Abstract
Background
Cratoxylum formosum (Jack) Dyer ssp. pruniflorum (Kurz) Gogel. (Hóng yá mù) (CF) has been used for treatment of fever, cough, and peptic ulcer. Previously, a 50% ethanol-water extract from twigs of CF was shown highly selective in cytotoxicity against cancer cells. This study aims to investigate the molecular mechanisms underlying the apoptosis-inducing effect of CF.
Methods
The cytotoxicity of CF was evaluated in the human hepatocellular carcinoma (HCC) HepG2 cell line in comparison with a non-cancerous African green monkey kidney epithelial cell line (Vero) by a neutral red assay. The apoptosis induction mechanisms were investigated through nuclear morphological changes, DNA fragmentation, mitochondrial membrane potential alterations, and caspase enzyme activities.
Results
CF selectively induced HepG2 cell death compared with non-cancerous Vero cells. A 1.5-fold higher apoptotic effect compared with melphalan was induced by 120 μg/mL of the 50% ethanol-water extract of CF. The apoptotic cell death in HepG2 cells occurred via extrinsic and intrinsic caspase-dependent pathways in dose- and time-dependent manners by significantly increasing the activities of caspase 3/7, 8, and 9, decreasing the mitochondrial membrane potential, and causing apoptotic body formation and DNA fragmentation.
Conclusions
CF extract induced a caspase-dependent apoptosis in HepG2 cells.
Similar content being viewed by others
Background
Hepatocellular carcinoma (HCC) is the fifth leading cancer worldwide and the third cause of cancer death [1] with a poor prognosis owing to the ineffectiveness of therapy [2]. The treatment options for HCC patients, such as surgical resection and liver transplantation [2], are only appropriate for the early stages of tumor development. A second primary site can be caused by a malignant clone even if the first cancer site has been diagnosed and resected [1]. Chemoprevention is considered to be another important strategy for both primary and secondary HCC prevention of HCC by phytochemicals or non-nutritive plant compounds. The development and research of new anticancer agents has decreased HCC-associated mortality [2].
Some anti-cancer agents induced apoptosis of cancer cells [3]. Apoptotic cancer cells are characterized by morphological changes, including chromatin condensation, membrane blebbing, and apoptotic body formation, which are related to DNA fragmentation observed as a ladder on gel electrophoresis [2]. Apoptosis comprises two major pathways, the intrinsic and extrinsic pathways [4]. The caspase enzymes are proteases playing an important role in the classification of apoptosis. Several studies have screened potential compounds that can trigger apoptosis via measurement of caspase activity [5–7]. Currently, anticancer drugs such as cisplatin, paclitaxel, adriamycin, and melphalan have been reported to eradicate cancer via the induction of apoptosis [5, 7–9].
Cratoxylum formosum (Jack) Dyer ssp. pruniflorum (Kurz) Gogel. (CF) (Hóng yá mù or commonly known as “Kuding Tea” in Southwest mainland China [10] and “Tiew kon” in Thailand [11]) is widely found in Southeast Asia [12] and belongs to the family of Guttiferae. Five other Cratoxylum species were found in Thailand and Southwest China, namely Cratoxylum arborescens, Cratoxylum cochinchinense, Cratoxylum maingayi, Cratoxylum sumatranum ssp. neriifolium, and Cratoxylum formosum (Jack) Dyer [13].
CF has been used in Chinese medicine for treatment of fever, cough, stomach ache, flatulence, diarrhea, food poisoning, internal bleeding, and peptic ulcer [10, 13]. It has also been used as a diuretic and for its tonic effects [10]. CF produces various secondary metabolites including xanthones, anthraquinones [10, 14], triterpenoids, flavonoids [13–16], and phenolic compounds [6]. These compounds are widely found in plants and some of them have been shown to exert cytotoxic effects toward some cancer cell lines and other biological activities such as antibacterial effects and antimicrobial effects [10].
Fractions obtained from dichloromethane crude extract of CF roots was reported to exhibit anticancer activity against MCF-7 human breast cancer cells, human cervical cancer HeLa cells, colon cancer HT-29 cells, and human oral cancer KB cells [13]. The cytotoxic effect of the CF extract was selectively high against human leukemia cancer cells (U937) compared with normal cells (Vero). Our group previously reported the apoptotic effect of CF on U937 cells, based on observations of DNA laddering and nuclear morphological changes [8]. This study aims to investigate the molecular mechanisms underlying the apoptosis-inducing effect.
Materials and methods
Reagents and chemicals
The organic solvents used for extraction were of analytical grade and obtained from Fisher Scientific (Loughborough, UK) and Labscan (Bangkok, Thailand). The reagents used in the cell assays were of molecular biological grade. Dulbecco’s modified Eagle’s medium (DMEM), fetal bovine serum (FBS), penicillin, and streptomycin were purchased from GIBCO® (Invitrogen Corporation, Carlsbad, CA, USA). Standard reagents, melphalan, and 4′,6-diamidino-2-phenylindole (DAPI) were obtained from Sigma-Aldrich Inc. (St. Louis, MO, USA). Dimethylsulfoxide (DMSO) was purchased from United States Biological (Salem, MA, USA). The reagents for cytotoxicity testing included neutral red (NR) and sodium bicarbonate (NaHCO3). We used a FlexiGene DNA Kit (Qiagen GmbH, Germany), isopropyl alcohol biotechnology grade (Bio Basic Inc., USA), and methanol analytical grade (BDH, England). Agarose molecular grade was purchased from Bio-Rad (Hercules, CA, USA). We purchased a 100 bp + 1.5 kb DNA ladder stained with loading dye from SibEnzyme Ltd. (Russia). Caspase-Glo® 3/7, 8, and 9 Assay Kits were purchased from Promega (Madison, WI, USA). We obtained 3,3′-dihexyloxacarbocyanine iodide (DiOC6) from Sigma-Aldrich Inc. Pure concentrated hydrogen peroxide (35%, v/v) was purchased from Qrec Co. Ltd. (New Zealand).
Cell culture
An African green monkey kidney cell line (Vero) at passage 26 and a human HCC cell line (HepG2) at passage 40 were maintained in DMEM supplemented with 10% FBS and 1% (w/v) penicillin/streptomycin. All cells were cultured at 37°C in a humidified atmosphere containing 5% CO2.
Plant material and extraction
Twigs of CF were collected in 2008 from the northeastern region of Thailand. The plants were visually authenticated according to taxonomy [17] and voucher specimens (TT-OC-SK-862) were deposited at the Medicinal Herbarium of the Faculty of Pharmaceutical Sciences, Khon Kaen University, Thailand. Briefly, 1 kg of dried plants was cut and macerated with 6 L of 50% ethanol and water for 7 days. The solvent was filtered, distilled in vacuo by a rotary evaporator below 40°C, and freeze-dried, yielding a 3.78% (w/w) crude extract. The crude extract was stored at 4°C in an airtight container. The stock solution of CF extract was freshly prepared and dissolved in 100% DMSO, and the concentration of DMSO in all experiments was restricted to ≤ 1% to maintain the cytotoxicity at < 10%.
Cytotoxicity assay
Cell viability was measured by an NR colorimetric assay. Human HepG2 cells and normal African green monkey kidney Vero cells were used as cell models to determine the percent cytotoxicity. Briefly, cells (4 × 106 cells/mL) were treated with CF extract at various concentrations (10, 25, 50, 100, 250, and 500 μg/mL) and the standard anticancer drug melphalan as a positive control, while the untreated groups were exposed to fresh medium at equal volumes to those used for the treated groups. After 24 h of exposure, the cells were incubated for 2 h with NR dye (50 μg/mL) dissolved in DMEM, and then washed with phosphate-buffered saline (PBS; pH 7.4). Finally, the cells were lysed with 0.33% HCl/isopropanol. A microplate reader (Tecan, France) was used to measure the absorbances of NR dye in the viable cells at 520 and 650 (reference) nm. The percentage of cell cytotoxicity of the CF extract was calculated as described by Machana et al.[18] and compared with that of melphalan. The selectivity index (SI) was calculated to determine the selective cancer cell line cytotoxicity [9]. The SI of the test compound was compared with the IC50 for the Vero cells divided by the IC50 for the HepG2 cells.
Apoptosis induction assay
DAPI staining assay
Cells (1 × 106 cells/well) were treated with CF extract and the melphalan for 24 h at 2 × IC50, and the morphological changes of the induced apoptotic cells were determined by DAPI staining. The concentrations of CF extract and melphalan used in the following apoptosis experiments were 120 and 80 μg/mL, respectively. The cells were fixed in pure methanol at 4°C for 10 min, and then stained with DAPI (1 μg/mL) at room temperature in the dark. The excess dye was removed and glycerin was added to the cells with PBS at a 1:1 ratio and mixed well. Next, the cells were fixed on a slide and the apoptotic bodies observed at 40× objective magnification under an inverted fluorescence microscope (Eclipse TS100; Nikon, UK). The percentage of apoptotic cells was calculated as described by Machana et al.[18]. The average% apoptotic cells were calculated from three independent wells with 10 eye views in each well.
DNA fragmentation assay
HepG2 cells (1 × 106 cells/well) were treated with 120 μg/mL of CF extract or 80 μg/mL of melphalan for 24 h, harvested, and washed three times with sterile PBS. Each cell suspension was transferred to a microcentrifuge tube (1.5 mL) and centrifuged (DAIHAN scientific, Korea) at 300 × g for 5 min, and the cell pellet was collected. A FlexiGene DNA kit was used to extract the DNA in accordance with the manufacturer’s instructions. Subsequently, 2 μg of DNA was analyzed by electrophoresis (Gibthai, Taiwan) in a 1.8% agarose gel containing 0.1% (v/v) ethidium bromide. The DNA fragments were analyzed by an InGenius L Gel Documentation system (Syngene, MD, USA) and an image of the electrophoresis gel was taken.
Caspase activity assay
HepG2 cells (4 × 106 cells/mL) were seeded in a 96-well plate, incubated overnight, and then treated with CF extract at 0, 30, 60, 120, and 240 μg/mL for 24 h (caspase 3/7 activity test) or 12 h (caspase 8, 9 activity test). In addition, HepG2 cells were treated with CF extract (120 μg/mL) at 0, 3, 6, 12, and 24 h for caspase 3/7 activity and 0, 3, 6, 9, and 12 h for caspase 8 and 9 activity). The enzyme activity of caspase 3/7, 8, and 9 was determined by Caspase-Glo® 3/7, 8, and 9 Assay Kits in accordance with the manufacturer’s instructions. The caspase activities were expressed as relative luminescence units (RLU) in direct proportion to the luminescent light and were detected by a microplate luminometer (SpectraMax Gemini X, Sandy, UT, USA). Blank determinations were performed on a test plate containing only sample and medium. All of the experiments were performed in triplicate.
Detection of mitochondrial membrane potential (MMP)
Cells (1 × 106 cells/mL) were seeded into a 24-well plate, incubated at 37°C for 24 h, and then treated with 0, 30, 60, or 120 μg/mL of CF extract for 12 or 24 h. Concentrated hydrogen peroxide (35%, v/v) was used as a positive control. Next, the cells were washed, harvested by trypsinization, transferred to a microcentrifuge tube, and stained with a lipophilic fluorogenic dye, DiOC6, at 40 nM for 30 min in the dark at 37°C. After the incubation, the cells were immediately analyzed by flow cytometry (BD FACSCanto II, San Jose, CA, USA) for incorporation of the fluorochrome.
Determination of phenolics in the CF extract
The total phenolic content was determined by the Folin-Ciocalteu reagent as described by Kaisoon et al.[19]. Briefly, 300 μL of CF extract was mixed with 2.25 mL of Folin-Ciocalteu reagent (10-fold dilution with distilled water). After standing at room temperature for 5 min, 2.25 mL of saturated NaHCO3 (60 g/L) solution was added to the mixture. After 90 min, the absorbance of the reaction mixture was measured at 725 nm by a UV-spectrophotometer (Shimadzu, Japan). The results were expressed as milligrams of gallic acid equivalents per 1 g of dried sample (mg GAE/g dry weight).
Statistical analysis
The data were represented by the mean ± standard deviation (SD) (n = 3). The statistical significance of differences for multiple-group comparisons was evaluated by one-way ANOVA followed by a Tukey honestly significant difference using SPSS Windows version 11.5 (SPSS, USA). Evaluation with a two-tailed Student’s t-test was also performed for two-group comparisons. Values of P < 0.05 were considered statistically significant.
Results
CF extract induced human HCC cell death
The cytotoxicity (Table 1) and growth inhibition (Figure 1) results of the crude CF extract compared to melphalan showed cytotoxicity with a 50% inhibitory concentration (IC50 ± SD) of 55.9 ± 10.6 μg/mL for HepG2 cells while no effect on normal Vero cells at concentrations up to 500 μg/mL. Melphalan exhibited an IC50 of 37.7 ± 9.8 μg/mL for HepG2 cells, but showed low selectivity (SI = 1.6). The CF extract showed high selectivity (SI = 8.9) against HepG2 cells with a minimum effect on normal Vero cells. The CF extract possessed potential cytotoxic activity toward the HCC cell line in vitro.
CF extract induced apoptosis
CF caused nuclear morphological changes in HepG2 cells
Images of nuclear DNA staining by DAPI in untreated (control) and treated HepG2 cells were shown in Figure 2. Nuclei with condensed chromatin and apoptotic bodies, characteristic of late-stage apoptosis [20], were observed in HepG2 cells incubated with the CF extract and the melphalan. The results revealed that the CF extract caused a higher percentage of apoptosis (62.2 ± 4.5%) as compared to melphalan alone (41.6 ± 2.1%). Apoptotic bodies were not observed in the untreated HepG2 cells.
Nuclei morphological changes and percentage of apoptotic cells in HepG2 cells (A) after treatment with 80 μg/mL of melphalan, (B) 120 μg/mL of CF, and (C) in the control or untreated HepG2 cells. Apoptotic bodies are indicated by white arrows. Pictures captured using an inverted fluorescent microscope at 40 × objective magnification.
CF induced apoptotic DNA fragmentation in HepG2 cells
Apoptotic cell death leads to regulated nuclear fragmentation by endonuclease enzymes in a controlled manner that involves DNA fragmentation [20]. During the apoptotic process, the release of endonuclease enzymes associated with DNA fragmentation in apoptotic cells results in DNA laddering, which can be observed by agarose gel electrophoresis [21]. This phenomenon specifically indicates that treated cancer cells have undergone late-stage apoptosis [22]. Figure 3 shows that treatment of HepG2 cells with 120 μg/mL of CF extract led to DNA laddering after 24 h of exposure, and were similar to those for cells treated with melphalan alone.
Apoptosis induction mechanisms of the CF extract
CF induces apoptosis in HepG2 cells via activation of caspase activity
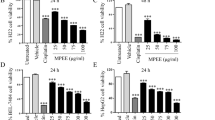
Apoptotic cell death occurs via major extrinsic and intrinsic pathways. Caspases 3/7, 8, and 9 are the major apoptotic caspases. Activation of caspase 8 indicates the extrinsic pathway, while activation of caspase 9 indicates the intrinsic pathway. When cells undergo apoptosis through either the extrinsic or intrinsic pathway, caspase 3 activity is activated at the final step prior to cell death [5–7]. The apoptosis induced in HepG2 cells by the CF extract through caspase activity was initially observed at 30 μg/mL after 6 h of incubation (Figure 4). The CF extract increased caspases activity with dose and time dependent manner. At 24 h the CF extract at all concentration tested significantly increased caspase 3/7 from the untreated cells (P < 0.001, one-way ANOVA). The cells treated with 30, 60, 120, and 240 μg/mL significantly increased caspase 8 activity (P = 0.018, P = 0.029, P < 0.001, P < 0.001, respectively, one-way ANOVA). However, no significant difference of caspase 9 activity was observed (one-way ANOVA) between the group of cells treated with 30 and 60 μg/mL and the untreated cells (P = 0.150 and P = 0.100). Whereas the group of cells treated with 120, and 240 μg/mL significantly increased caspase 9 activity compared to the untreated cells (P < 0.001, one-way ANOVA).
Activation of caspase-3/7, -8 and -9 in the HepG2 treated with the CF extract. Relative luminescent units (RLU) of (A) caspase 3/7, (B) 8 and (C) 9, respectively, after HepG2 cells exposed to CF extract at various concentrations for 12 (for caspase 8, 9) and 24 (for caspase 3) h. (D), (E), and (F) represent the caspase 3/7, 8, 9 activity after being treated with CF extract (120 μg/mL) at various points. Values represent a mean ± SD, based on triplicates. *P < 0.05, **P < 0.001 significantly different from untreated cells.
At 3 h the CF extract (120 μg/mL) did not significantly increase caspase 3/7 (P = 0.885), caspase 8 (P = 1.000), and caspase 9 activity (P = 1.000) when compared to the control (one-way ANOVA). The CF extract (120 μg/mL) significantly increased caspase 3/7 at 6, 12, and 24 h (P < 0.001, one-way ANOVA). The CF extract at the same concentration did not significantly increased caspase 8 at 6 and 9 h (P = 0.597, and P = 0.07) but significantly increase caspase 8 activity at 12 h (P < 0.001, one-way ANOVA). Moreover, the CF extract at the same concentration did not significantly increased caspase 9 at 6 and 9 h (P = 0.568, and P = 0.079) but significantly increase caspase 9 activity at 12 h (P < 0.001, one-way ANOVA).
CF induced apoptosis in HepG2 cells through the mitochondrial pathway
MMP changes after cells were exposed to the CF extract at various concentrations were observed. The alteration of MMP in the cells treated with 30 μg/mL CF extract compared to untreated cells was not significantly different at 12 and 24 h (P = 0.994 and P = 0.150, respectively). The MMP alteration in the cells treated with 60 and 120 μg/mL CF extract and 35% (v/v) H2O2 were significantly different from the untreated at 12 and 24 h (P < 0.001, one-way ANOVA) (Figure 5).
Total phenolic content of the CF extract
The total phenolic content was estimated by a Folin-Ciocalteu colorimetric method by gallic acid as the standard phenolic compound [19]. The calibration curve of gallic acid showed a linear correlation for gallic acid concentrations between 25–200 μg/mL, with an r2 of 0.997. The total phenolic content was 5.36 mg GAE/g dry weight of CF extract.
Discussion
The current treatment of HCC includes 5-fluorouracil [23], doxorubicin, mitomycin C [24–26], cisplatin [25], and melphalan [27, 28]. The 50% ethanol-water extract of CF inhibited the proliferation rate of HCC cells with a higher SI than melphalan. Based on published guidelines, any extract that possesses potentially toxic side effects must have an IC50 of less than 100 μg/mL [9]. Our laboratory previously found that CF induced apoptosis in leukemia cancer U937 cells [8] and this study aims to investigate the anticancer activity of 50% ethanol-water extract of CF.
Apoptosis was observed in the action mechanism of anti-cancer drugs [5–8]. The failure to trigger apoptosis was related to drug resistance and tumorigenesis [4]. Apoptosis includes both intrinsic and extrinsic pathways [29], and can be observed by investigating DNA condensation, membrane blebbing, and apoptotic body formation by staining cells with DAPI or Hoechst 3332 dye and detecting morphological changes under an inverted fluorescence microscope [29–31]. DNA fragmentation has also been investigated by 2D gel electrophoresis [30]. However, DNA fragmentation can also occur under non-apoptotic conditions [22].
Both apoptosis pathways are related to caspase activity, specifically caspase 3, 8, and 9 activity [29, 32, 33], that can be used to classify the extrinsic and intrinsic pathways [31]. The intrinsic or mitochondrial pathway is caused by nuclear DNA damage, resulting in alterations in the MMP, and release of cytochrome c into the cytosol. Ultimately, caspase 9 is also activated [31]. In the present study, the DNA condensation (Figure 2), MMP changes (Figure 5), and caspase 9 activation (Figure 4C and F) were observed after exposure of HepG2 cells to the CF extract.
The caspase 8 is the key caspase for the extrinsic pathway of apoptosis [34]. This extrinsic pathway of apoptosis occurred extrinsically from interactions of a death receptor with death ligands. In this study, the caspase 8 activity in HepG2 cell was increased after exposure to CF extract (Figure 4B and E).
Activation of caspase 8 and 9 also activates the executioner caspase 3/7, which can cleave several intracellular proteins and leads to cell disruption, including apoptotic body formation, chromosome breakage, and DNA fragmentation [30]. In the present study, CF induced the late stages of apoptosis characterized by an increase in caspase 3/7 activity (Figure 4A and D), as evidenced by apoptotic body formation (Figure 2) and DNA fragmentation (Figure 3). The induction of apoptosis by CF was clearly demonstrated by caspase activity in both the extrinsic and intrinsic pathways. Further studies of the crude CF extract effects on cancer cell lines are necessary to isolate the phytoconstituents contributing to the anticancer activity.
Previously, benzophenone glycosides that inhibited leukemia L1210 viability [35] were identified in a 60% ethanol extract from CF stems [10, 36]. Xanthones, anthraquinones, and cochinchinone were identified in dichloromethane extracts of CF roots and bark [13, 37], and quercetin, xanthones, and mangiferin were found in CF leaves and stem [38]. The bioactive compounds possessing anticancer activity in CF were cochinchinone A [39, 40], macluraxanthon [41], norcowanin [42, 43], and brasilixanthone [44]. Other activities were also reported for the dichloromethane extract, in which dulcisxanthone B had a DPPH-scavenging effect and inhibited lipid peroxidation [45].
The total phenolic content in the 50% ethanol-water extract of CF was 5.36 mg GAE/g dry weight of material. A previous report defined a high phenolic content as 20 mg GAE/g dry weight [46], based on the Folin-Ciocalteu method. Thus, the total content of the CF extract found in the present study would be considered low.
Conclusions
CF extract induced a caspase-dependent apoptosis in HepG2 cell.
Abbreviations
- CF:
-
Cratoxylum formosum (Jack) Dyer ssp. pruniflorum (Kurz) Gogel.
- DAPI:
-
4′,6-diamidino-2-phenylindole
- DiOC6:
-
3,3′-dihexyloxacarbocyanine iodide
- DMEM:
-
Dulbecco’s modified Eagle’s medium
- DMSO:
-
Dimethyl sulfoxide
- FBS:
-
Fetal bovine serum
- GC-MS:
-
Gas chromatography/mass spectrometry
- HCC:
-
Hepatocellular carcinoma
- MMP:
-
Mitochondrial membrane potential
- NR:
-
Reutral red
- PBS:
-
Phosphate-buffered saline
- SI:
-
Selectivity index
- UV:
-
Ultraviolet
- RLU:
-
Relative luminescence unit.
References
Stagos D, Amoutzias GD, Matakos A, Spyrou A, Tsatsakis AM, Kouretas D: Chemoprevention of liver cancer by plant polyphenols. Food Chem Toxicol. 2012, 50: 2155-2170. 10.1016/j.fct.2012.04.002.
Yeh CT, Yen GC: Induction of apoptosis by the Anthocyanidins through regulation of Bcl-2 gene and activation of c-Jun N-terminal kinase cascade in hepatoma cells. J Agric Food Chem. 2005, 53: 1740-1749. 10.1021/jf048955e.
Edinger AL, Thompson CB: Death by design: apoptosis, necrosis and autophagy. Curr Opin Cell Biol. 2004, 16: 663-669. 10.1016/j.ceb.2004.09.011.
Liu M-J, Wang Z, Li H-X, Wu R-C, Liu Y-Z, Wu Q-Y: Mitochondrial dysfunction as an early event in the process of apoptosis induced by woodfordin I in human leukemia K562 cells. Toxicol Appl Pharmacol. 2004, 194: 141-155. 10.1016/j.taap.2003.08.017.
Henkels KM, Turchi JJ: Cisplatin-induced apoptosis proceeds by caspase-3-dependent and -independent pathways in cisplatin-resistant and-sensitive human ovarian cancer cell lines. Cancer Res. 1999, 59: 3077-3083.
Koceva-Chyła A, Jedrzejczak M, Skierski J, Kania K, Jóźwiak Z: Mechanisms of induction of apoptosis by anthraquinone anticancer drugs aclarubicin and mitoxantrone in comparison with doxorubicin: relation to drug cytotoxicity and caspase-3 activation. Apoptosis. 2005, 10: 1497-1514. 10.1007/s10495-005-1540-9.
Mielgo A, Torres VA, Clair K, Barbero S, Stupack DG: Paclitaxel promotes a caspase 8-mediated apoptosis through death effector domain association with microtubules. Oncogene. 2009, 28: 3551-3562. 10.1038/onc.2009.210.
Machana S, Weerapreeyakul N, Barusrux S, Thumanu K, Tanthanuch W: FTIR microspectroscopy discriminates anticancer action on human leukemic cells by extracts of Pinus kesiya; Cratoxylum formosum ssp. pruniflorum and melphalan. Talanta. 2012, 93: 371-382.
Prayong P, Barusrux S, Weerapreeyakul N: Cytotoxic activity screening of some indigenous Thai plants. Fitoterapia. 2008, 79: 598-601. 10.1016/j.fitote.2008.06.007.
Duan YH, Dai Y, Wang GH, Zhang X, Chen HF, Chen JB, Yao X-S, Zhang X-K: Bioactive xanthones from the stems of Cratoxylum formosum ssp. pruniflorum. J Nat Prod. 2010, 73: 1283-1287. 10.1021/np1001797.
Kuvatanasuchati J, Laphookhieo S, Rodanant P: Antimicrobial activity against periodontopathic bacteria and cytotoxic study of Cratoxylum formosum and Clausena lansium. J Med Plants Res. 2011, 5: 5988-5992.
Iinuma M, Tosa H, Ito T, Tanaka T, Madulid DA: Two xanthones from roots of Cratoxylum formosanum. Phytochemistry. 1996, 42: 1195-1198. 10.1016/0031-9422(96)00111-2.
Boonnak N, Karalai C, Chantrapromma S, Ponglimanont C, Fun H-K, Kanjana-Opas A, Laphookhieo S: Bioactive prenylated xanthones and anthraquinones from Cratoxylum formosum ssp. pruniflorum. Tetrahedron. 2006, 62: 8850-8859. 10.1016/j.tet.2006.06.003.
Sun M, Sakakibara H, Ashida H, Danno G, Kanazawa K: Cytochrome P4501A1-inhibitory action of antimutagenic anthraquinones in medicinal plants and the structure-activity relationship. Biosci Biotechnol Biochem. 2000, 64: 1373-1378. 10.1271/bbb.64.1373.
Maisuthisakul P, Pongsawatmanit R, Gordon MH: Characterization of the phytochemicals and antioxidant properties of extracts from Teaw (Cratoxylum formosum Dyer). Food Chem. 2007, 100: 1620-1629. 10.1016/j.foodchem.2005.12.044.
Akao Y, Nakagawa Y, Iinuma M, Nozawa Y: Anti-cancer effects of xanthones from pericarps of mangosteen. Int J Mol Sci. 2008, 9: 355-370. 10.3390/ijms9030355.
Weerapreeyakul N, Nonpunya A, Barusrux S, Thitimetharoch T, Sripanidkulchai B: Evaluation of the anticancer potential of six herbs against a hepatoma cell line. Chin Med. 2012, 7: 15-10.1186/1749-8546-7-15.
Machana S, Weerapreeyakul N, Barusrux S, Nonpunya A, Sripanidkulchai B, Thitimetharoch T: Cytotoxic and apoptotic effects of six herbal plants against the human hepatocarcinoma (HepG2) cell line. Chin Med. 2011, 6: 39-10.1186/1749-8546-6-39.
Kaisoon O, Siriamornpun S, Weerapreeyakul N, Meeso N: Phenolic compounds and antioxidant activities of edible flowers from Thailand. J Funct Foods. 2011, 3: 88-99. 10.1016/j.jff.2011.03.002.
Priyadarsini Vidya R, Murugan Senthil R, Maitreyi S, Ramalingam K, Karunagaran D, Nagini S: The flavonoid quercetin induces cell cycle arrest and mitochondria-mediated apoptosis in human cervical cancer (HeLa) cells through p53 induction and NF-κB inhibition. Eur J Pharmacol. 2010, 649: 84-91. 10.1016/j.ejphar.2010.09.020.
Murillo G, Salti GI, Kosmeder JW, Pezzuto JM, Mehta RG: Deguelin inhibits the growth of colon cancer cells through the induction of apoptosis and cell cycle arrest. Eur J Cancer. 2002, 38: 2446-2454. 10.1016/S0959-8049(02)00192-2.
Czene S, Testa E, Nygren J, Belyaev I, Harms-Ringdahl M: DNA fragmentation and morphological changes in apoptotic human lymphocytes. Biochem Biophys Res Commun. 2002, 294: 872-878. 10.1016/S0006-291X(02)00588-0.
Eichhorst ST, Müller M, Li-Weber M, Schulze-Bergkamen H, Angel P, Krammer PH: A novel AP-1 element in the CD95 ligand promoter is required for induction of apoptosis in hepatocellular carcinoma cells upon treatment with anticancer drugs. Mol Cell Biol. 2000, 20: 7826-7837. 10.1128/MCB.20.20.7826-7837.2000.
Izumi R, Shimizu K, Iyobe T, Ii T, Yagi M, Matsui O, Nonomura A, Miyazaki I: Postoperative adjuvant hepatic arterial infusion of lipiodol containing anticancer drugs in patients with hepatocellular carcinoma. Hepatology. 1994, 20: 295-301. 10.1002/hep.1840200205.
Roayaie S, Frischer JS, Emre SH, Fishbein TM, Sheiner PA, Sung M, Miller CM, Schwartz ME: Long-term results with multimodal adjuvant therapy and liver transplantation for the treatment of hepatocellular carcinomas larger than 5 centimeters. Ann Surg. 2002, 235: 533-539. 10.1097/00000658-200204000-00012.
Varela M, Real MI, Burrel M, Forner A, Sala M, Brunet M, Ayuso C, Castells L, Montañá X, Llovet JM, Bruix J: Chemoembolization of hepatocellular carcinoma with drug eluting beads: efficacy and doxorubicin pharmacokinetics. J Hepatol. 2007, 46: 474-481. 10.1016/j.jhep.2006.10.020.
Alexander HR, Libutti SK, Bartlett DL, Puhlmann M, Fraker DL, Bachenheimer LC: A phase I-II study of isolated hepatic perfusion using melphalan with or without tumor necrosis factor for patients with ocular melanoma metastatic to liver. Clin Cancer Res. 2000, 6: 3062-3070.
Alexander HR, Bartlett DL, Libutti SK, Pingpank JF, Fraker DL, Royal R, Steinberg SM, Helsabeck CB, Beresneva TH: Analysis of factors associated with outcome in patients undergoing isolated hepatic perfusion for unresectable liver metastases from colorectal center. Ann Surg Oncol. 2009, 16: 1852–1859-
Yang J-S, Wu C-C, Kuo C-L, Lan Y-H, Yeh C-C, Yu C-C, Lien J-C, Hsu Y-M, Kuo W-W, Wood WG, Tsuzuki M, Chung J-G: Solanum lyratum extracts induce extrinsic and intrinsic pathways of apoptosis in WEHI-3 murine leukemia cells and inhibit allograft tumor. Evid Based Complement Altern Med. 2012, 2012: ID254960-
Jiang CP, Ding H, Shi DH, Wang YR, Li EG, Wu JH: Pro-apoptotic effects of tectorigenin on human hepatocellular carcinoma HepG2 cells. World J Gastroenterol. 2012, 18: 1753-1764. 10.3748/wjg.v18.i15.1753.
Yu H-Y, Jin C-Y, Kim K-S, Lee Y-C, Park S-H, Kim G-Y, Kim W-J, Moon H-I, Choi YH, Lee J-H: Oleifolioside A mediates caspase-independent human cervical carcinoma HeLa cell apoptosis involving nuclear relocation of mitochondrial apoptogenic factors AIF and EndoG. J Agric Food Chem. 2012, 60: 5400-5406. 10.1021/jf3014475.
Kamiya H, Kanno T, Fujita Y, Gotoh A, Nakano T, Nishizaki T: Apoptosis-related gene transcription in human A549 lung cancer cells via A(3) adenosine receptor. Cell Physiol Biochem. 2012, 29: 687-696. 10.1159/000312589.
Zhou J, Kong D, Zhang X, Wang Y, Feng Z, Zhang X, Zhang L, Wang Y, Xie Y, Chen X: Myoglobin-induced apoptosis: two pathways related to endoplasmic reticulum stress. Ther Apher Dial. 2012, 16: 272-280. 10.1111/j.1744-9987.2011.01057.x.
Ashkenazi A: Targeting the extrinsic apoptosis pathway in cancer. Cytokine Growth Factor Rev. 2008, 19: 325-331. 10.1016/j.cytogfr.2008.04.001.
Winarno H, Katrin WE: Benzophenone glucoside isolated from the ethyl acetate extract of the bark of mahkota dewa [Phaleria macrocarpa (Scheff.) Boerl.] and its inhibitory activity on leukemia L1210 cell line. Indo J Chem. 2010, 9: 142-145.
Duan YH, Dai Y, Wang GH, Chen HF, Gao H, Chen JB, Yao XS, Zhang XK: Xanthone and benzophenone glycosides from the stems of Cratoxylum formosum ssp. pruniflorum. Chem Pharm Bull (Tokyo). 2011, 59: 231-234. 10.1248/cpb.59.231.
Boonnak N, Karalai C, Chantrapromma S, Ponglimanont C, Kanjana-Opas A, Chantrapromma K, Fun H-K: Quinonoids from the barks of Cratoxylum formosum subsp. pruniflorum. Can J Chem. 2007, 85: 341-345. 10.1139/v07-031.
Nguyen LHD, Harrison LJ: Triterpenoid and xanthone constituents of Cratoxylum cochinchinense. Phytochemist. 1998, 50: 471-476.
Mahabusarakam W, Rattanaburi S, Phongpaichit S, Kanjana-Opas A: Antibacterial and cytotoxic xanthones from Cratoxylum cochinchinense. Phytochem Lett. 2008, 1: 211-214. 10.1016/j.phytol.2008.09.012.
Ren Y, Matthew S, Lantvit DD, Ninh TN, Chai H, Fuchs JR, Soejarto DD, de Blanco EJ C, Swanson SM, Kinghorn AD: Cytotoxic and NF-κB inhibitory constituents of the stems of Cratoxylum cochinchinense and their semisynthetic analogues. J Nat Prod. 2011, 74: 1117-1125. 10.1021/np200051j.
Lee BW, Gal SW, Park K-M, Park KH: Cytotoxic xanthones from Cudrania tricuspidata. J Nat Prod. 2005, 68: 456-458. 10.1021/np030481a.
Azebaze AG, Meyer M, Valentin A, Nguemfo EL, Fomum ZT, Nkengfack AE: Prenylated xanthone derivatives with antiplasmodial activity from Allanblackia monticola STANER L.C. Chem Pharm Bull (Tokyo). 2006, 54: 111-113. 10.1248/cpb.54.111.
Menasria F, Azebaze AGB, Billard C, Faussat AM, Nkengfack AE, Meyer M, Kolb JP: Apoptotic effects on B-cell chronic lymphocytic leukemia (B-CLL) cells of heterocyclic compounds isolated from Guttiferaes. Leukemia Res. 2008, 32: 1914–1926-
Li W, Ding Y, Quang TH, Ngan NTT, Sun YN, Yan XT, Yang SY, Choi CW, Lee EJ, Paek KY, Kim YH: NF-κB inhibition and PPAR activation by phenolic compounds from Hypericum perforatum L. adventitious root. Bull Korean Chem Soc. 2013, 34: 1407-1413. 10.5012/bkcs.2013.34.5.1407.
Phuwapraisirisan P, Udomchotphruet S, Surapinit S, Tip-Pyang S: Antioxidant xanthones from Cratoxylum cochinchinense. Nat Prod Res. 2006, 20: 1332-1337. 10.1080/14786410601102033.
Tawaha K, Alali FQ, Gharaibeh M, Mohammad M, El-Elimat T: Antioxidant activity and total phenolic content of selected Jordanian plant species. Food Chem. 2007, 104: 1372-1378. 10.1016/j.foodchem.2007.01.064.
Acknowledgements
AN was supported by the Research Fund for Supporting Lecturer to Admit High Potential Student to Study and Research on His Expert Program Year 2010, Khon Kaen University (KKU) and this research project was partially supported by the Plant Genetics Conservation Project under the Royal Initiation of Her Royal Highness Princess Maha Chakri Sirindhorn, KKU (550109). The authors would like to thank Mr. Bryan Roderick Hamman and Mrs. Janice Loewen-Hamman for assistance with the English-language presentation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no conflict of interests.
Authors’ contributions
NW conceived and designed the study. AN performed the experiments and statistical analysis. NW and AN wrote the manuscript. SB reviewed the literature, revised the manuscript and coordinated the study. All the authors read and approved the final version of the manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Nonpunya, A., Weerapreeyakul, N. & Barusrux, S. Cratoxylum formosum (Jack) Dyer ssp. pruniflorum (Kurz) Gogel. (Hóng yá mù) extract induces apoptosis in human hepatocellular carcinoma HepG2 cells through caspase-dependent pathways. Chin Med 9, 12 (2014). https://doi.org/10.1186/1749-8546-9-12
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1749-8546-9-12