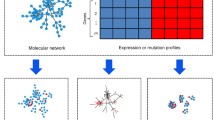
Abstract
Identification and validation of interaction networks and network biomarkers have become more critical and important in the development of disease-specific biomarkers, which are functionally changed during disease development, progression or treatment. The present review headlined the definition, significance, research and potential application for network biomarkers, interaction networks and dynamical network biomarkers (DNB). Disease-specific interaction networks, network biomarkers, or DNB have great significance in the understanding of molecular pathogenesis, risk assessment, disease classification and monitoring, or evaluations of therapeutic responses and toxicities. Protein-based DNB will provide more information to define the differences between the normal and pre-disease stages, which might point to early diagnosis for patients. Clinical bioinformatics should be a key approach to the identification and validation of disease-specific biomarkers.
Similar content being viewed by others
Introduction
The respiratory disease is a complex process from self-limiting to life-threatening entities such as from the chronic obstructive pulmonary disease (COPD) to respiratory failure, pulmonary embolism and lung cancer. For example, COPD will be one of the top five chronic diseases in terms of global mortality and morbidity by 2030 [1–3]. One of the major challenges in the respiratory medicine is the lack of disease-specific biomarkers for disease diagnosis, illness monitoring, therapy evaluation, and prognosis prediction. The biomarker should be a measurable indicator of normal biologic processes, pathogenic processes, or therapeutic responses, for the risk assessment, early diagnosis, and predicting and monitoring responses to therapies and toxicities [4, 5]. Disease-specific biomarkers are also expected to demonstrate the disease-associated specificity, sensitivity, traceability, stability, repeatability and reliability [6]. For example, somatic mutations in the tyrosine kinase domain of the epidermal growth factor receptor was shown to be a predictive marker for a greater efficacy of gefitinib in patients with non small cell lung cancer [7, 8]. However, only a few have been found to be useful clinically, although numbers of discovered and identified biomarkers are generated from preclinical research.
The development and progression of the disease may be caused from the interplay of a group of correlated molecules or a network, rather than from the malfunction of the individual gene, protein, or cell [9]. It is believed that dynamic alternations of complex interaction networks and molecular sub-networks can represent and influence responses of cells or organs to real-time changed microenvironment [10–12]. Thus, identification and validation of interaction networks and network biomarkers, especially at the protein level, become critical to develop disease-specific biomarkers for monitoring disease occurrence, progression or treatment efficacy [13–15]. The present review headlights network biomarkers, interaction networks, dynamical network biomarkers, with special focus on respiratory diseases, with an emphasis to integrate bioinformatics-based screening of biomarkers, network biomarker, dynamic network biomarkers with clinical informatics and phenotypes and establish a systems biomedicine-evidenced disease-specific dynamic network biomarkers
Network biomarkers
Gene or protein expression data and other high-dimensional profile data with over thousands of measurements in each sample can be generated from transcriptome, proteome and metabolome studies. The mining and analysis of such high-throughput data have led the current omics-based research from studying individual components to understanding functional modules or networks for biomolecular systems [16, 17]. It requires multi-dimensional views and the integration of molecular interaction networks in the analysis of high-throughput data to define the variation of disease severity and progression, drug sensitivity and resistance, cell growth and differentiation, and pathogenesis elaboration. A new concept of Systems Clinical Medicine was introduced to integrate systems biology, clinical science, omics-based technology, bioinformatics and computational science to improve diagnosis, therapies and prognosis of diseases [18]. Proteomics-based bioinformatics is a critical part of the systems clinical medicine and the core approach to carry out the investigation for pathogenesis, to explore the potential of clinical applications and to improve the outcomes of patients with certain diseases.
The concept of network biomarkers was proposed as a new type of biomarkers, including a set of biomarkers and their interactions [19]. A set of high-confident biomarkers from cardiovascular-related network were identified as candidate network biomarkers and used to classify two groups of patients more accurately than current single ones without consideration of biological molecular interaction [19]. Some molecular interactions in such network could be activated under specific conditions, indicating the dysfunctional process underlying the corresponding disease phenotypes and making the detection of network biomarkers possible. Microarray analysis and human protein-protein interaction (PPI) network were combined to identify more reproducible sub-network markers than individual markers in breast cancer and achieved higher accuracy in interpretation of molecular mechanisms and classification of metastatic versus non-metastatic tumors [20].
Network biomarkers have been widely studied for early diagnosis, prognosis prediction and efficacy prediction for cancer. For example, gene expression profiling was combined with functional genomic and proteomic data from various species to generate a network containing 118 genes linked by 866 potential functional associations for breast cancer [21]. One component within the network, HMMR gene, which encodes a centrosome subunit, was discovered to be associated with the breast cancer-associated gene BRCA1 and with a higher risk of breast cancer. This network-based strategy may be used to discover additional network biomarkers for early diagnosis.
Interaction networks
Interaction networks include gene regulatory network (GRN), PPI network, RNA network, signaling pathway network, and metabolic network. Interaction networks can provide models of cellular networks based on the integration of a large and heterogeneous dataset, e.g., from proteomics and high-throughput functional genomics studies [22]. For example, GRNs could be drawn from microarray data consisting of 62 primary tumors and 41 normal prostate tissues to explore the significant GRNs correlated with disease, severity and stage in the prostate cancer [23]. Notch1 signaling pathway was found to directly activate a feed-forward-loop transcriptional network and induce c-MYC gene expression to promot the growth of human T cell lymphoblastic leukemia cells, using an integrative systems biology approach which integrated gene expression array and ChIP-on-chip data [24]. A strategy of metabonomics based on rapid resolution liquid chromatography/tandem mass spectrometry, multivariate statistics and metabolic correlation networks was implemented to find biologically significant metabolite biomarkers in breast cancer [25]. A total of 12 metabolites were identified as potential biomarkers including amino acids, organic acids, and nucleosides. Statistical epistasis networks were inferred to characterize the global space of pairwise interactions among approximately 1500 Single Nucleotide Polymorphisms (SNPs) spanning nearly 500 cancer susceptibility genes in a large population-based study of bladder cancer [26]. The network was found to have a largest connected component of 39 SNPs that was absent in any other permuted-data networks. The observations suggested that the particular statistical epistasis network captured important features of the genetic architecture of bladder cancer that have not been described previously.
Dynamic network biomarkers
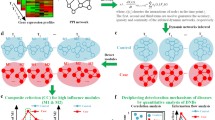
Dynamical network biomarkers (DNB) show time-dependent alterations of network biomarkers monitored and evaluated at different stages and time points during the development of diseases [12]. It is more than new nomenclature, although every molecular biomarker is embedded in a molecular network and this network will show dynamic properties over time. The common objective of developing biomarkers, network biomarkers and DNB is to discover disease-specific biomarker or a panel of biomarkers for monitoring disease occurrence, progression or treatment. The study on network biomarkers emphasizes the interaction among molecules, while DNB stresses dynamical alterations of biomarkers to provide more precise and intact view for biomarkers mining. The expression levels of biomarkers can provide one dimensional information, network biomarkers provide two dimensional information by adding interactions of biomarkers, and DNB provide a three dimensional image of biomarker-biomarker interactions by demonstrating the location and time of altered biomarkers, and time-dependent stronger or weaker interactions among biomarkers in the network [12]. The integration of bioinformatics-based dynamic network biomarkers with clinical informatics and phenotypes is expected to provide a four-dimensional image of systems biomedicine-evidenced disease-specific dynamic network biomarkers. The differences among molecular biomarkers, network biomarker and DNB are shown in Figure 1.
Biomarkers, network biomarkers and dynamical network biomarkers. Biomarkers provide one dimensional information, while network biomarkers provide two dimensional information by adding interactions. DNBs provide a three dimensional image of biomarker-biomarker interactions by showing time-dependent stronger or weaker interactions among biomarkers in the network. DNB: Dynamical network biomarkers.
We have emphasized to detect early-warning signals of the “pre-disease state” to prevent the critical transition from normal to disease state and achieve the early diagnosis and intervention for a complex disease. Biomarkers, network biomarkers, and DNB have respective functions in distinguishing normal, pre-disease, and disease stages during disease progression, as illustrated in Figure 2. Biomarkers or network biomarkers are mainly applied to distinguish the normal and disease situation, while DNB were proposed to be able to identify a pre-disease state even with small amounts of samples, provided that high throughput data were available for each sample [27, 28]. Tissue-specific early warning signals were identified using DNB theory during type 2 diabetes mellitus (T2DM) development and progression [29]. Other than two different critical states characterized as responses to insulin resistance and serious inflammation, a new T2DM-associated function, i.e. steroid hormone biosynthesis, was also discovered. DNB could signal the emergence of the critical transitions for early diagnosis of diseases, and provide the causal network of the transitions for revealing molecular mechanisms of disease initiation and progression at a network level.
Disease states and biomarkers. There are three stages during disease progression, i.e., a normal state, a pre-disease state and a disease state. A normal state is a relatively healthy stage including the chronic inflammation period or the stable period during which the disease is under control, whereas a pre-disease state is the limit of the normal state just before the critical transition into the disease state. And there are three types of biomarkers, i.e., traditional molecular biomarkers, network biomarkers, and newly developed dynamical network biomarkers (DNB). Both molecular and network biomarkers are static measurements on the disease and indicators on the disease state, whereas DNB are dynamical measurements on the pre-disease, thus providing the early-warning signals for the pre-disease state.
DNB have advantages over network biomarkers [16], e.g. make early diagnosis possible, develop biomarkers from a small number of samples based on a model-free method, or be relatively easy for clinical application. In addition, the concept of DNB will be generally applicable for detecting early-warning signals of critical transitions in any other biological process, e.g. cell differentiation processes, aging processes, and phase changes of cell cycle. Dynamic networks controlling the differentiation processes were monitored with a high throughput reverse phase protein microarray at 33 time points for 14 days to characterize adult adipose-derived stem cell differentiation [30]. Dynamic networks demonstrated key phosphoregulatory events in signal transduction pathways correlated with adipogenic differentiation. This is a novel way to understand the signaling architecture of mesenchymal stem cell differentiation and might be useful in developing network-targeted therapies for clinical applications. The dynamic protein phosphorylation networks, e.g. the insulin signaling network in adipocytes, were investigated by high-resolution mass spectrometry-based proteomics [31]. The integration of large-scale phosphoproteomics data predicted physiological substrates of several diverse insulin-regulated kinases, within which an Akt substrate, SIN1, a core component of the mTORC2 complex, was identified to regulate mTORC2 activity in response to growth factors, revealing topological insights into the Akt/mTOR signaling network. This particular study provides an excellent example to understand complex and dynamic signaling networks in tissues which contain numerous phosphorylation sites on proteins involved in diverse molecular functions.
Focus on respiratory diseases
Network biomarkers and interaction networks were most studied in lung cancer among all kinds of respiratory diseases. Transcription factor profiling of lung adenocarcinomas in c-myc-transgenic mice suggested a model of a transcriptional regulatory network where different transcription factors acted in concert upon c-Myc overexpression [32]. Molecular networks for transcriptional regulation could explain partly the carcinogenic effect seen in mice with over-expression of the c-Myc oncogene. Similarly, the GATA2 transcriptional network was discovered to be requisite for RAS oncogene-driven non-small cell lung cancer [33]. Novel functional view of the crocidolite asbestos-treated A549 human lung epithelial transcriptome revealed an intricate network of pathways with opposing functions including cell death, cancer, cell cycle, cellular growth, proliferation, and gene expression [34]. Network Component Analysis and Pathway Crosstalk Analysis was performed to construct a regulatory network in human lung cancer (A549) cells which were treated with motexafin gadolinium (MGd), a metal cation-containing chemotherapeutic drug for 4, 12, and 24 hours [35]. This dynamic network of transcription and pathway crosstalk clearly demonstrated molecular mechanism of MGd-treated human lung cancer cells. After downloading the preprocessed microarray expression dataset from Gene Expression Omnibus database, our group [36] applied a new computational strategy for the identification and biological interpretation of new candidate genes in lung cancer and smoking by coupling a network-based approach with gene set enrichment analysis. Panels of top ranked gene candidates, major gene hubs and commonly involved pathways in both the smoking and cancer related network were identified. This new approach of bioinformatics for biomarker identification can probe into deep genetic relationships between cigarette smoking and lung cancer, although further validation in clinical settings is needed.
For diagnosis of lung cancer, a systems biology approach integrating microarray gene expression profiles and protein-protein interaction information was proposed to develop a network-based biomarker [37]. In addition, the network-based biomarker, acting as the screening test. About 40 significant proteins in lung carcinogenesis were identified on the basis of the network-based biomarker principle. In addition, the network-based biomarker acting as a screening test was shown to be effective to diagnose cancer in smokers with signs of lung cancer. Artificial neural network model built with the six serum tumor markers was shown to be able to distinguish lung cancer, from lung benign disease and normal people as well as from three common gastrointestinal cancers [38]. A transcriptome network analysis method was used to construct gene regulation networks on published microarray data and select candidate genes for squamous lung cancer [39]. The genes of SPI1, FLI1, FOS, ETS2, EGR1 and PPARG were defined as candidate biomarkers, although further validation is needed for clinical screening or diagnosis. A transcriptional network classifier containing 25-gene network signature for distinguishing adenocarcinoma from squamous cell carcinoma was inferred from the molecular profiles of 111 human lung carcinomas to characterize different subtypes of lung cancer [40]. Network-based approach was also used to predict the prognosis for lung cancer. The combination of physical and biological factors with a graphical Bayesian network framework was found to improve the overall prediction for local failure following radiotherapy in lung cancer [41]. A systems biology-based network approach using lung tissues for analysis was used to identify a cell cycle gene module and three hub genes as predictor of overall survival in lung adenocarcinoma patients [42].
A number of studies on the role of interaction networks and network biomarkers have been performed to explore molecular mechanisms and identify potential biomarkers associated with other respiratory diseases, though the specificity and repeatability remain unclear and further validation is needed. Network inference algorithms elucidated nuclear factor erythroid 2-related factor regulation of mouse lung oxidative stress and showed the promise for operating on high-throughput gene expression data to identify transcriptional regulatory and other signaling relationships [43]. Response network analysis of differential gene expression in human epithelial lung cells was used to compare the response to H5N1 infection with a more benign infection with Respiratory Syncytial Virus [44]. Characteristics of H5N1 infection compared to respiratory syncytial virus infection showed several immune response factors specific for each of these infections. Metabolic network in Pseudomonas aeruginosa-infected chronic cystic fibrosis lung demonstrated how the bacterial metabolism adapted over time and how the tradeoffs between growth and other important cellular processes shifted during disease progression [45]. A successful application of DNB on respiratory diseases was on a murine acute lung injury model driven by carbonyl chloride inhalation [46]. By applying DNB theory on the time-course microarray data from lung tissue RNA, a group of observable molecules were screened out at 8 hour, which formed a strong correlated subnetwork just before the occurrences of lung injury and thus provided a reliable early-warning signal [27]. Although this was an animal study with limited samples, it validated the effectiveness of DNB for the identification of toxicity mechanisms and the early diagnosis of carbonyl chloride induced acute lung injury.
DNB with clinical bioinformatics
Clinical bioinformatics is an emerging science combining clinical informatics, bioinformatics, medical informatics, information technology, mathematics and omics science together [47]. There is an increasing need to emphasize clinical phenotypes and medical informatics in developing disease-specific DNB. Clinical bioinformatics has been suggested as a new way to integrate clinical symptoms, signs and measurements with human sample-generated bioinformatics for the development of disease-specific DNB [47]. It is challenging to adjust DNB to clinical symptoms and signs, disease development and progression, and therapeutic strategy. Clinical bioinformatics should be emphasized for DNB identification and validation to handle data preprocessing and consolidation, the data-driven search, verification, prioritization and biological interpretation of putative metabolic candidate biomarkers, as suggested by Baumgartner et al. [48]. It is also challenging to select proper data mining tools for analyzing clinical proteomic data, to design clinical studies like case–control or prospective cohort studies, or to translate new findings of disease-specific DNB to clinical application.
The importance of the integration of proteomic profiles and data with clinical bioinformatics was emphasized to bridge the gap between proteomics and diseases, e.g. acute and chronic organ injury, inflammation, and multiple organ dysfunction [49]. It is easy to understand that proteomics as a powerful tool help investigate the relationship between biological molecules and disease mechanisms, while difficult to integrate proteomics-based DNB with physiological and pathophysiological situations, with organ-, tissue-, type-, function-, disease-specific patterns, or with disease diagnosis, therapies and prognosis. A new protocol of disease-specific DNB identification and evaluation was developed by integrating proteomic profiles of inflammatory mediators with clinical informatics in patients with acute exacerbation of chronic obstructive pulmonary disease (AECOPD) [50, 51]. Serum protein profiles from patients with AECOPD were assessed by different strategies of proteomics on days 1 and 3 of the admission day and the discharging day and correlated with clinical informatics by a Digital Evaluation Score System for assessing severity of patients. A panel of inflammatory mediators with dynamical changes during disease progression was demonstrated to be COPD specific or AECOPD specific biomarkers and correlated well with clinical bioinformatics. Such protocol can integrate proteomics with clinical informatics to explore a new way to validate and optimize disease-special DNB, which might be used clinically after further validation with a larger sample size.
Conclusions
Disease-specific interaction networks, network biomarkers or DNB have the great impact in the understanding of mechanism interpretation, risk assessment, early diagnosis, illness monitoring, disease classification, stage and grade, effectiveness and toxicities prediction, as well as combinational therapies optimization. Protein-based DNB will provide more information to define the differences between the normal and pre-disease stages and point to early diagnosis. Network biomarkers, interaction networks and DNB have been applied in respiratory diseases and will face a number of opportunities and challenges. Clinical bioinformatics should be considered as a key approach to the identification and validation of disease-specific biomarkers.
Review
The last few years have seen a fast development of network medicine, especially in cancer research. A number of interaction networks or network biomarkers have been proposed to be related to disease development, progression and therapeutic effects. Thus, they are of great potential for early diagnosis, prognosis prediction and efficacy prediction. However, there are also several challenges to translate the research of network based approach into clinical application. Firstly, since many findings were from in vitro cell line studies or current database, further validation studies in clinical settings will be of great importance. Secondly, network biomarkers, interaction networks or DNB are more complex than single biomarker and more difficult to develop, optional computational methods or algorithms are needed. Thirdly, the generation of network biomarkers or DNB usually need a three dimensional image of biomarker-biomarker interactions showing time-dependent stronger or weaker interactions among biomarkers in the network, which needs rich and integral data collection. All these make sense in respiratory diseases, in which area network based approach will also progress and have a bright future.
References
Rabe KF, Hurd S, Anzueto A, Barnes PJ, Buist SA, Calverley P, Fukuchi Y, Jenkins C, Rodriguez-Roisin R, van Weel C, Zielinski J: Global strategy for the diagnosis, management, and prevention of chronic obstructive pulmonary disease: GOLD executive summary. Am J Respir Crit Care Med 2007, 176: 532–555.
Buist AS, McBurnie MA, Vollmer WM, Gillespie S, Burney P, Mannino DM, Menezes AM, Sullivan SD, Lee TA, Weiss KB, Jensen RL, Marks GB, Gulsvik A, Nizankowska-Mogilnicka E, BOLD Collaborative Research Group: International variation in the prevalence of COPD (the BOLD Study): a population-based prevalence study. Lancet 2007, 370: 741–750.
Fang X, Wang X, Bai C: COPD in China: the burden and importance of proper management. Chest 2011, 139: 920–929.
Simon R: Development and validation of therapeutically relevant multi-gene biomarker classifiers. J Natl Cancer Inst 2005, 97: 866–867.
Ludwig JA, Weinstein JN: Biomarkers in cancer staging, prognosis and treatment selection. Nat Rev Cancer 2005, 5: 845–856.
Wang X, Ward PA: Opportunities and challenges of disease biomarkers: a new section in the Journal of Translational Medicine. J Transl Med 2012, 10: 240.
Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG, Louis DN, Christiani DC, Settleman J, Haber DA: Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 2004, 350: 2129–2139.
Paez JG, Janne PA, Lee JC, Tracy S, Greulich H, Gabriel S, Herman P, Kaye FJ, Lindeman N, Boggon TJ, Naoki K, Sasaki H, Fujii Y, Eck MJ, Sellers WR, Johnson BE, Meyerson M: EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 2004, 304: 1497–1500.
Schadt EE: Molecular networks as sensors and drivers of common human diseases. Nature 2009, 461: 218–223.
Garraway LA, Lander ES: Lessons from the cancer genome. Cell 2013, 153: 17–37.
Barabasi AL, Gulbahce N, Loscalzo J: Network medicine: a network-based approach to human disease. Nat Rev Genet 2011, 12: 56–68.
Wang X: Role of clinical bioinformatics in the development of network-based Biomarkers. J Clin Bioinforma 2011, 1: 28.
Nibbe RK, Koyuturk M, Chance MR: An integrative-omics approach to identify functional sub-networks in human colorectal cancer. PLoS Comput Biol 2010, 6: e1000639.
Wang HQ, Wong HS, Zhu H, Yip TT: A neural network-based biomarker association information extraction approach for cancer classification. J Biomed Inform 2009, 42: 654–666.
Vilar S, Gonzalez-Diaz H, Santana L, Uriarte E: A network-QSAR model for prediction of genetic-component biomarkers in human colorectal cancer. J Theor Biol 2009, 261: 449–458.
Liu R, Wang X, Aihara K, Chen L: Early Diagnosis of Complex Diseases by Molecular Biomarkers, Network Biomarkers, and Dynamical Network Biomarkers. Med Res Rev 2013. doi:10.1002/med.21293. [Epub ahead of print] doi:10.1002/med.21293. [Epub ahead of print]
Liu ZP, Wang Y, Zhang XS, Chen L: Network-based analysis of complex diseases. IET Syst Biol 2012, 6: 22–33.
Wu D, Rice CM, Wang X: Cancer bioinformatics: a new approach to systems clinical medicine. BMC Bioinformatics 2012, 13: 71.
Jin G, Zhou X, Wang H, Zhao H, Cui K, Zhang XS, Chen L, Hazen SL, Li K, Wong ST: The knowledge-integrated network biomarkers discovery for major adverse cardiac events. J Proteome Res 2008, 7: 4013–4021.
Chuang HY, Lee E, Liu YT, Lee D, Ideker T: Network-based classification of breast cancer metastasis. Mol Syst Biol 2007, 3: 140.
Pujana MA, Han JD, Starita LM, Stevens KN, Tewari M, Ahn JS, Rennert G, Moreno V, Kirchhoff T, Gold B, Assmann V, Elshamy WM, Rual JF, Levine D, Rozek LS, Gelman RS, Gunsalus KC, Greenberg RA, Sobhian B, Bertin N, Venkatesan K, Ayivi-Guedehoussou N, Solé X, Hernández P, Lázaro C, Nathanson KL, Weber BL, Cusick ME, Hill DE, Offit K, et al.: Network modeling links breast cancer susceptibility and centrosome dysfunction. Nat Genet 2007, 39: 1338–1349.
Erler JT, Linding R: Network-based drugs and biomarkers. J Pathol 2010, 220: 290–296.
Yeh HY, Cheng SW, Lin YC, Yeh CY, Lin SF, Soo VW: Identifying significant genetic regulatory networks in the prostate cancer from microarray data based on transcription factor analysis and conditional independency. BMC Med Genomics 2009, 2: 70.
Palomero T, Lim WK, Odom DT, Sulis ML, Real PJ, Margolin A, Barnes KC, O'Neil J, Neuberg D, Weng AP, Aster JC, Sigaux F, Soulier J, Look AT, Young RA, Califano A, Ferrando AA: NOTCH1 directly regulates c-MYC and activates a feed-forward-loop transcriptional network promoting leukemic cell growth. Proc Natl Acad Sci U S A 2006, 103: 18261–18266.
Chen Y, Zhang R, Song Y, He J, Sun J, Bai J, An Z, Dong L, Zhan Q, Abliz Z: RRLC-MS/MS-based metabonomics combined with in-depth analysis of metabolic correlation network: finding potential biomarkers for breast cancer. Analyst 2009, 134: 2003–2011.
Hu T, Sinnott-Armstrong NA, Kiralis JW, Andrew AS, Karagas MR, Moore JH: Characterizing genetic interactions in human disease association studies using statistical epistasis networks. BMC Bioinformatics 2011, 12: 364.
Chen L, Liu R, Liu ZP, Li M, Aihara K: Detecting early-warning signals for sudden deterioration of complex diseases by dynamical network biomarkers. Sci Rep 2012, 2: 342.
Liu R, Li M, Liu ZP, Wu J, Chen L, Aihara K: Identifying critical transitions and their leading biomolecular networks in complex diseases. Sci Rep 2012, 2: 813.
Li M, Zeng T, Liu R, Chen L: Detecting tissue-specific early warning signals for complex diseases based on dynamical network biomarkers: study of type 2 diabetes by cross-tissue analysis. Brief Bioinform 2013, 15: 229–243.
Wilson B, Liotta LA, Petricoiniii E: Dynamic protein pathway activation mapping of adipose-derived stem cell differentiation implicates novel regulators of adipocyte differentiation. Mol Cell Proteomics 2013, 12: 2522–2535.
Humphrey SJ, Yang G, Yang P, Fazakerley DJ, Stockli J, Yang JY, James DE: Dynamic adipocyte phosphoproteome reveals that Akt directly regulates mTORC2. Cell Metab 2013, 17: 1009–1020.
Reymann S, Borlak J: Transcription profiling of lung adenocarcinomas of c-myc-transgenic mice: identification of the c-myc regulatory gene network. BMC Syst Biol 2008, 2: 46.
Kumar MS, Hancock DC, Molina-Arcas M, Steckel M, East P, Diefenbacher M, Armenteros-Monterroso E, Lassailly F, Matthews N, Nye E, Stamp G, Behrens A, Downward J: The GATA2 transcriptional network is requisite for RAS oncogene-driven non-small cell lung cancer. Cell 2012, 149: 642–655.
Hevel JM, Olson-Buelow LC, Ganesan B, Stevens JR, Hardman JP, Aust AE: Novel functional view of the crocidolite asbestos-treated A549 human lung epithelial transcriptome reveals an intricate network of pathways with opposing functions. BMC Genomics 2008, 9: 376.
Shao L, Wang L, Wei Z, Xiong Y, Wang Y, Tang K, Li Y, Feng G, Xing Q, He L: Dynamic network of transcription and pathway crosstalk to reveal molecular mechanism of MGd-treated human lung cancer cells. PLoS One 2012, 7: e31984.
Fang X, Netzer M, Baumgartner C, Bai C, Wang X: Genetic network and gene set enrichment analysis to identify biomarkers related to cigarette smoking and lung cancer. Cancer Treat Rev 2013, 39: 77–88.
Wang YC, Chen BS: A network-based biomarker approach for molecular investigation and diagnosis of lung cancer. BMC Med Genomics 2011, 4: 2.
Feng F, Wu Y, Wu Y, Nie G, Ni R: The effect of artificial neural network model combined with six tumor markers in auxiliary diagnosis of lung cancer. J Med Syst 2012, 36: 2973–2980.
Bai J, Hu S: Transcriptome network analysis reveals potential candidate genes for squamous lung cancer. Int J Mol Med 2012, 29: 95–101.
Chang HH, Dreyfuss JM, Ramoni MF: A transcriptional network signature characterizes lung cancer subtypes. Cancer 2011, 117: 353–360.
Oh JH, Craft J, Al LR, Vaidya M, Meng Y, Deasy JO, Bradley JD, El NI: A Bayesian network approach for modeling local failure in lung cancer. Phys Med Biol 2011, 56: 1635–1651.
Li Y, Tang H, Sun Z, Bungum AO, Edell ES, Lingle WL, Stoddard SM, Zhang M, Jen J, Yang P, Wang L: Network-based approach identified cell cycle genes as predictor of overall survival in lung adenocarcinoma patients. Lung Cancer 2013, 80: 91–98.
Taylor RC, Acquaah-Mensah G, Singhal M, Malhotra D, Biswal S: Network inference algorithms elucidate Nrf2 regulation of mouse lung oxidative stress. PLoS Comput Biol 2008, 4: e1000166.
Tatebe K, Zeytun A, Ribeiro RM, Hoffmann R, Harrod KS, Forst CV: Response network analysis of differential gene expression in human epithelial lung cells during avian influenza infections. BMC Bioinformatics 2010, 11: 170.
Oberhardt MA, Goldberg JB, Hogardt M, Papin JA: Metabolic network analysis of Pseudomonas aeruginosa during chronic cystic fibrosis lung infection. J Bacteriol 2010, 192: 5534–5548.
Sciuto AM, Phillips CS, Orzolek LD, Hege AI, Moran TS, Dillman JR: Genomic analysis of murine pulmonary tissue following carbonyl chloride inhalation. Chem Res Toxicol 2005, 18: 1654–1660.
Wang X, Liotta L: Clinical bioinformatics: a new emerging science. J Clin Bioinforma 2011, 1: 1.
Baumgartner C, Osl M, Netzer M, Baumgartner D: Bioinformatic-driven search for metabolic biomarkers in disease. J Clin Bioinforma 2011, 1: 2.
Zhao H, Adler KB, Bai C, Tang F, Wang X: Epithelial proteomics in multiple organs and tissues: similarities and variations between cells, organs, and diseases. J Proteome Res 2006, 5: 743–755.
Chen H, Song Z, Qian M, Bai C, Wang X: Selection of disease-specific biomarkers by integrating inflammatory mediators with clinical informatics in AECOPD patients: a preliminary study. J Cell Mol Med 2012, 16: 1286–1297.
Chen H, Wang Y, Bai C, Wang X: Alterations of plasma inflammatory biomarkers in the healthy and chronic obstructive pulmonary disease patients with or without acute exacerbation. J Proteomics 2012, 75: 2835–2843.
Acknowledgements
The work was supported by Shanghai Leading Academic Discipline Project (Project Number: B115), Zhongshan Distinguished Professor Grant (XDW), and The National Nature Science Foundation of China (91230204, 81270099), The Shanghai Committee of Science and Technology (12JC1402200, 12431900207, 11410708600).
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
XD Wu contributed to collection of information and writing of the manuscript. LNC contributed to revision of the manuscript. XD Wang contributed to the critical revision of the manuscript. All authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, X., Chen, L. & Wang, X. Network biomarkers, interaction networks and dynamical network biomarkers in respiratory diseases. Clin Trans Med 3, 16 (2014). https://doi.org/10.1186/2001-1326-3-16
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/2001-1326-3-16