Abstract
Background
General transcription factor IIi (GTF2I) mutations are very common in thymic epithelial tumors (TETs) and are related to a more favorable prognosis in TET patients. However, limited research has been conducted on the role of GTF2I in the tumor immune microenvironment (TIME). Further, long non-coding RNAs (lncRNAs) have been associated with the survival of patients with TETs. Therefore, this study aimed to explore the relationship between GTF2I mutations and TIME and build a new potential signature for predicting tumor recurrence in the TETs. Research data was downloaded from The Cancer Genome Atlas database and the CIBERSORT algorithm was used to evaluate TIME differences between GTF2I mutant and wild-type TETs. Relevant differentially expressed lncRNAs based on differentially expressed immune-related genes were identified to establish lncRNA pairs. We constructed a signature using univariate and multivariate Cox regression analyses.
Results
GTF2I is the most commonly mutated gene in TETs, and is associated with an increased number of early-stage pathological types, as well as no history of myasthenia gravis or radiotherapy treatment. In the GTF2I wild-type group, immune score and immune cell infiltrations with M2 macrophages, activated mast cells, neutrophils, plasma, T helper follicular cells, and activated memory CD4 T cells were higher than the GTF2I mutant group. A risk model was built using five lncRNA pairs, and the 1-, 3-, and 5-year area under the curves were 0.782, 0.873, and 0.895, respectively. A higher risk score was related to more advanced histologic type.
Conclusion
We can define the GTF2I mutant-type TET as an immune stable type and the GTF2I wild-type as an immune stressed type. A signature based on lncRNA pairs was also constructed to effectively predict tumor recurrence.
Similar content being viewed by others
Background
Thymic epithelial tumors (TETs) are tumors originating from thymic epithelial cells, with an incidence of approximately 1–3 cases/million, and are the most common type of anterior mediastinal tumor [1, 2]. According to the 2021 World Health Organization (WHO) classification, TETs can be classified into thymoma type A, AB, B1, B2, B3, and thymic carcinoma [3]. Type B2 and B3 thymoma and thymic carcinoma have a higher degree of malignancy compared with type A, AB and B1 thymomas, implying a less favourable prognosis [4, 5]. However, some studies have determined that type A, AB, and B1 thymomas with low grade oncological morphology were more likely to recur [6]. Masaoka stage has been identified as an independent prognostic factor with a strong ability to predict tumor recurrence and patient survival [7]. Nevertheless, neither Masaoka stage nor pathological classification are sufficiently accurate to predict tumor recurrence in TET patients, and there are no recognized biomarkers to predict TETs recurrence in clinical practice. General transcription factor IIi (GTF2I) mutations are reported at a high frequency in TETs, and these mutations have been associated with a favorable prognosis of patients with TETs [8, 9]. Recently, a growing number of studies have been conducted on tumor immune microenvironments (TIME) which increase our understanding of the immune mechanism during tumor development and metastasis, while also promoting the discovery of new methods for studying tumor development [10,11,12]. The relationship between GTF2I mutations and TIME has not previously been studied in TETs despite its importance to aid in understanding the molecular behavior of TETs to enable clinical application. Long non-coding RNA (lncRNA) are segments of non-coding RNA greater than 200 nucleotides in length that lack protein coding ability. An increasing number of studies are exploring the role of lncRNA during tumor development [13,14,15], for example, lncRNA H19 is a powerful prognostic biomarker of neuroendocrine prostate cancer that predicts the probability of tumor recurrence [16]. Several studies have also reported that lncRNA LOXL1-AS1, LINC00174, and XLOC_003810 play a key role in the development of TETs, which indicated the potential of lncRNA to predict the prognosis of TET patients [17,18,19]. As lncRNAs and GTF2I are both important in TETs, it is critical to determine whether lncRNAs associated with GTF2I mutations in TETs. Previous studies have established lncRNA signatures based on lncRNA expression levels to predict the prognosis of patients with TETs [20, 21]. However, the expression level of lncRNA depends on the detection platform; therefore, clinical utilization of these lncRNA signatures is limited by the accuracy of the detected lncRNA level. A prognostic gene or lncRNA signature using a gene or lncRNA pairs method has also been established, which is not restricted by the expression level of a gene or lncRNA and instead focuses on the expression level differences of a gene or lncRNA [22,23,24].
In this study, we first examined the TIME differences between GTF2I mutant and wild-type TETs based on The Cancer Genome Atlas (TCGA) patient data. Relevant differentially expressed lncRNAs were selected based on differentially expressed immune-related genes between GTF2I mutant and wild-type TETs. Next, we constructed a novel lncRNA signature to predict tumor recurrence in patients with TETs by applying the lncRNA pairs method.
Results
Characterizing TET mutations and GTF2I mutation relation to clinical features and survival
Gene mutations in all TET samples are shown in Fig. 1A, including gene mutation frequency and type. GTF2I (47%), muscle RAS oncogene homolog (MRAS) (7%), titin (TTN) (6%), and mucin 16 (MUC16) (4%) were four genes identified with the highest mutation frequency in the TET samples. Notably, GTF2I mutations were all missense mutations.
Based on the GTF2I mutations, we divided all TET patients into two groups: the GTF2I mutant and GTF2I wild-type. Survival curves showed that patients in the GTF2I mutant-type group had more favorable prognostic results with lower probabilities of tumor recurrence than the GTF2I wild-type group (Fig. 1B). The clinical features of the two groups are shown in Table 1. The GTF2I mutation was determined to be related to Masaoka stage I, more indolent pathological subtypes, and no history of myasthenia gravis (MG) or no radiotherapy treatment. GTF2I wild-type was associated with advanced pathological types, MG history, and radiotherapy treatment.
Estimation of immune cell infiltration and immune checkpoints between GTF2I mutant and wild-type TETs
There was no significant difference (p = 0.98) in estimate scores between the GTF2I mutant and wild-type groups (Fig. 2A). The stromal score and immune cell infiltrations with resting dendritic cells and monocytes in the GTF2I mutant group were higher than those in the GTF2I wild-type group (Fig. 2A-B). The immune score and immune cell infiltrations with M2 macrophages, activated mast cells, neutrophils, plasma cells, T helper follicular cells, and activated memory CD4 T cells in the GTF2I mutant group were lower than in the GTF2I wild-type group (Fig. 2A-B). The expression of PD-1, PD-L1, and CTLA4 in the GTF2I mutant group were lower than in the GTF2I wild-type group (Fig. 2C). Immune checkpoints such as CD2, C-C motif chemokine ligand 19 (CCL19), CD3 epsilon subunit of T-cell receptor complex (CD3E), heat shock protein family A (Hsp70) member 8 (HSPA8), CD3 delta subunit of T-cell receptor complex (CD3D), C-X-C motif chemokine ligand 12 (CXCL12), CD27, C-C motif chemokine receptor 7 (CCR7), galectin 9 (LGALS9), CD48, TP53, heat shock protein family A (Hsp70) member 2 (HSPA2), granzyme B (GZMB), perforin 1 (PRF1), baculoviral IAP repeat containing 5 (BIRC5), T cell immunoglobulin and mucin domain containing 4 (TIMD4), TNF receptor superfamily member 18 (TNFRSF18), and enhancer of zeste 2 polycomb repressive complex 2 subunit (EZH2) showed a higher level of expression in the GTF2I wild-type group than in the GTF2I mutant group (Fig. 2D).
GSEA
The top nine pathways with enrichment in the GTF2I mutant group were the adherens junction, dilated cardiomyopathy, arrhythmogenic right ventricular cardiomyopathy, vascular smooth muscle contraction, basal cell carcinoma, melanogenesis, Notch signaling pathway, transforming growth factor (TGF) beta signaling pathway, and wingless/integrated (WNT) signaling pathway (Fig. . 3A).
A GSEA results of KEGG analysis. B-C Volcano map of DEGs (B) and DERs (C) between GTF2I wild type group and GTF2I mutant group samples. D-E Heatmap of the top 40 genes (D) and lncRNAs (E) that were most significantly differentially expressed between GTF2I wild type group and GTF2I mutant group samples. F-G Barplots of GO analyses (F) and KEGG analyses (G)
Functional enrichment analysis of DEGs
We performed differential expression analysis in the GTF2I mutant and wild-type TET samples, resulting in 396 differentially expressed genes (DEGs) and 93 differentially expressed lncRNAs (DERs) visualized as volcano plots (Fig. . 3B-C). The heatmaps showed the top 40 genes and lncRNAs that were the most significantly differentially (p < 0.05) expressed between GTF2I wild-type and mutant TETs (Fig. . 3D-E). Gene ontology (GO) analysis revealed that extracellular matrix structural constituent, endopeptidase, signaling receptor activator activity, endopeptidase inhibitor, and receptor-ligand activity, were significantly enriched (Fig. . 3F). The Kyoto encyclopedia of genes and genomes (KEGG) analysis demonstrated that the staphylococcus aureus infection, cytokine-cytokine receptor interaction, and IL-17 signaling pathway were significantly enriched (p < 0.05) (Fig. . 3G).
Construction and validation of the risk assessment model
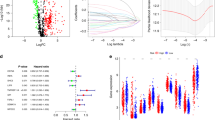
Eight immune-related differentially expressed genes (IRDEGs) and 53 immune-related differentially expressed lncRNAs (IRDERs) were identified by taking the intersection and performing Spearman’s correlation analysis. We used an iterative cycle among the 53 IRDERs and identified 1266 IRDER pairs. We performed univariate Cox regression analysis on all IRDER pairs and identified 19 IRDER pairs (Fig. 4A). Next, we performed multivariate Cox regression which resulted in five IRDER pairs to build the risk model (Fig. 4B). When the Akaike’s information criterion (AIC) value referred to 0.1, we identified the maximum knee point as the cut-off value and attained the largest area under the curve (AUC) value of 0.782 (Fig. 4C). The formula used to calculate the risk score for all TET patients was:
A The prognostic IRDER pairs extracted by univariate cox regression analysis. B The prognostic signature constructed by multivariate cox regression analysis. C AUC of the signature and the best cutoff value obtained by the AIC. D Risk score distribution of patients between high- and low-risk groups. E Tumor recurrent status of patients between high- and low-risk groups. F Kaplan–Meier survival curve of patients between high- and low-risk groups
Risk score = − 0.700794392573432× CCDC144NL-AS1|LINC01748–1.60567899345149× AC074389.2|LINC02542+ 1.19279714426103× AC131902.1|AL365356.5–0.725120180351969× AC005383.1|AC023906.2–0.755363912198633× AC009041.2|LINC02384. We divided the groups into high- and low-risk using the cut-off value of 0.1 (Fig. 4D). We observed that patients with a high-risk score had a greater chance of tumor recurrence and a shorter time to recurrence (Fig. 4E). Kaplan-Meier analysis showed that patients in the low-risk group exhibited better prognostic results with lower odds of tumor recurrence than in the high-risk group (Fig. 4F). In addition, the clinicopathologic characteristics such as age (Fig. 5A), gender (Fig. 5B), GTF2I mutation (Fig. 5C), histologic subtype (Fig. 5D), MG history (Fig. 5E) and Masaoka stage (Fig. 5F) also demonstrated that patients in the low-risk group had lower odds of tumor recurrence compared with the high-risk group.
Validation of the risk assessment model
Time-dependent receiver operating characteristic (ROC) curves were used to assess the specificity and sensitivity of the risk model and the 1-, 3-, and 5-year AUCs were 0.782, 0.873, and 0.895, respectively (Fig. 6A). Univariate Cox regression analysis showed that the hazard ratio (HR) of the histologic type and 95% confidence interval (CI) were 1.470 and 1.075–2.012 (p = 0.016), respectively. HR of the risk score and 95% CI were 3.351 and 2.161–5.196 (p < 0.001), respectively (Fig. 6B). Results of multivariate Cox regression analysis showed that risk score (HR = 3.923; 95% CI = 2.067–7.444; p < 0.001,) was an independent prognostic predictor (Fig. 6C). The AUC values for Masaoka stage, age, gender, and histologic subtype in 1-year survival were 0.468, 0.464, 0.483, and 0.637, respectively (Fig. 6D). The strip charts (Fig. 6E) and consequent scatter diagrams formed from a Wilcoxon signed-rank test revealed that a higher risk score was significantly related (p < 0.05) to more aggressive histologic subtypes (Fig. 7A). There was no significant association between the risk score and age (Fig. 7B), gender (Fig. 7C), GTF2I mutation (Fig. 7D), Masaoka stage (Fig. 7E), and MG history (Fig. 7F).
A Time-dependent ROC curve analyses of risk score. B Univariate Cox analyses of clinical factors and risk score. C Multivariate Cox analyses of clinical factors and risk score. D One-year ROC curve analyses of Masaoka stage, Age, Gender, Histologic type and risk score. E Heatmap showed the comparison of the relationship between the clinical characteristics of patients between high- and low-risk groups
Construction and validation of the nomogram
A nomogram was established using risk score and common clinical factors of patients with TETs, including age, GTF2I mutation, gender, Masaoka stage, and histologic subtype (Fig. 8A). The 1-, 3-, and 5-year calibration curves showed a strong ability to predict tumor recurrence (Fig. 8B).
Discussion
Through the analysis of thymoma mutation data from TCGA, we determined that GTF2I mutations were highly common in thymoma and were mostly missense mutations, which was consistent with previous findings [8, 9, 25, 26]. Additionally, we found that the GTF2I mutant type had a lower chance of tumor recurrence, and a longer disease-free survival than the wild-type TET. Previous studies have also shown that type B1, B2, B3, and thymic carcinoma were more aggressive pathological subtypes with a worse prognosis, whereas thymoma type A and type AB exhibited more favorable prognosis [4, 5]. Mutant type was more concentrated in thymoma type A and AB, whereas the wild-type occurred more often in type B1, B2, B3, and thymic carcinoma. This demonstrates to some extent that the prognosis relevant to GTF2I in TET provides a potential molecular mechanism for the prognosis relevant to pathological subtypes. However, some studies showed inconsistent results whereby more indolent subtypes of TETs, such as type A and AB thymomas, were more likely to recur [6].Further research is still needed to determine the intrinsic truth.
Due to the limited studies exploring the relationship between GTF2I mutations and TIME worldwide, we investigated the differences in the TIME between TET patients with GTF2I mutant thymoma and wild-type, with the aim to illustrate prognostic differences between the two groups. According to the TIME infiltration characterization of patients with mutant and wild-type TETs, we defined GTF2I mutant type as the most immune stable type because the peak of immune response passed and the immune response was more stable. We defined the GTF2I wild-type as the more immune stressed type because the immune response had not yet reached the peak, the immune response to tumor antigen stimulation was strong, the immune response continued to enhance in the development to the peak, and various immune checkpoints were highly expressed with great immunotherapy potential [27,28,29]. Including three star immune checkpoints, PD-1, PD-L1, and CTLA4, a total of 21 immune checkpoints were identified as upregulated in GTF2I wild-type TET samples in our study. Therefore, once we find that a patient with advanced TET is GTF2I wild-type is diagnosed, indicating that the patient has a poor prognosis, better efficacy may be achieved by using PD-1, PD-L1, or CTLA4 inhibitors for treatment. Some previous studies have shown that anti-PD-L1 or anti-PD-1 antibodies such as pembrolizumab and avelumab can have a therapeutic effect on recurrent and metastatic advanced thymoma or thymic carcinoma and can control tumor growth [30,31,32]. We recommend that patients with advanced TETs should be tested for GTF2I gene mutation status, to more accurately predict patient prognosis and predict immune checkpoint inhibitor responses.
For the GTF2I wild-type TETs, multiple immune cells were highly infiltrated. The immune response was in a state of continuous activation, which was too strong and would attack their own normal tissues and organs, hence, the patients were in a state of continuous stress. Therefore, GTF2I wild-type TET patients were more likely to form autoimmune diseases as well as develop symptoms of MG than patients with a mutant type. At the same time, our results suggested that the use of PD-1, PD-L1 or CTLA4 inhibitors in the treatment of patients with advanced TETs required close attention on the generation of immune-related adverse reactions such as MG while treating tumors. Anti-PD-L1 or anti-PD-1 antibodies have been reported to cause adverse reactions such as MG, myocarditis and pneumonia in patients with TETs [32,33,34,35]. Further, our study provides multiple potential immunotherapeutic targets such as CD2. The CD2 monoclonal antibody DANYELZA may be an option for TET patients if they exhibit low responsiveness or poor tolerability to PD-1 or PD-L1 inhibitors. Our study has identified immune checkpoints requiring further investigation.
The Notch, TGF beta and WNT signaling pathways were enriched in the GTF2I mutant group. Previous studies have reported that these signaling pathways play an important role in regulating TIME and cancer progression [36,37,38,39,40,41]. The results of GO and KEGG enrichment analyses illustrated that IRDEGs were closely related with cytokine receptor interactions, receptor-ligands interactions, and signaling receptor mutual activation.
Several studies have shown that lncRNAs could be used as prognostic marker in multiple cancers [42,43,44,45,46], and could build signatures that can predict tumor recurrence [47,48,49]. However, due to inconsistent accuracy when detecting lncRNA expression level, which could deeply influence clinical practice, we constructed a lncRNA signature using a novel lncRNA method that can reduce the errors caused by the detection platform [22,23,24]. To our knowledge, the establishment of the lncRNA pairs method to construct a predictive signature in TETs to predict the prognosis of TET patients before has not yet been conducted. Moreover, good efficacy of this signature in predicting the prognosis of TET patients further verifies the importance of GTF2I mutations in TETs. We also determined that some lncRNAs associated with GTF2I mutations may play an important role in TETs, which to our knowledge has not been previously reported. The lncRNA CCDC144NL-AS1 was significantly upregulated in gastric cancer tissues and was related to poor prognosis [50]. Overexpression of lncRNA CCDC144NL-AS1 has also been associated with poor prognosis of patients suffering from non-small cell lung cancer [51]. The lncRNA LINC01748 acts as an independent predictor of poor prognosis in non-small cell lung cancer patients [52]. Our study showed that the lncRNAs CCDC144NL-AS1 and LINC01748 may also play an important role in the development of thymoma and are prognostic markers requiring further investigation.
Despite the implications of our results, this study still has several limitations. The number of TET patient samples used to compare clinical features and TIME between GTF2I mutant-types and wild-types and used to establish the lncRNA pairs signature was small (119) and external validation was not available.
Conclusion
We analysed TIME differences between patients with GTF2I mutant and wild-type TETs, and defined GTF2I mutant-type as immune stable and the GTF2I wild-type as immune stressed. In addition, we established a signature based on IRDEGs to predict tumor recurrence, and a risk score was determined as an independent clinical prognostic factor. Our study improves the understanding of GTF2I mutations in the TIME and provides more insight into effective immunotherapy strategies.
Methods
Data processing
We downloaded the tumor mutation and transcription data of 119 patients with TETs and their clinicopathological information from TCGA database, however, clinical information was missing from some patients. Three hundred and thirty-two immune related genes were obtained from the immune system process gene set (systemic name: M13664) in the Gene Set Enrichment Analysis (GSEA) (http://www.gsea-msigdb.org/gsea/index.jsp).
Evaluation of gene mutation in TETs
Gene mutation in TETs was visualized using a waterfall plot with the R package “maftools” and oncoplot function. Differences such as gender, age, race, MG history, Masaoka stage, histologic type, radiation therapy history, and vital status were compared using a Chi-square test between GTF2I mutant and wild-type TET patients.
We performed Kaplan-Meier analysis to assess the disease-free survival difference between GTF2I mutant and wild-type TET patients and visualized this via survival curves using the “survival” and “survminer” R packages, and the ggsurvplot function.
Investigation of the TIME and immune checkpoints
CIBERSORT (http://cibersort.stanford.edu/) can evaluate immune cell infiltration levels on the basis of the gene expression profiles from complex tissues [53]. Using the transcriptional profile of TETs and this software, we calculated the relative percentage of infiltration of 22 immune infiltrating cells and used wilcoxTest function, the “limma” and “ggpubr” R packages to assess the differences between GTF2I mutant and wild-type TET patients. We calculated the immune score of each sample using the R package “ESTIMATE” and analyzed the difference between GTF2I mutant and wild-type TET patients [54]. We identified 178 immune checkpoints by reviewing existing literatures. Then we performed gene expression differential analysis between GT2I mutant samples and wild-type samples, and obtained 65 differentially expressed immune checkpoint genes. Finally, we removed genes with a median immune checkpoint gene expression (log2) less than 1, and obtained 18 immune checkpoints.
GSEA
We utilized gene sets (c2. cp. kegg.v6.2.-symbols) with GSEA software (https://www.gsea-msigdb.org/gsea/login.jsp) to investigate differences across pathway activity between GTF2I mutant and wild-types (p < 0.05).
Differential expression analysis between GTF2I mutant and wild-type TET tissues and enrichment analysis of the differentially expressed genes
We identified the DEGs and lncRNAs by comparing the TET tissues of 56 patients with GTF2I mutation and 63 patients without GTF2I mutation from TCGA dataset with a threshold for false discovery rate of < 0.05, along with |log2 FC (fold-change) | > 2 using the R package “limma”. We performed KEGG and GO enrichment analyses of these DEGs using the R package “clusterProfiler”, “org.Hs.eg.db” and “enrichplot” to investigate the biological processes difference between GTF2I mutant and wild-type TETs.
Pairing lncRNAs methods
We obtained IRDEGs by taking the intersection of immune genes and DEGs. Using Spearman’s correlation analysis between the IRDEGs and DERs, we received IRDERs with correlation coefficients > 0.4 and p < 0.05. We compared all IRDERs pairwise with ifelse function; in a lncRNA pair, one lncRNA was defined as a lncRNA a and the other is defined as lncRNA b. When lncRNA a was larger than lncRNA b, its value was defined as 1. When lncRNA a was smaller than lncRNA b, it was defined as 0.
Construction of the lncRNA pairs prognostic risk signature
First, we performed a univariate Cox regression analysis with p < 0.01 in all IRDER pairs. Then, we performed multivariate Cox regression and identified five IRDER pairs to build the risk model for predicting TET recurrence. The formula used to calculate the risk score is as follows:
where βi is the expression quantity of the IRDER pairs and Si is the coefficient of correlation of the IRDER pairs.
The maximum knee was determined by the AIC value of the 1-year ROC curve, which was used as a cut-off point of the risk score for classifying patients into the high- and low-risk groups. We used the R package “survivalROC” and polygon function to plot the 1-, 3-, 5-year ROC curves. We performed Kaplan-Meier analysis to assess the disease-free survival difference between the high- and low-risk groups and was visualized through survival curves by using the R packages “survival” and “survminer”, and ggsurvplot function. Univariate and multivariate Cox regression analyses were performed between risk score and clinical features such as age, gender, histologic type, Masaoka stage, and GTF2I mutation. We also conducted a Wilcoxon signed-rank test to compare the risk score differences between groups with different clinical features.
Construction and validation of the nomogram
Common clinical features in patients with TETs, such as age, GTF2I mutation, gender, Masaoka stage, and histologic type, were used to construct a nomogram using the R package “rms” and calibrate function. The time-dependent calibration curves from the R package “rms” were used to determine prediction accuracy of the nomogram [55, 56].
Availability of data and materials
The entire RNA-seq profile data, mutation data and the clinical data of ESCC patients in this study come from The Cancer Genome Atlas (TCGA, https://cancergenome.nih.gov/) database.
Abbreviations
- AIC:
-
Akaike’s information criterion
- BIRC5:
-
Baculoviral IAP repeat containing 5
- CCL19:
-
C-C motif chemokine ligand 19
- CCR7:
-
C-C motif chemokine receptor 7
- CD3D:
-
CD3 delta subunit of T-cell receptor complex
- CD3E:
-
CD3 epsilon subunit of T-cell receptor complex
- CI:
-
Confidence interval
- CXCL12:
-
C-X-C motif chemokine ligand 12
- DEG:
-
Differentially expressed gene
- DER:
-
Differentially expressed lncRNA
- EZH2:
-
Enhancer of zeste 2 polycomb repressive complex 2 subunit
- GO:
-
Gene Ontology
- GSEA:
-
Gene Set Enrichment Analysis
- GTF2I:
-
General transcription factor IIi
- GZMB:
-
Granzyme B
- HR:
-
Hazard ratio
- HSPA2:
-
Heat shock protein family A (Hsp70) member 2
- HSPA8:
-
Heat shock protein family A (Hsp70) member 8
- IRDEG:
-
Immune-related differentially expressed gene
- IRDER:
-
Immune-related differentially expressed lncRNA
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LGALS9:
-
Galectin 9
- LncRNA:
-
Long non-coding RNA
- MG:
-
Myasthenia gravis
- MRAS:
-
Muscle RAS oncogene homolog
- MUC16:
-
Mucin 16
- PRF1:
-
Perforin 1
- ROC:
-
Receiver operating characteristic
- TCGA:
-
The Cancer Genome Atlas
- TET:
-
Thymic epithelial tumor
- TGF:
-
Transforming growth factor
- TIMD4:
-
T cell immunoglobulin and mucin domain containing 4
- TIME:
-
Tumor immune microenvironment
- TNFRSF18:
-
TNF receptor superfamily member 18
- TTN:
-
Titin
- WHO:
-
World Health Organization
- WNT:
-
Wingless/integrated
References
Engels EA. Epidemiology of thymoma and associated malignancies. J Thorac Oncol. 2010;5(10 Suppl 4):S260–5.
de Jong WK, Blaauwgeers JL, Schaapveld M, Timens W, Klinkenberg TJ, Groen HJ. Thymic epithelial tumours: a population-based study of the incidence, diagnostic procedures and therapy. Eur J Cancer. 2008;44(1):123–30.
Marx A, Chan JKC, Chalabreysse L, Dacic S, Detterbeck F, French CA, et al. The 2021 WHO classification of tumors of the thymus and mediastinum: what is new in thymic epithelial, germ cell, and mesenchymal tumors? J Thorac Oncol. 2022;17(2):200–13.
Kondo K, Yoshizawa K, Tsuyuguchi M, Kimura S, Sumitomo M, Morita J, et al. WHO histologic classification is a prognostic indicator in thymoma. Ann Thorac Surg. 2004;77(4):1183–8.
Rea F, Marulli G, Girardi R, Bortolotti L, Favaretto A, Galligioni A, et al. Long-term survival and prognostic factors in thymic epithelial tumours. Eur J Cardiothorac Surg. 2004;26(2):412–8.
Lamarca A, Moreno V, Feliu J. Thymoma and thymic carcinoma in the target therapies era. Cancer Treat Rev. 2013;39(5):413–20.
Masaoka A. Staging system of thymoma. J Thorac Oncol. 2010;5(10 Suppl 4):S304–12.
Feng Y, Lei Y, Wu X, Huang Y, Rao H, Zhang Y, et al. GTF2I mutation frequently occurs in more indolent thymic epithelial tumors and predicts better prognosis. Lung Cancer. 2017;110:48–52.
Petrini I, Meltzer PS, Kim IK, Lucchi M, Park KS, Fontanini G, et al. A specific missense mutation in GTF2I occurs at high frequency in thymic epithelial tumors. Nat Genet. 2014;46(8):844–9.
Hinshaw DC, Shevde LA. The tumor microenvironment innately modulates cancer progression. Cancer Res. 2019;79(18):4557–66.
Pitt JM, Marabelle A, Eggermont A, Soria JC, Kroemer G, Zitvogel L. Targeting the tumor microenvironment: removing obstruction to anticancer immune responses and immunotherapy. Ann Oncol. 2016;27(8):1482–92.
Ozga AJ, Chow MT, Luster AD. Chemokines and the immune response to cancer. Immunity. 2021;54(5):859–74.
Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer Res. 2017;77(15):3965–81.
Chi Y, Wang D, Wang J, Yu W, Yang J. Long Non-Coding RNA in the Pathogenesis of Cancers. Cells. 2019;8(9):1015.
Liu Q, Huang J, Zhou N, Zhang Z, Zhang A, Lu Z, et al. LncRNA loc285194 is a p53-regulated tumor suppressor. Nucleic Acids Res. 2013;41(9):4976–87.
Singh N, Ramnarine VR, Song JH, Pandey R, Padi SKR, Nouri M, et al. The long noncoding RNA H19 regulates tumor plasticity in neuroendocrine prostate cancer. Nat Commun. 2021;12(1):7349.
Wang J, Huang H, Zhang X, Ma H. LOXL1‑AS1 promotes thymoma and thymic carcinomaprogression by regulating miR‑525‑5p‑HSPA9. Oncol Rep. 2021;45(6):117.
Tito C, Ganci F, Sacconi A, Masciarelli S, Fontemaggi G, Pulito C, et al. LINC00174 is a novel prognostic factor in thymic epithelial tumors involved in cell migration and lipid metabolism. Cell Death Dis. 2020;11(11):959.
Hu B, Niu L, Jiang Z, Xu S, Hu Y, Cao K. LncRNA XLOC_003810 promotes T cell activation and inhibits PD-1/PD-L1 expression in patients with myasthenia gravis-related thymoma. Scand J Immunol. 2020;92(1):e12886.
Shao J, Lyu W, Zhou J, Xu W, Wang D, Liang S, et al. A panel of five-lncRNA signature as a potential biomarker for predicting survival in gastric and thoracic cancers. Front Genet. 2021;12:666155.
Su Y, Chen Y, Tian Z, Lu C, Chen L, Ma X. lncRNAs classifier to accurately predict the recurrence of thymic epithelial tumors. Thorac Cancer. 2020;11(7):1773–83.
Zhou R, Liang J, Tian H, Chen Q, Yang C, Liu C. Development of a Ferroptosis-related lncRNA signature to predict the prognosis and immune landscape of bladder cancer. Dis Markers. 2021;2021:1031906.
Li B, Cui Y, Diehn M, Li R. Development and validation of an individualized immune prognostic signature in early-stage nonsquamous non-small cell lung cancer. JAMA Oncol. 2017;3(11):1529–37.
Wang Y, Zhang X, Dai X, He D. Applying immune-related lncRNA pairs to construct a prognostic signature and predict the immune landscape of stomach adenocarcinoma. Expert Rev Anticancer Ther. 2021;21(10):1161–70.
Jain D, Guleria P, Singh V, Parshad R, Kumar S, Gaiser T, et al. GTF2I mutation in Thymomas: Independence from racial-ethnic backgrounds. An Indian/German comparative study. Pathol Oncol Res. 2021;27:1609858.
Wells K, Lamrca A, Papaxoinis G, Wallace A, Quinn AM, Summers Y, et al. Unique correlation between GTF2I mutation and spindle cell morphology in thymomas (type A and AB thymomas). J Clin Pathol. 2021;20(7):837.
Zhang B, Wu Q, Li B, Wang D, Wang L, Zhou YL. m(6)A regulator-mediated methylation modification patterns and tumor microenvironment infiltration characterization in gastric cancer. Mol Cancer. 2020;19(1):53.
Tekpli X, Lien T, Røssevold AH, Nebdal D, Borgen E, Ohnstad HO, et al. An independent poor-prognosis subtype of breast cancer defined by a distinct tumor immune microenvironment. Nat Commun. 2019;10(1):5499.
Lin A, Wei T, Meng H, Luo P, Zhang J. Role of the dynamic tumor microenvironment in controversies regarding immune checkpoint inhibitors for the treatment of non-small cell lung cancer (NSCLC) with EGFR mutations. Mol Cancer. 2019;18(1):139.
Cafaro A, Bongiovanni A, Di Iorio V, Oboldi D, Masini C, Ibrahim T. Pembrolizumab in a patient with heavily pre-treated squamous cell Thymic carcinoma and cardiac impairment: a case report and literature review. Front Oncol. 2020;10:1478.
Shen L, Chen H, Wei Q. Immune-therapy-related toxicity events and dramatic remission after a single dose of pembrolizumab treatment in metastatic thymoma: a case report. Front Immunol. 2021;12:621858.
Rajan A, Heery CR, Thomas A, Mammen AL, Perry S, O'Sullivan Coyne G, et al. Efficacy and tolerability of anti-programmed death-ligand 1 (PD-L1) antibody (Avelumab) treatment in advanced thymoma. J Immunother Cancer. 2019;7(1):269.
Konstantina T, Konstantinos R, Anastasios K, Anastasia M, Eleni L, Ioannis S, et al. Fatal adverse events in two thymoma patients treated with anti-PD-1 immune check point inhibitor and literature review. Lung Cancer. 2019;135:29–32.
Zhang Y, Li W, Hu X, Zhang J. A case report of advanced thymoma re-treated with PD-1 inhibitor after initial immune-related pneumonitis. Ann Palliat Med. 2021;10(9):10083–90.
Portolés Hernández A, Blanco Clemente M, Escribano García D, Velasco Calvo R, Núñez García B, Oteo Domínguez JF, et al. Checkpoint inhibitor-induced fulminant myocarditis, complete atrioventricular block and myasthenia gravis-a case report. Cardiovasc Diagn Ther. 2021;11(4):1013–9.
Meurette O, Mehlen P. Notch signaling in the tumor microenvironment. Cancer Cell. 2018;34(4):536–48.
Itatani Y, Kawada K, Sakai Y. Transforming growth factor‑β signaling pathway in colorectal cancer and its tumor microenvironment. Int J Mol Sci.2019;20(23):5822.
Wang K, Qiu X, Zhao Y, Wang H, Chen L. The Wnt/β‑catenin signaling pathway in the tumor microenvironment of hepatocellular carcinoma. Cancer Biol Med. 2021;19(3):305–18.
Taciak B, Pruszynska I, Kiraga L, Bialasek M, Krol M. Wnt signaling pathway in development and cancer. J Physiol Pharmacol. 2018;69(2):185–96.
Colak S, Ten Dijke P. Targeting TGF-β signaling in cancer. Trends Cancer. 2017;3(1):56–71.
Tyagi A, Sharma AK, Damodaran C. A review on notch signaling and colorectal cancer. Cells. 2020;9(6):1549.
Liang Y, Song X, Li Y, Chen B, Zhao W, Wang L, et al. LncRNA BCRT1 promotes breast cancer progression by targeting miR-1303/PTBP3 axis. Mol Cancer. 2020;19(1):85.
Shigeyasu K, Toden S, Ozawa T, Matsuyama T, Nagasaka T, Ishikawa T, et al. The PVT1 lncRNA is a novel epigenetic enhancer of MYC, and a promising risk-stratification biomarker in colorectal cancer. Mol Cancer. 2020;19(1):155.
Wang X, Yang J, Guo G, Feng R, Chen K, Liao Y, et al. Novel lncRNA-IUR suppresses Bcr-Abl-induced tumorigenesis through regulation of STAT5-CD71 pathway. Mol Cancer. 2019;18(1):84.
Liu Z, Liu L, Weng S, Guo C, Dang Q, Xu H, et al. Machine learning-based integration develops an immune-derived lncRNA signature for improving outcomes in colorectal cancer. Nat Commun. 2022;13(1):816.
Zhu H, Liu Q, Yang X, Ding C, Wang Q, Xiong Y. LncRNA LINC00649 recruits TAF15 and enhances MAPK6 expression to promote the development of lung squamous cell carcinoma via activating MAPK signaling pathway. Cancer Gene Ther. 2022;29(8-9):1285–95.
Chen F, Li Z, Deng C, Yan H. Integrated analysis identifying new lncRNA markers revealed in ceRNA network for tumor recurrence in papillary thyroid carcinoma and build of nomogram. J Cell Biochem. 2019;120(12):19673–83.
Cheng S, Guo J, Wang D, Fang Q, Liu Y, Xie W, et al. A novel three-LncRNA signature predicting tumor recurrence in nonfunctioning pituitary adenomas. Front Genet. 2021;12:754503.
Zhang X, Han J, Du L, Li X, Hao J, Wang L, et al. Unique metastasis-associated lncRNA signature optimizes prediction of tumor relapse in lung adenocarcinoma. Thorac Cancer. 2020;11(3):728–37.
Fan H, Ge Y, Ma X, Li Z, Shi L, Lin L, et al. Long non-coding RNA CCDC144NL-AS1 sponges miR-143-3p and regulates MAP3K7 by acting as a competing endogenous RNA in gastric cancer. Cell Death Dis. 2020;11(7):521.
Zhang L, Chi B, Chai J, Qin L, Zhang G, Hua P, et al. LncRNA CCDC144NL-AS1 serves as a prognosis biomarker for non-small cell lung cancer and promotes cellular function by targeting miR-490-3p. Mol Biotechnol. 2021;63(10):933–40.
Tan Y, Xu F, Xu L, Cui J. Long non‑coding RNA LINC01748 exerts carcinogenic effects in non‑small cell lung cancer cell lines by regulating the microRNA‑520a‑5p/HMGA1 axis. Int J Mol Med. 2022;49(2):22.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12(5):453–7.
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612.
Eagle KA, Lim MJ, Dabbous OH, Pieper KS, Goldberg RJ, Van de Werf F, et al. A validated prediction model for all forms of acute coronary syndrome: estimating the risk of 6-month postdischarge death in an international registry. Jama. 2004;291(22):2727–33.
Lin JX, Wang ZK, Hong QQ, Zhang P, Zhang ZZ, He L, et al. Assessment of clinicopathological characteristics and development of an individualized prognostic model for patients with Hepatoid adenocarcinoma of the stomach. JAMA Netw Open. 2021;4(10):e2128217.
Acknowledgments
We acknowledge the TCGA databases for their public platforms and the contributors for uploading their valuable datasets.
Data availability statement
The data are available from the corresponding author upon reasonable request.
Funding
This work was supported by the Natural Science Foundation of Guangdong Province (No. 2021A1515010908).
Author information
Authors and Affiliations
Contributions
(1) Data curation, Methodology, Investigation, Writing-original draft, Writing-review and Editing: Wei Liu. (2) Editing: Hao-Shuai Yang, Shao-Yi Zheng and Jian-Hao Weng. (3) Supervision: Hong-He Luo. (4) Conceptualization, Writing-review, Editing and Supervision: Yan-Fen Feng and Yi-Yan Lei. The author(s) read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All methods in this research were carried out in accordance with relevant guidelines and regulations of The First Affiliated Hospital of Sun Yat-sen University, Guangzhou, China. Data of our present study was downloaded from the publicly available dataset TCGA, so our study did not require approval from an ethics committee and did not involve human or animal subjects.
Consent for publication
Not Applicable.
Competing interests
The authors declare that there are no conflicts of interest regarding the publication of this article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Liu, W., Yang, HS., Zheng, SY. et al. Thymic epithelial tumors: examining the GTF2I mutation and developing a novel prognostic signature with LncRNA pairs to predict tumor recurrence. BMC Genomics 23, 656 (2022). https://doi.org/10.1186/s12864-022-08880-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-022-08880-3