Abstract
Background
Commelinaceae (Commelinales) comprise 41 genera and are widely distributed in both the Old and New Worlds, except in Europe. The relationships among genera in this family have been suggested in several morphological and molecular studies. However, it is difficult to explain their relationships due to high morphological variations and low support values. Currently, many researchers have been using complete chloroplast genome data for inferring the evolution of land plants. In this study, we completed 15 new plastid genome sequences of subfamily Commelinoideae using the Mi-seq platform. We utilized genome data to reveal the structural variations and reconstruct the problematic positions of genera for the first time.
Results
All examined species of Commelinoideae have three pseudogenes (accD, rpoA, and ycf15), and the former two might be a synapomorphy within Commelinales. Only four species in tribe Commelineae presented IR expansion, which affected duplication of the rpl22 gene. We identified inversions that range from approximately 3 to 15 kb in four taxa (Amischotolype, Belosynapsis, Murdannia, and Streptolirion). The phylogenetic analysis using 77 chloroplast protein-coding genes with maximum parsimony, maximum likelihood, and Bayesian inference suggests that Palisota is most closely related to tribe Commelineae, supported by high support values. This result differs significantly from the current classification of Commelinaceae. Also, we resolved the unclear position of Streptoliriinae and the monophyly of Dichorisandrinae. Among the ten CDS (ndhH, rpoC2, ndhA, rps3, ndhG, ndhD, ccsA, ndhF, matK, and ycf1), which have high nucleotide diversity values (Pi > 0.045) and over 500 bp length, four CDS (ndhH, rpoC2, matK, and ycf1) show that they are congruent with the topology derived from 77 chloroplast protein-coding genes.
Conclusions
In this study, we provide detailed information on the 15 complete plastid genomes of Commelinoideae taxa. We identified characteristic pseudogenes and nucleotide diversity, which can be used to infer the family evolutionary history. Also, further research is needed to revise the position of Palisota in the current classification of Commelinaceae.
Similar content being viewed by others
Introduction
Commelinaceae Mirb., commonly known as the dayflower and spiderwort family, are the largest family of Commelinales Mirb. ex Bercht. & J. Presl, including four other families: Haemodoraceae, Hanguanaceae, Philydraceae, and Pontederiaceae, [1, 2]. Commelinaceae consist of 41 genera and approximately 730 species, widely distributed in both the Old and New Worlds, except in Europe [2,3,4]. Genus Callisia Loefl. and Tradescantia L. emend. M. Pell. are commonly used as ornamentals, while Commelina L. is used as vegetables and more commonly known as troublesome weeds. The species of Commelinaceae are usually succulent herbs with closed leaf-sheaths, raphide-canals, and three-celled glandular microhairs [3, 4]. Additionally, flowers of Commelinaceae are mainly insect-pollinated, have short blooming times, and lack any kind of nectaries [5, 6]. The flowering unit (inflorescence) of Commelinaceae is a many-branched thyrse, with each branch generally consisting of a many-flowered cincinnus. The cincinni can sometimes be 1-flowered or, more rarely, the whole inflorescence can be reduced to a single flower [4, 7].
Previous classifications of Commelinaceae emphasized floral and anatomical characters. In the first classification, Commelinaceae were divided into two tribes, Commelineae and Tradescantieae, based on the number of stamens and their fertility [8]. Then, Bruckner [9] used flower symmetry, and Pichon [10] used anatomical characters to exclude Cartonema R. Br. from Commelinaceae. In 1966, 15 genera of Commelinaceae were defined using various floral characters [11]. In the current classification, Commelinaceae were divided into two subfamilies, Cartonematoideae (Pichon) Faden ex G. C. Tucker and Commelinoideae Faden & D. R. Hunt, based on the presence of raphide-canals and glandular microhairs [4]. Cartonematoideae consists of two genera (Cartonema and Triceratella Brenan), whereas Commelinoideae includes 39 genera, divided into two tribes, Commelineae (Meisn.) Faden & D. R. Hunt and Tradescantieae (Meisn.) Faden & D. R. Hunt, based on palynological characters. The latter tribe was arranged into seven subtribes based on morphological and cytological characters [4, 12]. However, it is difficult to interpret relationships among genera due to their morphological variation. The morphology-based phylogeny was highly homoplasy and incongruent with the current classification [13]. In order to clarify the relationships within Commelinaceae, several phylogenetic studies have been conducted [14,15,16,17,18,19,20]. Based solely on the plastidial rbcL marker, Cartonema was recovered in a basal clade, and both Commelineae and Tradescantieae were monophyletic, except for the position of Palisota Rchb., which had low support values [15]. Furthermore, the plastidial ndhF suggested that subtribe Tradescantiinae was paraphyletic, whereas Thyrsantheminae and Dichorisandrinae were polyphyletic [16]. Combined data of nuclear 5S NTS and plastid trnL-F regions resulted in a well-supported relationship between Commelineae and Tradescantieae. However, the position of Palisota and Spatholirion Ridl. were ambiguous [17].
Chloroplast genome or plastid genome (cpDNA) is highly conserved and has a typical quadripartite structure containing a large single copy (LSC) and a small single copy (SSC) separated by two inverted repeats (IRs). The size of cpDNA ranges from 19,400 bp (Cytinus hypocistis) to 242,575 bp (Pelargonium transvaalense) and generally contains 120–130 genes, which perform important roles in photosynthesis, translation, and transcription [21, 22]. The rapid development of next-generation sequencing (NGS) has enabled many studies with high-quality complete plastid genomes with raw reads at low costs. Due to its conserved characteristics, chloroplast protein-coding genes were used to reconstruct the phylogenetic relationships in other monocot groups [23,24,25]. Furthermore, these data are useful to infer biogeography, molecular evolution, and age estimation [26,27,28]. The aims of this study are to 1) explore the genome evolution in Commelinaceae subfamily Commelinoideae through analyses of sequence variation, and gene content and order; 2) find latent phylogenetically informative genes through high nucleotide diversity; 3) reconstruct the phylogenetic relationships among members of Commelinoideae with other monocot groups using 77 chloroplast protein-coding genes data, especially the relationships among the six subtribes of Tradescantieae.
Results
Chloroplast genome assembly and annotation
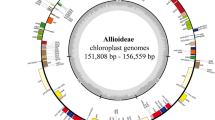
We completed 15 new plastid genomes in this study listed in Table 1 through 9 to 21 million raw reads for each species (Fig. S1, Table S1). A total of 16 plastid genomes, including Belosynapsis ciliata, exhibit the typical quadripartite structure containing LSC and SSC regions separated by two inverted repeats (Fig. 1). Plastid genome sequences of Murdannia edulis and B. ciliata are over 170 kb in length whereas that of Commelina communis is 160,116 bp in length (Table 1). In addition, M. edulis has the lowest GC content (34.4%), whereas Palisota barteri has the highest GC content (36.2%) (Table 1). The highest length difference (about 8801 bp) was observed in the LSC region, between B. ciliata and C. communis. GC content in the SSC region was about 3.4% between Dichorisandra thyrsiflora and M. edulis (Table 1). Plastid genomes of Commelinoideae have 131 genes, of which 111 are unique, and 20 are duplicated in the IR regions (Table 2), except for the rpl22 gene, which was not duplicated in tribe Tradescantieae. There are 77 protein-coding genes (CDS), 30 transfer RNA (tRNA) genes and four ribosomal RNA (rRNA) genes in examined Commelinoideae taxa (Table 2). In these genes, three CDS (rps12, clpP, and ycf3) have two introns, while nine CDS (atpF, ndhA, ndhB, petB, petD, rpl2, rpl16, rpoC1, and rps16) and six tRNA (trnK-UUU, trnG-UCC, trnL-UAA, trnV-UAC, trnI-GAU, and trnA-UGC) have one intron (Table 2). The rps12 gene was trans-spliced, which has the 5′ exon in the LSC region and the 3′ exon and intron in the IR regions. Three pseudogenes (accD, rpoA, and ycf15) were identified from all Commelinoideae species, one (ycf15) of which was duplicated in the IR regions (Table 2). These three genes contained several internal stop codons due to insertions and deletions, thus are identified as pseudogenes. Also, we identified ndhB as a pseudogene in two species (Pollia japonica and Rhopalephora scaberrima) due to point mutation.
Comparative chloroplast genome structure and nucleotide diversity
The aligned data of whole plastid genomes showed high similarities in coding genes, and high variations in non-coding genes (Fig. 2). We found several genome structure variations among Commelinoideae species. M. edulis and Streptolirion volubile had one inversion from rbcL to psaI intergenetic spacer (approximately 3 kb) and petN to trnE-UUC (approximately 2.8 kb), respectively. Amischotolype hispida and B. ciliata had two large inversions from trnV-UAC to rbcL and psbJ to petD about approximately 5 kb and 16 kb, respectively. The IR-SSC boundary was similar among species of Commelinoideae (Fig. 3). All plastid genomes have an incompletely duplicated ycf1 gene in the IRB-SSC junctions. We also found an expansion of IR regions in tribe Commelineae, which resulted in the duplication of the rpl22 genes (Fig. 3).
We analyzed nucleotide divergences of CDS, tRNA, and rRNA to explain variant characteristics among the 16 Commelinoideae plastid genomes (Fig. 4, Table S3). Nucleotide diversity (Pi) for each CDS ranges from 0.00427 (psbL) to 0.09543 (ycf1) with an average of 0.03473. Nine CDS (rps3, ndhG, ndhD, ccsA, rps15, rpl32, ndhF, matK, and ycf1) have remarkably high values (Pi > 0.05) and seven CDS (psbL, rpl23, rps19, ndhB, rpl2, rps7, rps12) have low values (Pi < 0.01; Fig. 4). Compared with tribe Tradescantieae, Commelineae have higher values in 59 out of 77 CDS (Fig. 4). The rpl22 gene has the highest difference of values between Commelineae (Pi = 0.01499) and Tradescantieae (Pi = 0.04655). In the tRNA and rRNA regions, Pi values range from 0 (trnT-UGU, trnH-GUG, trnV-GAC, and trnI-GAU) to 0.02697 (trnQ-UUG), with an average of 0.006. Commelineae has the highest value in the trnL-UAA (Pi = 0.02941), while Tradescantieae has no value in this gene. We tried to find latent phylogenetically informative genes for the Commelinoideae by checking individual CDS with high values (Pi > 0.045) and over 500 bp length. Ten CDS (ndhH, rpoC2, ndhA, rps3, ndhG, ndhD, ccsA, ndhF, matK, and ycf1) were checked with a ML analysis and compared positions among 16 genera of Commelinoideae (Fig. 5). Four CDS (ndhH, rpoC2, matK, and ycf1) have similar topology in Commelinoideae even though the other monocot groups were unclear.
The Maximum Likelihood tree of 42 monocots inferred from 77 chloroplast protein-coding genes. Numbers indicate support (maximum parsimony bootstrap (PBP)/maximum likelihood bootstrap (MBP)/posterior probability (PP)). Only support under PBP = 90/MBP = 100/PP = 1.00 is shown. The dashes “-” indicate incongruence between MP and ML/BI trees
Phylogenetic analysis
The aligned 77 chloroplast protein-coding genes had 65,481 bp, of which 16,380 were parsimony informative. The MP analysis produced single most-parsimonious tree (tree length = 72,586, CI = 0.488, RI = 0.626). The tree topologies of the MP, ML, and BI analyzes were found to be congruent with 100% bootstrap (PBP, MBP) values and 1.00 Bayesian posterior probabilities (PP) in almost all nodes, except for Palisota, which was unresolved in MP analysis (not shown) (Fig. 5). The result suggested that Palisota was sister to the group consisting of the rest of Commelinoideae (Fig. 5). In Tradescantieae, Streptoliriinae was positioned at the basal node. Then, Dichorisandrinae divided into two clades ((Dichorisandra, Siderasis), (Cochliostema, Geogenanthus)) with relatively low support values in both MP and ML analysis (PBP = 77, MBP = 84, PP = 1) (Fig. 5). Among the remaining three subtribes, where two clades ((Coleotrypinae and Cyanotinae), (Tradescantiinae)) were formed with high support values (PBP = 100, MBP = 100, PP = 1), respectively (Fig. 5).
Discussion
Chloroplast genome structure
In this study, we completed 15 new plastid genomes of Commelinoideae taxa (Table 1). Plastid genomes have typical quadripartite structures, including LSC, SSC and two IR regions. Plastid genomes of Commelinoideae have variable total length and GC content. The LSC and SSC regions are relatively longer and higher AT-content than the IR region (Table 1). The functions of AT-rich sequences in the plastid genome were known as enhancing gene transfer success by making stable transcripts [29]. However, AT-rich sequences caused structural variations like inversions by their weak hydrogen bonding. In this study, we identified small to large inversions in four species (Fig. 2). There is one inversion in M. edulis and S. volubile, and two inversions in A. hispida and B. ciliata (Fig. 2). Inversions are known as common genomic rearrangement events and provide informative infrageneric relationships. In previous studies, inversions were caused by microhomology-driven recombination via short repeats and suggested the monophyly of tribe Desmodieae (Fabaceae) [30]. Our results also suggest that both Amischotolype and Belosynapsis have two large inversions in the same loci and formed a clade sister to subtribe Dichorisandrinae (Fig. 5).
We identified an IR expansion in members of Commelineae (Commelina, Murdannia, Pollia, and Rhopalephora). Four species have one more rpl22 gene, which is duplicated in the terminal IR regions (Fig. 3). Although IR expansion affected gene composition, the IR region’s total length is similar among 16 Commelinoideae species. IR expansion and contraction are important events in several families. In Ranunculaceae, IR expansion was detected as a synapomorphy of tribe Anemoneae [31]. Likewise, IR expansion lent further support to the relationship between two subfamilies Ehrhartoideae and Pooideae (Poaceae) [32]. This event also may be phylogenetically informative in Commelinoideae since only members of tribe Commelineae sharing this genome variation (Fig. 5).
Within Commelinoideae plastid genomes, three protein-coding genes (accD, rpoA, and ycf15) were classified as pseudogenes (Fig. S2). The ycf15 gene has several abnormal stop codons caused by insertions and deletions (indel) of bases similar to other monocots. We also identified that all examined species have indels at the frontal part of the accD gene (until 400 bp) and the terminal part of the rpoA gene (after 700 bp; Fig. S2). The accD gene, encoding the beta-carboxyl transferase subunit of acetyl-CoA carboxylase, is found in most flowering plants and synthesizes fatty acids within the chloroplast. It was suggested as an essential gene associated with maintaining chloroplast structure [33]. However, it was reported as a gene loss or pseudogenization in Acoraceae and Poaceae [34, 35]. Recent studies suggested that the accD gene was found to be nuclear originated in several eudicots [36, 37]. The rpoA gene, which encodes the alpha subunit of RNA polymerase, is also found in most flowering plants but was recorded to having been lost in the chloroplast genome of mosses [38]. In one species, Physcomitrella patens (Funariaceae), the rpoA gene was transferred to the nucleus [39]. We need further studies to confirm whether these two genes have been transferred to the nucleus or not in Commelinaceae. We identified that the pseudogened accD and rpoA only appeared in Commelinoideae among Commelinales. It might be a specific character of gene composition in Commelinales. We also found a point mutated base in the third codon of the ndhB gene in P. japonica and R. scaberrima, which formed a clade in this study (Fig. 5).
We measured the nucleotide diversity of CDS, tRNA, and rRNA to identify the genetic divergence between 16 Commelinoideae plastid genomes. We found that the CDS in the IR regions have lower nucleotide diversity than that of the LSC and SSC regions (Fig. 4). This result has also been identified in the other monocots [40,41,42]. It is possibly attributed to a copy correction of the IR regions via gene conversion [43]. Especially, we can see this result in the rpl22 gene. Only Commelineae species present a duplicated rpl22 gene due to the above-mentioned IR expansion, while the remaining 12 taxa have one gene in the LSC or LSC-IR junction (Fig. 3). Difference of nucleotide diversity in this gene between Commelineae (Pi = 0.015) and Tradescantieae (Pi = 0.0466) is 0.0316. It might be phylogenetically useful information for Tradescantieae only.
Implications of plastomes data for phylogenetic reconstructions
The first phylogenetic analysis of Commelinaceae based on rbcL marker revealed a relationship of 32 species representing 30 genera of Commelinaceae [15]. Cartonematoideae was in a basal clade, sister to Commelinoideae and all remaining species [15]. Aside from Palisota, Commelinoideae was divided into two tribes, Commelineae and Tradescantieae, with low bootstrap support values due to insufficient information [15]. Although several phylogenetic studies were conducted, the relationships between the genera of Commelinaceae have remained unresolved. The position of Palisota had been problematic, being recovered as: 1) sister to all genera of Commelinoideae with high bootstrap values [15]; 2) low bootstrap support value with other members of Tradescantieae [16]; or 3) sister to tribe Commelineae [19]. Subtribe Streptoliriinae was recovered as sister to tribe Commelineae in the trnL-trnF analysis [17]. Finally, subtribe Dichorisandrinae seemed polyphyletic in the previous studies [15, 16, 19, 44]. These results are most likely due to limited taxon sampling and/or used few informative genetic markers. The aligned 77 chloroplast coding genes in this study suggest a more well-supported relationship between the genera (Fig. 5). We identified that Commelinoideae divided into two clades, tribe Commelineae and Tradescantieae, with high support values (Fig. 5). However, Palisota, which belongs to Tradescantieae in the current classification [3], is recovered by us as sister to tribe Commelineae (Fig. 5). The ML and BI results present high support values, even though this relationship is unresolved in MP (data not shown). Compared with the current classification, it seems like that subsidiary cells in the stomata and exine morphology are homoplastic to divide two tribes in Commelinoideae [3]. In the Commelinaceae, Palisota is unusual genus for its unique morphological characters like a fleshy berry as a fruit, stamen and staminode arrangement, complex reproductive system, and a basic number of chromosome (x = 20) [13, 45]. Zygomorphic androecium character places Palisota within the Commelineae clade in the morphological cladistic analysis [13]. However, this character is also homoplastic within Commelinoideae. Further research is needed to suggest appropriate characters for Palisota. The four species of Commelineae sampled by us are recovered with a relationship similar to previous studies [15]: (Murdannia, (Commelina, (Pollia, Rhopalephora))). Within Tradescantieae, Streptoliriinae diverged first, followed by Dichorisandrinae divided into two clades with relatively low support values (PBP = 77/MBP = 84/PP = 1) (Fig. 5). The clade composed by Coleotrypinae and Cyanotinae is recovered following the diversion of subtribe Dichorisandrinae, which is sister to Tradescantiinae sensu Pellegrini, [46]. Interestingly, the Asian and African subtribe Coleotrypinae and Cyanotinae were nested well within the New World subtribes (Fig. 5). This result is similar to previous studies and support the hypothesis that one shift from the Old World to the New World followed by dispersal back to the Old World [15, 16].
Conclusions
Our study revealed genome structural characteristics, nucleotide diversity, improved relationships between genera using 15 newly complete chloroplast genomes of Commelinoideae. Compared with other Commelinales, we found two characteristic pseudogenes in all members of Commelinoideae, which might be a synapomorphy within the order. We also reconstruct the phylogenetic relationships using 77 chloroplast protein-coding genes. Although not being able to address the Commelinaceae as a whole, due to not sampling of subfamily Cartonematoideae, we have been able to recover well-supported relationships for the taxa of Commelinoideae, especially between the subtribes of Tradescantieae. One interesting result was that Palisota (subtribe Palisotinae) is more closely related to tribe Commelineae than the remaining members of tribe Tradescantieae. In the current classification, Palisota is a member of Tradescantieae according to the number of subsidiary cells in stomata and pollen exine lacking spines [3]. However, it seems like that these characters are homoplastic, so we need a further study to suggest appropriate characters for two tribes in Commelinoideae. We resolved the ambiguous position of Streptoliriinae which was placed with Commelineae group [17]. Also, Dichorisandrinae was monophyletic in this study which was polyphyletic in the previous studies [15, 16, 19]. Four genes (ndhH, rpoC2, matK, and ycf1) are congruent with the tree estimated from the 77 protein-coding genes. These genes will be helpful to reconstruct relationships of the whole Commelinaceae in the future. Future studies might use the information of chloroplast genomes, relationship between genera to define new classification of Commelinaceae that we provided in this study. These data will make sure the historical biogeography and genome evolution of Commelinaceae.
Materials and methods
Taxon sampling and DNA extraction
Fresh leaf samples were collected in the field and dried directly with silica gel in room temperature until DNA extraction (Table 1). The samples covered four out of 14 genera in tribe Commelineae and 11 out of 25 genera, including six subtribes of tribe Tradescantieae. We prepared the voucher specimens for all used samples and deposited them in the Gachon University Herbarium (GCU) with their accession numbers. We used a modified CTAB method to extract total DNA [47] and checked quality using a spectrophotometer (Biospec-nano; Shimadzu) and assessed by agarose gel electrophoresis.
Genome sequencing, assembly, and annotation
Next-generation sequencing (NGS) was conducted using the Illumina MiSeq sequencing system (Illumina, Seoul, Korea). We imported NGS raw data and trimmed the ends limited to a 5% error probability to remove poor quality reads using Geneious prime 2020.1.2 [48]. Then, we performed ‘map to reference’ using the Hanguana malayana chloroplast genome (GenBank accession = NC_029962.1) as a reference to isolate cpDNA reads. De novo assembly was implemented to reassemble reads using Geneious prime 2020.1.2 [48]. We used newly generated sequences as a reference to reassemble raw reads. We repeated this step until quadripartite structures were completed. Gaps were filled by Sanger sequencing using specific primers. Gene content and order were annotated using H. malayana as a reference using 80% similarity to identify genes in Geneious. All tRNAs were checked by tRNAScan-SE [49] with default search mode. Illustrations of plastomes were produced using OGDraw [50].
Comparative genome analysis
We compared genome structure, size, gene content across all 16 species including B. ciliata (GenBank accession = MK133255.1), subtribe Cyanotinae. The GC content was calculated and compared using Geneious. The whole chloroplast genome sequences of Commelinoideae species were aligned using MUSCLE embedded in Geneious and visualized using LAGAN mode in mVISTA [51, 52]. For the mVISTA plot, we used the annotated cpDNA of H. malayana as a reference. We also examined the nucleotide diversity (Pi) of chloroplast protein-coding genes, transfer RNA genes and ribosomal RNA genes among the 16 Commelinoideae species through a sliding window analysis using DnaSP v. 6.0 [53]. For the sequence divergence analysis, we applied the window size of 100 bp with a 25 bp step size. The IR and SC boundaries of the 16 Commelinoideae species were compared and illustrated using IRscope [54].
Phylogenetic analysis
A total of 42 chloroplast genome sequences (including 15 new chloroplast genomes of Commelinoideae) were used (Table S2). We extracted 77 protein-coding genes and aligned them using the MUSCLE embedded in Geneious prime 2020.1.2 [48]. For the data set, Acorus calamus (Acoraceae) was designated as an outgroup. We performed maximum parsimony (MP), maximum likelihood (ML), and Bayesian inference (BI) to infer relationships of Commelinoideae and related taxa. The MP analyses were carried out in PAUP* v4.0a [55] with all characters equally weighted and unordered. Gaps were treated as missing data. Searches of 1000 random taxon addition replicates used tree-bisection-reconnection (TBR) branch swapping, and MulTrees permitted ten trees to be held at each step. Bootstrap analyses (PBP, parsimony bootstrap percentages, 1000 pseudoreplicates) were conducted to examine internal support with the same parameters. We used jModelTest version 2.1.7 [56, 57] to find the best model with Akaike’s information criterion (AIC) before running the ML and BI analyses. The GTR + I + G was the best model for the concatenated data sets. We used the IQ-TREE web server (http://iqtree.cibiv.univie.ac.at/) to make the ML searches [58]. Support value (MBP, mean bootstrap percentage) was calculated with 1000 replicates of ultrafast bootstrap [59]. MrBayes v3.2.7 [60] was used for BI analyses. Two simultaneous runs were performed starting from random trees for at least 1,000,000 generations. One tree was sampled every 1000 generations. In total, 25% of trees were discarded as burn-in samples. The remaining trees were used to construct a 50% majority-rule consensus tree, with the proportion bifurcations found in this consensus tree given as posterior probability (PP) to estimate the robustness of half of the BI tree. The effective sample size values (ESS) were then checked for model parameters (at least 200). The phylogenetic trees were edited using FigTree v1.4.4 program [61].
Availability of data and materials
The 15 chloroplast genomes sequences we obtained from this study were archived in NCBI. The accession numbers are presented in Table 1.
References
Chase MW, Christenhusz M, Fay M, Byng J, Judd WS, Soltis D, et al. An update of the angiosperm phylogeny group classification for the orders and families of flowering plants: APG IV. Bot J Linn Soc. 2016;181(1):1–20.
Christenhusz MJM, Byng JW. The number of known plants species in the world and its annual increase. Phytotaxa. 2016;261(3):201. https://doi.org/10.11646/phytotaxa.261.3.1.
Faden RB, Hunt D. The classification of the Commelinaceae. Taxon. 1991;40(1):19–31. https://doi.org/10.2307/1222918.
Faden RB. Commelinaceae. In: Flowering Plants· Monocotyledons. Washington: Springer; 1998;4:109–28.
Owens S. Self-incompatibility in the Commelinaceae. Ann Bot. 1981;47(5):567–81. https://doi.org/10.1093/oxfordjournals.aob.a086054.
Faden RB. Floral attraction and floral hairs in the Commelinaceae. Ann Mo Bot Gard. 1992;79(1):46–52. https://doi.org/10.2307/2399808.
Panigo E, Ramos J, Lucero L, Perretta M, Vegetti A: The inflorescence in Commelinaceae. Flora - Morphology, Distribution, Functional Ecology of Plants 2011, 206(4):294–299, doi: https://doi.org/10.1016/j.flora.2010.07.003.
Meisner CF. CCLXI Commelinaceae. Plantarum vascularium genera. 1842;1:406–7.
Bruckner G. Beiträge zur anatomie morphologie und systematik der Commelinaceae; 1926.
Pichon MJNS. Sur les Commelinaces. 1946;12:217–42.
Brenan JP. The classification of Commelinaceae. Bot J Linn Soc. 1966;59(380):349–70. https://doi.org/10.1111/j.1095-8339.1966.tb00068.x.
Hardy CR, Faden RB. Plowmanianthus, a new genus of Commelinaceae with five new species from tropical America. Syst Bot. 2004;29(2):316–33. https://doi.org/10.1600/036364404774195511.
Evans TM, Faden RB, Simpson MG, Sytsma KJ. Phylogenetic relationships in the Commelinaceae: IA cladistic analysis of morphological data. Syst Bot. 2000;25(4):668–91. https://doi.org/10.2307/2666727.
Bergamo S. A phylogenetic evaluation of Callisia Loefl. (Commelinaceae) based on molecular data. Athens: uga; 2003.
Evans TM, Sytsma KJ, Faden RB, Givnish TJ. Phylogenetic relationships in the Commelinaceae: II. A cladistic analysis of rbcL sequences and morphology. Syst Bot. 2003;28(2):270–92.
Wade DJ, Evans TM, Faden RB. Subtribal relationships in tribe Tradescantieae (Commelinaceae) based on molecular and morphological data. Aliso. 2006;22(1):520–6. https://doi.org/10.5642/aliso.20062201.40.
Burns JH, Faden RB, Steppan SJ. Phylogenetic studies in the Commelinaceae subfamily Commelinoideae inferred from nuclear ribosomal and chloroplast DNA sequences. Syst Bot. 2011;36(2):268–76. https://doi.org/10.1600/036364411X569471.
Zuiderveen GH, Evans TM, Faden RB. A phylogenetic analysis of the African plant genus Palisota (family Commelinaceae) based on chloroplast DNA sequences; 2011.
Hertweck KL, Pires JC. Systematics and evolution of inflorescence structure in the Tradescantia Alliance (Commelinaceae). Syst Bot. 2014;39(1):105–16. https://doi.org/10.1600/036364414X677991.
Kelly SM, Evans TM. A phylogenetic analysis of the African plant genus Aneilema (family Commelinaceae) based on chloroplast DNA sequences; 2014.
Dyer TA. The chloroplast genome: its nature and role in development. Topics in photosynthesis. 1984;5:23–69.
Sugiura M. The chloroplast genome. Plant Mol Biol. 1992;19(1):149–68.
Kim JH, Kim DK, Forest F, Fay MF, Chase MW. Molecular phylogenetics of Ruscaceae sensu lato and related families (Asparagales) based on plastid and nuclear DNA sequences. Ann Bot. 2010;106(5):775–90. https://doi.org/10.1093/aob/mcq167.
Barrett CF, Baker WJ, Comer JR, Conran JG, Lahmeyer SC, Leebens‐Mack JH, Li J, Lim GS, Mayfield‐Jones DR, Perez L. Plastid genomes reveal support for deep phylogenetic relationships and extensive rate variation among palms and other commelinid monocots. New Phytol. 2016;209(2):855–70.
Do HDK, Kim C, Chase MW, Kim JH. Implications of plastome evolution in the true lilies (monocot order Liliales). Mol Phylogenet Evol. 2020;148:106818. https://doi.org/10.1016/j.ympev.2020.106818.
Jones SS, Burke SV, Duvall MR. Phylogenomics, molecular evolution, and estimated ages of lineages from the deep phylogeny of Poaceae. Plant systematics and evolution. 2014;300(6):1421–36.
Li Q-Q, Zhou S-D, Huang D-Q, He X-J, Wei X-Q. Molecular phylogeny, divergence time estimates and historical biogeography within one of the world’s largest monocot genera. AoB Plants. 2016;8:plw041.
Kim C, Kim S-C, Kim J-H. Historical biogeography of Melanthiaceae: a case of out-of-North America through the Bering land bridge. Front Plant Sci. 2019;10:396.
Stegemann S, Bock R. Experimental reconstruction of functional gene transfer from the tobacco plastid genome to the nucleus. Plant Cell. 2006;18(11):2869–78.
JJin D-P, Choi I-S, Choi B-H. Plastid genome evolution in tribe Desmodieae (Fabaceae: Papilionoideae). PloS one. 2019;14(6):e0218743.
He J, Yao M, Lyu R-D, Lin L-L, Liu H-J, Pei L-Y, Yan S-X, Xie L, Cheng J. Structural variation of the complete chloroplast genome and plastid phylogenomics of the genus Asteropyrum (Ranunculaceae). Sci Rep. 2019;9(1):1–13.
Guisinger MM, Chumley TW, Kuehl JV, Boore JL, Jansen RK. Implications of the plastid genome sequence of Typha (Typhaceae, Poales) for understanding genome evolution in Poaceae. J Mol Evol. 2010;70(2):149–66.
Kode V, Mudd EA, Iamtham S, Day A. The tobacco plastid accD gene is essential and is required for leaf development. Plant J. 2005;44(2):237–44.
Goremykin VV, Holland B, Hirsch-Ernst KI, Hellwig FH. Analysis of Acorus calamus chloroplast genome and its phylogenetic implications. Mol Biol Evol. 2005;22(9):1813–22.
HHarris ME, Meyer G, Vandergon T, Vandergon VO. Loss of the acetyl-CoA carboxylase (accD) gene in Poales. Plant Mol Biol Rep. 2013;31(1):21–31.
Rousseau-Gueutin M, Huang X, Higginson E, Ayliffe M, Day A, Timmis JN. Potential functional replacement of the plastidic acetyl-CoA carboxylase subunit (accD) gene by recent transfers to the nucleus in some angiosperm lineages. Plant Physiol. 2013;161(4):1918–29.
Li J, Gao L, Chen S, Tao K, Su Y, Wang T. Evolution of short inverted repeat in cupressophytes, transfer of accD to nucleus in Sciadopitys verticillata and phylogenetic position of Sciadopityaceae. Sci Rep. 2016;6(1):1–12.
Goffinet B, Wickett NJ, Shaw AJ, Cox CJ. Phylogenetic significance of the rpoA loss in the chloroplast genome of mosses. Taxon. 2005;54(2):353–60.
Sugiura C, Kobayashi Y, Aoki S, Sugita C, Sugita M. Complete chloroplast DNA sequence of the moss Physcomitrella patens: evidence for the loss and relocation of rpoA from the chloroplast to the nucleus. Nucleic Acids Res. 2003;31(18):5324–31.
Lee SR, Kim K, Lee BY, Lim CE. Complete chloroplast genomes of all six Hosta species occurring in Korea: molecular structures, comparative, and phylogenetic analyses. BMC Genomics. 2019;20(1):833. https://doi.org/10.1186/s12864-019-6215-y.
Huang J, Yu Y, Liu YM, Xie DF, He XJ, Zhou SD. Comparative Chloroplast Genomics of Fritillaria (Liliaceae), Inferences for Phylogenetic Relationships between Fritillaria and Lilium and Plastome Evolution. Plants (Basel). 2020;9(2):133.
Smidt EC, Paez MZ, Vieira LDN, Viruel J, de Baura VA, Balsanelli E, et al. Characterization of sequence variability hotspots in Cranichideae plastomes (Orchidaceae, Orchidoideae). PLoS One. 2020;15(1):e0227991. https://doi.org/10.1371/journal.pone.0227991.
Khakhlova O, Bock R. Elimination of deleterious mutations in plastid genomes by gene conversion. Plant J. 2006;46(1):85–94. https://doi.org/10.1111/j.1365-313X.2006.02673.x.
Pellegrini MOO, Faden RB. Recircumscription and taxonomic revision of Siderasis, with comments on the systematics of subtribe Dichorisandrinae (Commelinaceae). PhytoKeys. 2017;83(83):1–41. https://doi.org/10.3897/phytokeys.83.13490.
Tomlinson P. Anatomical data in the classification of Commelinaceae. Bot J Linn Soc. 1966;59(380):371–95. https://doi.org/10.1111/j.1095-8339.1966.tb00069.x.
Pellegrini MOO. Morphological phylogeny of Tradescantia L. (Commelinaceae) sheds light on a new infrageneric classification for the genus and novelties on the systematics of subtribe Tradescantiinae. PhytoKeys. 2017;89(89):11–72. https://doi.org/10.3897/phytokeys.89.20388.
Doyle JJ, Doyle JL. CTAB DNA extraction in plants. Phytochemical Bulletin. 1987;19:11–5.
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinform. 2012;28(12):1647–9.
Chan PP, Lowe TM. tRNAscan-SE: searching for tRNA genes in genomic sequences. Gene Prediction. 2019. p. 1–14.
Greiner S, Lehwark P, Bock R. OrganellarGenomeDRAW (OGDRAW) version 1.3. 1: expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 2019;47(W1):W59–64.
Brudno M, Do CB, Cooper GM, Kim MF, Davydov E, Green ED, Sidow A, Batzoglou S. LAGAN and Multi-LAGAN: efficient tools for large-scale multiple alignment of genomic DNA. Genome Res. 2003;13(4):721–31.
Frazer KA, Pachter L, Poliakov A, Rubin EM, Dubchak I. VISTA: computational tools for comparative genomics. Nucleic Acids Res. 2004;32(suppl_2):W273–9.
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol. 2017;34(12):3299–302.
Amiryousefi A, Hyvönen J, Poczai P. IRscope: an online program to visualize the junction sites of chloroplast genomes. Bioinform. 2018;34(17):3030–1.
Swofford DJPaupS, Sunderland: PAUP* 4.0 b. 4a. 2000.
Guindon S, Gascuel O. A simple, fast and accurate method to estimate large phylogenies by maximum-likelihood. Syst Biol. 2003;52:696–704.
Darriba D, Taboada GL, Doallo R, Posada DJ. 2: More models, new heuristics and high-performance computing. Nat Methods. 2012;9:772.
Trifinopoulos J, Nguyen L-T, von Haeseler A, Minh BQ. W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016;44(W1):W232–5.
Stamatakis A, Hoover P, Rougemont J. A rapid bootstrap algorithm for the RAxML web servers. Syst Biol. 2008;57(5):758–71.
Ronquist F, Teslenko M, Van Der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. 2012;61(3):539–42.
Rambaut A. FigTree V: 1.4. 4; 2018.
Acknowledgments
We would like to thank Gerardo A. Salazar at Universidad Nacional Autónoma de México (Mexico), Claudia T. Hornung-Leoni at Autonomous University of Hidalgo (Mexico), Manuel González Ledesma at Autonomous University of Hidalgo (Mexico), Kenneth M. Cameron at University of Wisconsin–Madison (United States of America), Chien-Ti Chao at National Taiwan Normal University (Taiwan), David Warmington at Cairns Botanic Gardens (Australia), Carlos Gustavo Espejo Zurita at Jardín Botánico Histórico La Concepción (Spain) for collecting and providing the plant material for this study.
Funding
This work was supported by the Gachon University research fund of 2019(GCU-2019-0821) and the National Research Foundation of Korea (NRF) Grant Fund (NRF-2017R1D1A1B06029326).
Author information
Authors and Affiliations
Contributions
JJ performed the experiments, analyzed the data, prepared figures and/or tables, authored or reviewed drafts of the paper, approved the final draft. CK authored or reviewed drafts of the paper, approved the final draft. JHK conceived and designed the experiments, contributed reagents/ materials/ analysis tools, authored or reviewed drafts of the paper, approved the final draft.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
We received fresh leaf materials of Callisia repens (Jardín Botánico Histórico La Concepción, Spain), Cochliostema odoratissimum (Cairns Botanic Gardens, Australia), and Siderasis fuscata (Ghent University Botanical Garden, Belgium) for this study. We prepared the voucher specimens for all used samples and deposited them in the Gachon University Herbarium (GCU) with their accession numbers (Table 1). The study including plant samples complies with relevant institutional, national, and international guidelines and legislation. No specific permits were required for plant collection. The study did not require ethical approval or consent, as no endangered or protected plant species were involved.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Table S1.
List of sampling taxa from 15 species of Commelinaceae and assembly information. Table S2. List of species used for phylogenomic analyses. Table S3. Nucleotide diversity (Pi) of 16 Commelinoideae species. Figure S1. Complete chloroplast genome of 15 Commelinaceae taxa in this study. Figure S2. Amino acid alignment of plastid accD and rpoA genes within 22 Commelinales taxa.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Jung, J., Kim, C. & Kim, JH. Insights into phylogenetic relationships and genome evolution of subfamily Commelinoideae (Commelinaceae Mirb.) inferred from complete chloroplast genomes. BMC Genomics 22, 231 (2021). https://doi.org/10.1186/s12864-021-07541-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-021-07541-1