Abstract
We examined the gene expression profiles in arthroscopic biopsies retrieved from 10 rheumatoid arthritis patients before and after anti-TNF treatment with infliximab to investigate whether such profiles can be used to predict responses to the therapy, and to study effects of the therapy on the profiles. Responses to treatment were assessed using European League Against Rheumatism response criteria. Three patients were found to be good responders, five patients to be moderate responders and two patients to be nonresponders. The TNF-α status of the biopsies from each of the patients before treatment was also investigated immunohistochemically, and it was detected in biopsies from four of the patients, including all three of the good responders. The gene expression data demonstrate that all patients had unique gene expression signatures, with low intrapatient variability between biopsies. The data also revealed significant differences between the good responding and nonresponding patients (279 differentially expressed genes were detected, with a false discovery rate < 0.025). Among the identified genes we found that MMP-3 was significantly upregulated in good responders (log2 fold change, 2.95) compared with nonresponders, providing further support for the potential of MMP-3 as a marker for good responses to therapy. An even more extensive list of 685 significantly differentially expressed genes was found between patients in whom TNF-α was found and nonresponders, indicating that TNF-α could be an important biomarker for successful infliximab treatment. Significant differences were also observed between biopsies taken before and after anti-TNF treatment, including 115 differentially expressed genes in the good responding group. Interestingly, the effect was even stronger in the group in which TNF-α was immunohistochemically detected before therapy. Here, 1,058 genes were differentially expressed, including many that were novel in this context (for example, CXCL3 and CXCL14). Subsequent Gene Ontology analysis revealed that several 'themes' were significantly over-represented that are known to be affected by anti-TNF treatment in inflammatory tissue; for example, immune response (GO:0006955), cell communication (GO:0007154), signal transduction (GO:0007165) and chemotaxis (GO:0006935). No genes reached statistical significance in the moderately responding or nonresponding groups. In conclusion, this pilot study suggests that further investigation is warranted on the usefulness of gene expression profiling of synovial tissue to predict and monitor the outcome of rheumatoid arthritis therapies.
Similar content being viewed by others
Introduction
Rheumatoid arthritis (RA) is a chronic inflammatory disease currently defined by clinical criteria [1], but knowledge is scarce on the pathogenetic pathways involved, and the variations in this respect between different patients [2]. Since the main inflammatory processes occur in the inflamed synovial tissue, it is assumed that detailed analysis of molecular events in inflamed synovia may provide additional clues regarding the pathways involved and may identify new biomarkers for RA [3, 4]. A series of new targeted therapies have been introduced in recent years for the treatment of RA, and several more are anticipated [5–10]. The clinical response to currently available therapies is quite variable between patients, and although several of these treatments have well-defined molecular targets, such as TNF-α, our knowledge of the molecules and pathways that are affected in vivo is still limited [11]. Improved knowledge of individual patterns in the joints of patients and the effects of targeted therapies would provide new opportunities to elucidate the mode of action and to facilitate the design of individualized treatment strategies.
Current knowledge about interindividual differences in molecular patterns in inflamed RA joints originates from immunomorphological studies, which have shown both considerable heterogeneity between different patients [12] and certain striking common features, such as an abundance of macrophages and macrophage-derived proinflammatory cytokines in inflamed joints of RA patients [13, 14]. Few studies, however, have presented clear evidence that immunomorphologically recognized molecular patterns may reflect important pathogenetic differences between patients, or that such differences could be used to predict responses to therapy to specific drugs [12, 15, 16]. New high-throughput approaches are therefore needed that can analyze molecular patterns in many samples taken repeatedly during the course of a disease process, before and after specific therapies. A highly promising approach is microarray analysis, which allows the expression levels of thousands of genes to be monitored simultaneously and can therefore increase out understanding of the processes involved. Microarrays have previously been used to investigate various aspects of RA [2, 4, 17–22] and its potential uses within the field were recently reviewed by Jarvis [23].
Using a microarray approach we have previously demonstrated that gene expression varies less between biopsies sampled from one patient than between biopsies sampled from different patients [24]. These findings indicated that it would be meaningful to analyze synovial samples taken from individuals before and after a specific treatment, and to investigate possible correlations between changes in molecular patterns and the successfulness of the therapy. In the study presented here we used microarrays to investigate mRNA expression patterns in arthroscopically obtained synovial biopsies before and after 2 months of treatment with infliximab. Ten patients that were treated at 0 weeks, 2 weeks and 6 weeks were investigated. The clinical responses of the patients were assessed using the EULAR (European League Against Rheumatism) response criteria [25], and synovial tissue from each patient had previously been immunohistologically analyzed [16] for the presence of detectable levels of TNF-α before treatment (except for patient 10, see Materials and methods). Real-time PCR was used to verify the transcriptional data.
Materials and methods
Patients
Ten patients (eight females and two males, median age 54 years, range 25–69 years) meeting the American College of Rheumatology criteria for RA were recruited for the study (Table 1). All patients received infliximab infusions of 3 mg/kg at week 0 and again after 2 weeks and 6 weeks. Five of these patients were receiving prednisolone in doses lower than 7.5 mg/day, and all 10 patients were treated with methotrexate to a maximum of 20 mg/week. Patients were assessed for the overall activity of the disease using DAS28 (disease activity score including a 28-joint count) values before and after 12 weeks of treatment. EULAR response criteria were used to determine the patient responses to therapy after 12 weeks of treatment.
Synovial biopsies were obtained from all patients during arthroscopy before and after a median of 9 weeks of treatment (with one exception, patient 7, from whom the second biopsy was obtained during open surgery). The biopsies obtained were taken from the site of inflammation, which was classified as being either close to cartilage or not close to cartilage (defined as <1.5 cm or >1.5 cm away from cartilage, respectively). All biopsies were snap-frozen in precooled isopentane within 2 minutes to ensure high RNA quality. One biopsy was taken before and after treatment from patients 1, 5, 7, 9 and 10, while multiple biopsies were retrieved before and/or after treatment from patients 2, 3, 4, 6 and 8. Since only small amounts of RNA were available for patient 9, the RNA from this patient was only used in the second series of hybridizations (see below). The average biopsy weight was 22 mg. The ethical committee at the Karolinska Hospital approved all experiments on human cells and tissues. Informed consent was obtained from all study participants.
Additional immunohistochemical data for patient 10
Synovial biopsies from patient 10 were stained and scored for the presence of TNF-α as described by Ernestam and colleagues [16]. Briefly, TNF monoclonal antibodies 1 and 11 were scored using a semiquantitative four-point scale as follows: 0 = no positive cells, 1 = 1–10 positive cells, 2 = 11–100 positive cells, and 3 => 100 positive cells. The score for patient 10 was 1 (Table 1).
RNA extraction
RNA was successfully extracted from all biopsies essentially as previously described [24]. Briefly, RNA was extracted from the biopsies using steel-bead matrix tubes and a tabletop FastPrep homogenizer (Qbiogene, Irvie, California, USA). After homogenization, the tubes were centrifuged and the water phase from each sample was collected and transferred to a Qiashredder (Qiagen, Venlo, Netherlands) columns to ensure complete homogenization. The RNA in the samples was purified on RNeasy spin columns (Qiagen) and was treated with DNAse H. The extracted RNA was eluted in RNase-free water. For further details see the KTH microarray core facility webpage [26] ('Preparation of RNA from tissues' under 'Protocols'). The average biopsy weight used for RNA extraction was 22 mg and the average RNA yield was 387 ng/μl in a volume of 30 μl. RNA quality was assured using the RNA 6000 Nano LabChip kit of the Bioanalyzer system from Agilent Technologies (Santa Clara, California, USA).
RNA amplification
The extracted RNA was amplified by T7-based in vitro transcription using an Arcturus (Arcturus, Mountain View, California, USA) RiboAmp RNA amplification kit following the manufacturer's instructions. This procedure generates amplified RNA that consists mainly of sequences 250–1,800 bp long. Between 300 ng and 1 μg total RNA was used in each RNA amplification, and the average yield was 469 ng/μl in 11 μl H2O.
RNA reference
Universal Human Reference RNA from Stratagene (La Jolla, California, USA) was used as reference RNA, amplified in the same manner as the sample RNA. The reference RNA was pooled before it was used for hybridization.
Labeling and cDNA synthesis
Amplified RNA was labeled as previously described [24]. Briefly, amplified RNA was labeled using random hexamer primers in a cDNA synthesis reaction using Superscript III (Invitrogen, Carlsbad, California, USA) and an amino allyl-dUTP + dNTP mix. The mixture was incubated at 46°C for 2 hours. All purification steps were performed using a MinElute Reaction Cleanup Kit (Qiagen). The eluate from each sample was mixed with a dried portion of either Cy3 or Cy5 monoreactive esters (Amersham-Biosciences, Chalfont St Giles, UK) and was incubated for 30 minutes in a dark container at room temperature. After spin-column purification the samples were eluted and were ready for hybridization. For further information about the preparation of NHS-ester fluorophores and indirect labeling of cDNA, see the KTH microarray core facility webpage [26] (SOP 001 and SOP 002 at 'Protocols').
cDNA microarray
The cDNA arrays used in this study have been previously described [24]. Briefly, the clones on the array (spotted inhouse) originate from the first 310 96-well plates of a commercial clone collection containing 46,000 sequence-verified human cDNA clones (Research Genetics, now Invitrogen). According to UniGene mapping performed in September 2005, based on GenBank accession numbers (29,717 on the whole chip), 23,300 out of the 30,000 spots on the chip had UniGene IDs, and 13,426 of these were unique. For more information about the chip, see the KTH microarray core facility webpage [26].
Hybridization
The hybridization procedure has been previously described [24]. Briefly, after prehybridization, washing and centrifugation, the two samples (one Cy5-labeled and one Cy3-labeled) were pooled and hybridized in a water bath at 42°C for 14–18 hours. After hybridization, the slides were washed, centrifuged dry and scanned. For further information about the hybridization, see the KTH microarray core facility webpage [26] (SOP 003 under 'Protocols').
Scanning and image processing
An Agilent G2565BA scanner was used to scan the slides and acquire 50 MB TIFF images of them. The scanner resolution was set at 10 μm. GenePix 5.1.0.0 (Axon Instruments, Union City, California, USA) was used to extract the raw signals from the TIFF images and to assign each spot an ID. Spots defined as 'Not Found' by GenePix were flagged with a negative flag (-50) and were removed downstream in the analysis. Spots with a clearly abnormal morphology due to the presence of dust particles or other factors were manually flagged as bad (-100) and were also removed in downstream analysis. No further processing of the slides was performed in GenePix. The data are available at ArrayExpress, a public repository for microarray data (accession number E-TABM-104) [27].
Experimental design
Two series of hybridizations were performed. For each hybridization either RNA from two different samples or from one sample and reference RNA were labeled using different fluorophores (cy3 or cy5) and were hybridized onto the same array. Each hybridization was carried out using a dye-swapped technical replicate, which is a replicate with the same amplified RNA but with reversed fluorophores hybridized onto a separate array. The average correlation between the log2 fold changes (M values) of technical replicates was 0.8. In the first series of hybridizations, biopsies taken before treatment were hybridized in a common reference design to enable comparisons between patients. In the second series of hybridizations, biopsies taken before and after treatment were hybridized on the same chip in a direct design. A direct design has higher sensitivity since it provides better approximations of standard errors in comparison with the more common reference design, but the latter reference design allows more flexibility in downstream analysis [28].
Data analysis
Data were analyzed as previously described [24]. Briefly, the data were mainly analyzed with the help of software packages [26, 29–31] available in the R language [32]. After importing the result files (gpr files) produced by GenePix into R language, unreliable spots with abnormal physical properties were removed using four filters (described in [24] – more information can be found at the KTH package webpage [26]). On average 3,826 spots were removed by the filters, leaving approximately 26,174 spots for downstream analysis. After filtering, the slides were normalized using print-tip Loess normalization [33].
A moderated t test implemented in the LIMMA (Linear Models for Microarray Data) package [29, 34] was used to identify differentially expressed (DE) genes. The probability of falsely identifying DE genes due to problems arising from multiple testing was controlled using the false discovery rate approach [35], in which a q value (analogous to a P value) is assigned to each gene. A cutoff value was set at 0.025, which means that the expected proportion of false positives should be less than 0.025% among differentially expressed genes. When making comparisons instead of merely taking the average of replicates, the duplicateCorrelation function [36] available in LIMMA [29] was used to acquire an approximation of gene-wise variance. This retains valuable information about the variance when fitting a linear model to the data in order to identify differentially expressed genes.
Biopsies were taken both close to cartilage and not close to cartilage from patients 2, 3, 4 and 6. Data from these biopsies were not averaged. These data were instead treated as different biological replicates in the tests for differentially expressed genes since these two site types are known to represent different cellular populations [37, 38]. Functions available in Bioconductor [39] were used, with the help of Gene Ontology (GO) [40] to find 'biological process' GO terms over-represented in sets of DE genes. Only GO terms represented by more than five genes in the set of DE genes investigated were used to reduce the amount of false positives. Each GO category is given a P value, essentially utilizing Fisher's exact test [41]. The P values were corrected for multiple testing using the false discovery rate approach [35]. Several hierarchical clusterings were performed [42], in which 1 – Pearson correlation was used as the distance measure (for more detailed information see [24]).
Real-time PCR
Up to 1 μg total RNA was used in the first-strand synthesis reaction depending on the amount of material left, to which 2 μl 20TVN primer (4 μg/μl; Operon, Huntsville, Alabama, USA) was added to prime the reaction. The volume was adjusted to 15.5 μl using RNase-free water, was mixed and was then incubated for 10 minutes at 70°C to denature the RNA. The samples were then incubated for another 2 minutes on ice and were spun briefly. A cDNA synthesis mixture (12.5 μl) consisting of 6 μl of 5 × first-strand buffer, 3 μl of 0.1 M dithiothreitol, 2 μl Superscript III (Invitrogen) and 1.5 μl of 10 mM dNTP mix (Amersham) were added to each sample. The whole mixture was then incubated at 46°C for first-strand synthesis. After 1 hour the temperature was increased to 70°C for 15 minutes to terminate the reaction. Then 2 U RNase H (Invitrogen) was added to degrade the RNA. After RNAse treatment the temperature was increased to 70°C to inactivate the RNAse. All samples were then diluted with RNase-free water to a final volume of 200 μl.
Real-time PCR was performed using the iCycler system from Biorad (Hercules, California, USA). Each reaction was performed with 3.0 μl template, 12.5 μl iQSYBR Green Supermix (Biorad), 300 nM primer and water to adjust the final volume to 25 μl. The PCR amplification steps were applied in the following conditions: 3 minutes at 95°C, and then 40 cycles of 20 seconds at 94°C, 30 seconds at 60°C and 1 minute at 72°C. This was followed by melt curve analysis to ensure specific amplification. The signal was calculated in all experiments using the 'PCR base line subtracted RFU' and a cutoff value of 300 was used to establish threshold cycle values.
All primers were designed using Primer Express software (PE Applied Biosystems, Foster City, California, USA) and amplicons were designed to span exon-exon junctions to minimize contamination by genomic DNA. Primer sequence information is available in Additional file 1. Differences in relative expression levels between samples were calculated using the ΔΔCt method [43] and all samples were analyzed in triplicate. β-Actin was used as a reference gene. The following genes (with GenBank IDs indicating the sequences present on the microarray) were assayed in the real-time PCR: CCL19 (GenBank AA680186), CCL3 (GenBank AA677522), CXCL10 (GenBank AA878880), CXCL9 (GenBank AA131406), CXCL1 (GenBank W42723), CXCL3 (GenBank AA935273), IL1RN (GenBank T72877), FCGR1A (GenBank AA453258), CXCL1 (GenBank W42723), CXCL14 (GenBank AI016051) and TNF-α (GenBank AA699697).
Results
Two series of hybridizations were performed to investigate the differences between patients before treatment and to determine possible correlations between gene expression patterns that could be used to predict clinical responses, and to investigate gene expression patterns before and after treatment to identify potential molecular explanations for variations in the clinical responses. To facilitate these two tasks we determined and analyzed genes that were DE in different comparisons obtained from the microarray experiments. As already described in Materials and methods, a gene was considered DE if q < 0.025. EULAR response criteria were used to assess the clinical responses.
Hierarchical clustering of biopsies before treatment
Before comparing patients with differing clinical responses, a hierarchical clustering was performed on the 876 genes that had M values (sample versus universal RNA reference) with interquartile ranges > 1 (Figure 1) in the first series of hybridizations. This clustering was performed to obtain an overview of the differences between the technical replicates (that is, replicates of the same amplified RNA aliquot with reversed fluorophores, denoted A and B), the resampled biopsies (denoted 1 or 2) and the biological replicates in the form of different biopsy types (denoted close to cartilage or not close to cartilage) from the patients. The data show that all patients had a unique gene expression signature with low patient intravariability between biopsies, in accordance with previous work [24], but the clustering did not separate the patients clearly into groups correlating with their EULAR responses.
Hierarchical cluster dendrogram of the biopsies taken before treatment. A hierarchical cluster dendrogram of the 876 genes with M values (log2 fold change) with interquartile range > 1 in the first series of hybridizations. The clinical response of each patient is given in the plot (European League Against Rheumatism response criteria). pat, patient; cc, close to cartilage; ncc, not close to cartilage; biopsy 1 or 2 for each biopsy type, technical replicate A or B.
Differential gene expression between different responder groups
To further investigate whether the differential gene expression data retrieved from the biopsies taken before treatment correlated with the clinical responses, series of comparisons were performed between good responders (patients 5, 6 and 10), moderate responders (patients 2, 3, 7, 8 and 9) and nonresponders (patients 1 and 4). A total of 279 DE genes were obtained when good responders were compared with nonresponders. No genes reached statistical significance if good responders were compared with moderate responders and nonresponders. In contrast, 382 DE genes were statistically significant when the nonresponders were compared with those that displayed a response (moderate or good). Twelve genes were found to be DE between the patients in whom TNF-α was detected before treatment immunohistochemically (patients 3, 5, 6 and 10, hereafter denoted TNF-positive patients) and the TNF-negative patients. Interestingly, the highest number of DE genes (685) was found in the comparison of TNF-positive patients and nonresponders (Figure 2a). These results indicate that there were similarities within the defined responder groups and clear distinctions between the pretreatment gene signatures of good responders and nonresponders (Table 2).
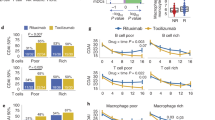
Volcano plots displaying differential expression. The y axis corresponds to the negative false discovery rate on a log10 scale. The x axis displays the M value (log2 fold change). Differential expression (proportion of false positives < 0.025): (a) between TNF-positive patients and nonresponders (685 differentially expressed (DE) genes) before treatment, (b) before and after treatment in the good responders (115 DE genes), (c) before and after treatment in the moderate responders (0 DE genes), (d) due to treatment in the nonresponders (no DE genes), (e) before and after treatment in all patients (974 DE genes), and (f) before and after treatment in the TNF-positive group of patients (1058 DE genes). TNF-α is highlighted in (e) and (f). FDR, false discovery rate.
Effect of treatment with infliximab
The effect of treatment with infliximab was investigated in the second series of hybridizations, in which RNA from biopsies taken before and after treatment was hybridized in a direct design. The results are presented in Figure 2b–f. These plots display the distribution of genes obtained with different statistical thresholds (y axis, negative false discovery rate on a log10 scale) in correlation to their M value (change due to treatment). In the good responders 115 genes were DE (Figure 2b), whereas no genes reached significance in the moderate responders or the nonresponding patients (Figure 2c,d). A total of 974 genes were DE (Figure 2e) when testing for the effect of treatment using all patients, but the largest effect (1,058 DE genes) was observed in the TNF-positive group of patients (Figure 2f, Additional file 8).
Over-representation analysis
To investigate whether any biological processes were over-represented among the genes that were DE before and after treatment in the TNF-positive patients and good responders, GO analysis was performed as described in Materials and methods [39]. For the TNF-positive patients (Figure 3) this revealed significant themes related to inflammatory disease and relevant for anti-TNF treatment, such as immune response (GO:0006955), cell communication (GO:0007154), signal transduction (GO:0007165) and chemotaxis (GO:0006935). Similar themes were also significant for good responders (see Additional files 9, 10 and 11).
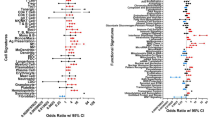
Gene Ontology analysis of the genes differentially expressed before and after treatment in the TNF-positive group of patients. The 20 most significant biological processes (Gene Ontology categories) and their parent terms are shown. The color of each node illustrates significance and can be interpreted in the scale bar, which displays the false discovery rate on a log10 scale. The arrows indicate the parent to child direction. The numbers of each arrow demonstrate the amount of unique Entrez Gene IDs [57] among the DE genes that are mapped to the parent term, and the number of unique Entrez Gene IDs among all genes present on the chip that are mapped to the parent term.
Verification
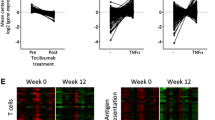
Real-time PCR was performed on unamplified total RNA. Three DE genes were verified before treatment between the TNF-positive patients and the nonresponders. As shown in Figure 4a, TNF-α was not DE according to results obtained using the microarray platform. The real-time PCR results indicated that TNF-α was upregulated in the patients in whom TNF-α was detectable before treatment, but the variation was high. The results from both approaches agreed in the direction of changes in CXCL14 and CXCL1 levels. The change of CXCL14 was higher according to results obtained with the microarray platform, but this is probably due to the high variation in the real-time PCR data for this gene. Verification experiments using real-time PCR were also performed on nine genes that were DE before and after treatment in TNF-positive patients. As displayed in Figure 4, the microarray and real-time PCR results agreed on the expression direction (either up-regulated or downregulated) of all genes but, as shown previously [44, 45], the changes were underestimated by the microarray relative to the real-time PCR results.
Real-time PCR results. (a) Real-time results for four genes in the first series of hybridizations comparing TNF-positive patients and nonresponders. (b) Real-time results for nine genes in the second series of hybridizations in the TNF-positive patients. The whiskers display the standard deviations in both (a) and (b).
Discussion
This investigation was a pilot study of the feasibility of using gene expression profiling to predict responses to TNF-blocking and to assess the effects of TNF-blocking on gene expression patterns in synovial tissues of RA patients. The results reveal significant differences between different response groups before treatment and identify genes that were significantly affected by treatment in patients who responded well to therapy and in patients where TNF could be detected immunohistochemically before treatment.
The first objective of this study was to investigate whether the gene expression signatures of the arthroscopic biopsies taken before treatment could predict the outcome of the treatment. In the hierarchical cluster (Figure 1) based on the gene expression signature before treatment, the patients with moderate/good EULAR responses tend to group together. A possible explanation for the limited overall degree of separation may be that the selection of the subset of genes chosen for clustering was based on variations between all biopsies, instead of being based on response groups where it was not clear a priori if the gene expression signature related to the TNF response would be relatively strong or weak. We therefore evaluated another strategy – investigating genes that are differentially expressed in groups of patients, classified on the basis of their response to therapy according to EULAR response criteria. Clustering of DE genes between, for example, nonresponders and good/moderate responders resulted in a clear distinction in the dendrogram between the two groups (see Additional file 2).
We also investigated differences in gene expression between the groups in which TNF was present and those in which TNF was absent in the inflamed joint tissue immunohistochemically. The obvious assumption was that TNF-dependent pathways can be assumed to play a more active role in the inflammation of patients when this molecule is present in substantial amounts prior to therapy. Previous research has suggested that the number of TNF-α-producing cells detected immunohistochemically in the rheumatoid synovium may indicate the response to infliximab therapy [12, 16]. In nonresponding patients, other molecules, and thus different pathogenic pathways, might be sustaining the chronic inflammation. A particularly large difference in genes expressed before and after therapy was seen in those patients who showed both TNF expression before therapy and a good clinical response, providing indirect additional support for this hypothesis. Analysis of differentially expressed genes before and after treatment in all patients simultaneously resulted almost as many significant genes were detected (974 DE genes) as in the TNF-positive group, but as exemplified by TNF-α in figures 2e and 2f important observations can be missed if confounding cohorts used to test for a specific effect with unaffected/moderately effected patients. Thus, even though one would be able to detect differences due to treatment using all patients, the heterogeneity of the group will make it difficult to differentiate small differences from noise (TNF-α, q = 0.09 using all patients). Another obvious conclusion is that other targets need to be identified for the treatment of nonresponders.
Several interesting GO terms were over-represented in the genes differentially expressed before and after treatment in the TNF-positive group (Figure 3). The genes mapped to significant GO terms are available in Additional file 10. Previous work has documented several mechanisms of action of TNF antagonists in RA, the most prominent being downregulation of synovial inflammation [12, 46], reduction of cell migration into the synovium [47], modulation of cartilage and bone destruction markers [48–50] and induction of apoptosis [51]. It is relevant to note in this context that our GO analysis identified modulation of chemotaxis, immune functions, inflammatory responses and signal transduction as potentially relevant mechanisms. Our findings therefore confirm the relevance of the previously described mechanisms of action for this type of RA therapy. Moreover, these findings may facilitate the selection of better target molecules for further studies regarding the mechanisms of action of TNF antagonists (such as the FcgRs, Toll-like receptors and the IL18 receptor). This study might also aid in suggesting new pathways to be studied in a more focused approach with confirmation at the protein levels and investigation of the clinical significance.
Many of the chemokines that were downregulated due to treatment have previously been detected in the synovial membrane and analyzed by immunohistochemical techniques. Some of them are novel in this context, however; for example, CXCL3, which was down-regulated following treatment, and CXCL14, which had a higher level of expression in nonresponders before treatment. The downregulation of CXCL3, a chemoattractant of neutrophils, natural killer cells and cytotoxic T cells [52], is consistent with earlier studies showing that the influx of inflammatory cells is downregulated by TNF therapy [53]. CXCL14 has not been reported to be significant in the context of rheumatic tissue but has been shown by Shurin and coworkers [54] to be a potent chemoattractant and activator of dendritic cells, which have been proposed to play a role in the initiation and perpetuation of RA by presentation of arthritigenic antigens to autoreactive T cells [55]. CXCL1, known to be an angiogenic protein, is particularly interesting since its expression was higher in good responders than in nonresponders before treatment, and was also found to be downregulated in good responders following treatment. CXCL1 has previously been shown [56] to be induced by TNF-α and IL-1β. In this study CXCL1 was found to be downregulated, as was TNF-α. IL-1β was also downregulated (q value = 0.026), but did not reach our threshold for statistical significance (q value = 0.025).
Another interesting finding is related to the expression of MMP-3 at the synovium level. We have previously shown that high levels of MMP-3 in serum before treatment may indicate good responses to therapy, as evaluated by the change in DAS28 (disease activity score including a 28-joint count) values [48]. Our analysis of differential expression between good and poor responders revealed an upregulation of MMP-3 in the synovial tissue of good responders relative to nonresponders. These findings suggest that further confirmatory studies on larger numbers of patients are warranted, with the ultimate aim of identifying a biological marker that could be used to predict responses to treatment.
Conclusion
The present paper describes the use of an mRNA expression array technique to analyze synovial biopsies arthroscopically obtained before and after treatment with infliximab. Three major results were obtained. First, we confirmed our previous observations that intrapatient variations in biopsies are smaller than interpatient variations, reflecting unique mRNA signatures of each patient rather than tissue heterogeneity. Secondly, we demonstrated significant pretreatment differences in gene expression between patient groups, the largest being between the patients with positive immunohistochemical scores for TNF-α and nonresponding patients. Thirdly, we also observed genes differentially expressed before and after infliximab therapy among the good responders, but the largest differences were found when testing for effect of treatment in the patients where TNF-α could be detected before treatment.
In summary, these results show the potential of using gene expression profiling to elucidate the effects of anti-TNF treatment in RA patients using synovial biopsies and indicate the possibility of identifying gene expression signatures predictive of good responses.
Abbreviations
- bp:
-
= base pair
- DE:
-
= differentially expressed
- EULAR:
-
= European League Against Rheumatism
- GO:
-
= Gene Ontology
- IL:
-
= interleukin
- LIMMA:
-
= Linear Models for Microarray Data
- M value:
-
= log2 fold change
- PCR:
-
= polymerase chain reaction
- RA:
-
= rheumatoid arthritis
- TNF:
-
= tumor necrosis factor.
References
Arnett FC, Edworthy SM, Bloch DA, McShane DJ, Fries JF, Cooper NS, Healey LA, Kaplan SR, Liang MH, Luthra HS, et al: The American Rheumatism Association 1987 revised criteria for the classification of rheumatoid arthritis. Arthritis Rheum. 1988, 31: 315-324.
van der Pouw Kraan TC, van Gaalen FA, Kasperkovitz PV, Verbeet NL, Smeets TJ, Kraan MC, Fero M, Tak PP, Huizinga TW, Pieterman E, et al: Rheumatoid arthritis is a heterogeneous disease: evidence for differences in the activation of the STAT-1 pathway between rheumatoid tissues. Arthritis Rheum. 2003, 48: 2132-2145. 10.1002/art.11096.
Ulfgren AK, Lindblad S, Klareskog L, Andersson J, Andersson U: Detection of cytokine producing cells in the synovial membrane from patients with rheumatoid arthritis. Ann Rheum Dis. 1995, 54: 654-661.
van der Pouw Kraan TC, van Gaalen FA, Huizinga TW, Pieterman E, Breedveld FC, Verweij CL: Discovery of distinctive gene expression profiles in rheumatoid synovium using cDNA microarray technology: evidence for the existence of multiple pathways of tissue destruction and repair. Genes Immun. 2003, 4: 187-196. 10.1038/sj.gene.6363975.
Maini R, St Clair EW, Breedveld F, Furst D, Kalden J, Weisman M, Smolen J, Emery P, Harriman G, Feldmann M, et al: Infliximab (chimeric anti-tumour necrosis factor alpha monoclonal antibody) versus placebo in rheumatoid arthritis patients receiving concomitant methotrexate: a randomised phase III trial. ATTRACT Study Group. Lancet. 1999, 354: 1932-1939. 10.1016/S0140-6736(99)05246-0.
Moreland LW, Baumgartner SW, Schiff MH, Tindall EA, Fleischmann RM, Weaver AL, Ettlinger RE, Cohen S, Koopman WJ, Mohler K, et al: Treatment of rheumatoid arthritis with a recombinant human tumor necrosis factor receptor (p75)-Fc fusion protein. N Engl J Med. 1997, 337: 141-147. 10.1056/NEJM199707173370301.
Klareskog L, van der Heijde D, de Jager JP, Gough A, Kalden J, Malaise M, Martin Mola E, Pavelka K, Sany J, Settas L, et al: Therapeutic effect of the combination of etanercept and methotrexate compared with each treatment alone in patients with rheumatoid arthritis: double-blind randomised controlled trial. Lancet. 2004, 363: 675-681. 10.1016/S0140-6736(04)15640-7.
Weinblatt ME, Keystone EC, Furst DE, Moreland LW, Weisman MH, Birbara CA, Teoh LA, Fischkoff SA, Chartash EK: Adalimumab, a fully human anti-tumor necrosis factor alpha monoclonal antibody, for the treatment of rheumatoid arthritis in patients taking concomitant methotrexate: the ARMADA trial. Arthritis Rheum. 2003, 48: 35-45. 10.1002/art.10697.
Furst DE, Schiff MH, Fleischmann RM, Strand V, Birbara CA, Compagnone D, Fischkoff SA, Chartash EK: Adalimumab, a fully human anti tumor necrosis factor-alpha monoclonal antibody, and concomitant standard antirheumatic therapy for the treatment of rheumatoid arthritis: results of STAR (Safety Trial of Adalimumab in Rheumatoid Arthritis). J Rheumatol. 2003, 30: 2563-2571.
Kavanaugh A: Anakinra (interleukin-1 receptor antagonist) has positive effects on function and quality of life in patients with rheumatoid arthritis. Adv Ther. 2006, 23: 208-217.
Feldmann M, Brennan FM, Foxwell BM, Taylor PC, Williams RO, Maini RN: Anti-TNF therapy: where have we got to in 2005?. J Autoimmun. 2005, 25 (Suppl): 26-28. 10.1016/j.jaut.2005.09.006.
Ulfgren AK, Andersson U, Engstrom M, Klareskog L, Maini RN, Taylor PC: Systemic anti-tumor necrosis factor alpha therapy in rheumatoid arthritis down-regulates synovial tumor necrosis factor alpha synthesis. Arthritis Rheum. 2000, 43: 2391-2396. 10.1002/1529-0131(200011)43:11<2391::AID-ANR3>3.0.CO;2-F.
Firestein GS: Inhibiting inflammation in rheumatoid arthritis. N Engl J Med. 2006, 354: 80-82. 10.1056/NEJMcibr054344.
Kinne RW, Brauer R, Stuhlmuller B, Palombo-Kinne E, Burmester GR: Macrophages in rheumatoid arthritis. Arthritis Res. 2000, 2: 189-202. 10.1186/ar86.
Bresnihan B, Baeten D, Firestein GS, Fitzgerald OM, Gerlag DM, Haringman JJ, McInnes IB, Reece RJ, Smith MD, Ulfgren AK, et al: Synovial tissue analysis in clinical trials. J Rheumatol. 2005, 32: 2481-2484.
Ernestam S, Af Klint E, Catrina AI, Sundberg E, Engstrom M, Klareskog L, Ulfgren AK: Synovial expression of IL-15 in rheumatoid arthritis is not influenced by blockade of tumour necrosis factor. Arthritis Res Ther. 2006, 8: R18-10.1186/ar1871.
Batliwalla FM, Baechler EC, Xiao X, Li W, Balasubramanian S, Khalili H, Damle A, Ortmann WA, Perrone A, Kantor AB, et al: Peripheral blood gene expression profiling in rheumatoid arthritis. Genes Immun. 2005, 6: 388-397. 10.1038/sj.gene.6364209.
Devauchelle V, Marion S, Cagnard N, Mistou S, Falgarone G, Breban M, Letourneur F, Pitaval A, Alibert O, Lucchesi C, et al: DNA microarray allows molecular profiling of rheumatoid arthritis and identification of pathophysiological targets. Genes Immun. 2004, 5: 597-608. 10.1038/sj.gene.6364132.
Gallagher J, Howlin J, McCarthy C, Murphy EP, Bresnihan B, FitzGerald O, Godson C, Brady HR, Martin F: Identification of Naf1/ABIN-1 among TNF-alpha-induced expressed genes in human synoviocytes using oligonucleotide microarrays. FEBS Lett. 2003, 551: 8-12. 10.1016/S0014-5793(03)00823-8.
Huh SJ, Paik DJ, Chung HS, Youn J: Regulation of GRB2 and FLICE2 expression by TNF-alpha in rheumatoid synovium. Immunol Lett. 2003, 90: 93-96. 10.1016/j.imlet.2003.07.002.
Olsen N, Sokka T, Seehorn CL, Kraft B, Maas K, Moore J, Aune TM: A gene expression signature for recent onset rheumatoid arthritis in peripheral blood mononuclear cells. Ann Rheum Dis. 2004, 63: 1387-1392. 10.1136/ard.2003.017194.
Tsubaki T, Arita N, Kawakami T, Shiratsuchi T, Yamamoto H, Takubo N, Yamada K, Nakata S, Yamamoto S, Nose M: Characterization of histopathology and gene-expression profiles of synovitis in early rheumatoid arthritis using targeted biopsy specimens. Arthritis Res Ther. 2005, 7: R825-R836. 10.1186/ar1751.
Jarvis JN: Diagnostic and prognostic potential of gene microarrays in rheumatoid arthritis. Expert Rev Mol Diagn. 2005, 5: 655-659. 10.1586/14737159.5.5.655.
Lindberg J, af Klint E, Ulfgren AK, Stark A, Andersson T, Nilsson P, Klareskog L, Lundeberg J: Variability in synovial inflammation in rheumatoid arthritis investigated by microarray technology. Arthritis Res Ther. 2006, 8: R47-10.1186/ar1903.
van Gestel AM, Prevoo ML, van 't Hof MA, van Rijswijk MH, van de Putte LB, van Riel PL: Development and validation of the European League Against Rheumatism response criteria for rheumatoid arthritis. Comparison with the preliminary American College of Rheumatology and the World Health Organization/International League Against Rheumatism Criteria. Arthritis Rheum. 1996, 39: 34-40.
KTH microarray core facility homepage. [http://www.ktharray.se/]
Parkinson H, Sarkans U, Shojatalab M, Abeygunawardena N, Contrino S, Coulson R, Farne A, Lara GG, Holloway E, Kapushesky M, et al: ArrayExpress – a public repository for microarray gene expression data at the EBI. Nucleic Acids Res. 2005, D553-D555. 33 Database
Yang YH, Speed T: Design issues for cDNA microarray experiments. Nat Rev Genet. 2002, 3: 579-588.
Smyth GK: Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004, 3: Article3-
Aroma – an R object-oriented microarray analysis environment. [http://www.maths.lth.se/help/R/aroma]
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, et al: Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004, 5: R80-10.1186/gb-2004-5-10-r80.
R: A language and environment for statistical computing. [http://cran.r-project.org]
Smyth GK, Speed TP: Normalization of cDNA microarray data. Methods. 2003, 31: 265-273. 10.1016/S1046-2023(03)00155-5.
Lonnstedt I, Speed TP: Replicated microarray data. Stat Sin. 2002, 12: 31-46.
Reiner A, Yekutieli D, Benjamini Y: Identifying differentially expressed genes using false discovery rate controlling procedures. Bioinformatics. 2003, 19: 368-375. 10.1093/bioinformatics/btf877.
Smyth GK, Michaud J, Scott HS: The use of within-array replicate spots for assessing differential expression in microarray experiments. Bioinformatics. 2005, 21: 2067-75. 10.1093/bioinformatics/bti270.
Smeets TJ, Kraan MC, Galjaard S, Youssef PP, Smith MD, Tak PP: Analysis of the cell infiltrate and expression of matrix metalloproteinases and granzyme B in paired synovial biopsy specimens from the cartilage-pannus junction in patients with RA. Ann Rheum Dis. 2001, 60: 561-565. 10.1136/ard.60.6.561.
Youssef PP, Kraan M, Breedveld F, Bresnihan B, Cassidy N, Cunnane G, Emery P, Fitzgerald O, Kane D, Lindblad S, et al: Quantitative microscopic analysis of inflammation in rheumatoid arthritis synovial membrane samples selected at arthroscopy compared with samples obtained blindly by needle biopsy. Arthritis Rheum. 1998, 41: 663-669. 10.1002/1529-0131(199804)41:4<663::AID-ART13>3.0.CO;2-L.
GOstats: Tools for Manipulating GO and Microarrays. [http://www.bioconductor.org]
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al: Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000, 25: 25-29. 10.1038/75556.
Fleiss JL: Statistical Methods for Rates and Proportions. 1981, New York: John Wiley
Eisen MB, Spellman PT, Brown PO, Botstein D: Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA. 1998, 95: 14863-14868. 10.1073/pnas.95.25.14863.
Livak KJ, Schmittgen TD: Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001, 25: 402-408. 10.1006/meth.2001.1262.
Yuen T, Wurmbach E, Pfeffer RL, Ebersole BJ, Sealfon SC: Accuracy and calibration of commercial oligonucleotide and custom cDNA microarrays. Nucleic Acids Res. 2002, 30: e48-10.1093/nar/30.10.e48.
Dallas PB, Gottardo NG, Firth MJ, Beesley AH, Hoffmann K, Terry PA, Freitas JR, Boag JM, Cummings AJ, Kees UR: Gene expression levels assessed by oligonucleotide microarray analysis and quantitative real-time RT-PCR – how well do they correlate?. BMC Genom. 2005, 6: 59-. 10.1186/1471-2164-6-59.
Smeets TJ, Kraan MC, van Loon ME, Tak PP: Tumor necrosis factor alpha blockade reduces the synovial cell infiltrate early after initiation of treatment, but apparently not by induction of apoptosis in synovial tissue. Arthritis Rheum. 2003, 48: 2155-2162. 10.1002/art.11098.
Tak PP, Taylor PC, Breedveld FC, Smeets TJ, Daha MR, Kluin PM, Meinders AE, Maini RN: Decrease in cellularity and expression of adhesion molecules by anti-tumor necrosis factor alpha monoclonal antibody treatment in patients with rheumatoid arthritis. Arthritis Rheum. 1996, 39: 1077-1081.
Catrina AI, Lampa J, Ernestam S, af Klint E, Bratt J, Klareskog L, Ulfgren AK: Anti-tumour necrosis factor (TNF)-alpha therapy (etanercept) down-regulates serum matrix metalloproteinase (MMP)-3 and MMP-1 in rheumatoid arthritis. Rheumatology (Oxford). 2002, 41: 484-489. 10.1093/rheumatology/41.5.484.
Catrina AI, af Klint E, Ernestam S, Catrina SB, Makrygiannakis D, Botusan IR, Klareskog L, Ulfgren AK: Anti-tumor necrosis factor therapy increases synovial osteoprotegerin expression in rheumatoid arthritis. Arthritis Rheum. 2006, 54: 76-81. 10.1002/art.21528.
Brennan FM, Browne KA, Green PA, Jaspar JM, Maini RN, Feldmann M: Reduction of serum matrixmetalloproteinase 1 and matrix metalloproteinase 3 in rheumatoid arthritis patients following anti-tumour necrosis factor-alpha (cA2) therapy. Br J Rheumatol. 1997, 36: 643-650. 10.1093/rheumatology/36.6.643.
Catrina AI, Trollmo C, af Klint E, Engstrom M, Lampa J, Hermansson Y, Klareskog L, Ulfgren AK: Evidence that anti-tumor necrosis factor therapy with both etanercept and infliximab induces apoptosis in macrophages, but not lymphocytes, in rheumatoid arthritis joints: extended report. Arthritis Rheum. 2005, 52: 61-72. 10.1002/art.20764.
Yang SK, Eckmann L, Panja A, Kagnoff MF: Differential and regulated expression of C-X-C, C-C, and C-chemokines by human colon epithelial cells. Gastroenterology. 1997, 113: 1214-1223. 10.1053/gast.1997.v113.pm9322516.
Tak PP, Smeets TJ, Daha MR, Kluin PM, Meijers KA, Brand R, Meinders AE, Breedveld FC: Analysis of the synovial cell infiltrate in early rheumatoid synovial tissue in relation to local disease activity. Arthritis Rheum. 1997, 40: 217-225.
Shurin GV, Ferris R, Tourkova IL, Perez L, Lokshin A, Balkir L, Collins B, Chatta GS, Shurin MR: Loss of new chemokine CXCL14 in tumor tissue is associated with low infiltration by dendritic cells (DC), while restoration of human CXCL14 expression in tumor cells causes attraction of DC both in vitro and in vivo. J Immunol. 2005, 174: 5490-5498.
Jongbloed SL, Lebre MC, Fraser AR, Gracie JA, Sturrock RD, Tak PP, McInnes IB: Enumeration and phenotypical analysis of distinct dendritic cell subsets in psoriatic arthritis and rheumatoid arthritis. Arthritis Res Ther. 2006, 8: R15-10.1186/ar1864.
Koch AE, Kunkel SL, Shah MR, Hosaka S, Halloran MM, Haines GK, Burdick MD, Pope RM, Strieter RM: Growth-related gene product alpha. A chemotactic cytokine for neutrophils in rheumatoid arthritis. J Immunol. 1995, 155: 3660-3666.
Maglott D, Ostell J, Pruitt KD, Tatusova T: Entrez Gene: gene-centered information at NCBI. Nucleic Acids Res. 2005, D54-D58. 33 Database
Acknowledgements
This work was supported by the Swedish Knowledge Foundation through the Industrial PhD program in Medical Bioinformatics at the Centre for Medical Innovations (CMI) at the Karolinska Institute. The work was also supported by the Swedish Rheumatism Association, King Gustav V's 80 years Foundation, the Åke Wiberg Foundation, the Swedish Research Council, the insurance company AFA, and the Freemason Lodge 'Barnhuset' in Stockholm. Marianne Engström and Anna Westring are thanked for their technical assistance and Annelie Walden is thanked for chip production.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
JL performed parts of the microarray-related laboratory work, contributed to the data analysis and wrote parts of the article. EaK performed the arthroscopic surgery and contributed to the data analysis. AIC participated in data analysis and writing of the article. PN was responsible for chip production. LK was involved in planning the project, data analysis and writing of the article. A-KU contributed to the planning and design of the project and participated in both the data analysis and writing of the article. JL was involved in project design, analysis of laboratory results, data analysis and writing of the article.
Electronic supplementary material
13075_2006_1955_MOESM2_ESM.pdf
Additional file 2: A pdf file containing figures for Table 2. (PDF 87 KB)
13075_2006_1955_MOESM3_ESM.xls
Additional file 3: An Excel file containing a list of DE genes of EULAR good responders versus nonresponders for Table 2. (XLS 90 KB)
13075_2006_1955_MOESM4_ESM.xls
Additional file 4: An Excel file containing DE genes of TNF-positive patients versus TNF-negative patients for Table 2. (XLS 284 KB)
13075_2006_1955_MOESM5_ESM.xls
Additional file 5: An Excel file containing DE genes of EULAR nonresponders versus responders for Table 2. (XLS 102 KB)
13075_2006_1955_MOESM6_ESM.xls
Additional file 6: An Excel file containing DE genes of TNF-positive patients versus EULAR nonresponders for Table 2. (XLS 428 KB)
13075_2006_1955_MOESM7_ESM.xls
Additional file 7: An Excel file containing GO analysis of DE genes between TNF-positive patients and nonresponders for Table 2. (XLS 40 KB)
13075_2006_1955_MOESM8_ESM.xls
Additional file 8: An Excel file containing a list of the DE genes due to treatment in the TNF-positive group of patients. (XLS 518 KB)
13075_2006_1955_MOESM9_ESM.pdf
Additional file 9: A pdf file containing an image of GO analysis of the DE genes due to treatment in EULAR good responders. (PDF 106 KB)
13075_2006_1955_MOESM10_ESM.xls
Additional file 10: An Excel file containing a list of GO categories (biological process) of DE genes due to treatment in TNF-positive patients. Each category is assigned a q value to establish over-representation of the specific category among the DE genes. (XLS 126 KB)
13075_2006_1955_MOESM11_ESM.xls
Additional file 11: An Excel file containing a list of GO categories (biological process) of DE genes due to treatment in EULAR good responders. Each category is assigned a q value to establish over-representation of the specific category among the DE genes. (XLS 28 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Lindberg, J., af Klint, E., Catrina, A.I. et al. Effect of infliximab on mRNA expression profiles in synovial tissue of rheumatoid arthritis patients. Arthritis Res Ther 8, R179 (2006). https://doi.org/10.1186/ar2090
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1186/ar2090