Abstract
Background
So far many algorithms have been proposed towards the detection of significant genes in microarray analysis problems. Several of those approaches are freely available as R-packages though their engagement in gene expression analysis by non-bioinformaticians is usually a frustrating task. Besides, only some of those packages offer a complete suite of tools starting from initial data import and ending to analysis report. Here we present an R/Bioconductor package that implements a hybrid gene selection method along with a bunch of functions to facilitate a thorough and convenient gene expression profiling analysis.
Results
mAPKL is an open-source R/Bioconductor package that implements the mAP-KL hybrid gene selection method. The advantage of this method is that selects a small number of gene exemplars while achieving comparable classification results to other well established algorithms on a variety of datasets and dataset sizes. The mAPKL package is accompanied with extra functionalities including (i) solid data import; (ii) data sampling following a user-defined proportion; (iii) preprocessing through several normalization and transformation alternatives; (iv) classification with the aid of SVM and performance evaluation; (v) network analysis of the significant genes (exemplars), including degree of centrality, closeness, betweeness, clustering coefficient as well as the construction of an edge list table; (vi) gene annotation analysis, (vii) pathway analysis and (viii) auto-generated analysis reporting.
Conclusions
Users are able to run a thorough gene expression analysis in a timely manner starting from raw data and concluding to network characteristics of the selected gene exemplars. Detailed instructions and example data are provided in the R package, which is freely available at Bioconductor under the GPL-2 or later license http://www.bioconductor.org/packages/3.1/bioc/html/mAPKL.html.
Similar content being viewed by others
Background
The advent of microarray technology gave rise to the development of a plethora of analytical approaches, where each of them try to detect those significant genes among thousands of genes that are enough to summarize all the necessary statistical and biological information for successful classification and pathway analysis. Several of those methods are implemented as open-source R-packages with the anticipation to become broadly engaged by bioscientists in microarray analysis problems. Though, several factors hinder their extensive application particularly from biologists. In some cases the required computational skills while on other the necessity to integrate results of different formats stemming from several packages are some of those factors.
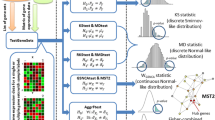
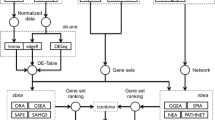
The mAP-KL is a hybrid gene selection method, which is based on the hypothesis that among the statistically significant genes in a ranked list, there should be clusters of genes that share similar biological functions related to the investigated disease [1]. The output of this method is a subset of genes consisting of one exemplar per cluster that best describes the phenotypes’ characteristics. The method has been compared against twelve other well-established feature selection methods (eBayes, SAM, RF, BGA, cat, maxT, PLS-CV, ODP, PCA, SNR, t-test, HykGene), where all of them applied on a variety of diseases including gene expression data of six neuromuscular diseases and four types of cancer. The overall evaluation results suggest that mAP-KL implements an efficient sampling of the ranked gene list, selecting those genes that are necessary for accurate classification while at the same time reflect biological relevance to the respective disease, thus providing a reasonable basis for further biological insights [1].
To provide the research community with the capability to apply mAP-KL in any given gene expression dataset, we have implemented this methodology to an open-source R/Bioconductor package accompanied with extra functionalities such as data sampling preprocessing, classification, network analysis, gene annotation analysis, pathway analysis and reporting. The centric idea during the package’s design was to build functions that can either shape an extensive analysis pipeline or used as standalone modules. For instance, a user may import any dataset of raw gene expression data and apply with a single command eight at maximum different preprocessing methods. Then, may analyze any of the preprocessed data with the mAP-KL method and conclude to lists of significant genes (exemplars). Classification assessment, annotation analysis, pathway analysis and network characteristics are some of the possible analyses that a user may apply on these exemplars. On the other hand, a user may as well employ any of the available functions to exploit a particular functionality for example, to partition a dataset into train and validation sets, to obtain annotation info for a given list of probe ids, and so on.
In the rest of this article, we will present the key design characteristics that influenced the implementation of this package, and next we will present a thorough workflow highlighted the functional modules of mAPKL. A discussion on other competing packages and how they resemble or differ to mAPKL will follow, and finally the future directions of our package will be outlined.
Implementation
The mAPKL has been implemented in R as an S4 package that takes advantage of the rich functionality of the ‘ExpressionSet’ (eSet) class [2]. This type of class is designed to accommodate a variety of information including expression data from microarray experiments (assayData), ‘meta-data’ describing samples in the experiment (phenoData), annotations and meta-data about the features on the chip (featureData, annotation), information about the protocol used for processing the samples (protocolData), and a flexible structure to describe the experiment (experimentData). All those different sources of information are handled by class-methods thus the proper manipulation is guaranteed. Besides, using this class objects throughout this package we make feasible any collaboration with other Bioconductor packages hence, extending the meta-analysis options.
From a functional perspective, we have designed and implemented functions that each one of those addresses a particular operation. Indeed, the loadFiles function imports gene expression data and convert them into eSet class objects. Hence, this function can be used as a standalone importing and conversion mechanism to eSet class objects by a user who wishes to engage other packages that use those class objects. Similarly, a user may engage the preprocess function to apply at maximum eight different normalization/transformation methods on microarray data and then insert the preprocessed data to another algorithm of statistical inference. In the same way, we may exploit separately each of the available modules. On the other hand, the implemented functions combine a pipeline to assist particularly non-bioinformatician users to run a thorough microarray analysis as we will present in the following section.
Results
We will present an analysis scenario displaying the functional modules of the mAPKL (see Additional file 1) with the aid of the ‘mAPKLData’ Bioconductor experiment data package (see Additional file 2). During the development of the mAPKL package we also built the mAPKLData package as a necessary supplement to display all different functionalities. It provides the GSE5764 dataset, which is available at the NCBI Gene Expression Omnibus and includes gene expression data from a breast cancer study published by Turashvili et al. [3] that contains 30 samples related to breast cancer (20 normal and 10 tumor samples), based on Affymetrix HG-U133_Plus_2 microarray platform.
Data import and manipulation
We developed the loadFiles function that takes as input text tab-delimited files and converts them into eSet class objects. Typically, a gene expression analysis requires a pair of files for training and optionally another pair for validation purposes. Each pair includes a file with the gene expressions in a matrix format, where rows represent probe ids and columns represent samples, and a file with the relevant class labels. In our case, the class label file should have a minimum of two columns with headers, for example ‘title’ and ‘type’. In the ‘type’ column we assign the class labels of the samples where ‘0’ stands for control and ‘1’ stands for ‘treatment’ samples. The loadFiles function, supports importing and converting both, training and validation files, in a single command line.
A further data manipulation attribute offered in the mAPKL package is the dataset partition into train and validation sets according to a user defined percentage. In particular, the sampling function takes as input an eSet class objects describing a cohort of samples and split into separate train and test sets following a user’s predefined percentage. The minimum percentage of samples that might be kept for validation purposes is 10 % and the user is always confident that the function creates stratified sets in relation to their class analogy within the initial dataset. Moreover, the samples’ selection follows a random pick based on a pre-defined seed number, which on the one hand guarantees reproducibility while on the other eliminate any bias during the selection. In the current scenario, we loaded the mAPKLData, and then with the aid of the sampling function we kept 40 % of the samples for validation purposes, which produced a train set with twelve normal and six tumor samples, and a validation set with eight normal and four disease samples. Those scenario steps require the following R commands.
-
library(mAPKL)
-
library(mAPKLData)
-
data(mAPKLData)
-
breast < − sampling(Data = mAPKLData, valPercent = 40, classLabels = “type”, seed = 135)
Data normalization and transformation
The package accommodates a practical preprocessing functional unit that supports log2 transformation along with four different normalization methods including mean-centering, z-score, quantile and cyclic loess. In particular, the preprocess function produces an S3 class object, i.e. a list with maximum nine available options, Table 1, as well as a multi-graph image with the density plots of the preprocessed data for each method, Fig. 1. Hence, the user may proceed with a gene selection analysis for any of the preprocessed gene expression values. In the following section, we run eight different scenarios and assess their classification results before reaching to a decision about the most appropriate normalization scheme for the mAPKLData. The following two lines of code apply the described normalization\transformation options instantly to the train and test sets respectively.
-
normTrainData < − preprocess(breast$trainData)
-
normTestData < − preprocess(breast$testData)
Gene selection analysis
The core function of this package is the mAPKL that implements the hybrid feature selection methodology. It takes as input an eSet class object with the training data and several predefined parameters necessary for the intrinsic statistical analysis and clustering. It may also accept a validation eSet object to directly apply on it the results of the mAP-KL analysis. This function returns an object of ‘mAPKLRes’ S4 class where its slots embody the matrix of the top N ranked genes, the clusters and their respective exemplars, the training and validation eSet objects of the exemplars, along with statistical information such as p-value, adjusted p-value and fold-change for all genes. In relation to the gene expression analysis of the mAPKLData, we carried out eight different mAP-KL analyses and concluded to eight different subsets of exemplars, which will be assessed for their discrimination power. In the next lines of code, we employ the gene expression values resulted from the cyclic loess normalization on log2 transformed data, and then apply the mAP-KL method. Similarly we may engage any preprocessed set of gene expression values and ask for their exemplars.
-
exprs(breast$trainData) < − normTrainData$clL2.normdata
-
exprs(breast$testData) < − normTestData$clL2.normdata
-
out.clL2 < − mAPKL(trObj = breast$trainData, classLabels = “type”,
-
valObj = breast$testData, dataType = 7)
Classification
The following functional module provides classification estimates for the selected genes. In particular, it utilizes the exemplars’ eSet objects from the ‘mAPKLRes’ class to run an SVM based cross-validation classification test to quantify the discrimination power of the gene exemplars. The necessary parameters for running the SVM classifier are computed dynamically with the tune.svm function of the ‘e1071’ R-package [4]. The classification measures are calculated through a mAPKL’s function called metrics and include the Area Under the Curve (AUC), the Matthews correlation coefficient (MCC), the accuracy (ACC), the true negative rate (TNR) or specificity and the true positive rate (TPR) or sensitivity. This function performs classification through the SVM algorithm and produces a classification result either on the training set or on a validation set. During this analysis we assessed the performance on the validation set using the following SVM parameters: ‘linear’ kernel and 5-folds cross-validation (although other options are feasible) and the R command is structured as follows:
-
clasPred < − classification(out.clL2@exemplTrain, “type”, out.clL2@exemplTest).
According to the classification results, Table 2, the exemplars’ list produced after log2 transformation and cyclic loess normalization achieved the best discrimination results and will be further explored from a pathway and a network-topology perspectives.
Gene annotation and pathway analysis
The next functional module exploits the microarray chip annotation file, when available, to collect necessary genome oriented information so as to facilitate other types of genome analysis such as pathway analysis. The ‘Annot’ S4 class provides slots for gene ‘symbol’, ‘entrezId’, ‘ensembleId’ and chromosomal location info of the exemplars. Thus, the user not only has at hand a valid conversion mechanism between probes and genes but also several additional meta-data for other types of analysis like pathway or Gene Ontology. For this purpose we run as follows the annotate function with the argument ‘chip’ equal to ‘hgu133plus2.db’ since this is the relevant microarray chip platform for the mAPKLData dataset.
-
gene.info < − annotate(out.clL2@exemplars,“hgu133plus2.db”)
Then, we may easily exploit the ‘Annot’ class object produced by the annotate function to perform a pathway analysis utilizing the ‘Reactome’ pathway database [5] through the probes2pathways function. Particularly, we first remove the ‘NA’ entries and then map the ‘ENTREZID’ to the ‘Reactome’ pathway identifiers, Fig. 2. Thus, for any given set of exemplars a user may effortlessly identify the relevant pathways.
Network characteristics
The netwAttr function deals with the network characteristics of the top N ranked genes but more importantly of the gene exemplars. Three different types of centralities (degree, closeness, betweenness) and a measure for clustering coefficient, called transitivity, are estimated with this function. The degree centrality of a node refers to the number of connections or edges of that node to other nodes. The closeness centrality describes the reciprocal accumulated shortest length distance from a node to all other connected nodes. The betweeness centrality depicts the number of times a node intervenes along the shortest path of two other nodes. Transitivity measures the degree of nodes to create clusters within a network. For all four network measures we provide both global and local values. Moreover, the netwAttr provides a weighted edge list (Node1-Node2-weight) based on the top N ranked genes, as a front end to network and graph packages for further analysis and visualization. Particularly, the netwAttr function computes the network degree, closeness, betweenness and transitivity. We compose an edge list (Node1-Node2-weight) based on the top N ranked genes (200 in this example) to interface with other network related R-packages like igraph [6] or with software tools outside the R platform like Cytoscape [7]. Indeed, we plot a network graph to present the relations-connections among the top N ranked genes, Fig. 3, through Cytoscape. For both significance and illustration purposes we display only the nodes with local weighted degree greater than the global weighted degree plus 2 times the standard deviation. The calculations of those network characteristics are based on the “clr” [8] network reconstruction method. However, there are two more available options, including the “aracne.a” and “aracne.m’ [9] algorithms. The basic syntax for this function is the following, although some additional R commands are necessary to create the described output values for the graph. Details are included in the mAPKL’s vignette, net.attr < − netwAttr(out.clL2).
Report generation
With regard to the report function, the user may produce a summarized report in html format that presents the results in all different stages of analysis. Specifically, the first section depicts the samples' names and their respective class labels, the second section presents the exemplars along with their statistical results and network characteristics, the third section deals with the classification results and illustrates the performance metrics achieved in either cross-validation or hold-out validation. Finally, the fourth section of this report cites the annotation info per exemplar relevant to the chip technology.
Discussion
Comparison with other microarray data analysis software
There are several software tools related to differential gene expression and samples classification. All of those tools can be roughly classified based on two criteria; the software technology used and the scope of the analysis they offer. Regarding the first criterion, software applications can be further divided into Web applications, desktop java applications and R-packages. Concerning the second criterion, there are methods for gene selection, for classification of samples, and methods that engage clustering either for gene selection or for unknown samples categorization. mAPKL package is the implementation of a hybrid gene selection algorithm under the R/Bioconductor framework, which also includes additional functionalities, aiming at providing a thorough gene expression analysis pipeline. In this context, we compared the mAPKL package against the R implementations of the feature selection methods included in the previously published mAP-KL evaluation [1] as well as against other R-packages that offer functionalities related to data sampling, gene selection and classification.
Regarding the R-packages included in the following comparisons, there are seven feature selection and five classification software tools. Particularly the feature selection packages represent the following feature selection approaches: the ‘edge’ [10] package implements the Optimal Discovery Procedure (ODP) [11] feature selection method, the ‘limma’ [12] includes the eBayes [13] approach, the ‘multtest’ [14] provides the maxT [15] procedure, the ‘plsgenomics’ [16] contain the Partial Least Squares (PLS) [17] method, the ‘randomForest’ [18] implements the RF [19] algorithm, the ‘samr’ [20] carries out the Significance Analysis of Microarrays (SAM) [21] method, and the ‘st’ [22] package implements the cat score [23]. As far as the classification packages that most resemble to mAPKL are the ‘caret’ [24], the ‘ClassifyR’ [25], the ‘CMA’ [26], the ‘MCRestimate’ [27], and the ‘MLInterfaces’ [28].
Firstly, the packages have been compared in terms of several functional criteria, Table 3, including ‘Data’ manipulation, ‘Significance analysis’, ‘Classification’, ‘Annotation analysis’, computation of ‘Network characteristics’ and ‘Reporting’. In addition, we have included subcategories in some of those criteria so as to provide an in-depth comparison. Thus, the ‘Data’ category falls into four subcategories related to functions for data importing, normalization and transformation, dataset sampling, and eSet class utilization. Regarding the ‘Classification’ criterion we have distinguished the prediction results from the available performance metrics.
Secondly, we employed the mAPKLData package to compare the classification performance of mAPKL against the other seven feature selection packages. For equality purposes we engaged the same preprocessing approach (the cyclic loess normalization on log2 transformed data), and we decided to keep subsets of genes of the same length based on the 15 “exemplars” of the mAPKL. Besides, we utilized the classification function included in our package to evaluate the top-15 gene subsets from the other seven approaches, and the classification results are presented in Table 4.
The accumulated comparison results suggest that the mAPKL is more than an R implementation of an effective feature selection method or a classification package. It is a complete and comprehensive gene expression analysis package offering a set of functions to assist the genomics researchers to perform a rigorous analysis starting from raw gene expression data and ending to annotated significant genes accompanied with certain evaluation metrics and networking properties, assembling the information in an auto-generated analysis report.
As part of the Bioconductor project, the mAPKL package is freely available under the GPL-2 or later license accompanied with detailed help pages per class and function. Besides, an elaborate vignette introduces all available functionalities through a case study scenario that is based on the ‘mAPKLData’ Bioconductor experiment data package. Thus, the user can see both illustrated codes and executed outputs to become easily accustomed to the package. Moreover, the Bioconductor project guarantees the easy implementation and platform independence, the versioning of the forthcoming package releases and the obliged that the package will be maintained by the author, which includes response to bug reports or queries from other users as well as checking periodically the functionality of the package.
Future directions
In relation to the potential expansions of the mAPKL package, we intend to include alternative methods for gene selection as well methods related to functional enrichment analysis.
Conclusions
We have presented an R/Bioconductor package named mAPKL concerning the implementation of the mAP-KL method for efficient gene selection from gene expression data. Additionally, it incorporates several features including data sampling to create train and validation sets, various preprocessing methods, functions for measuring the classification performance of the selected genes and gene annotation analysis to facilitate other types of analysis like pathway analysis. Moreover, it constructs networks based on the top N genes of our methodology and exploits several network characteristics of the ‘exemplars’ to produce graphical representations of the cellular network topology. In general, it is a user friendly R/Bioconductor package for gene expression data analysis that includes a comprehensive suite of functions. Those functions include default values to facilitate users without advanced computational or statistical background and collaborate smoothly to integrate into a custom analysis pipeline.
Availability and requirements
Project name: mAPKL
Project home page: http://www.bioconductor.org/packages/3.1/bioc/html/mAPKL.html
Operating system(s): Platform independent
Programming language: R
Other requirements:R (> = 3.2.0), Biobase
License:GPL (> = 2)
Any restrictions to use by non-academics: None
References
Sakellariou A, Sanoudou D, Spyrou G. Combining multiple hypothesis testing and affinity propagation clustering leads to accurate, robust and sample size independent classification on gene expression data. BMC Bioinformatics. 2012;13:270.
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004;5(10):R80.
Turashvili G, Bouchal J, Baumforth K, Wei W, Dziechciarkova M, Ehrmann J, et al. Novel markers for differentiation of lobular and ductal invasive breast carcinomas by laser microdissection and microarray analysis. BMC Cancer. 2007;7:55.
Dimitriadou E, Hornik K, Leisch F, Meyer D, Weingessel A. e1071: Misc Functions of the Department of Statistics (e1071). In. TU Wien; 2010. Available from: http://CRAN.R-project.org/package=e1071.
Vastrik I, D’Eustachio P, Schmidt E, Gopinath G, Croft D, de Bono B, et al. Reactome: a knowledge base of biologic pathways and processes. Genome Biol. 2007;8(3):R39.
Csardi G, Nepusz T. The igraph software package for complex network research. InterJournal 2006, Complex Systems:1695.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–504.
Faith JJ, Hayete B, Thaden JT, Mogno I, Wierzbowski J, Cottarel G, et al. Large-scale mapping and validation of Escherichia coli transcriptional regulation from a compendium of expression profiles. PLoS Biol. 2007;5(1):e8.
Margolin AA, Nemenman I, Basso K, Wiggins C, Stolovitzky G, Dalla Favera R, et al. ARACNE: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinformatics. 2006;7 Suppl 1:S7.
Storey JD, Leek JT, Bass AJ. edge: Extraction of Differential Gene Expression. In. Bioconductor version 2.0.0; 2015.
Storey JD. The optimal discovery procedure: a new approach to simultaneous significance testing. J R Stat Soc Ser B. 2007;69(3):347–68.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47.
Smyth GK. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Statistical applications in genetics and molecular biology. 2004;3:Article3.
Pollard KS, Dudoit S, Laan MJvd. Multiple Testing Procedures: the multtest Package and Applications to Genomics. In: Bioinformatics and Computational Biology Solutions Using R and Bioconductor. 2005: 249–271.
van der Laan MJ, Dudoit S, Pollard KS. Multiple testing. Part II. Step-down procedures for control of the family-wise error rate. Statistical applications in genetics and molecular biology. 2004;3:Article14.
Boulesteix A-L, Durif G, Lambert-Lacroix S, Peyre J, Strimmer K. plsgenomics: PLS Analyses for Genomics. In. Cran-R package version 1.3/r13; 2015.
Boulesteix AL. PLS dimension reduction for classification with microarray data. Statistical applications in genetics and molecular biology. 2004;3:Article33.
Liaw A, Wiener M. Classification and Regression by randomForest. R News. 2002;2(3):18–22.
Breiman L. Random forests. Mach Learn. 2001;45(1):5–32.
Tibshirani R, Chu G, Narasimhan B, Li J. samr: SAM: Significance Analysis of Microarrays. R package version 2.0. Available from: http://CRAN.R-project.org/package=samr.
Tusher VG, Tibshirani R, Chu G. Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci U S A. 2001;98:5116–21.
Opgen-Rhein R, Zuber V, Strimmer K: st: Shrinkage t Statistic and Correlation-Adjusted t-Score. In. Cran-R package version 1.2.4; 2015.
Zuber V, Strimmer K. Gene ranking and biomarker discovery under correlation. Bioinformatics. 2009;25:2700–7.
Kuhn. M, Wing CfJ, Weston S, Williams A, Keefer C, Engelhardt A, et al. caret: Classification and Regression Training. In. Cran-R package version 6.0-47; 2015.
Strbenac D, Mann GJ, Ormerod JT, Yang JY. ClassifyR: an R package for performance assessment of classification with applications to transcriptomics. Bioinformatics. 2015;31(11):1851–3.
Slawski M, Daumer M, Boulesteix AL. CMA: a comprehensive Bioconductor package for supervised classification with high dimensional data. BMC Bioinformatics. 2008;9:439.
Johannes M, Ruschhaupt M, Froehlich H, Mansmann U, Buness A, Warnat P, Huber W, Benner A, Beissbarth T: MCRestimate: Misclassification error estimation with cross-validation. In. Bioconductor-R package version 2.24.0; 2010.
Carey V, Gentleman R, Mar J, Vertrees J, Gatto L. MLInterfaces: Uniform interfaces to R machine learning procedures for data in Bioconductor containers. In. Bioconductor-R package version 1.48.0.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
AS analyzed the data. AS wrote the paper. AS and GS designed the computational pipeline. AS wrote the software. GS supervised the project. Both authors read and approved the final manuscript.
Additional files
Additional file 1:
Package ‘mAPKL_1.1.1.tar.gz’ including source code, documentation. (GZ 1979 kb)
Additional file 2:
Package ‘mAPKLData_1.0.0.tar.gz’ including microarray data and documentation. (GZ 9281 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Sakellariou, A., Spyrou, G. mAPKL: R/ Bioconductor package for detecting gene exemplars and revealing their characteristics. BMC Bioinformatics 16, 291 (2015). https://doi.org/10.1186/s12859-015-0719-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12859-015-0719-5