et al. [28].
Analysis of genetic correlation
Genetic correlation is a key population parameter that describes the shared genetic architecture of complex traits and diseases [29]. Due to the genetic correlation between traits, MR estimates might violate causal effects [30]. To ensure that the causal effects are not confounded by the genetics of exposure and outcome, we used linkage disequilibrium score regression (LDSC) to examine the genetic correlation between selected metabolites and IBS.
Analysis of confounding and reverse causation
Despite conducting an array of sensitivity analyses to assess the horizontal pleiotropy of the MR results and identify any SNPs that contravened the MR assumptions, some residual confounding SNPs may persist. We examined the metabolites in the IVs using the Phenoscanner V2 website (http://www.phenoscanner.medschl.cam.ac) to evaluate the association of each SNP with known IBS risk factors, including psychological factors, such as anxiety [31]. To explore whether IBS has any causal effect on the identified important blood metabolites, we used IBS-related SNPs as IVs for reverse MR analysis (i.e., IBS as the exposure and identified blood metabolites as the outcome for MR analysis) [10].
Results
Utilizing the IVW method as the primary evaluative approach, we identified 34 metabolites exhibiting a nominally significant causal association with IBS (P< 0.05). This cohort comprises 14 metabolites whose identities are currently unknown, in addition to 20 known metabolites. However, it is regrettable to note that no metabolites were found to retain significant association following correction for the FDR (P< 0.05). These mainly include lipids, such as 1-palmitoyl glycerophosphocholine; amino acids, such as serotonin; polypeptides; and sugars. Upon further analysis, and following the integration of complementary and sensitivity analyses, we identified seven metabolites that conformed to stringent selection criteria, thereby establishing them as candidate substances. Overall, the direction and scale of the estimates for these seven metabolites remained consistent across the three analytical methods (IVW, MR-Egger, and WM), and the IVW estimates were significant (P < 0.05). Additionally, the MR-PRESSO (P > 0.05), Cochran’s Q test (P > 0.05), and MR-Egger intercept test (P > 0.05) indicated a lack of heterogeneity and pleiotropy. The LOO analysis outcomes reinforced the idea that the MR estimation was not biased by a single SNP. Therefore, these seven blood metabolites were selected for subsequent analysis (Figs. 2 and 3, Table 1).
Replication and meta-analysis
We used GWAS data for IBS from another cohort downloaded from the FinnGen Alliance R9 and repeated the above MR analysis. We used GWAS data for IBS from another cohort downloaded from FinnGen Alliance R9 and replicated the above-mentioned MR analyses. We found that the replication results for Methionine, 1-palmitoylglycerophosphocholine, and Valine showed considerable differences, while Stearate, Arginine, 1-palmitoylglycerol, and 1-palmitoylglycerophosphoinositol yielded similar results, although they did not reach statistical significance. Meta-analysis was performed using random effects in Review Manager software (version 5.3, [32]. Our results showed similar outcomes. A previous meta-analysis has identified four metabolites associated with IBS. Methionine, 1-palmitoylglycerophosphocholine, and valine were excluded because of insignificant results (Figs. 4 and 5).
Analysis of genetic correlation
According to the results of the genetic correlation analysis, evidence of a genetic correlation between stearate and others in IBS is weak. This indicates that the shared genetic components did not confound the MR results (Table 2).
Analysis of confounding and reverse causation
For each SNP of the identified 20 metabolites, we searched for phenotypes associated with exposure and the resulting variables. In the IVs of arginine and N-acetyl aspartic acid, we found that rs837763 and rs11030392 seemed to be related to mental state, but this did not affect our results. After excluding these confounding IVs, the results of arginine were still significant (odds ratio [OR]:1.41, 95% confidence interval [CI]:1.10–1.80), and the final meta-analysis results were significant (OR:1.36, 95% CI:1.08–1.70). In the reverse MR analysis, none of the 20 known metabolites showed a reverse causal relationship with IBS. Only the unknown metabolites X-11438, X-11521, X-14374, and X-13741 may have a reverse causal relationship with IBS (Fig. 6).
Discussion
IBS has a complex pathophysiology affected by a wide range of variables, including changes in the gut-brain axis, visceral hypersensitivity, abnormalities in intestinal secretion, and intestinal permeability [33]. IBS, as a common gastrointestinal functional disorder, presents significant challenges due to the lack of objective diagnostic methods. Currently, it remains a diagnosis based on symptoms, and there is a notable absence of sensitive biological markers. At present, the international common is the Rome IV diagnostic criteria: Recurrent abdominal pain or discomfort occurring at least 3 days per week over the last 3 months, with symptom onset at least 6 months prior to diagnosis, and associated with two or more of the following conditions: (1) Related to defecation, (2) Associated with a change in the frequency of stool, (3) Associated with a change in the form (appearance) of stool [34]. The quality of life of IBS patients is often severely impacted by pain, functional disturbances, and accompanying psychological conditions such as depression and anxiety [35].
Various metabolites can act as signaling molecules that affect biological processes, and changes in host metabolites may be associated with the emergence of IBS symptoms [36]. IBS is exacerbated and maintained by changes and variations in intestinal metabolites. Han et al. highlighted the significant alterations in serum metabolites that aid in the diagnosis of IBS patients, proposed a potential role for metabolic dysregulation in its pathophysiology, and offered fresh information on the co-occurrence of IBS and depression [4]. In our study, we endeavor to provide new insights into the causal role of blood metabolites in the risk of IBS. This is pivotal as it suggests that screening for certain blood metabolites could be a viable strategy for identifying individuals at high risk for IBS. Future research could focus on validating these metabolites as potential biomarkers for IBS, which would significantly advance our understanding and management of this condition.
The current study is the first in-depth MR investigation to examine the connection between IBS and human blood metabolites. For the first time, our study used rigorous MR methods to set strict screening criteria. It verified the GWAS data of large IBS cohorts in two different queues, ultimately determining a causal relationship between the four metabolites and IBS and providing potential inspiration for further precision treatments. These molecules included three lipids and one amino acid. The lipids include stearate, 1-monopalmitin, and 1-palmitoylglycerophosphoinositol, and the amino acid was arginine. Among these, 1-monopalmitin and arginine may potentially be associated with an increased risk for IBS, while stearate and 1-palmitoylglycerophosphoinositol appear to be potential protective factors against IBS.
The role of bioactive lipids in the gut-brain axis and the ability of palmitoylglycolamide to lower inflammatory markers in mouse models of inflammatory bowel disease (IBD) and butyrate produced by the gut microbiota to effectively reduce inflammation and pain in animal models of IBS and IBD have been demonstrated in studies by Russo [37]. Numerous animal and plant fats contain stearate, an 18-carbon saturated fatty acid. Stearate is crucial for the growth of different tumors because it alters mitochondrial morphology, rendering cancer cells vulnerable to oxidative stress and cell death [38]. Compared with other saturated fatty acids, stearate is associated with reduced low-density lipoprotein cholesterol, and low low-density lipoprotein (LDL) levels seem to be related to the manifestation of IBS symptoms [39, 40]. Additionally, the stearate content is significantly decreased in diarrhea caused by chemotherapy [41]. Stearates are also associated with intestinal permeability, which may explain their causal association with IBS. Current evidence indicates that stearate, a saturated fatty acid, serves as an agonist of Toll-like receptor 4, initiating an inflammatory response and leading to alterations in the gut microbiota [42].
Recent research has shown that lipid metabolism disorder is present in IBS and is an important pathophysiological characteristic of this disease [43]. Although the roles of 1-monopalmitin and 1-palmitoylglycerophosphoinositol in IBS have not yet been reported, studies have verified their potential to regulate the immune system [44]. This could be a promising direction for further investigation.
The urea cycle and metabolism of arginine and proline include the semi-essential amino acid arginine. In humans, arginine metabolism mainly produces urea and ornithine via the arginase pathway and nitric oxide (NO) and citrulline via the nitric oxide synthase (NOS) pathway [45]. In the digestive system, NO serves as the primary inhibitory nonadrenergic and noncholinergic neurotransmitter [46]. Gastrointestinal motility is directly regulated by inhibitory and excitatory motor neurons in the smooth muscle layer. NO released from nerve stimulation in the myenteric plexus causes smooth muscle relaxation and plays a critical role in regulating esophageal, gastric, and intestinal peristalsis [47]. NO may play a role in the pathogenesis of visceral hypersensitivity in IBS, according to Kuiken et al. [48]. Higher blood levels of l-arginine and endogenous methylarginines have also been linked to adult IBS, according to the available research [49].
The MR analysis employed in this study had several advantages. This is the most exhaustive and methodological probe for determining a causal relationship between blood metabolites and IBS. Second, we adopted rigorous criteria to circumvent inevitable pitfalls encountered in previous studies, such as reverse causation and confounding biases. Specifically, several strategies were implemented to ensure the eradication of elements that contravened the MR assumptions, thereby engendering reliable results. The uniformity across the three MR estimation directions and sensitivity analyses indicated the robustness of the findings. Furthermore, the veracity of the results was corroborated by the replication and meta-analysis of GWAS data from a separate IBS cohort. Fourth, we evaluated the genetic association between metabolites and IBS using LDSC, which made the MR Estimate more convincing.
There are certain limitations to the present study. First, few SNPs can be identified at the whole-genome level. To address this issue, we implemented a slightly loosened threshold for the MR analysis, a strategy frequently used in previous studies. Second, most of the metabolite data was of European populations, which restricts the applicability of our findings to other racial groups. Meanwhile, we conducted a meta-analysis combining the European and FinnGen cohorts, yet the ancestral differences between Finland and other parts of Europe might introduce some heterogeneity in the association effects between these cohorts, impacting the reliability of the results. Third, although a relatively wide range of metabolites was included in this study, several of their roles and disease-related processes remain unknown, making it difficult to interpret the results of the MR analysis. Finally, although this study identified several metabolites linked to IBS, further investigations are needed to clarify their functions in the pathogenesis of the disease.
Conclusions
In this study, we determined a causal relationship between the four metabolites and IBS through MR analysis, providing preliminary evidence of the pathogenesis of IBS. Our results provide novel insights into the potential biomarkers of IBS. However, further studies are required to validate these findings.
Availability of data and materials
The datasets used in this study are accessible through online repositories. The article/Supplementary Material includes repository name(s) and accession number(s). Genetic association estimates for IBS were sourced from published GWAS and the FinnGen database. We are deeply appreciative of all participants and researchers who shared these valuable datasets.
Abbreviations
- GWAS:
-
Genome-wide association study
- IBD:
-
Inflammatory bowel disease
- IV:
-
Instrumental variable
- IVW:
-
Inverse variance weighting
- IBS:
-
Irritable Bowel Syndrome
- FDR:
-
False Discovery Rate
- LOO:
-
Leave-one-out analysis
- LD:
-
Linkage disequilibrium
- LDSC:
-
Linkage disequilibrium score regression
- MR:
-
Mendelian randomization
- LDL:
-
Low-density lipoprotein
- NOS:
-
Nitric oxide synthase
- WM:
-
Weighted median
References
Sultan S, Malhotra A. Irritable bowel syndrome. Ann Intern Med. 2017;166:ITC81.
Chey WD, Kurlander J, Eswaran S. Irritable bowel syndrome: a clinical review. JAMA. 2015;313
Zamani M, Alizadeh-Tabari S, Zamani V. Systematic review with meta-analysis: the prevalence of anxiety and depression in patients with irritable bowel syndrome. Aliment Pharmacol Ther. 2019;50:132–43.
Han L, Zhao L, Zhou Y, Yang C, Xiong T, Lu L, et al. Altered metabolome and microbiome features provide clues in understanding irritable bowel syndrome and depression comorbidity. ISME J. 2022;16:983–96.
Valeur J, Undseth R, Jakobsdottir G, Nyman M, Berstad A. Low serum levels of short-chain fatty acids after lactulose ingestion may indicate impaired colonic fermentation in patients with irritable bowel syndrome. CEG. 2015;303
Bongiorno D, Bivona DA, Cicino C, Trecarichi EM, Russo A, Marascio N, et al. Omic insights into various ceftazidime-avibactam-resistant Klebsiella pneumoniae isolates from two southern Italian regions. Front Cell Infect Microbiol. 2023;12:1010979.
Johnson CH, Ivanisevic J, Siuzdak G. Metabolomics: beyond biomarkers and towards mechanisms. Nat Rev Mol Cell Biol. 2016;17:451–9.
Laíns I, Gantner M, Murinello S, Lasky-Su JA, Miller JW, Friedlander M, et al. Metabolomics in the study of retinal health and disease. Prog Retin Eye Res. 2019;69:57–79.
Sekula P, Del Greco MF, Pattaro C, Köttgen A. Mendelian randomization as an approach to assess causality using observational data. J Am Soc Nephrol. 2016;27:3253–65.
Davey Smith G, Hemani G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet. 2014;23:R89–98.
Sanderson E, Glymour MM, Holmes MV, Kang H, Morrison J, Munafò MR, et al. Mendelian randomization. Nat Rev Methods Primers. 2022;2:6.
Bowden J, Holmes MV. Meta-analysis and Mendelian randomization: a review. Res Synth Methods. 2019;10:486–96.
Boef AGC, Dekkers OM, le Cessie S. Mendelian randomization studies: a review of the approaches used and the quality of reporting. Int J Epidemiol. 2015;44:496–511.
Shin S-Y, Fauman EB, Petersen A-K, Krumsiek J, Santos R, Huang J, et al. An atlas of genetic influences on human blood metabolites. Nat Genet. 2014;46:543–50.
Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 2012;40:D109–14.
Eijsbouts C, Zheng T, Kennedy NA, Bonfiglio F, Anderson CA, Moutsianas L, et al. Genome-wide analysis of 53,400 people with irritable bowel syndrome highlights shared genetic pathways with mood and anxiety disorders. Nat Genet. 2021;53:1543–52.
Guo Z, Zhang T, Yun Z, Jin Q, Cao X, Kong D, et al. Assessing the causal relationships between human blood metabolites and the risk of NAFLD: a comprehensive mendelian randomization study. Front Genet. 2023;14:1108086.
Burgess S, Thompson SG. CRP CHD genetics collaboration. Avoiding bias from weak instruments in Mendelian randomization studies. Int J Epidemiol. 2011;40:755–64.
Brion M-JA, Shakhbazov K, Visscher PM. Calculating statistical power in Mendelian randomization studies. Int J Epidemiol. 2013;42:1497–501.
Burgess S, Davey Smith G, Davies NM, Dudbridge F, Gill D, Glymour MM, et al. Guidelines for performing Mendelian randomization investigations: update for summer 2023. Wellcome Open Res. 2023;4:186.
Pierce BL, Burgess S. Efficient design for Mendelian randomization studies: subsample and 2-sample instrumental variable estimators. Am J Epidemiol. 2013;178:1177–84.
Bowden J, Del Greco MF, Minelli C, Davey Smith G, Sheehan NA, Thompson JR. Assessing the suitability of summary data for two-sample Mendelian randomization analyses using MR-egger regression: the role of the I2 statistic. Int J Epidemiol. 2016;45:1961–74.
Burgess S, Thompson SG. Interpreting findings from Mendelian randomization using the MR-egger method. Eur J Epidemiol. 2017;32:377–89.
Bowden J, Davey Smith G, Burgess S. Mendelian randomization with invalid instruments: effect estimation and bias detection through egger regression. Int J Epidemiol. 2015;44:512–25.
Greco MFD, Minelli C, Sheehan NA, Thompson JR. Detecting pleiotropy in Mendelian randomisation studies with summary data and a continuous outcome. Stat Med. 2015;34:2926–40.
Huang T, Long Y, Ou Y, Li J, Huang Y, Gao J. Association between circulating fatty acid metabolites and asthma risk: a two-sample bidirectional Mendelian randomization study. BMC Med Genet. 2023;16:112.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Methodol. 1995;57:289–300.
Cai J, Li X, Wu S, Tian Y, Zhang Y, Wei Z, et al. Assessing the causal association between human blood metabolites and the risk of epilepsy. J Transl Med. 2022;20:437.
Ni G, Moser G, Wray NR, Lee SH, Ripke S, Neale BM, et al. Estimation of genetic correlation via linkage disequilibrium score regression and genomic restricted maximum likelihood. Am J Hum Genet. 2018;102:1185–94.
O’Connor LJ, Price AL. Distinguishing genetic correlation from causation across 52 diseases and complex traits. Nat Genet. 2018;50:1728–34.
Kamat MA, Blackshaw JA, Young R, Surendran P, Burgess S, Danesh J, et al. PhenoScanner V2: an expanded tool for searching human genotype-phenotype associations. Bioinformatics. 2019;35:4851–3.
Zhai C, Guyatt G. Fixed-effect and random-effects models in meta-analysis. Chin Med J. 2023;2023:10–1097. https://doi.org/10.1097/CM9.0000000000002814.
Nee J, Lembo A. Review article: current and future treatment approaches for IBS with diarrhoea (IBS-D) and IBS mixed pattern (IBS-M). Aliment Pharmacol Ther. 2021;54
Lacy BE, Mearin F, Chang L, Chey WD, Lembo AJ, Simren M, et al. Bowel Disorders. Gastroenterology. 2016;150:1393–1407.e5.
Lee C, Doo E, Choi JM, Jang S, Ryu H-S, Lee JY, et al. The increased level of depression and anxiety in irritable bowel syndrome patients compared with healthy controls: systematic review and Meta-analysis. J Neurogastroenterol Motil. 2017;23:349–62.
Xiao L, Liu Q, Luo M, Xiong L. Gut microbiota-derived metabolites in irritable bowel syndrome. Front Cell Infect Microbiol. 2021;11
Russo R, Cristiano C, Avagliano C, De Caro C, La Rana G, Raso GM, et al. Gut-brain Axis: role of lipids in the regulation of inflammation. Pain and CNS Diseases CMC. 2018;25:3930–52.
Chu Q, Liu P, Song Y, Yang R, An J, Zhai X, et al. Stearate-derived very long-chain fatty acids are indispensable to tumor growth. EMBO J. 2023;42:e111268.
Blom-Høgestøl IK, Aasbrenn M, Chahal-Kummen M, Brunborg C, Eribe I, Kristinsson J, et al. Irritable bowel syndrome-like symptoms and health related quality of life two years after roux-en-Y gastric bypass - a prospective cohort study. BMC Gastroenterol. 2019;19:204.
Hunter JE, Zhang J, Kris-Etherton PM. Cardiovascular disease risk of dietary stearic acid compared with trans, other saturated, and unsaturated fatty acids: a systematic review. Am J Clin Nutr. 2010;91:46–63.
Melichar B, Dvorák J, Krcmová L, Hyspler R, Urbánek L, Solichová D. Intestinal permeability and vitamin a absorption in patients with chemotherapy-induced diarrhea. Am J Clin Oncol. 2008;31:580–4.
Rocha DM, Caldas AP, Oliveira LL, Bressan J, Hermsdorff HH. Saturated fatty acids trigger TLR4-mediated inflammatory response. Atherosclerosis. 2016;244:211–5.
Karpe AV, Liu J-W, Shah A, Koloski N, Holtmann G, Beale DJ. Utilising lipid and, arginine and proline metabolism in blood plasma to differentiate the biochemical expression in functional dyspepsia (FD) and irritable bowel syndrome (IBS). Metabolomics. 2022;18:38.
O’Donnell VB, Rossjohn J, Wakelam MJO. Phospholipid signaling in innate immune cells. J Clin Investig. 2018;128:2670–9.
Morris SM. Arginine metabolism: boundaries of our knowledge. J Nutr. 2007;137:1602S–9S.
Bult H, Boeckxstaens GE, Pelckmans PA, Jordaens FH, Maercke YMV, Herman AG. Nitric oxide as an inhibitory non-adrenergic non-cholinergic neurotransmitter. Nature. 1990;345:346–7.
Stanek A, Gadowska-Cicha A, Gawron K, Wielkoszynski T, Adamek B, Cieslar G, et al. Role of nitric oxide in physiology and pathology of the gastrointestinal tract. MRMC. 2008;8:1549–60.
Kuiken SD, Klooker TK, Tytgat GN, Lei A, Boeckxstaens GE. Possible role of nitric oxide in visceral hypersensitivity in patients with irritable bowel syndrome. Neurogastroenterol Motil. 2006;18:115–22.
McEvoy MA, Attia JR, Oldmeadow C, Holliday E, Smith WT, Mangoni AA, et al. Serum L-arginine and endogenous methylarginine concentrations predict irritable bowel syndrome in adults: a nested case-control study. United European Gastroenterol J. 2021;9:809–18.
Acknowledgements
Genetic instruments for studied exposures were obtained from Metabolomics GWAS Server. Genetic association estimates for IBS were obtained from the published GWASs. We thank all participants and investigators for sharing these data.
Funding
This research did not receive any specific grants from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
YZ, HL, and CL conceived and designed the study. YZ and RL collected the GWAS data and conducted the primary data analysis. ZP and ZL organized the data and reviewed the statistical analyses. CL provided valuable suggestions and revised the manuscript accordingly. All authors have read and agreed to the final version of the manuscript and unanimously consent to its publication.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests..
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Zeng, Y., Liu, H., Pei, Z. et al. Evaluation of the causal effects of blood metabolites on irritable bowel syndrome: Mendelian randomization. BMC Gastroenterol 24, 19 (2024). https://doi.org/10.1186/s12876-023-03111-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12876-023-03111-9