Abstract
Background
Gene expression studies require appropriate normalization methods. One such method uses stably expressed reference genes. Since suitable reference genes appear to be unique for each tissue, we have identified an optimal set of the most stably expressed genes in human blood that can be used for normalization.
Methods
Whole-genome Affymetrix Human 2.0 Plus arrays were examined from 526 samples of males and females ages 2 to 78, including control subjects and patients with Tourette syndrome, stroke, migraine, muscular dystrophy, and autism. The top 100 most stably expressed genes with a broad range of expression levels were identified. To validate the best candidate genes, we performed quantitative RT-PCR on a subset of 10 genes (TRAP1, DECR1, FPGS, FARP1, MAPRE2, PEX16, GINS2, CRY2, CSNK1G2 and A4GALT), 4 commonly employed reference genes (GAPDH, ACTB, B2M and HMBS) and PPIB, previously reported to be stably expressed in blood. Expression stability and ranking analysis were performed using GeNorm and NormFinder algorithms.
Results
Reference genes were ranked based on their expression stability and the minimum number of genes needed for nomalization as calculated using GeNorm showed that the fewest, most stably expressed genes needed for acurate normalization in RNA expression studies of human whole blood is a combination of TRAP1, FPGS, DECR1 and PPIB. We confirmed the ranking of the best candidate control genes by using an alternative algorithm (NormFinder).
Conclusion
The reference genes identified in this study are stably expressed in whole blood of humans of both genders with multiple disease conditions and ages 2 to 78. Importantly, they also have different functions within cells and thus should be expressed independently of each other. These genes should be useful as normalization genes for microarray and RT-PCR whole blood studies of human physiology, metabolism and disease.
Similar content being viewed by others
Background
Gene expression analysis is widely used to study various biological processes. Different transcript quantification methodologies exist, all of which rely on utilization of proper normalization techniques. Such analysis requires several variables need to be accounted for, including amount and quality of RNA, enzymatic efficiencies, and differences between tissues or cells in overall transcriptional activity. The current, most universally utilized normalization method for PCR and Northern blotting relies on the use of constitutively expressed endogenous reference genes, often termed ''house-keeping genes.'' It has been shown, however, that the expression of such reference genes can vary significantly between different tissues or conditions, thus making it impossible to use the same reference genes for various tissues [1–5]. Moreover, the identification of appropriate control genes presents a circular problem, because in order to find suitable genes for normalization, prior knowledge of their stable gene expression is needed, which also relies on properly normalized data.
The conventional use of a single gene for normalization can introduce a significant error in the quantification of transcript levels [6]. It is recommended that an optimal set of reference genes be identified for each individual experimental setting or tissue. In this study, we used whole-genome human microarrays and surveyed the expression of over 39,500 genes in 526 whole-blood samples from control subjects and patients with Tourette syndrome, stroke, migraine, muscular dystrophy, and autism to identify the best candidate control genes. The 10 best candidate control genes, as well as commonly used house-keeping controls and PPIB, were validated with independent samples of whole-blood RNA from both patients with multiple sclerosis (MS) and age- and gender-matched controls by qRT-PCR. GeNorm was used to identify an optimal set of the most stably expressed genes for normalization in whole-blood expression studies [6].
Methods
Patients and samples
We used a large cohort of available human Affymetrx array data that we have generated over the last several years from previous and on-going studies (Table 1). Many, but not all, of the subjects included in our analyses have previously been reported in the following studies: stroke [7–10], Tourette syndrome [11–13], controls [14–18], migraine [19], muscular dystrophy [20, 21] and autism [22, 23].
RNA Isolation and Quality Control
Whole blood was collected and total RNA was isolated using PAXgene tubes and kits (Qiagen, Valencia, CA, USA). RNA samples were examined for concentration and purity using a Nanodrop ND-1000 spectrophotometer, and integrity was checked using the Agilent 2100 Bioanalyzer. High quality RNA with a RIN number above 8.0 was used for the qRT-PCR experiment. The RIN number takes into consideration not only the conventional ratio of 28 S to 18 S ribosomal RNA, but also additional critical regions of the entire RNA electrophoregram.
Affymetrix experiments
Affymetrix Human 2.0 Plus arrays (Affymetrix, Santa Clara, CA, USA), surveying over 54,000 probe-sets, were used in this study. The 54,000 probe-sets represent over 39,500 potential human genes (Affymetrix Manual). The standard Affymetrix protocol was followed for the sample labeling, hybridization, and image scanning [23].
qRT-PCR cDNA synthesis
The cDNA synthesis and real-time PCR was performed at the Lucy Whittier Molecular and Diagnostic Core Facility (University of California, Davis, U.S.A.). The following cDNA synthesis protocol was validated through the Lucy Whittier Molecular and Diagnostic Core Facility using a variety of sample types and species. 10 μl (750 ng) of template RNA per sample, 1 μl gDNA WipeOut Buffer, 1 μl RNase-free water was incubated at 42°C for two minutes and briefly centrifuged. A 1 μL aliquot was TaqMan® analyzed with human GAPDH to confirm all gDNA had been digested. Then 8 μl from the reverse transcription reaction mix (0.5 μl Quantitect Reverse Transcriptase, 2 μl 5× Quantitect RT buffer, 0.5 μl RT Primer Mix, 0.5 μl 20 pmol Random Primers, 4.5 μl RNase free water) was added to each well, incubated at 42°C for 40 minutes, inactivated at 95°C for 3 minutes and briefly centrifuged. Finally, 100 μl of water was added to each well and mixed thoroughly.
Selection of TaqMan Assays
TaqMan assays were selected based on several criteria: 1) only pre-developed and pre-validated gene expression assays were selected from Applied Biosystems (ABI, Foster City, CA, U.S.A.), for which the company guarantees amplification efficiency over 96%. 2) Only _m1 assays (spanning exon -exon boundaries) were selected to ensure no genomic DNA will be amplified. 3) If available, the assays were preferentially selected to amplify the same target sequence where the Affymetrix probe-set was located. 4) If a gene selected for qRT-PCR validation was represented by multiple probe-sets on the Affymetrix array, the standard deviation (SD) of the expression of all probe-sets was inspected and if some were found to have a high standard deviation, their position in the reference sequence was avoided.
Real-Time TaqMan®PCR
Pre-validated gene expression assays were purchased from Applied Biosystems. See Table 1 for the Assay and Gene IDs. An aliquot of the cDNA from a subset of samples was TaqMan® analyzed with human GAPDH prior to profiling. TaqMan® analysis was done in duplicate reactions using the low density array format as described by Osman et al [24]. This was validated and processed through the Lucy Whittier Molecular and Diagnostic Core Facility (1). The samples were placed in a 384 well plate and amplified in an automated fluorometer (ABI PRISM 7900 HTA FAST, ABI). ABI's standard amplification conditions were used: 2 min at 50°C, 10 minutes at 95°C, 40 cycles of 15 seconds at 95°C and 60 seconds at 60°C. Fluorescent signals were obtained during the annealing temperature and Ct values exported with a threshold of 0.1 and a baseline of 3–10.
Data Analysis
Affymetrix Chips
All 526 arrays were summarized using the GC-RMA algorithm [25]. They were normalized within GeneSpring 7.3 software (Agilent Technologies, Palo Alto, CA) using a three-step normalization procedure, including data transformation, per chip and per gene normalization (GeneSpring Manual).
qRT-PCR GeNorm Analysis
All samples for which the standard deviation (SD) of the technical duplicates was ≥ 1.41 (or higher than 2-fold change) were considered missing data points and were not included in the analysis. The GeNorm tool was used to calculate the internal control genes stability measures (M) [6]. In short, input data was generated using the comparative Ct-method [26], where relative expression levels were calculated by setting the lowest Ct value of a particular gene to 1, and the relative expression levels for the same gene in the rest of the samples were calculated. Then for every control gene, the pair-wise variation with all other control genes is estimated in GeNorm as the SD of the log-transformed expression ratios. The internal control gene stability measure M is defined as the average pair-wise variation of a particular control gene with all other control genes. Stepwise exclusion of the gene with the highest M value is performed. Because a pair of control genes is needed to calculate M, the two most stably expressed genes cannot be ranked any further. The optimal number of control genes for normalization is determined by calculating the pair-wise variation Vn/Vn+1 between each set of two sequential normalization factors [6]. Consistent with the Vandesompele recommendations, the minimum number of genes for which the V was smaller than 0.15, was used to define the optimal set of control genes for normalization. A normalization factor is calculated based on the geometric mean of the optimal set of control genes.
qRT-PCR NormFinder Analysis
The NormFinder Excel plug-in [27] was used as an alternative algorithm to the GeNorm algorithm for ranking the expression stability of the control genes. It ranks the candidate control genes by estimating their expression stability while also taking into account the experimental design (control and disease groups). The input qRT-PCR results were the linearized quantities (2-ΔCt). The main differences between the GeNorm algorithm [6] and the model-based approach in NormFinder [27] is that the latter takes into account the inter- and intra-group variation, as well as the estimation of variances.
Analysis of Variance (ANOVA)
Partek Genomics Suite (St. Louis, MI, USA) was used to perform an ANOVA using the Diagnosis as fixed and the Plate as a random factor (μ + Diagnosis + Plate + (Diagnosis × Plate) + ε). The input data were the normalized data to the geometric average of the optimal set of control genes (TRAP1, DECR1, PPIB, and HMBS) using the comparative Ct method.
ΔΔCt Method
The ΔΔCt method [28] was used to calculate the normalized calibrated data. TRAP1, identified by GeNorm as one of the two most stably expressed genes was used as a normalizer. The average of the healthy controls was used as the calibrator. In short, the Ct value of the normalizer was subtracted from the Ct value of each sample (ΔCt). Then the value of the calibrator was subtracted (ΔΔCt) and the linearized values were calculated (2-(ΔΔCt)). A prerequisite for using this method of analysis is that the normalizer and the gene of interest have very similar amplification efficiency. The ABI assays used in this study have been pre-validated by the company to have over 96% amplification efficiency.
Results
Patients and samples
Of the 526 samples, the mean ages of the subjects for each study varied from age 4 (autism study patients and controls) to age 68 (stroke study patients) with a range of 2 to 78 years of age (Table 1). There was a predominance of males (387) to females (139). Control subjects had no known medical, surgical, or psychiatric illnesses. Study subjects had a variety of medical illnesses with varying pathophysiologies, including neurodevelopmental and neurological disorders (autism, mental retardation, Tourette syndrome), stroke (ischemic brain injury), muscular dystrophy (genetic muscle disease), migraine headaches, as well as a presumed autoimmune disease, multiple sclerosis (Table 1).
Selection of Reference Genes from Whole-Genome Affymetrix Arrays
Our goal was to identify stably expressed genes with a broad range of expression levels from our Affymetrix data on the 526 samples (Table 1). We chose the 20 most stably expressed genes from each of five intensity intervals (n = 100 genes): log2-transformed intensity values between 5–6, 6–7, 7–8, 8–9 and 9–10 (Figure 1). For a list of the most stable genes over all of the expression levels, see Additional file 1. Based on our selection criteria, these 100 most stably expressed genes covered a wide range of raw expression values from 32 to 1024. Functional categorization using the Database for Annotation and Visualization, and Integrated Discovery (DAVID, http://david.abcc.ncifcrf.gov/) showed that 50% of the annotated genes are involved in primary metabolic processes. For the list of the 100 most stably expressed genes on the 526 microarrays see Additional file 1.
The Average log 2 - intensity (x-axis) of 39,500 genes from a total of 526 Affymetrix Human 2.0 Plus microarrays is plotted against the standard deviation of the log 2 - intensity (y-axis) of those genes. Probe-sets highlighted in red are the 100 probe-sets with the lowest standard deviation within an interval of the log2-transformed intensity values between 5 and 10.
Selection of Candidate Control Genes for Validation by qRT-PCR
The set of 100 stably expressed genes on microarrays was filtered down to probe-sets with available gene annotation, as well as to probe-sets detecting only a single known gene (Affymetrix annotation *_at). Based on these criteria, 23 probe-sets were identified (data not shown). From them, 10 candidate genes were selected for validation with, on average, two genes per intensity interval. These 10 candidate genes included TRAP1, DECR1, FPGS, FARP1, MAPRE2, PEX16, GINS2, CRY2, CSNK1G2 and A4GALT. In the validation, we also included four commonly employed reference genes, ACTB, GAPDH, B2M and HMBS. In addition, PPIB was included because it was previously reported to be stably expressed in blood [3]. Each gene and ABI assay information is presented in Table 2. The average expression values from the Affymetrix array data of the genes selected for validation by qRT-PCR are shown in Figure 2. The range of the SDs of the the log2-transformed intensity values on the microarrays was between 0.10 and 0.42 for the 10 candidate reference genes on the microarrays (solid bars, Figure 2). The SDs for the remainder genes (open bars, B2M, ACTB, GAPDH, PPIB and HMBS) ranged between 0.47 and 0.72 on the microarrays (Figure 2). Based on the Affymetrix array data alone, FARP1 was the most stably expressed. A general trend of increasing SD with increasing intensity was noted for the candidate reference genes (Figure 2).
Average expression calculated from the 526 Affymetrix Human 2.0 Plus microarrays for the candidate control genes selected for qRT-PCR validation. Average of the log2-transformed intensity values ± standard deviation (y-axis) is plotted for each gene. Gray columns represent the candidate control genes selected from Figure 1. White columns represent commonly used house-keeping genes and PPIB.
qRT-PCR Analysis and Expression of Candidate Control Genes
The analysis flowchart presented in Figure 3 describes the steps used to analyze the qRT-PCR data. The subset of 15 candidate reference genes (Table 2), validated on qRT-PCR, exhibited a broad range of expression levels from 17.93 to 36.66 cycle threshold (Ct) values (Table 3, Figure 4). Samples for which the SD on the technical replicates was higher than 1.4 (a fold-change higher than 2) were considered unreliable data and were not included in the downstream analysis, thus reducing the sample size for some assays to lower than 40 (Table 3). Genes that had SD ≤ 1.0 on the average raw Ct values of all samples were considered to be stably expressed. Based on that criteria, TRAP1, B2M, DECR1, PPIB, FPGS, HMBS and FARP1 (Table 3, Figure 4, below the dotted line) showed stable expression over all 40 samples. In contrast, MAPRE2, PEX16, GINS2, ACTB, GAPDH, CRY2, CSNK1G2 and A4GALT did not show stable expression over the samples (Table 3, Figure 4 above the dotted line). The stably expressed genes covered a wide range of expression levels, from a high level of expression of 17.9 Ct for B2M, to low levels of 35.71 Ct for FARP1 (Table 3, Figure 4).
Expression (average raw Ct, x axis) is plotted versus Standard Deviation of average raw Ct (y axis) for all of the candidate control genes. qRT-PCR was performed on RNA samples from 20 multiple sclerosis (MS) patients and 20 age and gender matched controls. The average expression and its standard deviation was calculated for each gene for all of the patients combined (n = 40). (Ct = Cycle threshold).
Expression Stability and Ranking Analysis
To identify the most suitable set of genes for normalization in whole-blood, the average expression stability (M) for those reference genes which showed SD of raw Ct-values ≤ 1.0 on all samples was calculated using the GeNorm tool (see Methods). Those genes included TRAP1, FPGS, B2M, DECR1, PPIB and HMBS (Figure 4). All of these genes also had a SD of the Ct-values for the technical duplicates of ≤ 1.41 and a SD of the raw Ct values of all samples ≤ 1.0. The FARP1 gene was omitted from the analysis of the expression stability because GeNorm does not allow missing values. However, re-analysis where FARP1 was included with the rest of the five stably expressed genes (with n = 35) did not change the final results (data not shown).
An important pre-requisite for estimating the Expression Stability Measure is that the genes are not coordinately regulated. The Gene Ontology terms of the six genes, analyzed in GeNorm, are presented in Table 4. These genes are involved in different biological process, molecular functions and/or are located in different cellular compartments.
The GeNorm algorithm calculates the M value of a gene based on the average pair-wise variation between all genes in the analysis. Figure 5A plots the average expression stability of the six reference genes. This curve was generated by a step-wise exclusion of the least stable reference gene. It shows that B2M was the least stable, while TRAP1 and FPGS were the two most stably expressed genes in whole blood of patients with MS and gender and age matched controls.
Identification of the most stably expressed reference genes using GeNorm analysis. RNA samples from whole-blood from 20 MS patients and 20 matched controls were analyzed using qRT-PCR. A. Average expression stability values (M) of the reference genes during step-wise exclusion of the least stable control gene. The x- axis shows the gene with the highest M value (the lowest stability) for this set of genes. The least stable gene was excluded after each iteration. B. Determination ofthe optimal number of reference genes for normalization. V is the pair-wise variation of two sequential normalization factors. The least number of genes for each V<0.15 was selected as the optimal set of genes for normalization [6]. In this study, the four most stably expressed genes (TRAP1, FPGS, DECR1 and PPIB) represent the optimal set of genes for normalization.
Determination of the optimal number of reference genes for normalization of the dataset uses a metric called V. It is the pair-wise variation of two sequential normalization factors. For consistency with the recommendations of Vandesompele et al [6], the least number of genes for which V<0.15 should be selected as the optimal set of genes for normalization. In the study, the four most stably expressed genes, TRAP1, FPGS, DECR1 and PPIB, represent the optimal set of genes for normalization (Figure 5B). We estimated the gene expression stability and ranking of the control genes using an alternative algorithm (NormFinder). The relative ranking of the genes was essentially the same as the one obtained using GeNorm. The most stable gene was TRAP1 (stability value 0.066), followed by FPGS (0.067), DECR1 (0.078), PPIB (0.079), HMBS (0.095) and B2M (0.108). The main differences between the GeNorm algorithm (reference) and the model-based variance estimation approach in NormFinder [27] is that the latter takes into account the inter- and intra-group variation and uses direct estimation of variances as opposed to pair-wise variation.
Analysis of Variance (ANOVA)
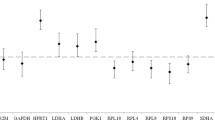
To identify genes that showed differential expression between the MS subjects and the healthy controls, an ANOVA was performed on the qRT-PCR data. Diagnosis and each PCR-plate were included as factors. CSNK1G2 showed significant differences in expression between MS and control samples (Table 5, p < 0.05). However, if multiple-comparison correction is applied to this p-value, it is not statistically significant. Expression of the remaining genes did not differ significantly between the matched pairs (Table 5). No significant p-values (p ≤ 0.05) for plate, nor for the Diagnosis-versus-plate interaction were found, suggesting no significant technical bias in the study (data not shown). The fold-change of expression between the MS and the healthy controls for the stably expressed genes (Figure 4, below the dotted line) was closest to 1.0 for FPGS (Table 5). ACTB and A4GALT fold-changes could not be estimated in Partek due to multiple missing values. The greatest deviation from the 1.0 ratio for all the genes was observed for CSNK1G2. We also used the ΔΔCt method [28], as described in the Methods section, to calculate fold-change. B2M showed the smallest deviation from the 1.0 ratio for the most stably expressed genes (Table 5). TRAP1, identified by GeNorm as one of the most stably expressed genes, was used as the Normalizer; therefore, its ratio is not relevant. The highest deviation from the 1.0 ratio for all the genes was observed for CSNK1G2, as expected.
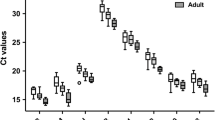
Data on the four genes, which constitute the optimal set of normalizers (TRAP1, FPGS, DECR1 and PPIB) on the MS and healthy control samples are shown separately as box-and-whisker plots (Figure 6). Data for both the ΔΔCt and Comparative Ct methods for MS versus control patients are shown in Table 5, with similar results being obtained with both methods.
RNA expression values are shown for the four most stably expressed genes (TRAP1, FPGS, DECR1 and PPIB) for multiple sclerosis (MS) and controls subjects. Circles represent individual qRT-PCR Ct average raw values for MS samples (blue, n = 20) and age and gender matched controls (red, n = 20). Boxes represent the 25th and 75th quartile and lines represent the median. Whiskers represent the range of data for the 10th and 90th quartile.
Discussion
Gene expression techniques such as qRT-PCR, microarrays and Northern blotting require accurate normalization methods in order to obtain reliable, quantitative data. A common approach is to use an endogenous reference gene. The purpose of the reference gene(s) is to remove differences not attributable to real biological variation. However, numerous reports indicate that differences in the expression levels of commonly used endogenous reference genes vary considerably between different tissues and between different experimental conditions. Thus, there are no universal reference genes for all tissues or experimental conditions [2, 3, 29–35].
To address this problem, the best reference genes must be determined for each individual tissue or experimental setting. Moreover, a combination of several endogenous control genes produces more reliable normalization than any single control gene [6]. We therefore examined the expression of over 39,000 genes in 526 samples of whole blood from men, women and children that included healthy controls and individuals with a variety of different diseases. The 100 most stably expressed probe-sets were identified based upon having the least variance across all of these samples over a broad range of expression values. These 100 genes could also be useful as endogenous internal controls for microarray studies – an approach not commonly used at present for microarray studies. Fifty percent of the annotated genes were involved in primary metabolic processes. Thus these genes appear to be good ''housekeeping genes'' for human blood because of their stable expression across various ages, genders, and diseases.
qRT-PCR validation was then performed on 10 candidate reference genes, four commonly used reference genes (ACTB, GAPDH, B2M, and HMBS) and PPIB, which is reported to be stably expressed in blood [3]. The two frequently used control reference genes in nervous tissue, ACTB and GAPDH, were not stably expressed under our experimental conditions and are therefore not suitable reference genes for human whole blood. They have also been shown to vary in a variety of experimental settings and tissues [2, 33, 36–39], including in whole blood [3]. B2M and PPIB showed stable expression in our study. They were not identified as stably expressed in our Affymetrix data because their expression was higher than our upper limit for selection. PPIB was also reported to be stably expressed in whole blood of patients with local and systemic inflammatory syndromes and healthy controls [3]. HMBS had a high SD based on our Affymetrix arrays on the 526 samples (SD of log-2-transformed values = 0.72). For comparison, the most stably expressed genes in the same intensity interval had a standard deviation (SD) of the log-2-transformed values around 0.2. HMBS, however, showed stable expression when using qRT-PCR in the MS versus healthy controls experiment. Closer inspection of the expression stability on the 526 samples from the Affymetrix chips showed that the HMBS SD of the log-2-transformed values for most of the studies was around 0.5, but it was 0.94 for the stroke study. This result suggests that there were study-specific differences for stroke versus the other diseases, leading to its unstable expression. Our conclusion that HMBS was unstably expressed across our 526 samples is consistent with a previous study reporting that HMBS was unstably expressed in leukocytes [6]. Among the 10 candidate reference genes, TRAP1, DECR1, FPGS, and FARP1 showed the most stable expression.
In most research studies, the target genes or genes regulated within the study are expressed at different levels. In such situations, it is preferable if the comparable endogenous or "housekeeping genes" are expressed at comparable levels as the target gene. This is essential if the microarray, RT-PCR or Northern blotting platform is non-linear at very low or at very high levels. Thus, having a set of endogenous reference genes that are stably expressed at different levels makes it possible to help select those whose expression is comparable to the target genes.
An optimal set of reference genes was identified based on the expression stability measure M as utilized in the GeNorm algorithm. It is based on the concept that the ratio between two ideal reference genes should always be constant regardless of the experimental conditions used. An important assumption here is that the genes in the analysis are not coordinately regulated and have independent functions. The six genes analyzed in GeNorm, seem to be involved in different biological processes, molecular functions and/or are located in different cellular compartments (Table 4). TRAP1 and PPIB are involved in the same biological process however their cellular compartmentalization is different. TRAP1 is a mitochondrial molecular chaperone also known as heat shock protein 75 (Hsp75). PPIB, encoding the cyclophillin B protein, is a cytoplasmic peptide isomerase. The broad spectrum of functions of the stably expressed genes makes it less likely that they are coordinately regulated.
An advantage of the approach taken in this study is that we performed a whole-genome survey of a large number of subjects of different ages, genders and diseases to identify the most stably expressed genes in whole blood. Drawbacks to this approach include the fact that it is an array-based discovery method that is less likely to reliably detect very low expressing genes. In addition, the subjects chosen do reflect some bias related to the diseases studied, and reflect other biases in the subject selection – such as predominantly hospitalied and male patients.
It should be emphasized that the ultimate usefulness of the proposed endogenous reference genes for whole blood has not been demonstrated in this study. To do this, future studies must show that after correcting technical and individual variability according to the control genes, the discriminatory power of the diagnostic genes should be substantially improved.
It is important to point out the utility of the reference genes identified here for use in clinical human testing. Should the reference genes identified in this study provide sufficiently accurate normalization, then genes of interest could be normalized to these reference genes in healthy individuals and in individuals with a given disease or physiological condition. Once reference expression levels and standard deviations are derived in a large group of healthy individuals, deviations could be identified in target individuals using these endogenous genes for normalization without the need for repeated samples from normal, healthy individuals. For non-clinical research studies, such normalization to the same endogenous reference genes would allow for comparison across studies.
Conclusion
We took advantage of the large amount of expression data on 33,000 genes from 526 individuals to select genes that had stable expression in whole blood. This gave us an added advantage to assess not only the gene expression of commonly used housekeeping genes but to perform a whole genome survey for stably expressed genes over multiple ages, disease conditions and different expression levels. Ranking based on the expression stability from qRT-PCR data of the candidate control genes and estimation of the minimum number of genes needed for nomalization showed that the fewest, most stably expressed genes needed for acurate normalization in RNA expression studies of human whole blood is a combination of TRAP1, FPGS, DECR1 and PPIB. These genes should be useful as normalization genes for a wide-range of whole blood expression studies in humans.
References
Huggett J, Dheda K, Bustin S, Zumla A: Real-time RT-PCR normalisation; strategies and considerations. Genes Immun. 2005, 6 (4): 279-284. 10.1038/sj.gene.6364190.
Lee PD, Sladek R, Greenwood CM, Hudson TJ: Control genes and variability: absence of ubiquitous reference transcripts in diverse mammalian expression studies. Genome Res. 2002, 12 (2): 292-297. 10.1101/gr.217802.
Pachot A, Blond JL, Mougin B, Miossec P: Peptidylpropyl isomerase B (PPIB): a suitable reference gene for mRNA quantification in peripheral whole blood. J Biotechnol. 2004, 114 (1–2): 121-124. 10.1016/j.jbiotec.2004.07.001.
Schmittgen TD, Zakrajsek BA: Effect of experimental treatment on housekeeping gene expression: validation by real-time, quantitative RT-PCR. J Biochem Biophys Methods. 2000, 46 (1–2): 69-81. 10.1016/S0165-022X(00)00129-9.
Thellin O, Zorzi W, Lakaye B, De Borman B, Coumans B, Hennen G, Grisar T, Igout A, Heinen E: Housekeeping genes as internal standards: use and limits. J Biotechnol. 1999, 75 (2–3): 291-295. 10.1016/S0168-1656(99)00163-7.
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F: Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3 (7): RESEARCH0034-10.1186/gb-2002-3-7-research0034.
Tang Y, Xu H, Du X, Lit L, Walker W, Lu A, Ran R, Gregg JP, Reilly M, Pancioli A, et al: Gene expression in blood changes rapidly in neutrophils and monocytes after ischemic stroke in humans: a microarray study. J Cereb Blood Flow Metab. 2006, 26 (8): 1089-1102. 10.1038/sj.jcbfm.9600264.
Xu H, Tang Y, Liu DZ, Ran R, Ander BP, Apperson M, Liu XS, Khoury JC, Gregg JP, Pancioli A, et al: Gene expression in peripheral blood differs after cardioembolic compared with large-vessel atherosclerotic stroke: biomarkers for the etiology of ischemic stroke. J Cereb Blood Flow Metab. 2008, 28 (7): 1320-1328. 10.1038/jcbfm.2008.22.
Zhan X, Kim C, Sharp FR: Very brief focal ischemia simulating transient ischemic attacks (TIAs) can injure brain and induce Hsp70 protein. Brain Res. 2008, 1234: 183-197. 10.1016/j.brainres.2008.07.094.
Sharp FR, Xu H, Lit L, Walker W, Apperson M, Gilbert DL, Glauser TA, Wong B, Hershey A, Liu DZ, et al: The future of genomic profiling of neurological diseases using blood. Arch Neurol. 2006, 63 (11): 1529-1536. 10.1001/archneur.63.11.1529.
Du X, Tang Y, Xu H, Lit L, Walker W, Ashwood P, Gregg JP, Sharp FR: Genomic profiles for human peripheral blood T cells, B cells, natural killer cells, monocytes, and polymorphonuclear cells: comparisons to ischemic stroke, migraine, and Tourette syndrome. Genomics. 2006, 87 (6): 693-703. 10.1016/j.ygeno.2006.02.003.
Lit L, Enstrom A, Sharp FR, Gilbert DL: Age-related gene expression in Tourette syndrome. J Psychiatr Res. 2009, 43 (3): 319-330. 10.1016/j.jpsychires.2008.03.012.
Lit L, Gilbert DL, Walker W, Sharp FR: A subgroup of Tourette's patients overexpress specific natural killer cell genes in blood: a preliminary report. Am J Med Genet B Neuropsychiatr Genet. 2007, 144B (7): 958-963. 10.1002/ajmg.b.30550.
Tang Y, Schapiro MB, Franz DN, Patterson BJ, Hickey FJ, Schorry EK, Hopkin RJ, Wylie M, Narayan T, Glauser TA, et al: Blood expression profiles for tuberous sclerosis complex 2, neurofibromatosis type 1, and Down's syndrome. Ann Neurol. 2004, 56 (6): 808-814. 10.1002/ana.20291.
Tang Y, Lu A, Ran R, Aronow BJ, Schorry EK, Hopkin RJ, Gilbert DL, Glauser TA, Hershey AD, Richtand NW, et al: Human blood genomics: distinct profiles for gender, age and neurofibromatosis type 1. Brain Res Mol Brain Res. 2004, 132 (2): 155-167. 10.1016/j.molbrainres.2003.10.014.
Tang Y, Glauser TA, Gilbert DL, Hershey AD, Privitera MD, Ficker DM, Szaflarski JP, Sharp FR: Valproic acid blood genomic expression patterns in children with epilepsy – a pilot study. Acta Neurol Scand. 2004, 109 (3): 159-168. 10.1046/j.1600-0404.2003.00253.x.
Tang Y, Gilbert DL, Glauser TA, Hershey AD, Sharp FR: Blood gene expression profiling of neurologic diseases: a pilot microarray study. Arch Neurol. 2005, 62 (2): 210-215. 10.1001/archneur.62.2.210.
Tang Y, Lu A, Aronow BJ, Sharp FR: Blood genomic responses differ after stroke, seizures, hypoglycemia, and hypoxia: blood genomic fingerprints of disease. Ann Neurol. 2001, 50 (6): 699-707. 10.1002/ana.10042.
Hershey AD, Tang Y, Powers SW, Kabbouche MA, Gilbert DL, Glauser TA, Sharp FR: Genomic abnormalities in patients with migraine and chronic migraine: preliminary blood gene expression suggests platelet abnormalities. Headache. 2004, 44 (10): 994-1004. 10.1111/j.1526-4610.2004.04193.x.
Wong B, Gilbert DL, Walker WL, Liao IH, Lit L, Stamova B, Jickling G, Apperson M, Sharp FR: Gene expression in blood of subjects with Duchenne muscular dystrophy. Neurogenetics. 2008, 9: 9.
Walker WL, Liao IH, Gilbert DL, Wong B, Pollard KS, McCulloch CE, Lit L, Sharp FR: Empirical Bayes accomodation of batch-effects in microarray data using identical replicate reference samples: application to RNA expression profiling of blood from Duchenne muscular dystrophy patients. BMC Genomics. 2008, 9 (494): 494-10.1186/1471-2164-9-494.
Enstrom AM, Lit L, Onore CE, Gregg JP, Hansen RL, Pessah IN, Hertz-Picciotto I, Water Van de JA, Sharp FR, Ashwood P: Altered gene expression and function of peripheral blood natural killer cells in children with autism. Brain Behav Immun. 2009, 23 (1): 124-133. 10.1016/j.bbi.2008.08.001.
Gregg JP, Lit L, Baron CA, Hertz-Picciotto I, Walker W, Davis RA, Croen LA, Ozonoff S, Hansen R, Pessah IN, et al: Gene expression changes in children with autism. Genomics. 2008, 91 (1): 22-29. 10.1016/j.ygeno.2007.09.003.
Osman F, Leutenegger C, Golino D, Rowhani A: Comparison of low-density arrays, RT-PCR and real-time TaqMan RT-PCR in detection of grapevine viruses. J Virol Methods. 2008, 149 (2): 292-299. 10.1016/j.jviromet.2008.01.012.
Wu Z, Irizarry RA: Preprocessing of oligonucleotide array data. Nat Biotechnol. 2004, 22 (6): 656-658. 10.1038/nbt0604-656b.
Schmittgen TD, Livak KJ: Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc. 2008, 3 (6): 1101-1108. 10.1038/nprot.2008.73.
Andersen CL, Jensen JL, Orntoft TF: Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64 (15): 5245-5250. 10.1158/0008-5472.CAN-04-0496.
Livak KJ, Schmittgen TD: Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001, 25 (4): 402-408. 10.1006/meth.2001.1262.
Biederman J, Yee J, Cortes P: Validation of internal control genes for gene expression analysis in diabetic glomerulosclerosis. Kidney Int. 2004, 66 (6): 2308-2314. 10.1111/j.1523-1755.2004.66016.x.
Etschmann B, Wilcken B, Stoevesand K, Schulenburg von der A, Sterner-Kock A: Selection of reference genes for quantitative real-time PCR analysis in canine mammary tumors using the GeNorm algorithm. Vet Pathol. 2006, 43 (6): 934-942. 10.1354/vp.43-6-934.
Hoerndli FJ, Toigo M, Schild A, Gotz J, Day PJ: Reference genes identified in SH-SY5Y cells using custom-made gene arrays with validation by quantitative polymerase chain reaction. Anal Biochem. 2004, 335 (1): 30-41. 10.1016/j.ab.2004.08.028.
Jung M, Ramankulov A, Roigas J, Johannsen M, Ringsdorf M, Kristiansen G, Jung K: In search of suitable reference genes for gene expression studies of human renal cell carcinoma by real-time PCR. BMC Mol Biol. 2007, 8 (47): 47-10.1186/1471-2199-8-47.
Ohl F, Jung M, Radonic A, Sachs M, Loening SA, Jung K: Identification and validation of suitable endogenous reference genes for gene expression studies of human bladder cancer. J Urol. 2006, 175 (5): 1915-1920. 10.1016/S0022-5347(05)00919-5.
Radonic A, Thulke S, Bae HG, Muller MA, Siegert W, Nitsche A: Reference gene selection for quantitative real-time PCR analysis in virus infected cells: SARS corona virus, Yellow fever virus, Human Herpesvirus-6, Camelpox virus and Cytomegalovirus infections. Virol J. 2005, 2 (7): 7-10.1186/1743-422X-2-7.
Silver N, Best S, Jiang J, Thein SL: Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol Biol. 2006, 7 (33): 33-10.1186/1471-2199-7-33.
Hamalainen HK, Tubman JC, Vikman S, Kyrola T, Ylikoski E, Warrington JA, Lahesmaa R: Identification and validation of endogenous reference genes for expression profiling of T helper cell differentiation by quantitative real-time RT-PCR. Anal Biochem. 2001, 299 (1): 63-70. 10.1006/abio.2001.5369.
Roge R, Thorsen J, Torring C, Ozbay A, Moller BK, Carstens J: Commonly used reference genes are actively regulated in in vitro stimulated lymphocytes. Scand J Immunol. 2007, 65 (2): 202-209. 10.1111/j.1365-3083.2006.01879.x.
Ross DT, Scherf U, Eisen MB, Perou CM, Rees C, Spellman P, Iyer V, Jeffrey SS, Rijn Van de M, Waltham M, et al: Systematic variation in gene expression patterns in human cancer cell lines. Nat Genet. 2000, 24 (3): 227-235. 10.1038/73432.
Warrington JA, Nair A, Mahadevappa M, Tsyganskaya M: Comparison of human adult and fetal expression and identification of 535 housekeeping/maintenance genes. Physiol Genomics. 2000, 2 (3): 143-147.
Pre-publication history
The pre-publication history for this paper can be accessed here:http://www.biomedcentral.com/1755-8794/2/49/prepub
Acknowledgements
These studies were supported by grants from: the NIH/NINDS (NS056302); an American Stroke Association Bugher Foundation Center for Stroke Prevention Research; Cure Autism Now/Autism Speaks; the UC Discovery Program and Pediatric Biosciences; the Tourette Syndrome Association; and a gift from the Ron and Darrin Mittelstaedt family supported Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
BSS, MA, WLW and FRS designed the study, analyzed the results and wrote the manucript. MA provided the samples from the MS patients for the RT-PCR and array studies. YT, HX, PA, XA, DL, BPA, IHL, JPG, RJT, GCJ and LL either helped collect samples, helped analyze the data from the microarray studies and/or read and proof read the manuscript. JPG supervised the processing of a number of the arrays used in these studies. All authors read and approved the final manuscript.
Electronic supplementary material
12920_2009_113_MOESM1_ESM.xls
Additional file 1: 100 Most Stable Genes and 10 Affy Candidate Reference Genes. Information on sheet 2 represents the 100 most stable genes including Affynetrix probe-set IDs, reference public IDs, gene symbols, and expression levels. The data on sheet 2 represents the 10 Affymetrix candidate reference genes with probe-sets, ABI assay, and expression levels information. (XLS 92 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Stamova, B.S., Apperson, M., Walker, W.L. et al. Identification and validation of suitable endogenous reference genes for gene expression studies in human peripheral blood. BMC Med Genomics 2, 49 (2009). https://doi.org/10.1186/1755-8794-2-49
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1755-8794-2-49