Abstract
Background
Array comparative genomic hybridization (CGH) has been repeatedly shown to be a successful tool for the identification of genomic variations in a clinical population. During the last decade, the implementation of array CGH has resulted in the identification of new causative submicroscopic chromosome imbalances and copy number variations (CNVs) in neuropsychiatric (neurobehavioral) diseases. Currently, array-CGH-based technologies have become an integral part of molecular diagnosis and research in individuals with neuropsychiatric disorders and children with intellectual disability (mental retardation) and congenital anomalies. Here, we introduce the Russian cohort of children with intellectual disability, autism, epilepsy and congenital anomalies analyzed by BAC array CGH and a novel bioinformatic strategy.
Results
Among 54 individuals highly selected according to clinical criteria and molecular and cytogenetic data (from 2426 patients evaluated cytogenetically and molecularly between November 2007 and May 2012), chromosomal imbalances were detected in 26 individuals (48%). In two patients (4%), a previously undescribed condition was observed. The latter has been designated as meiotic (constitutional) genomic instability resulted in multiple submicroscopic rearrangements (including CNVs). Using bioinformatic strategy, we were able to identify clinically relevant CNVs in 15 individuals (28%). Selected cases were confirmed by molecular cytogenetic and molecular genetic methods. Eight out of 26 chromosomal imbalances (31%) have not been previously reported. Among them, three cases were co-occurrence of subtle chromosome 9 and 21 deletions.
Conclusions
We conducted an array CGH study of Russian patients suffering from intellectual disability, autism, epilepsy and congenital anomalies. In total, phenotypic manifestations of clinically relevant genomic variations were found to result from genomic rearrangements affecting 1247 disease-causing and pathway-involved genes. Obviously, a significantly lesser part of them are true candidates for intellectual disability, autism or epilepsy. The success of our preliminary array CGH and bioinformatic study allows us to expand the cohort. According to the available literature, this is the first comprehensive array CGH evaluation of a Russian cohort of children with neuropsychiatric disorders and congenital anomalies.
Similar content being viewed by others
Background
Chromosomal imbalances and copy number variations (CNVs) are probably the most common genetic causes of intellectual disability (mental retardation) and congenital anomalies. Genome variations involving chromosomal and subchromosomal loci are frequently detected in a wide spectrum of neuropsychiatric disorders. Indeed, these patients seem to need in an evaluation by array comparative genomic hybridization (CGH) or molecular karyotyping. The latter is repeatedly shown to be a powerful tool for the identification of genomic variations (submicroscopic chromosomal imbalances and CNVs) in a clinical population. Moreover, to ensure an adequate diagnostic yield (i.e. >10-15%), molecular diagnosis of constitutional chromosomal and subchromosomal imbalances is recommended to be performed by molecular karyotyping or related array-CGH-based technologies, which has become an important genetic test for patients suffering from intellectual disability (neuropsychiatric diseases) and congenital anomalies. Whole-genome scanning technologies are unique for detecting losses or gains of genomic material and are consistently used for studying genetic causes of postnatal morbidity (i.e. dysmorphology, malformations, developmental delay, intellectual disability, autism, epilepsy, and schizophrenia) providing for the delineation of the molecular mechanisms and causative genes [1–10].
Here, we present the first Russian experience of an array CGH application to a clinical population. The cohort included 54 children with intellectual disability, autism, epilepsy and congenital anomalies highly selected from 2426 patients according to clinical criteria and molecular and cytogenetic data.
Results
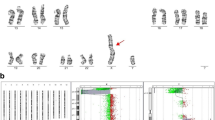
Fifty four members of the Russian cohort of children with intellectual disability, autism, epilepsy and congenital anomalies were studied by BAC array CGH with the resolution of 0.3-1 Mb. Some examples of array CGH graphical overviews are shown in Figure 1. Details of the cohort (array CGH results and patients) are summarized in Table 1. Figure 2 overviews the incidence of genomic imbalances in the cohort according to their type and to the diagnosing feasibility or, in other words, the difference between cytogenetically visible (patients re-evaluated by cytogenetic analysis after array CGH) and cytogenetically invisible chromosome aberrations. CNVs were evaluated by a bioinformatic approach towards the CNV/gene prioritization followed by genotype-phenotype correlations. Selected cases were confirmed by molecular cytogenetic and molecular genetic methods (Table 1).
Examples of array CGH graphical overviews. A. Case #8: a deletion of chromosome X short arm and a duplication of chromosome 3 short arm — arr Xp22.33p22.2(2,333,897-9,726,574)x1,3p26.3p22.3(200,000-36,550,871)x3 — due to an unbalanced maternal translocation t(X;3) (conventional karyotyping was performed after array CGH analysis). B. Case #27: a deletion of chromosome 7 long arm (subtelomeric 7q deletion) — arr 7q36.2q36.3(152,768,630-158,261,821)x1. C. Case #18: a duplication of chromosome 19 short arm — arr 19p13.3(260,000-4,953,188)x3. D. Case #20: a deletion of chromosome 10 long arm (subtelomeric 10q deletion) — arr 10q26.2q26.3(128,192,760-134,070,099)x1. E. Case #40: a duplication of chromosome 5 long arm — arr 5q13.2(68,931,140-72,690,180)x3.
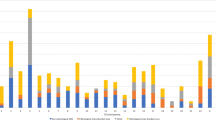
The incidence of chromosome imbalances (subdivided to cytogenetically detectable and undetectable abnormalities), CNVs and meiotic genome instability in the Russian cohort of children with intellectual disability with or without autism and congenital anomalies. Cytogenetically detectable cases were patients, who were cytogenetically re-evaluated. This has yielded the correct diagnosis (see also Table 1).
Chromosomal imbalances
Constitutive chromosomal imbalances were detected in 26 individuals (including case #1 exhibiting three presumably unrelated chromosome abnormalities). Chromosomal imbalances (including multiple chromosome abnormalities) observed in cases #1, #8, #11, #11, #12, #18, #26, #40, and #43 were found to be unique to this cohort (Table 1). Among them, we found a recurrent chromosome abnormality specific to the Russian cohort of children with intellectual disability, autism, epilepsy and congenital anomalies. This was referred to co-occurrence of chromosome 9 long arm deletion in 9q34.2q34.3 and chromosome 21 long arm deletion in 21q22.3 (cases #11, #26, #43; Table 1). One can suggest that a ~179 kbp interval on 9q34.2 and a ~187 kbp interval on 21q22.3 (deduced on the basis of breakpoint locations) are both specifically organized at the sequence level to produce a complex genomic rearrangement causing intellectual disability with autistic features, speech delay and facial dismorphisms.
CNVs
Cases #22 and #28 exhibited multiple CNVs (more than three submicroscopic duplications). All the duplications have been defined to contribute to the severe phenotypic outcome by bioinformatic analysis. Since similar cases were not found in the available literature, we have taken opportunity to designate the condition as meiotic (constitutional) genomic instability.
By bioinformatics, we have identified clinically relevant CNVs in 15 individuals. These were found to encompass genes, mutations in which cause developmental delays, congenital anomalies, intellectual disability, autism, epilepsy, or other neuropsychiatric disorders. Alternatively, CNV genes, which have not as yet been associated with a particular disease phenotype, were not necessarily those associated with benign genome variations. The bioinformatic analysis has demonstrated a significant proportion of such genes to be involved in functional pathways, which, if altered, can be disease-causing (see Additional file 1).
Comparative cohort characterization
To make a comparative characterization of the cohort, we have addressed it in a case-by-case manner. Previously unreported CNVs, exclusive chromosomal imbalances and meiotic genome instability were excluded. Case # 1 is the unique combination of known chromosomal abnormalities [13, 14]. Multiple chromosome abnormalities are extremely rare and are usually associated with severe phenotypes (i.e. prenatal mortality) [15, 16]. Severe congenital anomalies were eventually observed in this case.
Case #2 is a microdeletion of the chromosome 19 long arm that is clinically similar to 19q13.11 deletion syndrome [17]. Case #3 is a chromosome 20 long arm microdeletion associated with mild intellectual disability and facial dysmorphisms. The chromosomal region was involved in larger chromosome 20 long arm deletions, which demonstrated more severe phenotypes [18].
Ring chromosome 11 (case #4) characterized in the present study is phenotypically similar to previous cases [19], exhibiting, however, additional clinical features (autism), which are attributed to the presence of CNVs. Case #6 has turned to be a recurrent translocation between chromosomes 4 and 8 causing the Wolf-Hirschhorn syndrome, being, however, more complex as those described previously [20].
Case #8 was a chromosome imbalance identified in a distant patient (immediate cytogenetic analysis was not available). After array CGH, additional further molecular cytogenetic studies and cytogenetic re-evaluation, it was defined as an unbalanced translocation between short arms of chromosomes X and 3. Several similar cases have been described [21, 22].
Case #12 is a deletion of chromosome 3 short arm in a child with Asperger syndrome. Chromosomal abnormalities are occasional in this neurobehavioral disorder. However, a study has shown positive Asperger syndrome linkage to 3p21-24 [23].
In case #13, a CNV (duplication) in Xp11.23 was found. Recently, a duplication of the same chromosomal region was shown to cause similar phenotype [24]. Case #15 was another chromosome imbalance identified in a distant patient that has been defined as an unbalanced translocation between long arms of chromosomes 10 and 13 after cytogenetic re-evaluation.
Chromosomal rearrangements in the chromosome 19 short arm are extremely rare [25]. Case #18 is the largest ever reported 19p subtelomeric duplication. Case #20 is a 10q subtelomeric deletion; similar cases were previously described, but the clinical manifestations are usually variable and a specific phenotype is not associated [2]. Case #23 exhibits a number of clinical features of the Xq28 duplication syndrome [26]; the phenotypic discrepancies have been attributed to CNVs.
A 7q subtelomeric deletion was identified in case #27. Despite of characteristic clinical features (lumbosacral dysgenesis) [27], it was almost impossible to suggest this chromosomal imbalance prior to array CGH. Cases #30 and #52 have demonstrated chromosome Y long arm deletions. Similar cases were detected during a case–control array CGH study applied to prenatal diagnosis, but the outcome has remained uncertain [28].
Cases #32, #32, #49 and #50 were found to exhibit Xq28 deletions. A retrospective clinical analysis has shown that Rett-syndrome-like phenotypic manifestations do present in these girls. It is not surprising inasmuch as two cases of these deletions involved MECP2. Interestingly, there have been several attempts to characterize chromosomal rearrangements by molecular cytognetic techniques in Rett syndrome cohorts without apparent evidence for the presence of genomic variations involving MECP2[29–31].
In case #39, a CNV within RB1 gene was found and the phenotype was similar to interstitial 13q microdeletions [32]. An Xq28 CNV (involving RAB39B gene) was found in a child with intellectual disability and autism (case #48). It is to note, that RAB39B mutations cause X-linked mental retardation associated with autism, epilepsy, and macrocephaly [33].
Finally, case #53 was the third chromosome imbalance identified in a distant patient. Subsequently, it turned out to be an unbalanced translocation between the long arm of chromosome 1and the short arm of chromosome 16 after fluorescence in situ hybridization (FISH) and cytogenetic evaluation. Comparing the phenotype with previous reports [34, 35], we concluded that the clinical features in this case has similarities as with trisomy 16p as with 1q subtelomeric deletions.
Discussion
The present study has again provided evidence that array CGH is a powerful technique for uncovering chromosomal imbalances and genomic rearrangements. Previous case–control studies (reviewed in [2, 5, 8–10]) have demonstrated the high diagnostic yield of microarray-based whole-genome analysis and the validity of array-CGH-based technologies for detecting genomic imbalances in clinical populations. However, the detection rate (over 50%) seems to be significantly higher in our study comparing to the previous ones (5-20%) [1–10]. Since the incidence of genomic rearrangements detectable by array CGH is strongly influenced by selection bias of patients, the reported detection rate is likely to be higher inasmuch as our criteria were much more discriminating than previously applied (i.e. evaluating the diagnostic yield in “idiopathic” cohorts). Thus, we do not insist that the reported detection rate is characteristic for an array CGH control cohort study. Still, it can be expected that an application of array CGH to highly selected patients is able to demonstrate an impressive detection rate of structural genome variations.
It is generally recognized that numerical and structural chromosome abnormalities are the major genetic causes of postnatal morbidity including a wide spectrum of diseases associated with brain dysfunction [13, 14]. However, before the implementation of molecular cytogenetic techniques (array-CGH- and FISH-based technologies), the contribution of genome/chromosome imbalances to the etiology was usually considered as less than 5% [4, 7, 9, 10, 14]. Array CGH data are not only relevant to molecular diagnosis, but also to the discovery of genetic (genomic) mechanisms of brain diseases [1–10, 14]. Here, we were also able to demonstrate similar etiologic yield. In this instance, the present gene list (Additional file 1) possesses implications for the delineation of the cellular and molecular basis of disease and requires further bioinformatic analyses.
Although array CGH is a highly efficient technique for definition of chromosome/genome rearrangements in clinical populations, it has limitations. For example, it does not properly reflect somatic mosaicism [36–38], which is commonly associated with postnatal morbidity, including brain diseases [37–41]. The pathogenic value of submicroscopic genomic variations can be a matter of conjecture [2–10]. Therefore, array-CGH-based methods have to be combined with molecular cytogenetic, post-genomic and bioinformatic technologies for detailed studies of disease mechanisms. Nonetheless, array CGH assays are mandatory for such studies.
Apart from numerical constitutional and mosaic chromosome abnormalities (case #1) the genomic rearrangements were found to affect 1247 genes in total. Details about these genes are provided in Additional file 1. Accordingly, we speculate that the majority of these genes can be mutated in intellectual disability, autism or epilepsy. The latter can be easily seen if one addresses a molecular/clinical genetic database (i.e. OMIM — http://www.omim.org/; referenced in Additional file 1). Moreover, combinations of deletions/duplications affecting these genomic loci produce a “CNV burden”, which can be considered a possible genetic cause of postnatal morbidity. On the other hand, it is to recognize that a significantly lesser part of them are true candidates for the aforementioned neuropsychiatric diseases.
Array CGH does improve the etiologic yield across the spectrum of patients with neuropsychiatric disorders and congenital anomalies. This improvement has been achieved by systematic cohort studies. The available literature indicates that we report on the first comprehensive array CGH analysis of a Russian cohort of children with intellectual disability, autism, epilepsy and congenital anomalies. The efficiency of the approach (array CGH + bioinformatic strategy) allows us to expand the cohort. To this end, it is necessary again to point out that patient pre-selection provided for the discovery of up to 31% of previously unreported genomic rearrangements among detectable imbalances. In other words, this report describes novel disease genes uncovered in a relatively small cohort. Hence, our next study of the expanded cohort would certainly lead to discovering new candidate genes and shared molecular pathways of intellectual disability, autism, epilepsy and related neuropsychiatric disorders.
Methods
Patients
From November 2007 to May 2012, 2426 patients referred to our molecular cytogenetic facilities. All the patients were studied by conventional karyotyping (G- and C-banding analyses) at a G-banding resolution of about 550 bands according to ISCN (2013) [11]. The ages varied between 1 month and 18 years. Fifty four patients were highly selected to be the first participants of the Russian cohort of children with intellectual disability, autism, epilepsy and congenital anomalies based on clinical and cytogenetic/molecular cytogenetic data. Molecular cytogenetic techniques (FISH-based methods) were used to exclude cases of cytogenetically visible chromosome abnormalities (confirmation by molecular cytogenetic techniques) and microdeletion/microduplication syndromes. FISH was also used in cases of chromosomal mosaicism (especially, those children who were the participants of the Russian autism cohort and were shown to exhibit chromosomal mosaicism; for more details see [42–44]). Such cases have been excluded. Subtelomeric chromosomal rearrangements (commonly detected in children with intellectual disability [2]), addressed by a previously reported FISH analysis with original DNA probes [45], were excluded from the cohort, as well. Cases positive for mutations causing fragile X and Rett syndrome (common causes of intellectual disability and autistic features in children) addressed by molecular genetic techniques (PCR/QPCR and direct sequencing of the MECP2 gene) were ruled out. For three distant patients, immediate cytogenetic analysis was not available due to natural (distance) limitations. As a result, they were included in the cohort. Two cytogenetically positive cases (supernumerary chromosome X in a male child and ring chromosome 11) exhibiting extremely atypical phenotypic manifestations were also included. Written informed consent was obtained from the patients’ parents.
Array CGH
Array CGH was performed with the customized human genomic microarrays (slightly modified Constitutional Chip®4.0) containing about 5200 human BAC clones (Human BAC Array-System, Perkin Elmer, USA). The resolution of the whole genome scan was estimated to be 0.3-1 Mb. The microarrays contain more than 1200 BAC probes for the majority of "new" and "old" microdeletion/microduplication syndromes (for the description see [2, 3, 5, 7]), about 900 BAC probes — subtelomeric regions, about 100 BAC probes — percintromeric chromosomal regions, 621 BAC probes — chromosome X, and about 2000 BAC probes — remaining euchromatic chromosomal regions, allowing the whole genome to be scanned with a resolution of at least 1 Mb. DNA labelling, hybridization, detection and data analysis was made according to previously described protocols [46] and to manufacturers’ instructions.
Confirmation methods
Molecular cytogenetic (FISH) and molecular genetic (PCR/QPCR) methods were used for the confirmation of genomic rearrangements in a number of cases (n=18). FISH with DNA probes from our original collection (probes for heterochromatic and euchromatic (repetitive and unique) chromosomal regions) was performed as described previously [45, 47, 48]. The confirmation was performed in nine cases (Table 1).
PCR (or QPCR) was essentially used to confirm the rearrangements affecting chromosomes X and Y (i.e. deletions of AR in Xq12, MECP2 in Xq28, and DAZ loci in Yq11.223). The confirmation was performed in nine cases (Table 1).
Bioinformatics
In silico (bioinformatic) analyses were performed in part according to Iourov et al., 2009, 2010 [49, 50]. To determine the pathogenic value of CNVs, genotype-phenotype correlations and CNV or gene prioritization were done by a series of evaluations using clinical/cytogenetic and genomic variation databases as well as bioinformatic tools for genome, epigenome and pathway analysis.
Genotype-phenotype correlations and pathogenic value of CNVs were estimated using DECIPHER (database of unbalanced chromosome aberrations) — http://decipher.sanger.ac.uk/, OMIM (online Mendelian inheritance in Man) — http://www.omim.org/, The Phenotype-Genotype Integrator (PheGenI) — http://www.ncbi.nlm.nih.gov/gap/PheGenI, SFARI Gene/AutDB (web-based searchable database for autism research) — http://www.mindspec.org/autdb.html, and a Catalog of Published Genome-Wide Association Studies (NHGRI) — http://www.genome.gov/gwastudies/.
CNVs were also compared to the Database of Genomic Variants (http://dgvbeta.tcag.ca/dgv/app/home?ref=GRCh37/hg19). The prioritization of CNVs and genes was done using BioGPS (a gene annotation and expression) — http://biogps.org, Ensembl Genome Browser — http://www.ensembl.org/index.html, KEGG (Kyoto Encyclopedia of Genes and Genomes) — http://www.genome.jp/kegg/, NCBI BioSystems Database — http://www.ncbi.nlm.nih.gov/biosystems, NCBI Gene — http://www.ncbi.nlm.nih.gov/gene/, NCBI Build 37.1/NCBI Map Viewer — http://www.ncbi.nlm.nih.gov/projects/mapview/map_search.cgi?taxid=9606, REACTOME — http://www.reactome.org/, Pathway Commons — http://www.pathwaycommons.org/pc/, and UCSC Genome Browser — http://genome.ucsc.edu/.
Endnote
aThe data reported in the present article have been presented in part at the European Human Genetics Conference 2011, Amsterdam, The Netherlands, May 28–31, 2011.
Abbreviations
- BAC:
-
bacterial artificial chromosome
- CNVs:
-
copy number variations
- CGH:
-
comparative genomic hybridization
- FISH:
-
fluorescence in situ hybridization
- PCR:
-
polymerase chain reaction
- QPCR:
-
quantitative polymerase chain reaction.
References
Bejjani BA, Shaffer LG: Application of array-based comparative genomic hybridization to clinical diagnostics. J Mol Diagn 2006, 8: 528–533. 10.2353/jmoldx.2006.060029
Knight SJ, Regan R: Idiopathic learning disability and genome imbalance. Cytogenet Genome Res 2006, 115: 215–224. 10.1159/000095917
Emanuel BS, Saitta SC: From microscopes to microarrays: dissecting recurrent chromosomal rearrangements. Nat Rev Genet 2007, 8: 869–883. 10.1038/nrg2136
Stankiewicz P, Beaudet AL: Use of array CGH in the evaluation of dysmorphology, malformations, developmental delay, and idiopathic mental retardation. Curr Opin Genet Dev 2007, 17: 182–192. 10.1016/j.gde.2007.04.009
Lee C, Iafrate AJ, Brothman AR: Copy number variations and clinical cytogenetic diagnosis of constitutional disorders. Nat Genet 2007, 39(7 Suppl):S48-S54.
Bejjani BA, Shaffer LG: Clinical utility of contemporary molecular cytogenetics. Annu Rev Genomics Hum Genet 2008, 9: 71–86. 10.1146/annurev.genom.9.081307.164207
Iourov IY, Vorsanova SG, Yurov YB: Molecular cytogenetics and cytogenomics of brain diseases. Curr Genomics 2008, 9: 452–465. 10.2174/138920208786241216
Sagoo GS, Butterworth AS, Sanderson S, Shaw-Smith C, Higgins JP, Burton H: Array CGH in patients with learning disability (mental retardation) and congenital anomalies: updated systematic review and meta-analysis of 19 studies and 13,926 subjects. Genet Med 2009, 11: 139–146. 10.1097/GIM.0b013e318194ee8f
Vissers LE, de Vries BB, Veltman JA: Genomic microarrays in mental retardation: from copy number variation to gene, from research to diagnosis. J Med Genet 2010, 47: 289–297. 10.1136/jmg.2009.072942
Hochstenbach R, Buizer-Voskamp JE, Vorstman JA, Ophoff RA: Genome arrays for the detection of copy number variations in idiopathic mental retardation, idiopathic generalized epilepsy and neuropsychiatric disorders: lessons for diagnostic workflow and research. Cytogenet Genome Res 2011, 135: 174–202. 10.1159/000332928
ISCN: An International System of Human Cytogenetic Nomenclature, Shaffer LG, McGowan-Jordan J, Schmid M (eds); S. Basel: Karger; 2013. 2013
Slavotinek AM: Novel microdeletion syndromes detected by chromosome microarrays. Hum Genet 2008, 124: 1–17. 10.1007/s00439-008-0513-9
Gersen SL, Keagle MB: The principles of clinical cytogenetics. Humana press 2005.
Iourov IY, Vorsanova SG, Yurov YB: Chromosomal variation in mammalian neuronal cells: known facts and attractive hypotheses. Int Rev Cytol 2006, 249: 143–191.
Reddy KS: Double trisomy in spontaneous abortions. Hum Genet 1997, 101: 339–345. 10.1007/s004390050638
Vorsanova SG, Iourov IY, Demidova IA, Kirillova EA, Soloviev IV, Yurov YB: Chimerism and multiple numerical chromosome imbalances in a spontaneously aborted fetus. Tsitol Genet 2006, 40: 28–30.
Malan V, Raoul O, Firth HV, Royer G, Turleau C, Bernheim A, Willatt L, Munnich A, Vekemans M, Lyonnet S, Cormier-Daire V, Colleaux L: 19q13.11 deletion syndrome: a novel clinically recognisable genetic condition identified by array comparative genomic hybridisation. J Med Genet 2009, 46: 635–640. 10.1136/jmg.2008.062034
Hiraki Y, Nishimura A, Hayashidani M, Terada Y, Nishimura G, Okamoto N, Nishina S, Tsurusaki Y, Doi H, Saitsu H, Miyake N, Matsumoto N: A de novo deletion of 20q11.2-q12 in a boy presenting with abnormal hands and feet, retinal dysplasia, and intractable feeding difficulty. Am J Med Genet A 2011, 155A: 409–414.
Hansson KB, Gijsbers AC, Oostdijk W, Rehbock JJ, de Snoo F, Ruivenkamp CA, Kant SG: Molecular and clinical characterization of patients with a ring chromosome 11. Eur J Med Genet 2012, 55: 708–714. 10.1016/j.ejmg.2012.08.004
Zollino M, Murdolo M, Marangi G, Pecile V, Galasso C, Mazzanti L, Neri G: On the nosology and pathogenesis of Wolf-Hirschhorn syndrome: genotype-phenotype correlation analysis of 80 patients and literature review. Am J Med Genet C Semin Med Genet 2008, 148C: 257–269. 10.1002/ajmg.c.30190
Donnenfeld AE, Graham JM Jr, Packer RJ, Aquino R, Berg SZ, Emanuel BS: Microphthalmia and chorioretinal lesions in a girl with an Xp22.2-pter deletion and partial 3p trisomy: clinical observations relevant to Aicardi syndrome gene localization. Am J Med Genet 1990, 37: 182–186. 10.1002/ajmg.1320370205
Han DH, Chang JY, Lee WI, Bae CW: A case of partial trisomy 3p syndrome with rare clinical manifestations. Korean J Pediatr 2012, 55: 107–110. 10.3345/kjp.2012.55.3.107
Rehnström K, Ylisaukko-oja T, Nieminen-von Wendt T, Sarenius S, Källman T, Kempas E, von Wendt L, Peltonen L, Järvelä I: Independent replication and initial fine mapping of 3p21–24 in Asperger syndrome. J Med Genet 2006, 43: 6.
Alesi V, Bertoli M, Barrano G, Torres B, Pusceddu S, Pastorino M, Perria C, Nardone AM, Novelli A, Serra G: 335.4 kb microduplication in chromosome band Xp11.2p11.3 associated with developmental delay, growth retardation, autistic disorder and dysmorphic features. Gene 2012, 505: 384–387. 10.1016/j.gene.2012.05.031
Siggberg L, Olsén P, Näntö-Salonen K, Knuutila S: 19p13.3 aberrations are associated with dysmorphic features and deviant psychomotor development. Cytogenet Genome Res 2011, 132: 8–15. 10.1159/000320920
El-Hattab AW, Fang P, Jin W, Hughes JR, Gibson JB, Patel GS, Grange DK, Manwaring LP, Patel A, Stankiewicz P, Cheung SW: Int22h-1/int22h-2-mediated Xq28 rearrangements: intellectual disability associated with duplications and in utero male lethality with deletions. J Med Genet 2011, 48: 840–850. 10.1136/jmedgenet-2011-100125
Vorsanova SG, Iourov IY, Voinova-Ulas VY, Weise A, Monakhov VV, Kolotii AD, Soloviev IV, Novikov PV, Yurov YB, Liehr T: Partial monosomy 7q34-qter and 21pter-q22.13 due to cryptic unbalanced translocation t(7;21) but not monosomy of the whole chromosome 21: a case report plus review of the literature. Mol Cytogenet 2008, 1: 13. 10.1186/1755-8166-1-13
Park JH, Woo JH, Shim SH, Yang SJ, Choi YM, Yang KS, Cha DH: Application of a target array comparative genomic hybridization to prenatal diagnosis. BMC Med Genet 2010, 11: 102.
Vorsanova SG, Demidova IA, Ulas VY, Soloviev IV, Kazantzeva LZ, Yurov YB: Cytogenetic and molecular-cytogenetic investigation of Rett syndrome: analysis of 31 cases. Neuroreport 1996, 8: 187–189. 10.1097/00001756-199612200-00038
Vorsanova SG, Yurov YB, Ulas VY, Demidova IA, Sharonin VO, Kolotii AD, Gorbatchevskaia NL, Beresheva AK, Soloviev IV: Cytogenetic and molecular-cytogenetic studies of Rett syndrome (RTT): a retrospective analysis of a Russian cohort of RTT patients (the investigation of 57 girls and three boys). Brain Dev 2001, 23(Suppl 1):S196-S201.
Vorsanova SG, Iourov IY, Yurov YB: Neurological, genetic and epigenetic features of Rett syndrome. J Pediatr Neurol 2004, 2: 179–190.
Mitter D, Ullmann R, Muradyan A, Klein-Hitpass L, Kanber D, Ounap K, Kaulisch M, Lohmann D: Genotype-phenotype correlations in patients with retinoblastoma and interstitial 13q deletions. Eur J Hum Genet. 2011, 19: 947–958. 10.1038/ejhg.2011.58
Giannandrea M, Bianchi V, Mignogna ML, Sirri A, Carrabino S, D'Elia E, Vecellio M, Russo S, Cogliati F, Larizza L, et al.: Mutations in the small GTPase gene RAB39B are responsible for X-linked mental retardation associated with autism, epilepsy, and macrocephaly. Am J Hum Genet 2010, 86: 185–195. 10.1016/j.ajhg.2010.01.011
Sommer A, Pastore M, Wenger G: Trisomy 16p: a longitudinal profile and photo essay. Am J Med Genet A 2006, 140: 174–179.
Shimojima K, Okamoto N, Suzuki Y, Saito M, Mori M, Yamagata T, Momoi MY, Hattori H, Okano Y, Hisata K, et al.: Subtelomeric deletions of 1q43q44 and severe brain impairment associated with delayed myelination. J Hum Genet 2012, 57: 593–600. 10.1038/jhg.2012.77
Vorsanova SG, Yurov YB, Iourov IY: Human interphase chromosomes: a review of available molecular cytogenetic technologies. Mol Cytogenet 2010, 3: 1. 10.1186/1755-8166-3-1
Iourov IY, Vorsanova SG, Yurov YB: Chromosomal mosaicism goes global. Mol Cytogenet 2008, 1: 26. 10.1186/1755-8166-1-26
Iourov IY: Vorsanova SG. Yurov YB: Somatic Genome Variations. eLS; 2012.
Iourov IY, Vorsanova SG, Yurov YB: Intercellular genomic (chromosomal) variations resulting in somatic mosaicism: mechanisms and consequences. Curr Genomics 2006, 7: 435–446. 10.2174/138920206779116756
Iourov IY, Vorsanova SG, Yurov YB: Somatic genome variations in health and disease. Curr Genomics 2010, 11: 387–396. 10.2174/138920210793176065
Vorsanova SG, Yurov YB, Soloviev IV, Iourov IY: Molecular cytogenetic diagnosis and somatic genome variations. Curr Genomics 2010, 11: 440–446. 10.2174/138920210793176010
Vorsanova S, Yurov IY, Demidova I, Voinova-Ulas VY, Kravets V, Solov’ev I, Gorbachevskaya NL, Yurov YB: Variability in the heterochromatin regions of the chromosomes and chromosomal anomalies in children with autism: identification of genetic markers of autistic spectrum disorders. Neurosci Behav Physiol 2007, 37: 553–558. 10.1007/s11055-007-0052-1
Yurov YB, Vorsanova SG, Iourov IY, Demidova IA, Beresheva AK, Kravetz VS, Monakhov VV, Kolotii AD, Voinova-Ulas VY, Gorbachevskaya NL: Unexplained autism is frequently associated with low-level mosaic aneuploidy. J Med Genet 2007, 44: 521–525. 10.1136/jmg.2007.049312
Vorsanova SG, Voinova VY, Yurov IY, Kurinnaya OS, Demidova IA, Yurov YB: Cytogenetic, molecular-cytogenetic, and clinical-genealogical studies of the mothers of children with autism: a search for familial genetic markers for autistic disorders. Neurosci Behav Physiol 2010, 40: 745–756. 10.1007/s11055-010-9321-5
Vorsanova S, Koloti D, Sharonin V, Soloviev V, Yurov Y: FISH analysis of microaberrations at telomeric and subtelomeric regions in chromosomes of children with mental retardation. Am J Hum Genet 1998, 63(suppl 4):A154-A873.
Caserta D, Benkhalifa M, Baldi M, Fiorentino F, Qumsiyeh M, Moscarini M: Genome profiling of ovarian adenocarcinomas using pangenomic BACs microarray comparative genomic hybridization. Mol Cytogenet 2008, 1: 10. 10.1186/1755-8166-1-10
Yurov YB, Soloviev IV, Vorsanova SG, Marcais B, Roizes G, Lewis R: High resolution multicolor fluorescence in situ hybridization using cyanine and fluorescein dyes: rapid chromosome identification by directly fluorescently labeled alphoid DNA probes. Hum Genet 1996, 97: 390–398. 10.1007/BF02185780
Yurov Y, Vorsanova S, Soloviev I, Demidova I, Alexandrov I, Sharonin V, Beresheva A: Original collection of DNA probes for preimplanta tional, fetal prenatal and postnatal diagnosis of chromosomal analysis by FISH. Early prenatal diagnosis, fetal cells and DNA in mother, present state and per spectives/Eds M Macek, D Bianchi, H Cuckle–Prague 2002, 275–283.
Iourov IY, Vorsanova SG, Liehr T, Kolotii AD, Yurov YB: Increased chromosome instability dramatically disrupts neural genome integrity and mediates cerebellar degeneration in the ataxia-telangiectasia brain. Hum Mol Genet 2009, 18: 2656–2669. 10.1093/hmg/ddp207
Iourov IY, Vorsanova SG, Saprina EA, Yurov YB: Identification of candidate genes of autism on the basis of molecular cytogenetic and in silico studies of the genome organization of chromosomal regions involved in unbalanced rearrangements. Russ J Genet 2010, 46: 1190–1193. 10.1134/S102279541010011X
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
IYI, SGV and YBY conceived the research, designed the study, and wrote the manuscript. SGV and YBY referred the patients for the study. IYI, OSK and APS performed the array CGH analysis. IYI and MAZ performed the bioinformatic analysis and prepared the supplemental material (Additional file 1). IYI and OSK performed molecular cytogenetic and molecular genetic analyses. All authors have read and approved the final manuscript.
Electronic supplementary material
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Iourov, I.Y., Vorsanova, S.G., Kurinnaia, O.S. et al. Molecular karyotyping by array CGH in a Russian cohort of children with intellectual disability, autism, epilepsy and congenital anomalies. Mol Cytogenet 5, 46 (2012). https://doi.org/10.1186/1755-8166-5-46
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1755-8166-5-46