Abstract
Successful cultivation of rice (Oryza sativa L.) in many Asian countries requires submergence stress tolerance at the germination and early establishment stages. Two quantitative trait loci, Sub1 (conferring submergence tolerance) and AG1 (conferring anaerobic germination), were recently pyramided into a single genetic background, without compromising any desirable agronomic traits, leading to the development of Ciherang-Sub1 + AG1 (CSA). However, little research has been conducted to enhance plant tolerance to abiotic stress (submergence) and biotic stress (rice blast), which occur in a damp climate following flooding. The BC2F5 breeding line was phenotypically characterized using the AvrPi9 isolate. The biotic and abiotic stress tolerance of selected lines was tested under submergence stress and anaerobic germination conditions, and lines tolerant to each stress condition were identified through phenotypic and gene expression analyses. The Ciherang-Sub1 + AG1 + Pi9 (CSA-Pi9) line showed similar agronomic performance to its recurrent parent, CSA, but had significantly reduced chalkiness in field trials conducted in temperate regions. Unexpectedly, the CSA-Pi9 line also showed salinity tolerance. Thus, the breeding line newly developed in this study, CSA-Pi9, functioned under stress conditions, in which Sub1, AG1, and Pi9 play a role and had superior grain quality traits compared to its recurrent parent in temperate regions. We speculate that CSA-Pi9 will enable the establishment of climate-resilient rice cropping systems, particularly in East Asia.
Similar content being viewed by others
Background
Flooding is a frequent natural calamity in subtropical regions, and climate change can cause unexpected heavy rains and frequent inundation, which are expected to increase in Peninsular India and South Asia in the future (Mondal et al. 2020a, b). Rice (Oryza sativa L.) is the only plant species that can be grown in conditions prone to uncontrolled inundation or saline inundation; nonetheless, flooding is one of the major factors responsible for rice yield reduction (Hirabayashi et al. 2013; Kuanar et al. 2017). Rainfed lowland rice farms occupy 34–70% of the total rice production area in India and other South Asian countries, and more than 15 million ha of rainfed lowland rice systems are affected by flooding, decreasing rice productivity (Bailey-Serres et al. 2010; Emerick and Ronald 2019; Ali et al. 2006; Septiningsih et al. 2009). At least 16% of the rice yield is affected by flooding, accounting for a loss of more than one billion (USD) per year in South and Southeast Asia (Ram et al. 2002; Ali et al. 2006; Xu et al. 2006; Hirabayashi et al. 2013). Elevation in sea level, possibly due to global warming, has decreased rice cultivation near the coast, and the resultant frequent flooding has led to poor drainage, affecting the rice crop at all stages including germination and vegetative growth (Mackill et al. 2010). Most rice genotypes, including high-yielding varieties, cannot survive under deep standing water and cannot germinate under oxygen limiting conditions (Bailey-Serres et al. 2010; Baltazar et al. 2014). However, most modern varieties were developed without considering the need for germination and/or survival strategies under submerged conditions (Baltazar et al. 2019; Alam et al. 2020). A 34–83% yield reduction has been reported in modern varieties under stagnant flooding, but not fully submerged, conditions (Kato et al. 2019). In addition, compared to manual transplanting, direct seeding is becoming increasingly popular, as it is associated with reduced labor and operational costs, earlier flowering, shorter crop duration, and earlier maturity (by 7–10 days) (Baltazar et al. 2014; Kumar et al. 2015). However, one of the most serious problems in the open mechanized direct seeding system is early establishment, especially in wet seeding systems. Against those stresses, which are related with submergence in the germination and vegetative stages, two quantitative trait loci (QTLs) are required in modern indica rice varieties: Submergence 1 (Sub1), which confers submergence tolerance by Sub1A; and Anaerobic Germination-9-2 or AG1, which is associated with anaerobic germination and harbors the TREHALOSE-6-PHOSPHATE PHOSHPHATASE 7 (OsTPP7) gene (Xu et al. 2006; Toledo et al. 2015; Alam et al. 2020). Among the genes identified at the Sub1 QTL, Sub1A is found in a small collection of rice germplasm, whereas Sub1B and Sub1C are found in all rice varieties. Sub1A does not have any negative effects on rice yield and agronomic traits (Septiningsih and Mackill 2018). When Sub1 was transferred to the mega rice variety Swarna, which is grown on approximately 5 million ha, the resulting genotype (Swarna-Sub1) produced a yield of 5–6 tons per ha in the absence of flooding, similar to Swarna (Emerick and Ronald 2019). Furthermore, 80–95% of Swarna-Sub1 plants survived and produced 45% higher yield than Swarna in areas flooded for 10 days (Emerick and Ronald 2019). Introgression of the AG1 QTL, which improves germinative capacity under anaerobic conditions, is an effective approach for overcoming flooding-related problems at the germination stage including root establishment. IR64-AG1 near isogenic lines (NILs) showed 121% higher germination than IR64, irrespective of the flood duration (Lal et al. 2018). The AG1 QTL has been introgressed into several rice varieties, such as the variety Dongan (Kim et al. 2019a, b), without causing any negative effects on yield under normal conditions (Toledo et al. 2015). Similarly, the Sub1 QTL has been introgressed into various mega varieties including Samba Mahsuri, CR1009, Thadokkham, BR11, Bac Thom7, PSB Rc18, and M-202 (Septiningsih et al. 2009, 2013; Khanh et al. 2013; Xu et al. 2004). In addition, both Sub1 and AG1 QTLs have been pyramided together in Ciherang, IR64, Swarna, NSIC Rc222, and Bg358 (Toledo et al. 2015; Rafael et al. 2015; Sartaj et al. 2016), and combined with other abiotic stress tolerance QTLs/genes including AG2, Pup1 (phosphorous uptake under rainfed/upland condition), DTY (drought), and Saltol (salinity) in the rice varieties IR64, Swarna, PSB Rc82, and Mahsuri (Mondal et al. 2020a, b; Shin et al. 2020; Singh et al 2016; Dar et al. 2014).
The rice variety Ciherang-Sub1 + AG1 (CSA), which was developed by the introgression of Sub1 and AG1 into Ciherang, was used in this study (Septiningsih et al. 2015; Toledo et al. 2015). Ciherang, developed at the International Rice Research Institute (IRRI; Philippines), is the predominant variety of Indonesia with a high yield potential (5–7 tons/ha) (IRRI 2015; Arief et al. 2018). The breeding line CSA, which is morphologically highly similar to Ciherang and contains Sub1 and AG1 for tolerance to submergence and anaerobic germination conditions, was selected using the marker-assisted backcrossing (MABC) approach, with five markers for the Sub1 locus (RM8300, a simple sequence repeat [SSR] maker downstream of Sub1A, ART5, and three insertion/deletion [Indel] markers located within Sub1C) and four markers for the AG1 locus (TPP_GE5, HPP400_410_3, Drebups6bp, and Drebdws4bp) (Toledo et al. 2015). CSA is tolerant to submergence at all growth stages and has nearly the same plant height and tiller number as Ciherang-Sub1 (Toledo et al. 2015). Based on genotype, CSA is approximately 99% similar to Ciherang, thus there is minimal interaction between the genetic backgrounds of Ciherang and the QTL donor (Toledo et al. 2015). However, compared with Ciherang, CSA exhibits delayed flowering (by approximately 6 days) and higher grain yield in one season (Toledo et al. 2015). This variation in flowering period of Sub1-introgressed lines was previously reported by Septiningsih et al. (2013) and demonstrates the limitation of marker-assisted selection (MAS), when used alone, in the selection of identical genotypes by introgressing useful traits into high-yielding varieties. Nonetheless, MAS provides the opportunity to explore new traits by testing multiple genetic combinations of QTLs with the endogenous genes of high-yielding or mega varieties.
Submergence tolerance, anaerobic germination and salinity tolerance are crucial value-adding abiotic stress-related traits. In coastal areas, nutrient-poor and saline soil conditions promote biotic stresses, such as rice blast, which challenge rice production by affecting plants after flowering. The high-level accumulation of abscisic acid (ABA) in plants grown in saline areas affects their biotic stress response and makes them susceptible to fungal pathogens, such as the rice blast pathogen Magnaporthe oryzae (Couch and Kohn. 2002; Ton et al. 2009; Jiang et al. 2010; Asano et al. 2012). M. oryzae, a filamentous ascomycete, is distinct from Magnaporthe grisea, and can infect rice plants at all growth stages by entering through leaves, stems, nodes, and panicles (Wilson and Talbot 2009). Rice blast, caused by M. oryzae, is the most devastating disease of rice, responsible for 10–35% of yield loss (Fisher et al. 2012). More than 100 blast resistance (R) genes have been mapped to date, and most Pi genes, which have been characterized as 25 R genes, encode nucleotide-binding site leucine-rich repeat (NBS-LRR) proteins, which are constitutively expressed in blast-resistant plants (Wilson and Talbot 2009; Li et al. 2019). Exceptions are Pid2, Pi21 and Ptr which encode a receptor-like kinase protein, protein with heavy-metal-binding and proline-rich domains, and protein with four Armadillo repeats, respectively (Chen et al. 2006; Ballini et al. 2008; Zhao et al. 2018). Using Pi genes as part of a breeding process is slow and uneconomical because M. oryzae is a fast-evolving fungal pathogen that breaks the R gene-mediated resistance within 3–5 years (Miah et al. 2013; Li et al. 2019). Broad-spectrum resistance genes like Pi9, an R gene cloned from O. minuta J.Presl., a wild relative of cultivated rice, have significant potential to resist pests and diseases (Amante-Bordeos et al. 1992; Elgamal and Elshenawy 2018). The Pi9 gene is more useful than other Pi genes because it shows horizontal resistance against a diverse range of blast races (Qu et al. 2006). Recently, Zhou et al. (2020) identified 13 novel Pi9 alleles using 107 blast-resistant varieties. In addition, Pi9 is generally more effective when combined with other Pi genes in various rice varieties, without trade-off effects like decreased yield and delayed flowering (Khanna et al. 2015; Xiao et al. 2017; Wu et al. 2019). This implies that pyramiding Sub1, AG1 and Pi9 in a single genetic background will facilitate rice cultivation via direct seeding in large coastal areas including those in Korea.
Korea harbors 1.2 million registered foreigners (Statistics Korea 2020), who prefer indica rice over japonica rice for consumption, thus promoting the diversification of rice cultivation. Ciherang is one of the most famous indica varieties, especially in Indonesia. Furthermore, the rise in temperature during the growing season may require Koreans to identify or develop varieties adaptable to subtropical conditions. Thus, the development of new varieties suitable for direct seeding in temperate regions is important, and pyramiding genes that induce tolerance to multiple biotic and abiotic stresses in rice is in high demand. The objective of this study was to introgress the broad-spectrum blast resistance gene, Pi9, into the flood tolerant indica variety, CSA and evaluate the newly developed germplasm, Ciherang-Sub1 + AG1 + Pi9 (CSA-Pi9) in the saline western coastal area of Korea.
Results
Biotic and Abiotic Stress-Resistance Pyramiding Scheme
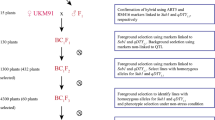
IRBL9-w, a monogenic line has the Pi9 resistance allele introgressed into the background of Lijiangxintuanheigu (LTH), a japonica variety which is extremely susceptible to blast, and CSA were crossed, and the resultant F1 individuals were backcrossed twice with CSA (Fig. 1). The backcross progeny that resembled the recurrent parent in plant type were selfed to generate the BC2F7 population. Screening for submergence tolerance and anaerobic germination was conducted in the BC2F5 generation. The genotype of BC2F5 and BC2F7 seeds was validated using QTL/gene-specific markers, and agronomic traits of BC2F5 and BC2F7 plants were tested under field conditions (Table 1). The BC2F7 plants were genotyped using GnS2 and DFR markers, which can specifically detect tolerance alleles of Sub1 and AG1 (Fig. 2a). Pi9-1477G and Nbs2-Pi9 (195), which are Pi9- and Nbs2-Pi9-specific markers, respectively, were used for the foreground genotyping of Pi9 (Fig. 2a, b). The Pi9-1477G marker, which could distinguish between blast-resistant and susceptible varieties, was developed in this study by aligning Pi9 alleles (Additional file 1: Fig. S1).
Analysis of the genetic background and foreground selection of CSA-Pi9. Foreground selection of CSA-Pi9 using Sub1A-, AG1- and Pi9-specific markers (a, b). Analysis of the genetic background of CSA-Pi9 using the KNU Axiom Oryza 580 K Genotyping Array containing SNPs designed on the basis of IRGSPv1.0 (left) and MH63 v.2 (right) (c). M: marker; bp: base pair; kbp: kilobase pair
The genetic background of CSA-Pi9 was analyzed using the KNU Axiom Oryza 580 K Genotyping Array, which consists of single nucleotide polymorphisms (SNPs) designed on the basis of IRGSP v1.0. and MH63 v2 (Fig. 2c). CSA-Pi9 showed 99.2% and 89.6% CSA-specific alleles, based on IRGSP v1.0 and MH63 v2, respectively. Compared with other chromosomes, the ratio of CSA-specific alleles was lower on chromosome 6, which harbors Pi9 (Additional file 3: Table S2).
Resistance of CSA-Pi9 to Rice Blast
To examine blast resistance induced by Pi9 derived from IRBL9-w, leaves of CSA-Pi9 and CSA (recurrent parent) were spot-inoculated with two rice blast isolates, PO6-6 (carrying the cognate effector of Pi9, Avrpi9; incompatible isolate) and RO1-1 (lacking AvrPi9; compatible isolate), as described previously (Kanzaki et al. 2002), and the size of lesions caused by the two isolates was compared at 9 days post-inoculation (dpi). When inoculated with PO6-6, lesions formed on CSA-Pi9 leaves were significantly smaller than those formed on CSA leaves (Fig. 3a, b), indicating that the introduced Pi9 gene was effective in provoking effector-triggered immunity. However, CSA-Pi9 and CSA leaves inoculated with RO1-1 showed no significant difference in lesion size (Fig. 3c, d). Taken together, these data suggest that the introgression of Pi9 into CSA induces resistance against AvrPi9-carrying isolates.
AvrPi9 phenotyping and validation of Pi9-mediated resistance. Photographs of lesions (a) and quantification of lesion length (b) in CSA-Pi9 and CSA leaves inoculated with PO6-6 (incompatible isolate). Photographs of lesions (c) and quantification of lesion length (d) in CSA-Pi9 and CSA leaves inoculated with RO1-1 (compatible isolate)
Independent Functions of Sub1 and AG1 in CSA-Pi9 Under Each Stress Condition
To assess the submergence tolerance of CSA-Pi9, 2-week-old plants of CSA-Pi9, CSA, IR64, and IR64-Sub1 were submerged in 70-cm deep tap water for 14 days and subsequently recovered in shade for 7 days (Fig. 4a–d). CSA-Pi9, CSA, and IR64-Sub1, which harbor Sub1, recovered from submergence stress and sprouted new leaves at 7 days after recovery (DAR) (Fig. 4a, b). However, IR64, the susceptible control, dried after de-submergence (Fig. 4a, b). To evaluate whether Sub1 restricts shoot elongation under submergence conditions (Xu et al. 2006), we measured the shoot length of plants before submergence and at 14 days after submergence (DAS) (Fig. 4c). CSA-Pi9 showed a submergence tolerant phenotype, with significantly reduced shoot elongation rate (18.6%), similar to IR64-Sub1 (13.6%). Shoot elongation rate was also reduced in CSA by 5.4%. However, IR64 showed the highest shoot elongation rate (47.7%). To determine the recovery of chlorophyll content, we measured the SPAD values of all genotypes pre- and post-submergence (Fig. 4d). The SPAD value of CSA-Pi9 was 34.6 after submergence and recovered to 97.4% of pre-submergence levels, which was higher than that observed in IR64-Sub1 (93.9%) and CSA (88.1%). Next, to determine the function of Sub1 and AG1 in CSA-Pi9, we examined the transcript levels of Sub1A and OsTPP7, the major genes underlying Sub1 and AG1 QTLs, respectively, and of alcohol dehydrogenase1 (adh1) (Fig. 4e–g). Expression levels of Sub1A in CSA-Pi9 and IR64-Sub1 were similar until 7 DAS. At 14 DAS, although the expression level of Sub1A in CSA-Pi9 decreased slightly compared with IR64-Sub1, it was still higher than that in CSA (Fig. 4e). Sub1A was expressed until 7 DAR in Sub1 lines (Fig. 4e). The expression level of adh1 in IR64-Sub1 was significantly higher than that in CSA-Pi9 and CSA at 1 DAS but decreased gradually until 1 DAR (Fig. 4f). The expression of adh1 was not strong in CSA-Pi9 at 1 DAS; however, it increased at 7 DAS and lasted until 14 DAS (Fig. 4f). After recovery, adh1 expression decreased, and showed no significant difference among the various genotypes. The OsTPP7 gene was expressed in CSA-Pi9 and CSA, and its expression level in CSA-Pi9 was similar to, or slightly higher than that observed in CSA under submergence conditions (Fig. 4g).
Characterization of the CSA-Pi9 breeding line treated with submergence stress for 14 days. Growth phenotype of CSA-Pi9 and other genotypes during submergence stress (a) and at 7 days after de-submergence (b). Shoot elongation rate [calculated as (shoot height after submergence (S)/shoot height before S) × 100] (c) and SPAD value (d) of CSA-Pi9 before and after submergence. Expression levels of Sub1A (e), adh1 (f) and OsTPP7 (g) under submergence stress. Data represent mean ± standard error (SE). Lowercase letters above bars and lines indicate statistically significant differences (P < 0.01; Duncan's multiple range test). IR64: submergence susceptible variety; IR64-Sub1: submergence tolerant variety; yellow arrow: newly emerging leaves; red arrow: no new leaves; DAS: days after submergence; DAR: days after recovery
To assess the function of AG1 in CSA-Pi9, the germination of CSA-Pi9, CSA, IR64, and IR64-AG1 seeds was tested under anaerobic conditions (Fig. 5). While CSA-Pi9, CSA, and IR64-AG1 seeds germinated under anaerobic conditions, none of the IR64 seeds germinated (Fig. 5a), as expected. Additionally, CSA-Pi9 seedlings showed a significantly higher survival rate (42%) than CSA (8%) and IR64-AG1 (20%) seedlings (Fig. 5b). Significant differences were neither observed in coleoptile length among the germinated lines (~ 28 mm) nor in shoot length between CSA-Pi9 (14.5 cm) and IR64-AG1 (17.1 cm) (Fig. 5c, d). We also analyzed the expression of OsTPP7 and adh1 in the root and shoot separately. The shoot of CSA and CSA-Pi9 expressed OsTPP7 and adh1 to higher levels than that of IR64-AG1, while CSA-Pi9 root expressed both genes to higher levels than CSA and IR64-AG1 roots (Fig. 5e, f). Thus, CSA-Pi9 recovered fully from submergence stress and sprouted new leaves, and also showed powerful germinability under anoxic conditions (Figs. 4, 5). Based on these results, we conclude that both Sub1 and AG1 are functional in CSA-Pi9 under each stress condition, regardless of the presence of Pi9.
Characterization of CSA-Pi9 under anaerobic germination conditions. Phenotype of CSA-Pi9 plants at 34 days after sowing (a), germination rate (b), coleoptile length (c) and shoot length (d). Expression levels of OsTPP7 (e) and adh1 (f). Data for IR64 were unavailable, as its seeds did not germinate. Lowercase letters above bars indicate statistically significant differences (P < 0.01; Duncan's multiple range test)
Increased Spikelet Number of CSA-Pi9 Helps Maintain Grain Yield in Paddy Fields
To investigate the agronomically important plant- and seed-related traits of CSA-Pi9 and CSA, plants of both genotypes were grown in paddy fields located in three different regions: field A (Suwon, normal growth conditions), field B (Seosan, normal growth conditions), and field C (Seosan, saline conditions). Plants in field A were tested for 2 years (2018 and 2019), whereas those in fields B and C were evaluated only for 1 year (2019). Plants in field A were grown for 146 days in 2018 and for 135 days in 2019. In field A, CSA-Pi9 and CSA plants flowered at the same date in 2018 (i.e., at 64 days after transplanting [DAT]); however, CSA-Pi9 flowered 15 days earlier than CSA plants in 2019 (76 and 91 DAT, respectively). In fields B and C, plants of both genotypes were grown for 135 and 128 DAT, respectively. In field B, CSA-Pi9 plants flowered 15 days earlier than CSA plants (77 and 92 DAT, respectively); however, in field C, CSA-Pi9 plants flowered at 91 DAT. No significant differences were observed between CSA-Pi9 and CSA in the length, number and architecture of panicles; however, the culm length of CSA-Pi9 plants was significantly shorter than that of CSA plants (Table 1, Fig. 6a).
In field A, CSA-Pi9 plants produced 187.22 and 182.67 spikelets per panicle in 2018 and 2019, respectively, which were 28.4% and 29.0% more, respectively, than those produced by CSA; however, in field B, the two genotypes showed no significant difference in spikelet number per panicle (Table 1). The fertility of CSA-Pi9 was the same or higher than that of CSA in fields A and B (Table 1). Compared with CSA, the hundred-grain weight of CSA-Pi9 was 22.3% lower in field A in 2018, and 3.0% and 6.4% lower in fields A and B, respectively, in 2019 (Table 1). However, the grain yield per plant of CSA-Pi9 was comparable with that of CSA in every field test, except in field A in 2018, where it was higher than that of CSA (Table 1). These results confirmed that the yield of CSA-Pi9 was comparable or preferable to CSA considering spikelet number per panicle and fertility, even though its hundred-grain weight was significantly lower than that of CSA. (Table 1).
In field C, which was located in a reclaimed saline region, CSA-Pi9 showed good yield performance, whereas none of the CSA plants survived. The flowering period of CSA-Pi9 in field C was 91 DAT, which was 14 days later than that in field B. The hundred-grain weight and grain yield of CSA-Pi9 was 1.86 g and 18.56 g per plant, respectively (Table 1). The tolerance of CSA-Pi9 to salinity stress was an unexpected finding.
Superior Quality of CSA-Pi9 Grains with Low Chalkiness
Next, we investigated the grain quality traits of CSA-Pi9 and CSA, including grain size, chalkiness, amylose content, alkali spreading value and protein content (Table 2). The width of CSA-Pi9 grains was similar to the CSA grains, whereas the length of CSA-Pi9 grains was 9.3% and 13.1% shorter than the CSA grains in fields A and B, respectively, thus the length to width ratio was lower in CSA-Pi9 (Table 2, Fig. 6b). In fields A and B, CSA-Pi9 showed brown rice recovery rates of 76.7% and 78.5%, respectively, which were 3.5% and 5.0% lower than those of CSA, respectively (Table 2). Chalkiness is a major index of grain quality. CSA-Pi9 grains showed 33.6% and 38.6% less chalkiness than CSA grains in fields A and B, respectively (Table 2). The amylose content and alkali spreading value of CSA-Pi9 grains were lower than those of CSA grains in fields A and B, and significant differences in amylose content and alkali spreading value were observed, respectively, between fields A and B for both genotypes (Table 2). No significant difference in protein content was detected between CSA-Pi9 and CSA in fields A and B (Table 2).
To evaluate the effect of salinity on grain quality in rice, we compared the grain quality of CSA-Pi9 between fields B and C (Table 2). The grain length and brown rice recovery rate of CSA-Pi9 in field C were slightly lower than those in field B (Table 2). The chalkiness of CSA-Pi9 grains showed no significant difference between the two fields. Compared with field B, the amylose content of CSA-Pi9 grains produced in field C was lower by 16.9%, whereas the alkali spreading value and protein content were significantly higher (100% and 67.0%, respectively) (Table 2). These data suggest that CSA-Pi9 exhibits superior grain quality traits compared with CSA, with decreased chalkiness under both normal and saline conditions, even though the grain size of CSA-Pi9 was smaller than that of CSA because of shortened grain length.
Discussion
Indica Rice and Its Availability in Temperate Regions
Owing to global warming, the climate of regions such as Korea is changing from temperate to subtropical; however, very few studies have evaluated rice germplasm as single plants across temperate and tropical regions (Chung et al. 2004; Ha et al. 2011; Huang et al. 2021; Navea et al. 2017; Takai et al. 2019). In addition, most of the stress tolerance QTLs/genes, such as Sub1, AG1, and Pi9, have been derived from rice germplasm adapted to tropical regions, and are rarely found in modern rice varieties (Ismail et al. 2013; Baltazar et al. 2019; Ning et al. 2020). Evaluation of rice plants adapted to tropical regions is difficult in temperate regions, because of variation in climate-sensitive traits such as plant type and flowering time (Ha et al. 2011). However, the indica rice accession developed in this study, CSA-Pi9, was well adapted to temperate regions. The introgressed Sub1, AG1, and Pi9 loci of CSA-Pi9 functioned normally under temperate climate conditions, without compromising any agronomic trait, and the tolerance of CSA-Pi9 to anaerobic and submergence conditions was similar to that of CSA under tropical climate conditions (Figs. 4, 5; Toledo et al. 2015). Previously, CSA showed a survival rate of approximately 47.3% under anaerobic conditions in a tropical region (Toledo et al. 2015); however, in this study conducted in temperate regions, the survival rate of CSA decreased to 8% (Fig. 5b). By contrast, CSA-Pi9 showed a survival rate of 42%, and expressed the OsTPP7 gene in roots germinated under anaerobic conditions in temperate regions (Fig. 5b). In addition, compared with CSA, CSA-Pi9 seedlings exhibited full recovery after de-submergence with less chlorosis, and showed significantly higher expression levels of three genes (Sub1A, OsTPP7, and adh1) than those of CSA during submergence stress in the temperate region (Fig. 4). Additionally, we noticed that Sub1 and AG1 were functional in temperate climate, and the tolerance levels were similar to those of Sub1 and AG1 in tropical conditions.
Advantages Acquired by a Molecular Breeding of CSA-Pi9
The development and utilization of adequate molecular markers for genotyping target loci and whole-genome background are essential for modern molecular breeding programs. However, the development of allele-specific markers for the Pi9 gene is difficult, because it contains repeat sequences. In this study, the SNP marker Pi9_1477G was used to discriminate among various Pi9 alleles during MABC. However, because of high sequence similarity between Nbs5 and Nbs2 at the primer-binding sites, Pi9-1477G amplified the Nbs5 region instead of Nbs2 in some steps of the selection process in the current study; for example, BC2F2-generation plants harboring the Nbs5-Pi9 allele were selected with Pi9_1477G (data not shown). Therefore, the application of multiple gene-specific markers is recommended to identify the Nbs2-Pi9 allele.
Unpredictable climate change imposes multiple biotic/abiotic stresses on plants, and most elite varieties are susceptible to these stresses. However, owing to the development of markers associated with several important traits, it is possible to pyramid multiple traits into one cultivar using the MABC approach (Das and Rao 2015; Das et al. 2018; Dixit et al. 2020). Several attempts have been made to develop multiple biotic/abiotic-stress resistant elite varieties (Chukwu et al. 2019; Chithrameenal et al. 2018; Feng et al. 2018; Sandhu et al. 2020). Nevertheless, only a few varieties have been developed that exhibit multi-stress resistance along with superior quality traits in the background of Tapaswini, Swarna and White Ponni as summarized in Table 3 (Das et al. 2018; Dixit et al. 2020; Muthu et al. 2020). Therefore, we introgressed Pi9 into CSA using the MABC method to improve not only abiotic/biotic stress tolerance but also agronomically important traits like culm length and flowering period in addition to quality related traits. The CSA-Pi9 breeding line developed in this study showed tolerance to submergence and anaerobic germination conditions and to blast, as shown by phenotypic and gene expression analyses. The yield of CSA-Pi9 was similar to that of CSA under normal field conditions, even though CSA showed higher hundred grain weight compared with CSA-Pi9 (Table 1). Moreover, the chalkiness of CSA-Pi9 was significantly lower than that of CSA in all field tests (Table 2), which implies that the quality of CSA-Pi9 grains is superior to that of CSA grains. However, CSA-Pi9 and CSA showed no polymorphic SNPs in Chalk5, GWC1, and FLR1, the representative genes affecting grain chalkiness (Guo et al. 2020; Li et al. 2014; Pu et al. 2017). Recently, breeders were challenged to develop a high yielding, climate change-ready rice variety, with good eating quality and nutritional composition as cooked rice (Collard et al. 2019; Manangkil et al. 2020). The CSA-Pi9 breeding line generated in this study represents an important genetic material that could be used not only in future rice breeding programs, but also for commercial cultivation in Asian countries as a multi-stress resistant cultivar with enhanced grain quality and agronomic performance.
Salinity Tolerance of CSA-Pi9
Traits not observed in the parental genotypes are occasionally observed in the progeny. In the current study, the CSA-Pi9 line showed salinity tolerance, which was a serendipitous finding in field trials. The CSA-Pi9 line was well adapted to the reclaimed land at the seashore, where growth conditions were humid and saline (Tables 1, 2). Because CSA was not adapted to the reclaimed land, we could assume that the salt tolerance of CSA-Pi9, which was derived from the introgression of Pi9 into CSA, was conferred by the Pi9 gene. However, the salt tolerance of O. minuta (Pi9 donor) and IRBL9-w has not been reported to date. To determine whether the salinity tolerance of CSA-Pi9 is derived from IRBL9-w, we examined the salinity tolerance of IRBL9-w using a hydroponic system, since IRBL9-w has not been tested in a reclaimed area. However, IRBL9-w was sensitive to salt concentrations of 0.6% and 1.0% (data not shown), suggesting that the salinity tolerance of CSA-Pi9 was not derived from IRBL9-w. Next, we compared the Saltol QTL region (10.8–16.4 Mb on chromosome 1) between CSA-Pi9 and CSA lines and identified 1192 SNPs (data not shown). Mansuri et al. (2020) reported candidate genes involved in salt tolerance, based on the integrative meta-analysis of RNA-seq and microarray data available at the National Center for Biotechnology Information (NCBI) database. Axiom Oryza 580 K Genotyping Array data was used to analyze the presence of polymorphic SNPs between CSA-Pi9 and CSA in gene listed in Mansuri et al. (2020). The results revealed 27 SNPs between CSA and CSA-Pi9 (Additional file 4: Table S3). One of these 27 SNPs was located in the pectinesterase gene, which belongs to the Saltol QTL region. However, because the Saltol QTL harbors many genes and has not been fully characterized, it is difficult to conclude which genes are the major contributors to salt tolerance in CSA-Pi9. The salinity tolerance of CSA-Pi9 could be caused by one or more of the candidate genes listed in Additional file 4: Table S3 or by the interactions among the introgressed QTL/genes. In the future, we plan to determine the mechanism of salinity resistance in CSA-Pi9. We speculate that the CSA-Pi9 line could be recommended to farmers who use the direct seeding method for cultivating rice in saline soil in East Asia under high humidity conditions, which promote rice blast.
Conclusions
In this study, the breeding line CSA-Pi9, was developed by introducing Pi9, a broad-spectrum blast resistance gene, from O. minuta into CSA. CSA-Pi9 harbors 89.6% of the CSA (recurrent parent)-specific alleles. The Sub1, AG1, and Pi9 QTLs/genes were expressed and functional in CSA-Pi9 under submergence stress and anaerobic germination conditions. The tolerance of CSA-Pi9 to salinity stress was a serendipitous finding. Compared with CSA, CSA-Pi9 showed similar yield, improved grain quality with less chalkiness when grown in a temperate region. Thus, the CSA-Pi9 breeding line developed in this study could be used as a useful donor in breeding programs aimed at developing multi-stress resistant varieties for cultivation in temperate regions.
Materials and Methods
Plant Materials
CSA (GID 4537744) harbors Sub1 and AG1 QTLs in the Ciherang background, and was introduced into Sejong University (SJU, Seoul, Korea) by a Seconded Special Material Transfer Agreement (seconded SMTA), via Hankyong National University (HKNU, Anseong, Korea), with the IRRI (Los Baños, Philippines). IRBL9-w (Entry no. I22), derived from O.minuta (accession no. 101141), seed was obtained from Kyunghee University seed stock (KHU, Yongin, Korea). Initially, five CSA and three IRBL9-w plants were grown for crossing form which the cross between CSA-4 and IRBL9-w-2 was selected to develop CSA-Pi9. IR64-Sub1 + AG1 (GID 4537760) was introduced into SJU by seconded SMTA via HKNU with the IRRI. The IR64-Sub1 (Entry no. 961209) and IR64-AG1 (Entry no. 961212) genotypes, derived from IR64-Sub1 + AG1, were used as positive controls in the submergence and anaerobic germination screening experiments, respectively. IR64 (Entry no. 961061; accession no. IRGC 117268), obtained as a seed stock from SJU, was used as a susceptible control in the submergence and anaerobic germination screening experiments.
Crossing Scheme for the Development of CSA-Pi9
CSA was crossed with IRBL9-w (Fig. 1), and 24 F1 plants were subjected to foreground genotyping to detect a functional allele of Sub1, AG1 and Pi9 using gene/QTL-specific markers: GnS2 for Sub1 tolerant allele, DFR for AG1 tolerant allele, ‘195’ for Nbs2-Pi9 resistance allele and ‘NBS5’ for Nbs5-Pi9 allele (Additional file 2: Table S1; Septiningsih et al. 2009; Kretzschmar et al. 2015; Qu et al. 2006). A single F1 plant showing the greatest phenotypic resemblance to CSA was backcrossed with CSA, and the resultant 94 BC1F1 plants were genotyped using foreground markers. Based on the genotyping results, a single plant was selected and backcrossed with CSA. Subsequently, 3 out of 288 BC2F1 plants were selected based on foreground genotyping results, and further characterized in a greenhouse for the heading date and phenotypic resemblance to CSA. Using the pedigree method, three BC2F2 breeding lines, morphologically resembling CSA and similar to CSA in agronomic traits including heading date, height and panicle, were selected, and the seeds of three plants of each line showing minimum segregation were propagated. BC2F3 plants were propagated in the field and advanced to the BC2F5 generation. Some of the BC2F5 seeds were used for the phenotypic screening of rice blast resistance, submergence tolerance and anaerobic germination. The BC2F5 plants were advanced further to the BC2F7 generation. The grain yield and quality of BC2F5 and BC2F7 plants were tested in the field in 2018 and 2019, respectively.
Primer Design and Genotyping
All Nbs sequences of Pi9 were compared, and two markers based on SNPs at nucleotide positions 659 and 1477 of Nbs2-Pi9 were developed (Additional file 1: Fig. S1a). However, since the sequence of Nbs5-Pi9 was identical to that of Nbs2-Pi9 at the SNP position, as shown in the sequence alignment, only the Pi9_SNP1477G marker was used for MAS. The Nbs2-Pi9-specific primer (Qu et al. 2006; Additional file 2: Table S1) was used to distinguish between Nbs2-Pi9 and Nbs5-Pi9.
Leaves were harvested from 2-week-old plants of CSA, CSA-Pi9, IR64, IR64-Sub1, and IR64-AG1 lines, and genomic DNA was extracted using the cetyltrimethylammonium bromide (CTAB) method (Murray and Thompson 1980). Sub1, AG1, and Pi9 were genotyped by PCR using foreground markers (Additional file 2: Table S1). PCR was performed on the SimpliAmp thermocycler (Thermo Scientific, Waltham, MA, USA) in a 20-µL reaction volume, containing 50 ng of DNA template, 0.5 µL each of 10 µM forward and reverse primers (Bioneer, Daejeon, Korea), 2 µL of 10X buffer, 0.5 µL of 2.5 mM dNTPs and 0.1 µL of Taq polymerase (IN5001-0500; Inclone, Yongin, Korea), under the following conditions: initial denaturation at 94 °C for 4 min, followed by 30–35 cycles of denaturation at 94 °C for 1 min, annealing at 55–61 °C for 30–60 s, and extension at 72 °C for 30–60 s, with a final extension at 72 °C for 7 min. The PCR products were electrophoresed (BioFACT, Daejeon, Korea) on 1% or 2% agarose gel at 100–160 V for 20–40 min in 0.5X TBE buffer. Gels were visualized using the gel-imager (Korea Lab Tech, Seongnam, Korea).
Background genotyping of CSA-Pi9 and CSA was performed using the KNU Axiom Oryza 580 K Genotyping Array (Kim et al. 2019a, b) at DNAlink (Seoul, Korea). The KNU Axiom Oryza 580 K Genotyping Array consists of markers based on IRGSP v1.0 and MH63 v2. The results of background genotyping were analyzed based on the allele type at DNAcare (Seoul, Korea). The background recovery ratio was calculated as the number of markers with CSA alleles within 0.5 Mb segments of each chromosome relative to the total number of markers.
Rice Blast Resistance Screening
CSA-Pi9 was challenged with M. oryzae isolates RO1-1 (compatible) and PO6-6 (incompatible). For blast resistance screening, the PO6-6 isolate was used to screen the plants of the early generations and both isolates (RO1-1 and PO6-6) were used to test the selected BC2F5-generation plants followed by the gene-specific markers of Pi9. Rice seedlings (F1 to BC2F3 generation) were planted individually in tubes. After 14–18 days of the growth, the seedlings were spray-inoculated with a spore suspension (1 × 106 spores per mL) and incubated in a humid chamber for 14 days before assessing the resistance phenotype (Wu et al. 2015). The plants scored as highly resistant and moderately resistant were selected. The fully expanded leaves of 6-week-old plants (BC2F5- generation) were spot-inoculated with a spore suspension (2 × 106 spores per mL), as described previously (Kanzaki et al. 2002), in 2018. The length of brown lesions on leaves was measured at 9 dpi.
Submergence Tolerance and Anaerobic Germination Screening
Plants of CSA-Pi9, CSA (recurrent parent), IR64 (susceptible control), and IR64-Sub1 (resistant control) lines were grown for 2 weeks in a greenhouse, and submergence tolerance was tested as described previously (Septiningsih et al. 2009), with slight modifications. Briefly, seeds were surface-sterilized with a disinfectant in a 50-mL tube for 24 h, and then rinsed with water. The sterilized seeds were transferred to Petri dishes containing a moderate amount of water and incubated at 28 °C in the dark for 3 days. Pots (6 cm radius and 11 cm height) filled with air-dried soil were placed in a water-containing plastic box (20 cm height, 58.5 cm width, and 35 cm depth) for 1 day to soak the soil. Three seeds of each genotype were sown on the soil surface and covered with a 1-cm layer of soil. Forty-five seeds of each genotype were tested in a total of 15 pots, and 60 pots were arranged in a tray in a completely randomized design. Plants were grown under normal conditions for 2 weeks, with a constant water supply. The height and SPAD value of each seedling (total 45 seedlings) were measured at 14 days after transplanting (DAT). Then, 14-day-old plants were transferred to a 1-ton tank filled with 70-cm deep tap water. Water was drained from the tank after 14 days, and plants were covered with a fabric for 1 day to avoid dehydration under direct sunlight. Plants recovered for 7 days in a greenhouse under ambient conditions, and water was supplied to prevent drying. The height and SPAD value were measured at 14 days after submergence (DAS) using 21 seedlings of each genotype (The experiment started with 45 seedlings of each genotype. RNA sampling was performed at 6 points with 4 samples (3 samples of each genotype were used in experiment and 1 sample was extra against of sample loss), and remaining 21 seedlings of each genotype were evaluated.) Submergence screening was performed twice under natural conditions in 2018.
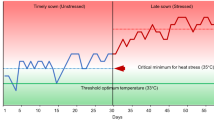
The germination of CSA-Pi9, CSA, IR64, and IR64-AG1 seeds was tested in 2018, as described previously (Angaji et al. 2010), with slight modifications. Two plastic boxes (20 cm height, 58.5 cm width, and 35 cm depth) were filled with an 8-cm layer of paddy field soil. Clods were broken apart with a trowel and sufficient volume of water. Each plastic box was divided into four sectors along its width using polyvinyl chloride (PVC) boards, and each sector was further divided into 10 sections along its length. To deoxidize the soil and water, one plastic box was placed in a chamber (anoxic conditions), while the other plastic box was placed in the greenhouse (hypoxic conditions) for 7 days. The chamber was maintained at a 33 °C day/28 °C night temperature under a 14-h light/10-h dark photoperiod, while the greenhouse was maintained under natural conditions. In each box, a total of 50 dried seeds of each genotype were sown in 10 rows (5 plants per row) at a soil depth of 1 cm. Water was added to each box every 3 days to maintain a 10-cm deep water layer throughout the experiment. The temperature of the water added to the box was approximately 30 °C to minimize oxygen provision. Seedlings that emerged above the water surface were counted, and the survival rate was calculated at 34 days after sowing. The coleoptile length and shoot length of three seedlings per genotype were measured at 3 days after the emergence of the last seedling. Anaerobic germination screening was conducted twice in the greenhouse and an environmentally controlled chamber, and genotypes were arranged in a completely randomized design in both experiments.
RNA Extraction and Gene Expression Analysis
To analyze the expression Sub1A, adh1, and OsTPP7 under submergence stress, three 14-day-old plants of each genotype (CSA-Pi9, CSA, IR64, and IR64-Sub1) were harvested at 0 (control; 14 days old plants), 1, 7, and 14 DAS. The harvested plants were rinsed with water in a tank, and excess water on the plants was soaked with a clean towel. The aerial and underground parts of each plant were separated and quickly frozen in liquid nitrogen. After 14 days, water was drained (de-submergence), and plant samples were collected at 1 and 7 DAR. To analyze the expression of OsTPP7 and adh1 during anaerobic germination, three seedlings of each genotype (CSA-Pi9, CSA, and IR64-AG1) were harvested at 34 days after sowing; IR64 seedlings could not be collected, as its seeds did not germinate.
Total RNA was extracted from the harvested aerial and underground parts of plants using the TRIzol Reagent (Thermo Fisher Scientific, MA, USA). First-strand cDNA (up to 1 µg/µL) was synthesized from the isolated total RNA using the Easy cDNA Synthesis Kit (NanoHelix, Daejeon, Korea), and diluted five times with distilled water. Quantitative PCR (qPCR) was performed on Mx3000 (Agilent, CA, USA) using the SensiFAST SYBR No-Rox Kit (Bioline, Meridian, London, UK). Each 20-µL qPCR reaction contained 200 ng of cDNA and 10 µM of sequence-specific forward and reverse primers (Additional file 2: Table S1; Bioneer, Daejeon, Korea); Sub1A-specific primers were designed in this study based on AAAA02037639.1, while primers used to amplify other genes have been published previously (Xu et al. 2006; Kretzschmar et al. 2015; Shin et al. 2020). The following conditions were used for qPCR: 95 °C for 10 min, followed by 40 cycles of 95 °C for 10 s, 61 °C for 15 s, and 72 °C for 20 s. Transcript levels of each gene were normalized to that of OsUBQ5 using the 2−ΔΔCt method (Livak and Schmittgen 2001).
Field Evaluation of Plant Agronomic Traits
Plants of CSA-Pi9 and CSA genotypes were grown in field A (Suwon, Korea; 37° 16′ 08.7″ N 126° 59′ 24.0″ E) in 2018 and 2019, and in fields B and C (western coastal reclaimed regions in Seosan, Korea; field B: 36° 39′ 59.6″ N, 126° 26′ 16.0″ E; field C: 36° 40′ 01.0″ N, 126° 22′ 30.6″ E) in 2019. Plants in field A were grown under normal conditions in both years (2018 and 2019). Pre-germinated seeds of the two genotypes were sown in the greenhouse on April 27, 2018, and April 26, 2019, and only healthy 40-day-old seedlings were transplanted into the field at a spacing of 30 cm × 15 cm (row-to-plant). Fertilizer (N–P–K, 21–11–21 kg/1000m2) was applied on the day of transplanting, and conventional pesticide control was applied. In 2019, plants in field B were grown under normal irrigation conditions, while those in field C were grown under high salinity conditions. To generate plant materials for fields B and C, seeds were sown in the greenhouse on April 21, 2019. Then, 35-day-old seedlings were transplanted into fields B and C at a spacing of 30 cm × 15 cm (row-to-plant). Conventional practice was used to fertilize the plants in field B; however, plants in field C were treated with excess N fertilizer (N–P–K, 30–7–9 kg/1000 m2). Because of severe soil salinity, an active water circulation practice was applied to field C. The salinity, electrical conductivity (EC) and pH of water in field C were measured using a water quality meter (WM-32EP, TOADKK, Japan). Plants in all fields were grown in a completely randomized block design (6 rows × 30 plants per row). Two blocks, each consisting of 12 rows, were tested together in the whole plot.
Five plants of each genotype were selected for agronomic evaluation. Culm length (distance from the surface of the soil to the neck of the panicle) and panicle length (distance between the neck and tip of the longest panicle) were measured using the tallest tiller of each plant. The number of productive tillers per plant was counted to determine the panicle number. The average values of spikelet number, fertility, and hundred-grain weight were calculated from three panicles per plant. Fertility was calculated as the ratio of the fertile spikelet number to the total spikelet number. Grain yield per plant was calculated from the grain weight of three selected plants. In field C, the plants were harvested in bulk, thus grain yield was calculated by dividing total grain weight into the number of bulked plants.
Ten grains each of CSA-Pi9 and CSA were selected to examine grain traits. Grain length and grain width were measured using the Vernier caliper (Mitutoyo, Japan). Brown rice recovery was estimated by the ratio of the weight of dehulled rice to that of hulled grain. Chalkiness was scored using ImageJ (https://imagej.net/ImageJ). Area of the chalky region, which was shown in white color against the clear region, was estimated using the scanned image of brown rice of each genotype. Alkali spreading value was analyzed using 1.4% potassium hydroxide and scored as previously described (Kim and Kim 2016). Brown rice was ground using the CT 293 Cyclotec (FOSS, Hilleroed, Denmark), and amylose content was estimated using a Lambda 650 UV/VIS spectrometer (Perkin Elmer Inc., Walatham, USA) with a linear regression curve (R2 = 0.995) at an absorbance of 620 nm. The crude protein content of brown rice was estimated using DA 7250 NIR Analyzer (PerkinElmer, UK). The statistical analysis of data was conducted using R version 3.6.3 (R-Studio, MA, USA) and SPSS 16.0 (IBM, NY, USA).
Availability of Data and Materials
Datasets generated in the current study are available from the corresponding author upon reasonable request.
Abbreviations
- ABA:
-
Abscisic acid
- adh1 :
-
Alcohol dehydrogenase 1
- Avr:
-
Avirulence
- CSA:
-
Ciherang-Sub1 + AG1
- CSA-Pi9:
-
Ciherang-Sub1 + AG1 + Pi9
- CTAB:
-
Cetyltrimethylammonium bromide
- DAR:
-
Days after recovery
- DAS:
-
Days after submergence
- DAT:
-
Days after transplanting
- dpi:
-
Days post-inoculation
- EC:
-
Electrical conductivity
- MABC:
-
Marker-assisted backcrossing
- MAS:
-
Marker-assisted selection
- NBS:
-
Nucleotide-binding site
- NBS-LRR:
-
Nucleotide-binding site leucine-rich repeat
- NIL:
-
Near isogenic line
- PVC:
-
Polyvinyl chloride
- qPCR:
-
Quantitative PCR
- QTL:
-
Quantitative trait locus
- SNP:
-
Single nucleotide polymorphism
References
Alam R, Hummel M, Yeung E, Locke AM, Ignacio JCI, Baltazar MD, Jia Z, Ismail AM, Septiningshi EM, Bailey-Serres J (2020) Flood resilience loci SUBMERGENCE 1 and ANAEROBIC GERMINATION 1 interact in seedlings established underwater. Plant Direct 4(7):e00240. https://doi.org/10.1002/pld3.240
Ali AJ, Xu JL, Ismail AM, Fu BY, Vijaykumar CHM, Gao YM, Domingo J, Maghirang R, Yu SB, Gregorio G, Yanaghihara S, Cohen M, Carmen B, Mackill DJ, Li ZK (2006) Hidden diversity for abiotic and biotic stress tolerances in the primary gene pool of rice revealed by a large backcross breeding program. Field Crop Res 97(1):66–76. https://doi.org/10.1016/j.fcr.2005.08.016
Amante-Bordeos A, Sitch LA, Nelson R, Dalmacio RD, Oliva NP, Aswidinnoor H, Leung H (1992) Transfer of bacterial blight and blast resistance from the tetraploid wild rice Oryza minuta to cultivated rice, Oryza sativa. Theor Appl Genet 84(3):345–354. https://doi.org/10.1007/BF00229493
Angaji SA, Septiningsih EM, Mackill DJ, Ismail AM (2010) QTLs associated with tolerance of flooding during germination in rice (Oryza sativa L.). Euphytica 172(2):159–168. https://doi.org/10.1007/s10681-009-0014-5
Arief RW, Asnawi R, Bakrie B (2018) Analysis of physical quality and nutrition of two rice varieties, with the implementation of various planting systems. Curr Agric Res J 6(1):20. https://doi.org/10.12944/CARJ.6.1.03
Asano T, Hayashi N, Kobayashi M, Aoki N, Miyao A, Mitsuhara I, Ichikawa H, Komatsu S, Hirochika H, Kikuchi S, Ohsugi R (2012) A rice calcium-dependent protein kinase OsCPK12 oppositely modulates salt-stress tolerance and blast disease resistance. Plant J 69(1):26–36. https://doi.org/10.1111/j.1365-313X.2011.04766.x
Bailey-Serres J, Fukao T, Ronald P, Ismail A, Heuer S, Mackill DJ (2010) Submergence tolerant rice: SUB1’s journey from landrace to modern cultivar. Rice 3(2):138–147. https://doi.org/10.1007/s12284-010-9048-5
Ballini E, Morel JB, Droc G, Price A, Courtois B, Notteghem JL, Tharreau D (2008) A genome-wide meta-analysis of rice blast resistance genes and quantitative trait loci provides new insights into partial and complete resistance. Mol Plant Microbe Interact 21(7):859–868. https://doi.org/10.1094/MPMI-21-7-0859
Baltazar MD, Ignacio JCI, Thomson MJ, Ismail AM, Mendioro MS, Septiningsih EM (2014) QTL mapping for tolerance of anaerobic germination from IR64 and the aus landrace Nanhi using SNP genotyping. Euphytica 197(2):251–260. https://doi.org/10.1007/s10681-014-1064-x
Baltazar MD, Ignacio JCI, Thomson MJ, Ismail AM, Mendioro MS, Septiningsih EM (2019) QTL mapping for tolerance to anaerobic germination in rice from IR64 and the aus landrace Kharsu 80A. Breed Sci 69(2):227–233. https://doi.org/10.1270/jsbbs.18159
Chen X, Shang J, Chen D, Lei C, Zou Y, Zhai W, Liu G, Xu J, Ling Z, Cao G, Ma B, Wang Y, Zhao X, Li S, Zhu L (2006) AB-lectin receptor kinase gene conferring rice blast resistance. Plant J 46(5):794–804. https://doi.org/10.1111/j.1365-313X.2006.02739.x
Chithrameenal K, Alagarasan G, Raveendran M, Robin S, Meena S, Ramanathan A, Ramalingam J (2018) Genetic enhancement of phosphorus starvation tolerance through marker assisted introgression of OsPSTOL1 gene in rice genotypes harbouring bacterial blight and blast resistance. PLoS ONE 13(9):e0204144. https://doi.org/10.1371/journal.pone.0204144
Chukwu SC, Rafii MY, Ramlee SI, Ismail SI, Hasan MM, Oladosu YA, Magaji UG, Akos I, Olalekan KK (2019) Bacterial leaf blight resistance in rice: a review of conventional breeding to molecular approach. Mol Biol Rep 46(1):1519–1532. https://doi.org/10.1007/s11033-019-04584-2
Chung YS, Yoon MB, Kim HS (2004) On climate variations and changes observed in South Korea. Clim Change 66(1):151–161. https://doi.org/10.1023/B:CLIM.0000043141.54763.f8
Collard BC, Gregorio GB, Thomson MJ, Islam MR, Vergara GV, Laborte AG, Nissila E, Kretzchmar T, Cobb JN (2019) Transforming rice breeding: re-designing the irrigated breeding pipeline at the International Rice Research Institute (IRRI). Crop Breed Genet Genomics 1(1):e190008. https://doi.org/10.20900/cbgg20190008
Couch BC, Kohn LM (2002) A multilocus gene genealogy concordant with host preference indicates segregation of a new species, Magnaporthe oryzae, from M. grisea. Mycologia 94(4):683–693. https://doi.org/10.1080/15572536.2003.11833196
Dar MH, Singh S, Singh US, Zaidi NW, Ismail AM (2014) Stress tolerant rice varieties-making headway in India. SATSA Mukhaptra Annual Technical Issue 18:1–14
Das G, Rao GJN (2015) Molecular marker assisted gene stacking for biotic and abiotic stress resistance genes in an elite rice cultivar. Front Plant Sci 6:698. https://doi.org/10.3389/fpls.2015.00698
Das G, Rao GJN, Varier M, Prakash A, Prasad D (2018) Improved Tapaswini having four BB resistance genes pyramided with six genes/QTLs, resistance/tolerance to biotic and abiotic stresses in rice. Sci Rep 8(1):1–16. https://doi.org/10.1038/s41598-018-20495-x
Dixit S, Singh UM, Singh AK, Alam S, Venkateshwarlu C, Nachimuthu VV, Yadav S, Abbai R, Selvaraj R, Devi MN, Ramayya PJ, Badri J, Ram T, Lakshmi J, Lakshmidevi G, Vidhya LRKJ, Phani A, Padmakumari LGS, Prasad MS, Seetalam M, Singh VK, Kumar A (2020) Marker assisted forward breeding to combine multiple biotic–abiotic stress resistance/tolerance in rice. Rice 13:1–15. https://doi.org/10.1186/s12284-020-00391-7
Elgamal WH, Elshenawy MM (2018) The Wild Oryza genomes. Springer, Cham, pp 193–202. https://doi.org/10.1007/978-3-319-71997-9
Emerick K, Ronald PC (2019) Sub1 rice: engineering rice for climate change. Cold Spring Harbor Perspect Biol 11(12):a034637
Feng B, Chen K, Cui Y, Wu Z, Zheng T, Zhu Y, Ali J, Wang B, Xu J, Zhang W, Li Z (2018) Genetic dissection and simultaneous improvement of drought and low nitrogen tolerances by designed QTL pyramiding in rice. Front Plant Sci 9:306. https://doi.org/10.3389/fpls.2018.00306
Fisher MC, Henk DA, Briggs CJ, Brownstein JS, Madoff LC, McCraw SL, Gurr SJ (2012) Emerging fungal threats to animal, plant and ecosystem health. Nature 484(7393):186–194. https://doi.org/10.1038/nature10947
Guo L, Chen W, Tao L, Hu B, Qu G, Tu B, Yuan H, Ma B, Wang Y, Zhu X, Qin P, Li S (2020) GWC1 is essential for high grain quality in rice. Plant Sci 296:110497. https://doi.org/10.1016/j.plantsci.2020.110497
Ha WG, Torollo GV, Kang KH, Lapiz M, Padolina TF (2011) Breeding temperate japonica rice MS11 for the tropics. Philipp J Crop Sci (Philipp) 36(1):111
Hirabayashi Y, Mahendran R, Koirala S, Konoshima L, Yamazaki D, Watanabe S, Kim H, Kanae S (2013) Global flood risk under climate change. Nat Clim Change 3(9):816–821. https://doi.org/10.1038/nclimate1911
Huang X, Jang S, Kim BK, Piao Z, Redona E, Koh HJ (2021) Evaluating genotype × environment interactions of yield traits and adaptability in rice cultivars grown under temperate. Subtrop Trop Environ Agric 11(6):558. https://doi.org/10.3390/agriculture11060558
IRRI (International Rice Research Institute) (2015) Growing rice, cultivating partnerships: 40 years of Indonesia-IRRI Collaboration. International Rice Research Institute, Philippines, p 32
Ismail AM, Singh US, Singh S, Dar MH, Mackill DJ (2013) The contribution of submergence-tolerant (Sub1) rice varieties to food security in flood-prone rainfed lowland areas in Asia. Field Crop Res 152:83–93. https://doi.org/10.1016/j.fcr.2013.01.007
Jiang SY, Ma Z, Ramachandran S (2010) Evolutionary history and stress regulation of the lectin superfamily in higher plants. BMC Evol Biol 10(1):1–24. https://doi.org/10.1186/1471-2148-10-79
Kanzaki H, Nirasawa S, Saitoh H, Ito M, Nishihara M, Terauchi R, Nakamura I (2002) Overexpression of the wasabi defensin gene confers enhanced resistance to blast fungus (Magnaporthe grisea) in transgenic rice. Theor Appl Genet 105(6):809–814. https://doi.org/10.1007/s00122-001-0817-9
Kato Y, Collard BC, Septiningsih EM, Ismail AM (2019) Increasing flooding tolerance in rice: combining tolerance of submergence and of stagnant flooding. Ann Bot 124(7):1199–1209. https://doi.org/10.1093/aob/mcz118
Khanh TD, Linh TH, Xuan TD (2013) Rapid and high-precision marker assisted backcrossing to introgress the SUB1 QTL into the Vietnamese elite rice variety. J Plant Breed Crop Sci 5(2):26–33. https://doi.org/10.5897/JPBCS12.052
Khanna A, Sharma V, Ellur RK, Shikari AB, Krishnan SG, Singh UD, Prakash G, Sharma TR, Rathour R, Variar M, Prashanthi SK, Nagarajan M, Vinod KK, Bhowmick PK, Rajashekhara H, Singh NK, Prabhu KV, Singh AK (2015) Marker assisted pyramiding of major blast resistance genes Pi9 and Pita in the genetic background of an elite Basmati rice variety, Pusa Basmati 1. Indian J Genet 75(4):417–425
Kim HY, Kim KM (2016) Mapping of grain alkali digestion trait using a Cheongcheong/Nagdong doubled haploid population in rice. J Plant Biotechnol 43(1):76–81. https://doi.org/10.5010/JPB.2016.43.1.76
Kim KW, Nam J, Chu SH et al (2019a) Development of KNU Axiom Oryza 580K Genotyping Array. In: KSBS & SABRAO International conference on plant breeding for sustainable development PCS02-58, p 158. https://www.breeding.or.kr/modules/setting/subPageManager/user/core/view/37/issuedBooks/inc/data/2019.pdf
Kim SM, Kim CS, Jeong JU, Reinke RF, Jeong JM (2019b) Marker-assisted breeding for improvement of anaerobic germination in japonica rice (Oryza sativa). Plant Breed 138(6):810–819. https://doi.org/10.1111/pbr.12719
Kretzschmar T, Pelayo MAF, Trijatmiko KR, Gabunada LFM, Alam R, Jimenez R, Septiningsih EM (2015) A trehalose-6-phosphate phosphatase enhances anaerobic germination tolerance in rice. Nature Plants 1(9):1–5. https://doi.org/10.1038/nplants.2015.124
Kuanar SR, Ray A, Sethi SK, Chattopadhyay K, Sarkar RK (2017) Physiological basis of stagnant flooding tolerance in rice. Rice Sci 24(2):73–84. https://doi.org/10.1016/j.rsci.2016.08.008
Kumar A, Kumar S, Dahiya K, Kumar S, Kumar M (2015) Productivity and economics of direct seeded rice (Oryza sativa L.). J Appl Nat Sci 7(1):410–416. https://doi.org/10.31018/jans.v7i1.625
Lal B, Gautam P, Nayak AK, Raja R, Shahid M, Tripathi R, Singh S, Septiningsih EM, Ismail AM (2018) Agronomic manipulations can enhance the productivity of anaerobic tolerant rice sown in flooded soils in rainfed areas. Field Crop Res 220:105–116. https://doi.org/10.1016/j.fcr.2016.08.026
Li Y, Fan C, Xing Y, Yun P, Luo L, Yan B, Peng B, Xie W, Wang G, Li X, Xiao J, Xu C, He Y (2014) Chalk5 encodes a vacuolar H+-translocating pyrophosphatase influencing grain chalkiness in rice. Nat Genet 46(4):398–404. https://doi.org/10.1038/ng.2923
Li W, Chern M, Yin J, Wang J, Chen X (2019) Recent advances in broad-spectrum resistance to the rice blast disease. Curr Opin Plant Biol 50:114–120. https://doi.org/10.1016/j.pbi.2019.03.015
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Mackill DJ, Ismail AM, Pamplona AM, Sanchez DL, Carandang JJ, Septiningsih EM (2010) Stress tolerant rice varieties for adaptation to a changing climate. Crop Environ Bioinform 7:250–259
Manangkil OE, Abdula SE, Quilang EJP (2020) Role of biotechnology in rice breeding strategy of the Philippines. Plant Biotechnol Rep 14(2):247–253. https://doi.org/10.1007/s11816-020-00618-5
Mansuri RM, Shobbar ZS, Jelodar NB, Ghaffari M, Mohammadi SM, Daryani P (2020) Salt tolerance involved candidate genes in rice: an integrative meta-analysis approach. BMC Plant Biol 20(1):1–14. https://doi.org/10.1186/s12870-020-02679-8
Miah G, Rafii MY, Ismail MR, Puteh AB, Rahim HA, Asfaliza R, Latif MA (2013) Blast resistance in rice: a review of conventional breeding to molecular approaches. Mol Biol Rep 40(3):2369–2388. https://doi.org/10.1007/s11033-012-2318-0
Mondal S, Hasan MJ, Ahmed T, Miah MG, Cruz PCS, Ismail AM (2020a) Effects of AG1 and AG2 QTLs on nonstructural carbohydrate and seed management options for rice seedling growth and establishment under flooding stress. Rice Sci 27(6):515–528. https://doi.org/10.1016/j.rsci.2020.09.008
Mondal S, Khan MIR, Entila F, Dixit S, Cruz PCS, Ali MP, Pittendrigh B, Septiningsih EM, Ismail AM (2020b) Responses of AG1 and AG2 QTL introgression lines and seed pre-treatment on growth and physiological processes during anaerobic germination of rice under flooding. Sci Rep 10(1):1–15. https://doi.org/10.1038/s41598-020-67240-x
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8(19):4321–4326. https://doi.org/10.1093/nar/8.19.4321
Muthu V, Abbai R, Nallathambi J, Rahman H, Ramasamy S, Kambale R, Thulasinathan T, Ayyenar B, Muthurajan R (2020) Pyramiding QTLs controlling tolerance against drought, salinity, and submergence in rice through marker assisted breeding. PLoS ONE 15(1):e0227421. https://doi.org/10.1371/journal.pone.0227421
Navea IP, Dwiyanti MS, Park JH, Kim BK, Lee SB, Huang X, Koh HJ, Chin JH (2017) Identification of quantitative trait loci for panicle length and yield related traits under different water and P application conditions in tropical region in rice (Oryza sativa L.). Euphytica 213(2):37. https://doi.org/10.1007/s10681-016-1822-z
Ning X, Yunyu W, Aihong L (2020) Strategy for use of rice blast resistance genes in rice molecular breeding. Rice Sci 27(4):263–277. https://doi.org/10.1016/j.rsci.2020.05.003
Pu CX, Han YF, Zhu S, Song FY, Zhao Y, Wang CY, Zhang YC, Yang Q, Wang J, Bu SL, Sun LJ, Zhang SW, Zhang SQ, Sun DY, Sun Y (2017) The rice receptor-like kinases DWARF AND RUNTISH SPIKELET1 and 2 repress cell death and affect sugar utilization during reproductive development. Plant Cell 29(1):70–89. https://doi.org/10.1105/tpc.16.00218
Qu S, Liu G, Zhou B, Bellizzi M, Zeng L, Dai L, Han B, Wang GL (2006) The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site–leucine-rich repeat protein and is a member of a multigene family in rice. Genetics 172(3):1901–1914. https://doi.org/10.1534/genetics.105.044891
Rafael AB, Barroga WV, Marcelo PNM, Septiningsih EM, Manangkil OE (2015) Introgression of Sub1 and AG1 tolerance genes to NSIC Rc222 through marker-assisted backcrossing. Philipp J Crop Sci (Philipp). https://agris.fao.org/agris-search/search.do?recordID=PH2016000155
Ram PC, Singh BB, Singh AK, Ram P, Singh PN, Singh HP, Boamfa L, Harren F, Santosa E, Jackson MB, Setter TL, Reuss J, Wade LJ, Singh VP, Singh RK (2002) Submergence tolerance in rainfed lowland rice: physiological basis and prospects for cultivar improvement through marker-aided breeding. Field Crop Res 76(2–3):131–152. https://doi.org/10.1016/S0378-4290(02)00035-7
Sandhu N, Yadav S, Kumar A (2020) Advances in developing multigene abiotic and biotic stress-tolerant rice varieties. In: Fahad S, Saud S, Chen Y, Wu C, Wang D (eds) Abiotic stress in plants. IntechOpen, London
Sartaj AB, Udawela KS, Herath HMVG (2016) Marker assisted backcross breeding of Bg 358 (Oryza sativa L.) for the anaerobic germination tolerant QTL AG1. Trop Agric Res 27(3):287–294. https://doi.org/10.4038/tar.v27i3.8207
Septiningsih EM, Mackill DJ (2018) Genetics and breeding of flooding tolerance in rice. In: Sasaki T, Ashikari M (eds) Rice genomics, genetics and breeding. Springer, Singapore, pp 275–295. https://doi.org/10.1007/978-981-10-7461-5
Septiningsih EM, Pamplona AM, Sanchez DL, Neeraja CN, Vergara GV, Heuer S, Ismail AM, Mackill DJ (2009) Development of submergence-tolerant rice cultivars: the Sub1 locus and beyond. Ann Bot 103(2):151–160. https://doi.org/10.1093/aob/mcn206
Septiningsih EM, Collard BC, Heuer S, Bailey-Serres J, Ismail AM, Mackill DJ (2013) Applying genomics tools for breeding submergence tolerance in rice. Transl Genomics Crop Breed 2:9–30
Septiningsih EM, Hidayatun N, Sanchez DL, Nugraha Y, Caranddang J, Pamplona AM, Collard BCY, Ismail AM, Mackil DJ (2015) Accelerating the development of new submergence tolerant rice varieties: the case of Ciherang-Sub1 and PSB Rc18-Sub1. Euphytica 202:259–268. https://doi.org/10.1007/s10681-014-1287-x
Shin NH, Han JH, Jang S, Song K, Koh HJ, Lee JH, Yoo SC, Chin JH (2020) Early vigor of a pyramiding line containing two quantitative trait loci, phosphorus uptake 1 (Pup1) and anaerobic germination 1 (AG1) in rice (O. sativa L.). Agriculture 10(10):453. https://doi.org/10.3390/agriculture10100453
Singh R, Singh Y, Xalaxo S, Verulkar S, Yadav N, Singh S, Singh N, Prasad KSN, Kondayya K, Rao PVR, Rani MG, Anuradha T, Suraynarayana Y, Sharma PC, Krishnamurthy SL, Sharma SK, Dwivedi JL, Singh AK, Singh PK, Nilanjay SNK, Kumar R, Chetia SK, Ahmad T, Rai M, Perraju P, Pande A, Singh DN, Mandal NP, Reddy JN, Singh ON, Katara JL, Marandi B, Swain P, Sarkar RK, Singh DP, Mohapatra T, Padmawathi G, Ram T, Kathiresan RM, Paramsivan K, Nadarajan S, Thirumeni S, Nagarajan M, Singh AK, Vikram P, Kumar A, Septiningsih E, Singh US, Ismail AM, Mackill D, Singh NK (2016) From QTL to variety-harnessing the benefits of QTLs for drought, flood and salt tolerance in mega rice varieties of India through a multi-institutional network. Plant Sci 242:278–287. https://doi.org/10.1016/j.plantsci.2015.08.008
Statistics Korea (2020) Korean Statistical Information Service. http://kostat.go.kr/portal/eng/pressReleases/8/3/index.board. Accessed 04 Mar 2021
Takai T, Lumanglas P, Simon EV, Arai-Sanoh Y, Asai H, Kobayashi N (2019) Identifying key traits in high-yielding rice cultivars for adaptability to both temperate and tropical environments. Crop J 7(5):685–693. https://doi.org/10.1016/j.cj.2019.06.004
Toledo AMU, Ignacio JCI, Casal C, Gonzaga ZJ, Mendioro MS, Septiningsih EM (2015) Development of improved Ciherang-Sub1 having tolerance to anaerobic germination conditions. Plant Breed Biotech 3(2):1–11. https://doi.org/10.9787/PBB.2015.3.2.077
Ton J, Flors V, Mauch-Mani B (2009) The multifaceted role of ABA in disease resistance. Trends Plant Sci 14(6):310–317. https://doi.org/10.1016/j.tplants.2009.03.006
Wilson RA, Talbot NJ (2009) Under pressure: investigating the biology of plant infection by Magnaporthe oryzae. Nat Rev Microbiol 7(3):185–195. https://doi.org/10.1038/nrmicro2032
Wu J, Kou Y, Bao J, Li Y, Tang M, Zhu X, Ponaya A, Xiao G, Li J, Li C, Song MY, Cumagun CJR, Deng Q, Lu G, Jeon JS, Naqvi NI, Zhou B (2015) Comparative genomics identifies the Magnaporthe oryzae avirulence effector Avr Pi9 that triggers Pi9-mediated blast resistance in rice. New Phytol 206(4):1463–1475. https://doi.org/10.1111/nph.13310
Wu Y, Xiao N, Chen Y, Yu L, Pan C, Li Y, Zhang X, Huang N, Ji H, Dai Z, Chen X, Li A (2019) Comprehensive evaluation of resistance effects of pyramiding lines with different broad-spectrum resistance genes against Magnaporthe oryzae in rice (Oryza sativa L.). Rice 12(1):1–13. https://doi.org/10.1186/s12284-019-0264-3
Xiao N, Wu Y, Pan C, Yu L, Chen Y, Liu G, Li Y, Zhang X, Wang Z, Dai Z, Liang C, Li A (2017) Improving of rice blast resistances in japonica by pyramiding major R genes. Front Plant Sci 7:1918. https://doi.org/10.3389/fpls.2016.01918
Xu K, Deb R, Mackill DJ (2004) A microsatellite marker and a codominant PCR-based marker for marker-assisted selection of submergence tolerance in rice. Crop Sci 44(1):248–253. https://doi.org/10.2135/cropsci2004.2480
Xu K, Xu X, Fukao T, Canlas P, Maghirang-Rodriguez R, Heuer S, Ismail AM, Bailey-Serres J, Ronald PC, Mackill DJ (2006) Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature 442(7103):705–708. https://doi.org/10.1038/nature04920
Zhao H, Wang X, Jia Y, Minkenberg B, Wheatley M, Fan J, Jia MH, Famoso A, Edwards JD, Wamishe Y, Valent B, Wang GL, Yang Y (2018) The rice blast resistance gene Ptr encodes an atypical protein required for broad-spectrum disease resistance. Nat Commun 9(1):1–12. https://doi.org/10.1038/s41467-018-04369-4
Zhou Y, Lei F, Wang Q, He W, Yuan B, Yuan W (2020) Identification of novel alleles of the rice blast-resistance gene Pi9 through sequence-based allele mining. Rice 13(1):1–15. https://doi.org/10.1186/s12284-020-00442-z
Dar MH, Singh S, Singh US, Zaidi NW, Ismail AM (2014) Stress tolerant rice varieties-making headway in India. SATSA Mukhaptra Annual Technical Issue 18:1-14
Acknowledgements
We thank Dr. Hong-Ryul Kim (Seoul National University) for guiding the field experiments at the Seoul National University Farm. We also thank Seok-Yong Yoon and Changhwan Yi (Saedlman Agriculture, Co., Ltd.) for testing the plant materials in the reclaimed area of Seosan. We appreciate Dr. Yeisoo Yu at DNACare Co., Ltd., for providing valuable comments and suggestions regarding the genotyping results, and Yeong Rok Kim (Gyeonggi-Do Agricultural Research & Extension Services) for assisting with grain quality analyses. We also acknowledge the valuable guidance provided by Dr. Casiana Vera Cruz (IRRI, Philippines) during the development of the Pi9-specific SNP marker.
Funding
This work was supported by a grant from the Cooperative Research Program for Agriculture Science & Technology Development (Project No. PJ015916), Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Contributions
JHC acquired the funding and initialized the project. JHC and JSJ designed the experiments, provided materials and supervised the project. NHS and JHH performed genomic analysis, collected abiotic stress related data and conducted field trials. NHS, JHH and JHS conducted statistical analysis and data visualization. KHXV and IPN conducted validation for biotic stress. SCY provided materials and conducted data analysis. NHS, JHH and JHS wrote the manuscript; JHC, JSJ and NHS revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Fig. S1.
Identification of a unique single nucleotide polymorphism (SNP), Nbs2-Pi9, by multiple sequence alignment of nucleotide-binding sites (Nbss) of the Pi9 gene cloned previously (Qu et al. 2006).
Additional file 2: Table S1.
List of primers used in this study.
Additional file 3: Table S2.
Statistics of CSA-Pi9-specific alleles on each chromosome.
Additional file 4: Table S3.
List of SNPs possibly associated with salinity tolerance in CSA-Pi9.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shin, NH., Han, JH., Vo, K.T.X. et al. Development of a Temperate Climate-Adapted indica Multi-stress Tolerant Rice Variety by Pyramiding Quantitative Trait Loci. Rice 15, 22 (2022). https://doi.org/10.1186/s12284-022-00568-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12284-022-00568-2