Abstract
Background
One facet of the complexity underlying the biology of HIV-1 resides not only in its limited number of viral proteins, but in the extensive repertoire of cellular proteins they interact with and their higher-order assembly. HIV-1 encodes the regulatory protein Tat (86–101aa), which is essential for HIV-1 replication and primarily orchestrates HIV-1 provirus transcriptional regulation. Previous studies have demonstrated that Tat function is highly dependent on specific interactions with a range of cellular proteins. However they can only partially account for the intricate molecular mechanisms underlying the dynamics of proviral gene expression. To obtain a comprehensive nuclear interaction map of Tat in T-cells, we have designed a proteomic strategy based on affinity chromatography coupled with mass spectrometry.
Results
Our approach resulted in the identification of a total of 183 candidates as Tat nuclear partners, 90% of which have not been previously characterised. Subsequently we applied in silico analysis, to validate and characterise our dataset which revealed that the Tat nuclear interactome exhibits unique signature(s). First, motif composition analysis highlighted that our dataset is enriched for domains mediating protein, RNA and DNA interactions, and helicase and ATPase activities. Secondly, functional classification and network reconstruction clearly depicted Tat as a polyvalent protein adaptor and positioned Tat at the nexus of a densely interconnected interaction network involved in a range of biological processes which included gene expression regulation, RNA biogenesis, chromatin structure, chromosome organisation, DNA replication and nuclear architecture.
Conclusion
We have completed the in vitro Tat nuclear interactome and have highlighted its modular network properties and particularly those involved in the coordination of gene expression by Tat. Ultimately, the highly specialised set of molecular interactions identified will provide a framework to further advance our understanding of the mechanisms of HIV-1 proviral gene silencing and activation.
Similar content being viewed by others
Background
HIV-1 encodes the nuclear regulatory protein Tat, which is essential for HIV-1 replication and which primarily orchestrates HIV-1 provirus transcriptional regulation. Tat transactivation from the viral promoter (LTR), is highly dependent on complex interactions between Tat, the short leader RNA present in the 5' region of all nascent HIV-1 transcripts, TAR (Trans-activation responsive element), and a number of host cellular proteins [1–4]. The molecular mechanisms whereby HIV-1 gene expression is regulated by Tat occurs at distinct levels. Initially, Tat enhances transcription initiation by promoting the assembly of the RNA polII complex by interacting with various transcription factors [2]. Subsequently, Tat activates elongation via two independent mechanisms: firstly, it enhances the processivity of RNA polII by interacting with elongation factors such as pTEF-b, which phosphorylates RNA polII C-terminal domain, and secondly, by recruiting histone acetyltransferase proteins which modify the chromatin template such as p300/CBP (CREB binding protein) and p300/CBP-associated factor (PCAF) and, as recently described, by interacting with BRM and BRG1, two chromatin remodellers[5–10]. Although the recruitment of these specific cellular factors by Tat to the HIV-1 LTR are crucial for Tat function, they only partially account for the intricate molecular mechanisms underlying the dynamics of proviral gene expression. Furthermore, Tat can be secreted by infected cells and extracellular Tat can exert autocrine or paracrine activities via interactions with cell surface receptors including integrins, CXCR4, CD26, HSPG and LRP[11].
While Tat is a small and compact protein, composed of only 86 or 101 amino acids, sequence and functional analysis reveals that Tat sequence encompasses a unique arrangement of five distinct and contiguous regions including the acidic, cysteine-rich, core, basic and glutamine-rich regions. Furthermore, Tat is subject to post-translational modifications, such as acetylation, methylation, phosphorylation and ubiquitination, thus increasing both the number and diversity of potential interfaces between Tat and cellular proteins [12–14]. Recently, a structural study employing nuclear magnetic resonance (NMR) spectroscopy has described Tat as a "natively unfolded" protein with fast dynamics lacking a well-structured three-dimensional fold. These characteristics would provide Tat the flexibility to interact with numerous cellular partners. Collectively these findings suggest that Tat is a potent, versatile protein suited for multiple interactions and highlights the concept that numerous protein-protein interactions underlie the molecular mechanisms of HIV-1 molecular pathogenesis [15–19].
In this report, we have attempted to further investigate the interplay of Tat with host cell proteins. Specifically, we have designed a proteomic strategy based on affinity chromatography (AC) coupled with mass spectrometry (MS) to purify Tat interacting proteins from T-cell nuclear extracts (Figure 1). Our approach has produced the in vitro Tat nuclear interactome, which includes a total of 183 individual nuclear components, most of which have not been previously identified as Tat partners. We subsequently applied in silico analysis, to validate our dataset and develop HIV-1 Tat interaction network maps. In this report, we have focused on the description of multi-protein complexes involved in gene expression regulation, which comprised the majority of our dataset and which clearly reflects Tat primary function.
Results
Experimental Design
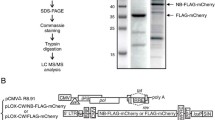
To identify multi-protein complexes associated with HIV-1 Tat, we employed the experimental strategy depicted in Figure 1. Our priority was to ensure a highly sensitive and specific methodology to identify both transient interactions and low-abundance proteins associated with complexes, while ensuring potential contaminants (false positives) remained as low as possible. In this study, we focused on nuclear protein interactions as Tat has been shown to primarily localise in the nucleus. Protein complexes were identified using in vitro "pull down" purification employing equivalent amounts of immobilised recombinant GST-Tat (bait) and GST (negative control) proteins, and incubation with T-cell nuclear extracts. Following extensive washes, captured protein complexes were eluted under denaturing conditions (Laemmli buffer) and resolved using a 1D SDS-PAGE gel. For protein identification, GST and GST-Tat interaction profiles within the entire separation range of each SDS-PAGE gel lane were systematically sliced into 2 mm gel pieces and subjected to in-gel tryptic digestion. Peptide mixtures were separated by liquid chromatography (LC) prior to tandem mass spectrometry analysis (MS/MS). The identity of selected proteins was validated by Western Blot (WB) analysis.
Tat Interaction Profile
Jurkat T-cell nuclear extracts were prepared as described in Materials and Methods and subjected to affinity chromatography (AC) with GST or GST-Tat. GST, GST-Tat and their respective associated proteins were eluted and separated by SDS-PAGE (Figure 2). A total of 164 gel slices from the GST and GST-Tat lanes were processed and the resulting tryptic peptides were analysed by LC-MS/MS. We successfully identified over 250 proteins with sizes ranging from 25 kDa to 400 kDa, which did not interact with GST alone. Proteins were identified with a minimum of two individual peptides (see Table 1 and Additional file 1). In effect, we obtained a moderate to high amino acid sequence coverage by matching tryptic peptides, ranging from 2.5% (MLL) up to 71% (prohibitin), which was inversely correlated with increasing protein size.
Interaction profile of Tat associated proteins, isolated from Jurkat nuclear fractions. T-cell nuclear extracts were incubated with immobilised GST (control) and GST-Tat (Bait). Specifically interacting proteins were subsequently eluted and resolved by SDS-PAGE and stained with Coomassie Blue. The resulting Tat interaction profile is specific and composed of bands of distinct size and intensity, representing putative proteins interacting with Tat. The purified recombinant proteins GST and GST-Tat are indicated by an arrowhead.
Dataset Curation Process
To eliminate potential contaminants from the dataset, we modified our preliminary dataset by retaining proteins known to exist in the nucleus and excluding non-nuclear components such as those associated with mitochondria, cytoskeletal proteins, and common contaminants such as keratin, and ribosomal and histone proteins. The resulting dataset contained 183 candidate proteins that could interact directly or indirectly with HIV-1 Tat. Remarkably, 10% of the selected proteins have been previously identified by other studies, demonstrating the effectiveness and robustness of our approach (Table 1)[5, 6, 20–38]. The remainder, not previously described, highlighted the potential of our approach to identify new interactions. We subsequently confirmed the identity of 11 proteins identified as new Tat interactors by Western-Blotting analysis (Figure 3). Furthermore, these interactions appear to be robust since some of them, like SIN3A or HDAC1 could tolerate washes containing up to 1 M NaCl (Figure 3A). The specificity of these interactions was further confirmed, employing a Tat-NLS deletion mutant, which still bound with 5 of them (SIN3A, HDAC1, SAP18, Ikaros and SPT16) (Figure 3B). Of note however, our study did not identify certain proteins known to interact with Tat, including cyclin T1, TIP60, P/CAF or BRM[6, 8, 9, 39–41]. However, when we performed GST pull-down with nuclear extract followed by Western-Blot (WB), we could specifically detect the presence of Cyclin T1 in the Tat-eluted fraction, confirming that our recombinant GST-Tat is competent to interact with this well characterised nuclear partner (Figure 3A).
Validation of the identity of selected proteins interacting with Tat. A. GST pull-downs were performed with immobilised GST or GST-Tat and Jurkat cell nuclear extracts (150 μg) followed by washes with increasing salt (NaCl) concentration (0.3 M, 0.5 M, 0.8 M and 1 M). Eluates were analysed by WB using the indicated antibodies. B. GST pull-downs were performed with immobilised GST or GST-Tat-NLS and Jurkat cell nuclear extract (150 μg) followed by washes with 300 mM NaCl. Expression levels of each endogenous protein are provided with the Input corresponding to 2 μg of nuclear extracts.
Dataset Validation Process
In the initial analysis of our dataset, we analysed the domain composition of each protein using CDD, Pfam and Smart databases[42–47]. The 10 most prevalent domains found within the entire dataset are listed in Table 2, where their frequency was compared against their expected frequency derived from the Nuclear Protein Database (NPD) Protein Domains[48, 49]. Interestingly, the dataset is highly enriched for interaction domains of RNA (RRM, DEXDc, DSRM) and DNA (HMG and SANT) recognition motifs [50–53]. Other enriched interaction domains are well known to be involved in mediating protein-protein interactions and include the PHD, WD40, RRM or Bromodomain motifs [51, 54–56]. Finally, the only enriched domains associated with a catalytic activity were the AAA domain, which is associated with diverse ATP-dependent functions and the DEXDc and HELICc domains, which have a helicase activity[50, 57]. Several of these have been previously shown to mediate interactions of cellular proteins with Tat. The protein DICER has been shown to interact with Tat via its DEXDc domain, in an RNA-dependent manner[58]. The WD40 domain of LIS1 also interacts with Tat, and the Bromodomain of P/CAF specifically recognises the acetyl-lysine (K50) in Tat[9, 59, 60].
When examined individually, these domains are versatile, and occur in a wide variety of proteins. However, collectively, they are frequently found in individual proteins or large complexes associated with key functions in gene expression regulation, and more specifically at the level of chromatin remodelling (HMG, PHD, BROMO, AAA, DEXDc, HELICc, SANT, WD40), gene transcription (RRM, HMG, DSRM, WD40), RNA processing (AAA, RRM, DEXDc, HELICc DSRM) and DNA replication/chromosome structure (AAA, SANT)[50–57, 61–67].
Overall, the protein domain analysis exhibited two distinct features: (i) our dataset appears to be specifically tailored to interact with molecules such as RNA, DNA and proteins; (ii) our dataset is highly specialised in gene expression regulation and DNA replication.
To examine functional composition, systematic gene annotation employing the online tool (G.O.) was carried out, and the entire dataset was organised according to the protein involvement in specific biological processes[68, 69]. This resulted in the distribution of the proteins over 8 categories, ranging from transcription to DNA replication (Figure 4). Hence, Tat interacts with specific cellular components associated with a range of distinct activities, which may account for the marked pleiotropic activities of the protein. The best represented biological processes include transcription, RNA processing and translation, which collectively accounted for 64% of our dataset. Other major biological processes include cell cycle (13%) and nucleus organisation (8%).
HIV-1 Tat Interaction Map
Construction and mapping of the Tat interaction network
While informative, linear analysis of the Tat interaction dataset is inadequate to fully appreciate the higher order organisation of the Tat interactome. As such, we constructed a network representation of the Tat interactome and subjected our dataset to in silico interaction analysis and employed Osprey as a visualisation tool[70]. We employed established PPI databases such as BIND and HPRD, complemented by extensive literature searches, to map previously characterised interactions between the candidate proteins and developed a detailed protein interaction network[71–74]. This is depicted in Figure 5[75–233].
Tat interaction network. Here we mapped, using Osprey as a visualization tool, previously established interactions between the individual components of the Tat interaction dataset employing publicly available protein-protein interaction databases (BIND and HPRD) combined with extensive literature search. The network reconstruction of the Tat interactome revealed the higher-order and collective behaviour of the Tat interacting proteins, which compose large but well defined biochemical entities, represented by coloured circles. Edges represent interactions and individual proteins are depicted as nodes. Names in bold and red indicate the previously known Tat interactors and names in bold and blue represent proteins which identity was validated by WB analysis.
Global network characteristics
Within the interaction map we included interactions involving a minimum of two proteins. Non-interacting and self-interacting proteins were excluded to simplify the network representation. The resulting network consists of 129 proteins linked by 299 interactions, with an average of 2.31 interactions per protein. Some proteins were highly connected, such as HDAC-1 and -2, which display a total of 23 and 23 interactions, respectively. A striking result of this mapping process was the identification of groups of proteins, which formed distinct and well-connected sub-networks corresponding to previously characterised multi-protein complexes (see below). These well-defined clusters are involved in complementary, consecutive and/or opposite steps of gene expression regulation, epigenetic control, chromosome and nuclear architecture. They include transcriptional repressors, such as SIN3/HDAC, NuRD, PRC2, MeCP1, and activators including SET, FACT, BAF53, SWI2/SNF2 and WICH; chromosome organisation factors including condensin, cohesin, toposome, minichromosome maintenance (MCM), and origin of replication complex (ORC); and nuclear structure including lamina, NPC and transport factors. Interestingly, the complexes could also be shown to be interconnected, which is a reflection of the fact that multiple proteins are shared between distinct complexes, as exemplified by RbAp46 and RbAp48, subunits of the SIN3/HDAC, NuRD and PRC2 complexes. Alternatively, other complexes (such as FACT) remain isolated with protein interactions solely restricted to the members of that complex.
Functional modules
Subdivision of the Tat interaction network into functional modules enabled us to gain insights into the functional properties of the multi-protein assembly. The depicted multi-subunit complexes are (i) chromatin modifying factors, which play central roles in the alteration of the structure and composition of chromatin, and are associated with the activation or repression of gene expression; (ii) chromosome organisation factors implicated in mitosis and DNA replication and (iii) nuclear structure components which participate in the nuclear architecture.
Activators of transcription
SET1
The Set1 histone methyltransferase complex includes, Setd1A, Ash2, CXXC1, RBBP5, WDR5 and Wdr83[234, 235]. The SET1 complex mediates the methylation of Lys4 in histone H3, which ultimately results in the activation of transcription. Three components (Setd1A, CXXC1, WDR5) of this complex were identified in the co-eluate by MS/MS and the presence of WDR5 was further confirmed by GST pull-down followed by WB analysis (Figure 3).
FACT
The heterodimeric FACT complex (SPT16-SSRP1) has been characterised as an elongation factor, which enables RNA polymerase II to progress through the chromatin template, once transcription has been initiated[236]. FACT acts as a histone chaperone and mediates the disassembly and reassembly of H2A/H2B dimers. Both SPT16 and SSRP1 were identified by AC-MS/MS. We further confirmed the presence of SPT16 in the co-eluate by WB (Figure 3).
BAF53, TRRAP/p400
TRRAP/p400 is a chromatin remodeling complex, part of the INO80 family, characterised by a unique subunit composition and the presence of a distinct ATPase[237]. The core of the p400/TRRAP complex, consists of BAF53A, P400, RUVBL1, RUVBL2, TRRAP. These components were identified by our AC-MS/MS approach, and the presence of BAF53A was further validated by WB analysis of GST-pull down products (Figure 3). Additional subunits, including YEATS4, DMAP1 and Eaf6, known to be part of the p400/TRRAP complex, were also identified by our approach. Intriguingly, the TIP60 protein, which has been described in distinct protein complexes harboring p400, BAF53A and TRRAP and is a well characterised interaction partner of Tat, was not detected in the co-eluate[40].
SWI2/SNF2
SWI2/SNF2 is another chromatin remodeling complex, part of the SWI/SNF family[238]. Here, we have identified most of the components of BAF (BRG1/BRM, BAF250, BAF170, BAF155, BAF60a, BAF53A, actin and InI) and PBAF (BRG1, BAF180, BAF170, BAF155, BAF60a, BAF53A, actin and InI) complexes except BRM, BAF155 and BAF57. Importantly, BRM, BRG1, InI1 and BAF170 were previously shown to interact with Tat[5, 6, 22].
WICH
The WICH complex, composed of WSTF and SNF2H, is a member of the ISWI-containing chromatin remodeling complexes[239]. In addition to its role in replication, it has been suggested that because of its association with various transcription factors, it may have a role in transcription. Of note, both subunits were identified by AC-MS/MS and the presence of SNF2H in the co-eluate following GST pull-down was confirmed by WB (Figure 3).
Repressors of transcription
SIN3/HDAC
SIN3/HDAC is composed of SIN3A, SAP30, SAP18, HDAC-1 AND -2 and RbAp46/48 and remarkably, all of these proteins except SAP30 were recovered and identified by our approach. Additionally, the presence of HDAC-1, RbAp46/48, SAP18 and SIN3A in the co-eluate following GST pull-down was confirmed by WB (Figure 3). SIN3/HDAC has been described as a global regulator of transcription; indeed, SIN3A mediates additional interactions with transcription factors and co-repressors, which direct SIN3/HDAC to specific promoters[240]. SIN3A deacetylase activity is mediated by the HDAC-1 and -2 proteins and results in transcriptional repression. Of note, while the presence of HDAC proteins, SIN3A at the level of the HIV-1 LTR has been previously demonstrated, this is the first report showing that Tat interacts with these proteins[241].
NuRD
NuRD shares with SIN3/HDAC four individual components which include HDAC-1 AND -2 and RbAp46/48 and additionally contains CDH4/Mi-2, MTA1/2, MBD3, and MINT. NuRD is recruited to target genes via DNA-binding proteins, such as Ikaros identified here by our approach and validated by WB (Figure 3)[242]. In addition to its HDAC activity, NuRD has an ATP-dependent nucleosome activity carried out by CHD4/Mi-2, a chromatin-remodelling ATPase protein, which encompasses a chromodomain of the SWI/SNF family. While we confirmed the presence of MTA1, we failed to detect NuRD in the co-eluate by WB analysis. Interestingly, the presence of MTA1 at the level of the HIV-1 LTR has been previously demonstrated[243].
MeCP1
MeCP1, also shares the HDAC-1/-2 and RbAp46/48 subunits and include the methyl-CpG-binding protein MBD2 and p66alpha identified by our approach[84, 244–247]. MeCP1 specifically recruits SIN3/HDAC or NuRD to DNA methylation sites recognised by MBD2, which represents an alternative mechanism mediating methylation-dependent transcriptional repression involving histone deacetylation and chromatin remodeling.
PRC2
RbAp46/48 are subunits of the Polycomb Repressive Complex 2 (PRC2), which also include EED, EZH1, EZH2, SUZ12. PRC2 can methylate lysine residues (K9 and K27) of histone H3[248]. This ultimately results in the repression of gene expression. Here we have identified SUZ12.
Replication and chromosome organisation factors
Condensin/Cohesin
The structural maintenance of chromosomes (SMC) proteins form the core of the cohesin and codensin complexes[249]. They are principally involved in chromosome condensation and cohesion and play an essential role into chromatid pairing and chromosome segregation during mitosis. Interestingly, recent studies have described their participation into transcriptional regulation and epigenetic processes. Here we have identified the following condensin I subunits, SMC2, SMC4, CAPG, CAPD2 and CAPH; and the following cohesin subunits: SMC3, PDS5A, and PDS5B to be part of the Tat nuclear interactome.
TOPOSOME
The toposome complex consists of the topoisomerase IIα associated with RNA helicase A (RHA), SSRP1, PRP8, hnRNP C and RHII/Gu[142]. This complex is involved in chromosome condensation and segregation and it has been suggested that topoisomerase functions in collaboration with the condensing complex[250]. We have identified topoisomerase IIα, RHA, PRP8 and RHII/Gu as part of our Tat interaction dataset.
MCM
The minichromosome maintenance (MCM) proteins are essential for DNA replication and include six members: MCM2–MCM7 which form an heterohexamer complex that binds to DNA replication origins[64, 251]. Additional complexes include MCM4/6/7 or MCM3/5. It has been suggested that these complexes play additional cellular roles such as transcriptional regulation and chromatin remodelling. Here we have identified three members of the MCM family, MCM3, MCM6 and MCM7
ORC
The origin of replication complex is essential for DNA replication initiation[252]. The binding of the ORC complex marks the origin of replication, where the MCMs proteins are subsequently recruited and uploaded to form the pre-replication complex. However, ORC localisation is not restricted to the origin of replication and has been implicated to a broader spectrum of activities such as silencing and transcriptional regulation, heterochromatin assembly, nucleosome remodeling and chromosome condensation[252].
Nuclear structure components
NPC and nuclear transport machinery
The Nuclear Pore Complex (NPC) located in the nuclear envelope (NE) is composed of the outer nuclear membrane (ONM) and the inner nuclear membrane (INM), and enables the selective transport of macromolecules in and out of the nucleus[253]. It is composed of over 30 nucleoporins. Here, we have identified several nucleoporins (Nup358, Nup205, Nup153, Nup98, Nup93, Nup88, Nup85) and NPC associated proteins including nuclear transport factors (KPNA2, KPNB1, XPO1, RANGAP1 and RAN) and microfilaments/tubule (TUBB3, TUBBA2, NUMA1 and DNCL1)
LAMINA
The nuclear lamina lines the INM and is composed of lamins and NE lamin binding proteins including Nesprin-1 alpha, MAN1, lamina-associated polypeptides-1 and 2 (LAP1, LAP2), emerin and Lamin B receptor[254, 255]. The four latter and Lamin B (LMNB) have been identified by our screen as components of the Tat nuclear interactome
These results further substantiate the concept that chromosome architecture, chromatin remodeling, epigenetic control and nuclear organisation constitute pivotal mechanisms in the regulation of HIV-1 provirus gene expression and underscore the diversity of essential biological tasks influenced by Tat interactions.
Discussion
While considerable efforts have been dedicated to characterise individual proteins or specific macromolecular complexes interacting with Tat, no comprehensive characterisation of the Tat interactome has yet been reported. To place Tat into a wider context of interacting systems and pathways, we systematically analysed protein complexes interacting with Tat within the nucleus, by performing subcellular fractionation followed by AC-MS/MS. This experimental approach is prone to introducing technical artifacts or false positives which can then bias the subsequent analysis. To reduce this, we filtered the raw list of proteins, removed potential contaminants and obtained a final dataset of 183 interaction candidates. Subsequently, we employed computational tools and in silico analysis to validate our interaction dataset and to generate a Tat-interaction network representation. This has resulted in the in vitro Tat nuclear interactome in Jurkat T-cells.
Our studies have revealed that the Tat nuclear interactome exhibits unique signature(s). The motif composition analysis underlines the enrichment for domains mediating protein, RNA and DNA interactions, which collectively are highly representated in transcription, chromatin remodelling, chromosome structure and RNA processing complexes. We also noted the enrichment of three crucial motifs (HELIC, DEXDc and AAA) associated with helicase and ATPase activities essential for RNA processing, chromatin remodeling and chromosome architecture, which constitute the basis of both DNA replication and gene expression regulation. In support of this, the functional analysis demonstrated that proteins involved in chromatin remodeling, transcription regulation and RNA processing constitute the greater part of our dataset. Finally, the network reconstruction of the Tat nuclear interactome revealed the higher-order and collective behaviour of the Tat-interacting proteins, which compose large but well defined biochemical entities, involved in critical pathways mediating gene expression regulation, chromosome/chromatin structure, and nuclear architecture. Taken together, the remarkable enrichment for essential proteins together with their corresponding macromolecular complexes, and their roles in both activation and repression of gene expression indicate that the described Tat-interactome might act as a modular switch committed to control HIV-1 gene expression.
The presence of numerous, previously identified Tat-interacting partners further validates our dataset. Conversely, critical Tat cellular partners previously identified were not identified by our experimental approach. This could be the result of the level of endogenous expression of these proteins in Jurkat cells and perhaps technical limitations including the following: (i) loss of a fraction of the proteins during the sub-cellular fractionation step; (ii) proteins resistant to trypsin digestion; (iii) proteins not detected by the MS/MS step; (iv) absence of specific Tat post-translational modifications on our recombinant bait, GST-Tat, which is produced by a bacterial expression system. Indeed, Tat interaction with its cellular partner has been shown to be regulated by the acetylation state of its lysines 28, 50 and 51. Interaction of cyclinT1 with Tat requires the acetylation on lysine 28, while acetylation on lysine 50 prevents Tat interaction with Brm but is necessary for its interaction with BRG1 [5–7, 256, 257]. Nevertheless, despite absence of CyclinT1 from our interaction dataset, we were able to detect it by WB, further establishing our recombinant GST-Tat as a suitable bait.
Our experimental approach does not enable us to distinguish direct from indirect interactions. Furthermore, while we treated our nuclear extract with the Benzonase® nuclease (included as part of the ProteoExtract® Subcellular Proteome Extraction Kit (Calbiochem)), which has both a DNase and an RNase activities, we cannot exclude that some of the observed interactions could be mediated by residual nucleic acids.
To explore the Tat interaction network in detail, we partitioned the interacting candidates into coherent functional modules, based on their reciprocal interactions. Our results further support the recently described role of SNF proteins in Tat-dependent transcription of the provirus[5–7, 22, 258, 259]. Indeed, we have identified BRG1 and the various components of its related complex, SWI2/SNF2. We have also identified for the first time, WICH, an ISWI complex as a candidate Tat interactor.
Other complexes with histone-modifying activities include TRRAP/p400, a histone acetylase, and SET1, a histone methylase, both of which have critical positive effects on transcription. In addition, we isolated the histone chaperone FACT, directly involved in promoting the processivity of RNA polII through the chromatin template, which appears to be of relevance for Tat function in transcription elongation[236].
Importantly, we have provided the first evidence describing a direct or indirect interaction of Tat with cellular proteins, including YY1, HDAC-1/-2, and the components of the SIN3/HDAC and NuRD complexes, previously reported to interact with the integrated HIV-1 LTR. Indeed, earlier studies have identified the presence of HDAC-1 and -2 at the HIV-1 LTR in HeLa cells containing an integrated HIV-1 LTR reporter, as well in latently HIV-1 infected cell lines (ACH2, U1, J-Lat 6.3)[260–266]. They mediate histone deacetylation of nuc-1, the nucleosome positioned immediately downstream of the transcription start site, and are believed to be one of the processes mediating HIV-1 provirus transcriptional silencing throughout the establishment and/or maintenance of HIV-1 latency. In addition, two recent studies have identified the presence of SIN3A and MTA1, a component of the NuRD complex, at the integrated HIV-1 LTR in Jurkat cells, respectively recruited by CBF-1 and CTIP2[241, 243]. While various studies have shown that several transcription factors (NF-κB (p50), AP-4, YY-1, c-myc, SP1, CBF-1 and LSF) can recruit HDAC-1 and -2, and SIN3A or MTA1 to the HIV-1 LTR, the molecular mechanism(s) regulating the activity and/or presence of HDAC-1/-2 and their associated complexes at the integrated HIV-1 LTR has not been fully elucidated[241, 243, 260–266]. The interaction of HIV-1 Tat with the SIN3/HDAC and NuRD complexes further implicates them as potential epigenetic regulators of HIV-1 post-integration latency and suggests how Tat might intersect with epigenetic pathways.
Alternatively, the enzymatic activities of the multi-protein complexes recruited by Tat, such as methylation, acetylation and deacetylation, could be directed to Tat itself and mediate post-translational modifications, as it has been shown previously with PCAF, p300/CBP, SIRT1, or SETDB1/2 [13, 14, 267, 268].
Accumulative evidence has recently described how cellular proteins belonging to the DNA replication and the mitotic chromosome condensation machineries have been shown to carry out additional activities in gene expression regulation and/or silencing by selectively affecting the chromatin/chromosome architecture during the interphase. The identification of Tat interactions with multiple components of the cohesin, condensin, toposome, MCM and ORC complexes provide us with a new prospective on how these pathways might also influence Tat function and HIV-1 provirus expression and silencing.
In addition to its role in regulating the access of specific regulatory factors to the nucleus, the nuclear architecture can affect the genome subnuclear organisation and chromatin structure. In general, there is a strong correlation between the nuclear periphery and heterochromatin establishment and/or maintenance and accordingly gene silencing, while the NPCs have been implicated in preserving euchromatin from such a process[269, 270]. More specifically, components of the inner nuclear envelop, lamina and NPC have been described to have an important role in regulating gene expression. LBR, part of our dataset, has been shown to interact with HP1 and consequently regulates heterochromatin formation[271]. Other lamin-associated proteins, such as emerin and Lap2, were also identified as Tat interactors in our studies, have been shown to associate with and sequester a number of transcriptional repressors, including HDACs, BAF, NCoR and beta-catenin[270, 272]. The identification of numerous crucial elements involved the nuclear architecture, as Tat interactors, suggest that they could be involved in regulating the transcriptional state of the provirus.
Conclusion
The results presented here, position the viral regulatory protein at the nexus of a range of interaction networks, which play essential and diverse roles in gene expression, RNA processing, chromatin organisation, chromosome structure and nuclear architecture, and provide the first insights into the modular network properties of the Tat interactome. Ultimately, the HIV-1 Tat rewiring of cellular networks could equip the provirus with a wide repertoire of tools to orchestrate HIV-1 gene expression and confer a remarkable adaptability to a continuously changing cellular environment. Overall, this confirms that Tat transactivation function appears to be the net result of complex interactions with distinct cellular complexes highly specialised in controlling gene expression and more specifically chromosome/chromatin structure. We anticipate that the data presented here will be useful for researchers investigating HIV-1 gene regulation and further studies will delineate the biological significance of these findings.
Methods
Cell culture
Jurkat cell line (Clone E6-1) was maintained in RPMI 1640 medium containing 10% fetal calf serum and supplemented with 0.3 mg/L of L-Glutamine (GIBCO) and antibiotics. Nuclear extracts were prepared from Jurkat cells with the ProteoExtract® Subcellular Proteome Extraction Kit (Calbiochem) according to the manufacturers instructions, which include the treatment of the nuclear fraction with Benzonase® nuclease.
Production of recombinant proteins
GST and GST-Tat (HIV-1 HXB2, 86 amino acids) recombinant proteins were produced in BL21 E. coli. and purified with gluthathione-Sepharose beads (Amersham) as described previously[273].
In vitroGST-pull down assays
To create high-density ligand surface, equivalent amounts of purified recombinant GST-Tat (bait) and GST (negative control) proteins were added in excess and immobilised on gluthathione-Sepharose 4 Fast flow (Amersham). The supernatant was discarded and following extensive washes in Binding Buffer (BB) (20 mM Tris, pH 7.4, 300 mM NaCl, 100 mM NaF, 1 mM DTT, 50 mM EDTA, 1% triton X100, 10% glycerol and protease inhibitor cocktail Complete, EDTA-free (Roche)), the beads were incubated with Jurkat cell nuclear extracts (300 μg), rotating at 4°C overnight. Following extensive washes in binding buffer, specifically retained protein complexes were eluted from the resin by incubating the beads in 2× Leamni sample buffer at 95°C for 5 min.
Gel electrophoresis and Coomassie staining
The eluate was resolved on a 10% SDS-PAGE and the resulting interaction profiles were sliced into 2 mm bands across the entire separation range without bias with respect to size and relatively abundance. A total of 164 gel slices were produced.
Sample preparation for Mass Spectrometric analysis
1D gel bands were excised from the control and experimental lanes and subjected to in-gel trypsin digestion. Briefly gel bands were washed with sequential additions of ammonium biocarbonate and acetonitrile (ACN) buffers and cysteine residues were reduced and alkylated using DTT and IAA. Samples were digested overnight with 4 ng/μl trypsin at 37°C. Peptides were extracted using 60% ACN, 0.2% TFA buffer and were dried and stored for subsequent MS analysis.
Protein identification by LC MSMS
LC MS/MS was carried out on a Finnigan LTQ mass spectrometer connected to a Surveyor chromatography system incorporating an autosampler. Tryptic peptides were resuspended in 0.1% formic acid and were separated by means of a modular CapaLC system (Finnigan) connected directly to the source of the LTQ. Each sample was loaded onto a Biobasic C18 Picofrit™ column (100 mm length, 75 μm ID) at a flow rate of 30 nL min-1. The samples were then eluted from the C18 Picofrit™ column by an increasing acetonitrile gradient. The mass spectrometer was operated in positive ion mode with a capillary temperature of 200°C, a capillary voltage of 46 V, a tube lens voltage of 140 V and with a potential of 1.8 kV applied to the frit. All data was acquired with the mass spectrometer operating in automatic data dependent switching mode. A zoom scan was performed on the 5 most intense ions to determine charge state prior to MS/MS analysis.
Data analysis
All MS/MS spectra were sequence database searched using TurboSEQUEST. The MS/MS spectra were searched against the redundant TREMBL database. The following search parameters were used, precursor-ion mass tolerance of 1.5, fragment ion tolerance of 1.0 with methionine oxidation and cysteine carboxyamidomethylation specified as differential modifications and a maximum of 2 missed cleavage sites allowed.
Western-Blotting analysis
To confirm the MS/MS identification of selected Tat-interacting proteins, we performed Western-Blotting analysis, using BioTrace ™ PVDF (Pall Corporation), on one fifth of the eluate resulting from GST-pull down as described above, but carried out with Jurkat cell nuclear extract (150 μg) and washes in BB including various salt (NaCl) concentrations (300 mM, 500 mM, 800 mM and 1 M). Similarly, GST-pull downs were performed with a Tat-deletion mutant GST-Tat-NLS described elsewhere[273]. The following primary antibodies and their corresponding dilutions were employed: CyclinT1 (H-245) at 1/1000 dilution, mSIN3a (K-20) at 1/5000 dilution, SAP18 (H-130) at 1/5000 dilution, SPT16 (H-300) at 1/1000 dilution (Santa Cruz Biotechnology); and BAF53A at 1/2000 dilution, Ikaros at 1/4000 dilution, HDAC1 at 1/5000 dilution, SNF2H at 1/1000 dilution, WDR51/500 dilution and RbAp46/48 1/1000 dilution (Abcam). the following secondary antibodies (GE Healthcare) and their corresponding dilutions were employed: ECL ™ Anti-mouse IgG at 1/5000 dilution and ECL ™ Anti-rabbit IgG at 1/10000 dilution.
References
Bannwarth S, Gatignol A: HIV-1 TAR RNA: the target of molecular interactions between the virus and its host. Curr HIV Res. 2005, 3: 61-71.
Brady J, Kashanchi F: Tat gets the "green" light on transcription initiation. Retrovirology. 2005, 2: 69-
Nekhai S, Jeang KT: Transcriptional and post-transcriptional regulation of HIV-1 gene expression: role of cellular factors for Tat and Rev. Future Microbiol. 2006, 1: 417-426.
Barboric M, Peterlin BM: A new paradigm in eukaryotic biology: HIV Tat and the control of transcriptional elongation. PLoS Biol. 2005, 3: e76-
Agbottah E, Deng L, Dannenberg LO, Pumfery A, Kashanchi F: Effect of SWI/SNF chromatin remodeling complex on HIV-1 Tat activated transcription. Retrovirology. 2006, 3: 48-
Treand C, du Chéné I, Bres V, Kiernan R, Benarous R, Benkirane M, Emiliani S: Requirement for SWI/SNF chromatin-remodeling complex in Tat-mediated activation of the HIV-1 promoter. Embo J. 2006, 25: 1690-1699.
Mahmoudi T, Parra M, Vries RG, Kauder SE, Verrijzer CP, Ott M, Verdin E: The SWI/SNF chromatin-remodeling complex is a cofactor for Tat transactivation of the HIV promoter. J Biol Chem. 2006, 281: 19960-19968.
Zhu Y, Pe'ery T, Peng J, Ramanathan Y, Marshall N, Marshall T, Amendt B, Mathews MB, Price DH: Transcription elongation factor P-TEFb is required for HIV-1 tat transactivation in vitro. Genes Dev. 1997, 11: 2622-2632.
Benkirane M, Chun RF, Xiao H, Ogryzko VV, Howard BH, Nakatani Y, Jeang KT: Activation of integrated provirus requires histone acetyltransferase. p300 and P/CAF are coactivators for HIV-1 Tat. J Biol Chem. 1998, 273: 24898-24905.
Gatignol A: Transcription of HIV: Tat and cellular chromatin. Adv Pharmacol. 2007, 55: 137-159.
Peruzzi F: The multiple functions of HIV-1 Tat: proliferation versus apoptosis. Front Biosci. 2006, 11: 708-717.
Hetzer C, Dormeyer W, Schnolzer M, Ott M: Decoding Tat: the biology of HIV Tat posttranslational modifications. Microbes Infect. 2005, 7: 1364-1369.
Van Duyne R, Easley R, Wu W, Berro R, Pedati C, Klase Z, Kehn-Hall K, Flynn EK, Symer DE, Kashanchi F: Lysine methylation of HIV-1 Tat regulates transcriptional activity of the viral LTR. Retrovirology. 2008, 5: 40-
Pagans S, Pedal A, North BJ, Kaehlcke K, Marshall BL, Dorr A, Hetzer-Egger C, Henklein P, Frye R, McBurney MW, et al: SIRT1 regulates HIV transcription via Tat deacetylation. PLoS Biol. 2005, 3: e41-
Fu W, Sanders-Beer BE, Katz KS, Maglott DR, Pruitt KD, Ptak RG: Human immunodeficiency virus type 1, human protein interaction database at NCBI. Nucleic Acids Res. 2008
Sorin M, Kalpana GV: Dynamics of virus-host interplay in HIV-1 replication. Curr HIV Res. 2006, 4: 117-130.
Takeuchi H, Matano T: Host factors involved in resistance to retroviral infection. Microbiol Immunol. 2008, 52: 318-325.
Tasara T, Hottiger MO, Hubscher U: Functional genomics in HIV-1 virus replication: protein-protein interactions as a basis for recruiting the host cell machinery for viral propagation. Biol Chem. 2001, 382: 993-999.
Van Duyne R, Kehn-Hall K, Klase Z, Easley R, Heydarian M, Saifuddin M, Wu W, Kashanchi F: Retroviral proteomics and interactomes: intricate balances of cell survival and viral replication. Expert Rev Proteomics. 2008, 5: 507-528.
Ammosova T, Jerebtsova M, Beullens M, Lesage B, Jackson A, Kashanchi F, Southerland W, Gordeuk VR, Bollen M, Nekhai S: Nuclear targeting of protein phosphatase-1 by HIV-1 Tat protein. J Biol Chem. 2005, 280: 36364-36371.
Ansari SA, Safak M, Gallia GL, Sawaya BE, Amini S, Khalili K: Interaction of YB-1 with human immunodeficiency virus type 1 Tat and TAR RNA modulates viral promoter activity. J Gen Virol. 1999, 80 (Pt 10): 2629-2638.
Ariumi Y, Serhan F, Turelli P, Telenti A, Trono D: The integrase interactor 1 (INI1) proteins facilitate Tat-mediated human immunodeficiency virus type 1 transcription. Retrovirology. 2006, 3: 47-
Berro R, Kehn K, de la Fuente C, Pumfery A, Adair R, Wade J, Colberg-Poley AM, Hiscott J, Kashanchi F: Acetylated Tat regulates human immunodeficiency virus type 1 splicing through its interaction with the splicing regulator p32. J Virol. 2006, 80: 3189-3204.
Cujec TP, Cho H, Maldonado E, Meyer J, Reinberg D, Peterlin BM: The human immunodeficiency virus transactivator Tat interacts with the RNA polymerase II holoenzyme. Mol Cell Biol. 1997, 17: 1817-1823.
Cujec TP, Okamoto H, Fujinaga K, Meyer J, Chamberlin H, Morgan DO, Peterlin BM: The HIV transactivator TAT binds to the CDK-activating kinase and activates the phosphorylation of the carboxy-terminal domain of RNA polymerase II. Genes Dev. 1997, 11: 2645-2657.
Garcia-Martinez LF, Mavankal G, Neveu JM, Lane WS, Ivanov D, Gaynor RB: Purification of a Tat-associated kinase reveals a TFIIH complex that modulates HIV-1 transcription. Embo J. 1997, 16: 2836-2850.
Hidalgo-Estevez AM, Gonzalez E, Punzon C, Fresno M: Human immunodeficiency virus type 1 Tat increases cooperation between AP-1 and NFAT transcription factors in T cells. J Gen Virol. 2006, 87: 1603-1612.
Kashanchi F, Piras G, Radonovich MF, Duvall JF, Fattaey A, Chiang CM, Roeder RG, Brady JN: Direct interaction of human TFIID with the HIV-1 transactivator tat. Nature. 1994, 367: 295-299.
Li YP: Protein B23 is an important human factor for the nucleolar localization of the human immunodeficiency virus protein Tat. J Virol. 1997, 71: 4098-4102.
Macian F, Rao A: Reciprocal modulatory interaction between human immunodeficiency virus type 1 Tat and transcription factor NFAT1. Mol Cell Biol. 1999, 19: 3645-3653.
Muller WE, Okamoto T, Reuter P, Ugarkovic D, Schroder HC: Functional characterization of Tat protein from human immunodeficiency virus. Evidence that Tat links viral RNAs to nuclear matrix. J Biol Chem. 1990, 265: 3803-3808.
Muller WE, Wenger R, Reuter P, Renneisen K, Schroder HC: Association of Tat protein and viral mRNA with nuclear matrix from HIV-1-infected H9 cells. Biochim Biophys Acta. 1989, 1008: 208-212.
Nekhai S, Zhou M, Fernandez A, Lane WS, Lamb NJ, Brady J, Kumar A: HIV-1 Tat-associated RNA polymerase C-terminal domain kinase, CDK2, phosphorylates CDK7 and stimulates Tat-mediated transcription. Biochem J. 2002, 364: 649-657.
Parada CA, Roeder RG: Enhanced processivity of RNA polymerase II triggered by Tat-induced phosphorylation of its carboxy-terminal domain. Nature. 1996, 384: 375-378.
Rohr O, Lecestre D, Chasserot-Golaz S, Marban C, Avram D, Aunis D, Leid M, Schaeffer E: Recruitment of Tat to heterochromatin protein HP1 via interaction with CTIP2 inhibits human immunodeficiency virus type 1 replication in microglial cells. J Virol. 2003, 77: 5415-5427.
Truant R, Cullen BR: The arginine-rich domains present in human immunodeficiency virus type 1 Tat and Rev function as direct importin beta-dependent nuclear localization signals. Mol Cell Biol. 1999, 19: 1210-1217.
Xiao H, Neuveut C, Benkirane M, Jeang KT: Interaction of the second coding exon of Tat with human EF-1 delta delineates a mechanism for HIV-1-mediated shut-off of host mRNA translation. Biochem Biophys Res Commun. 1998, 244: 384-389.
Yoder K, Sarasin A, Kraemer K, McIlhatton M, Bushman F, Fishel R: The DNA repair genes XPB and XPD defend cells from retroviral infection. Proc Natl Acad Sci USA. 2006, 103: 4622-4627.
Herrmann CH, Rice AP: Lentivirus Tat proteins specifically associate with a cellular protein kinase, TAK, that hyperphosphorylates the carboxyl-terminal domain of the large subunit of RNA polymerase II: candidate for a Tat cofactor. J Virol. 1995, 69: 1612-1620.
Kamine J, Elangovan B, Subramanian T, Coleman D, Chinnadurai G: Identification of a cellular protein that specifically interacts with the essential cysteine region of the HIV-1 Tat transactivator. Virology. 1996, 216: 357-366.
Yang X, Herrmann CH, Rice AP: The human immunodeficiency virus Tat proteins specifically associate with TAK in vivo and require the carboxyl-terminal domain of RNA polymerase II for function. J Virol. 1996, 70: 4576-4584.
Finn RD, Tate J, Mistry J, Coggill PC, Sammut SJ, Hotz HR, Ceric G, Forslund K, Eddy SR, Sonnhammer EL, Bateman A: The Pfam protein families database. Nucleic Acids Res. 2008, 36: D281-288.
Letunic I, Copley RR, Pils B, Pinkert S, Schultz J, Bork P: SMART 5: domains in the context of genomes and networks. Nucleic Acids Res. 2006, 34: D257-260.
Marchler-Bauer A, Anderson JB, Derbyshire MK, DeWeese-Scott C, Gonzales NR, Gwadz M, Hao L, He S, Hurwitz DI, Jackson JD, et al: CDD: a conserved domain database for interactive domain family analysis. Nucleic Acids Res. 2007, 35: D237-240.
Conserved Domains. [http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi]
SMART. [http://smart.embl-heidelberg.de/]
PFAM. [http://pfam.sanger.ac.uk/]
Dellaire G, Farrall R, Bickmore WA: The Nuclear Protein Database (NPD): sub-nuclear localisation and functional annotation of the nuclear proteome. Nucleic Acids Res. 2003, 31: 328-330.
Nuclear Protein Database (NPD). [https://npd.hgu.mrc.ac.uk/index.html]
Pyle AM: Translocation and unwinding mechanisms of RNA and DNA helicases. Annu Rev Biophys. 2008, 37: 317-336.
Stros M, Launholt D, Grasser KD: The HMG-box: a versatile protein domain occurring in a wide variety of DNA-binding proteins. Cell Mol Life Sci. 2007, 64: 2590-2606.
Clery A, Blatter M, Allain FH: RNA recognition motifs: boring? Not quite. Curr Opin Struct Biol. 2008, 18: 290-298.
Boyer LA, Latek RR, Peterson CL: The SANT domain: a unique histone-tail-binding module?. Nat Rev Mol Cell Biol. 2004, 5: 158-163.
Bienz M: The PHD finger, a nuclear protein-interaction domain. Trends Biochem Sci. 2006, 31: 35-40.
Mujtaba S, Zeng L, Zhou MM: Structure and acetyl-lysine recognition of the bromodomain. Oncogene. 2007, 26: 5521-5527.
Li D, Roberts R: WD-repeat proteins: structure characteristics, biological function, and their involvement in human diseases. Cell Mol Life Sci. 2001, 58: 2085-2097.
Tucker PA, Sallai L: The AAA+ superfamily – a myriad of motions. Curr Opin Struct Biol. 2007, 17: 641-652.
Bennasser Y, Jeang KT: HIV-1 Tat interaction with Dicer: requirement for RNA. Retrovirology. 2006, 3: 95-
Epie N, Ammosova T, Sapir T, Voloshin Y, Lane WS, Turner W, Reiner O, Nekhai S: HIV-1 Tat interacts with LIS1 protein. Retrovirology. 2005, 2: 6-
Mujtaba S, He Y, Zeng L, Farooq A, Carlson JE, Ott M, Verdin E, Zhou MM: Structural basis of lysine-acetylated HIV-1 Tat recognition by PCAF bromodomain. Mol Cell. 2002, 9: 575-586.
Bottomley MJ: Structures of protein domains that create or recognize histone modifications. EMBO Rep. 2004, 5: 464-469.
Gamsjaeger R, Liew CK, Loughlin FE, Crossley M, Mackay JP: Sticky fingers: zinc-fingers as protein-recognition motifs. Trends Biochem Sci. 2007, 32: 63-70.
Lunde BM, Moore C, Varani G: RNA-binding proteins: modular design for efficient function. Nat Rev Mol Cell Biol. 2007, 8: 479-490.
Maiorano D, Lutzmann M, Mechali M: MCM proteins and DNA replication. Curr Opin Cell Biol. 2006, 18: 130-136.
Racki LR, Narlikar GJ: ATP-dependent chromatin remodeling enzymes: two heads are not better, just different. Curr Opin Genet Dev. 2008, 18: 137-144.
Sclafani RA, Holzen TM: Cell cycle regulation of DNA replication. Annu Rev Genet. 2007, 41: 237-280.
Ruthenburg AJ, Li H, Patel DJ, Allis CD: Multivalent engagement of chromatin modifications by linked binding modules. Nat Rev Mol Cell Biol. 2007, 8: 983-994.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al: Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000, 25: 25-29.
Gene Ontology (G.O.). [http://www.geneontology.org/index.shtml]
Breitkreutz BJ, Stark C, Tyers M: Osprey: a network visualization system. Genome Biol. 2003, 4: R22-
Alfarano C, Andrade CE, Anthony K, Bahroos N, Bajec M, Bantoft K, Betel D, Bobechko B, Boutilier K, Burgess E, et al: The Biomolecular Interaction Network Database and related tools 2005 update. Nucleic Acids Res. 2005, 33: D418-424.
Peri S, Navarro JD, Amanchy R, Kristiansen TZ, Jonnalagadda CK, Surendranath V, Niranjan V, Muthusamy B, Gandhi TK, Gronborg M, et al: Development of human protein reference database as an initial platform for approaching systems biology in humans. Genome Res. 2003, 13: 2363-2371.
BIND Biomolecular Interaction Network Database. [http://www.bind.ca]
HPRD Human Proteins Reference Database. [http://www.hprd.org]
Adler HT, Chinery R, Wu DY, Kussick SJ, Payne JM, Fornace AJ, Tkachuk DC: Leukemic HRX fusion proteins inhibit GADD34-induced apoptosis and associate with the GADD34 and hSNF5/INI1 proteins. Mol Cell Biol. 1999, 19: 7050-7060.
Adzerikho RD, Aksentsev SL, Okun IM, Konev SV: [Letter: Change in trypsin sensitivity during structural rearrangements in biological membranes]. Biofizika. 1975, 20: 942-944.
Ashburner BP, Westerheide SD, Baldwin AS: The p65 (RelA) subunit of NF-kappaB interacts with the histone deacetylase (HDAC) corepressors HDAC1 and HDAC2 to negatively regulate gene expression. Mol Cell Biol. 2001, 21: 7065-7077.
Becker J, Melchior F, Gerke V, Bischoff FR, Ponstingl H, Wittinghofer A: RNA1 encodes a GTPase-activating protein specific for Gsp1p, the Ran/TC4 homologue of Saccharomyces cerevisiae. J Biol Chem. 1995, 270: 11860-11865.
Bharucha DC, Zhou M, Nekhai S, Brady JN, Shukla RR, Kumar A: A protein phosphatase from human T cells augments tat transactivation of the human immunodeficiency virus type 1 long-terminal repeat. Virology. 2002, 296: 6-16.
Bischoff FR, Klebe C, Kretschmer J, Wittinghofer A, Ponstingl H: RanGAP1 induces GTPase activity of nuclear Ras-related Ran. Proc Natl Acad Sci USA. 1994, 91: 2587-2591.
Block KL, Vornlocher HP, Hershey JW: Characterization of cDNAs encoding the p44 and p35 subunits of human translation initiation factor eIF3. J Biol Chem. 1998, 273: 31901-31908.
Bochar DA, Wang L, Beniya H, Kinev A, Xue Y, Lane WS, Wang W, Kashanchi F, Shiekhattar R: BRCA1 is associated with a human SWI/SNF-related complex: linking chromatin remodeling to breast cancer. Cell. 2000, 102: 257-265.
Bonifaci N, Moroianu J, Radu A, Blobel G: Karyopherin beta2 mediates nuclear import of a mRNA binding protein. Proc Natl Acad Sci USA. 1997, 94: 5055-5060.
Brackertz M, Boeke J, Zhang R, Renkawitz R: Two highly related p66 proteins comprise a new family of potent transcriptional repressors interacting with MBD2 and MBD3. J Biol Chem. 2002, 277: 40958-40966.
Brand M, Moggs JG, Oulad-Abdelghani M, Lejeune F, Dilworth FJ, Stevenin J, Almouzni G, Tora L: UV-damaged DNA-binding protein in the TFTC complex links DNA damage recognition to nucleosome acetylation. Embo J. 2001, 20: 3187-3196.
Chibi M, Meyer M, Skepu A, DJ GR, Moolman-Smook JC, Pugh DJ: RBBP6 Interacts with Multifunctional Protein YB-1 through Its RING Finger Domain, Leading to Ubiquitination and Proteosomal Degradation of YB-1. J Mol Biol. 2008
Cho H, Orphanides G, Sun X, Yang XJ, Ogryzko V, Lees E, Nakatani Y, Reinberg D: A human RNA polymerase II complex containing factors that modify chromatin structure. Mol Cell Biol. 1998, 18: 5355-5363.
Chou MY, Rooke N, Turck CW, Black DL: hnRNP H is a component of a splicing enhancer complex that activates a c-src alternative exon in neuronal cells. Mol Cell Biol. 1999, 19: 69-77.
Corsini L, Bonnal S, Basquin J, Hothorn M, Scheffzek K, Valcarcel J, Sattler M: U2AF-homology motif interactions are required for alternative splicing regulation by SPF45. Nat Struct Mol Biol. 2007, 14: 620-629.
Craig E, Zhang ZK, Davies KP, Kalpana GV: A masked NES in INI1/hSNF5 mediates hCRM1-dependent nuclear export: implications for tumorigenesis. Embo J. 2002, 21: 31-42.
Cramer P, Armache KJ, Baumli S, Benkert S, Brueckner F, Buchen C, Damsma GE, Dengl S, Geiger SR, Jasiak AJ, et al: Structure of eukaryotic RNA polymerases. Annu Rev Biophys. 2008, 37: 337-352.
Das BK, Xia L, Palandjian L, Gozani O, Chyung Y, Reed R: Characterization of a protein complex containing spliceosomal proteins SAPs 49, 130, 145, and 155. Mol Cell Biol. 1999, 19: 6796-6802.
Debernardi S, Bassini A, Jones LK, Chaplin T, Linder B, de Bruijn DR, Meese E, Young BD: The MLL fusion partner AF10 binds GAS41, a protein that interacts with the human SWI/SNF complex. Blood. 2002, 99: 275-281.
Delphin C, Guan T, Melchior F, Gerace L: RanGTP targets p97 to RanBP2, a filamentous protein localized at the cytoplasmic periphery of the nuclear pore complex. Mol Biol Cell. 1997, 8: 2379-2390.
Dhar SK, Delmolino L, Dutta A: Architecture of the human origin recognition complex. J Biol Chem. 2001, 276: 29067-29071.
Doyon Y, Selleck W, Lane WS, Tan S, Cote J: Structural and functional conservation of the NuA4 histone acetyltransferase complex from yeast to humans. Mol Cell Biol. 2004, 24: 1884-1896.
Dreger CK, Konig AR, Spring H, Lichter P, Herrmann H: Investigation of nuclear architecture with a domain-presenting expression system. J Struct Biol. 2002, 140: 100-115.
Ewing RM, Chu P, Elisma F, Li H, Taylor P, Climie S, McBroom-Cerajewski L, Robinson MD, O'Connor L, Li M, et al: Large-scale mapping of human protein-protein interactions by mass spectrometry. Mol Syst Biol. 2007, 3: 89-
Finch L, Hicks PE: Proceedings: Central hypertensive action of histamine in rats. Br J Pharmacol. 1975, 55: 274P-275P.
Fischer DD, Cai R, Bhatia U, Asselbergs FA, Song C, Terry R, Trogani N, Widmer R, Atadja P, Cohen D: Isolation and characterization of a novel class II histone deacetylase, HDAC10. J Biol Chem. 2002, 277: 6656-6666.
Fischle W, Dequiedt F, Fillion M, Hendzel MJ, Voelter W, Verdin E: Human HDAC7 histone deacetylase activity is associated with HDAC3 in vivo. J Biol Chem. 2001, 276: 35826-35835.
Fischle W, Dequiedt F, Hendzel MJ, Guenther MG, Lazar MA, Voelter W, Verdin E: Enzymatic activity associated with class II HDACs is dependent on a multiprotein complex containing HDAC3 and SMRT/N-CoR. Mol Cell. 2002, 9: 45-57.
Fleischer TC, Yun UJ, Ayer DE: Identification and characterization of three new components of the mSin3A corepressor complex. Mol Cell Biol. 2003, 23: 3456-3467.
Fornerod M, Ohno M, Yoshida M, Mattaj IW: CRM1 is an export receptor for leucine-rich nuclear export signals. Cell. 1997, 90: 1051-1060.
Franz C, Walczak R, Yavuz S, Santarella R, Gentzel M, Askjaer P, Galy V, Hetzer M, Mattaj IW, Antonin W: MEL-28/ELYS is required for the recruitment of nucleoporins to chromatin and postmitotic nuclear pore complex assembly. EMBO Rep. 2007, 8: 165-172.
Fuchs M, Gerber J, Drapkin R, Sif S, Ikura T, Ogryzko V, Lane WS, Nakatani Y, Livingston DM: The p400 complex is an essential E1A transformation target. Cell. 2001, 106: 297-307.
Fujita H, Fujii R, Aratani S, Amano T, Fukamizu A, Nakajima T: Antithetic effects of MBD2a on gene regulation. Mol Cell Biol. 2003, 23: 2645-2657.
Fujita M, Yamada C, Goto H, Yokoyama N, Kuzushima K, Inagaki M, Tsurumi T: Cell cycle regulation of human CDC6 protein. Intracellular localization, interaction with the human mcm complex, and CDC2 kinase-mediated hyperphosphorylation. J Biol Chem. 1999, 274: 25927-25932.
Fujita N, Jaye DL, Kajita M, Geigerman C, Moreno CS, Wade PA: MTA3, a Mi-2/NuRD complex subunit, regulates an invasive growth pathway in breast cancer. Cell. 2003, 113: 207-219.
Furukawa K, Kondo T: Identification of the lamina-associated-polypeptide-2-binding domain of B-type lamin. Eur J Biochem. 1998, 251: 729-733.
Gacesa P, Whish WJ: The immobilization of adenine nucleotides on polysaccharides by using glutaraldehyde coupling and borohydride reduction. Biochem J. 1978, 175: 349-352.
Gorlich D, Henklein P, Laskey RA, Hartmann E: A 41 amino acid motif in importin-alpha confers binding to importin-beta and hence transit into the nucleus. Embo J. 1996, 15: 1810-1817.
Griffis ER, Xu S, Powers MA: Nup98 localizes to both nuclear and cytoplasmic sides of the nuclear pore and binds to two distinct nucleoporin subcomplexes. Mol Biol Cell. 2003, 14: 600-610.
Grozinger CM, Hassig CA, Schreiber SL: Three proteins define a class of human histone deacetylases related to yeast Hda1p. Proc Natl Acad Sci USA. 1999, 96: 4868-4873.
Hakimi MA, Bochar DA, Chenoweth J, Lane WS, Mandel G, Shiekhattar R: A core-BRAF35 complex containing histone deacetylase mediates repression of neuronal-specific genes. Proc Natl Acad Sci USA. 2002, 99: 7420-7425.
Hakimi MA, Bochar DA, Schmiesing JA, Dong Y, Barak OG, Speicher DW, Yokomori K, Shiekhattar R: A chromatin remodelling complex that loads cohesin onto human chromosomes. Nature. 2002, 418: 994-998.
Hakimi MA, Dong Y, Lane WS, Speicher DW, Shiekhattar R: A candidate X-linked mental retardation gene is a component of a new family of histone deacetylase-containing complexes. J Biol Chem. 2003, 278: 7234-7239.
Harborth J, Weber K, Osborn M: GAS41, a highly conserved protein in eukaryotic nuclei, binds to NuMA. J Biol Chem. 2000, 275: 31979-31985.
Harris TE, Chi A, Shabanowitz J, Hunt DF, Rhoads RE, Lawrence JC: mTOR-dependent stimulation of the association of eIF4G and eIF3 by insulin. Embo J. 2006, 25: 1659-1668.
Hassig CA, Fleischer TC, Billin AN, Schreiber SL, Ayer DE: Histone deacetylase activity is required for full transcriptional repression by mSin3A. Cell. 1997, 89: 341-347.
Hassig CA, Tong JK, Fleischer TC, Owa T, Grable PG, Ayer DE, Schreiber SL: A role for histone deacetylase activity in HDAC1-mediated transcriptional repression. Proc Natl Acad Sci USA. 1998, 95: 3519-3524.
Hillig RC, Renault L, Vetter IR, Drell Tt, Wittinghofer A, Becker J: The crystal structure of rna1p: a new fold for a GTPase-activating protein. Mol Cell. 1999, 3: 781-791.
Hsieh YJ, Kundu TK, Wang Z, Kovelman R, Roeder RG: The TFIIIC90 subunit of TFIIIC interacts with multiple components of the RNA polymerase III machinery and contains a histone-specific acetyltransferase activity. Mol Cell Biol. 1999, 19: 7697-7704.
Ishimi Y, Ichinose S, Omori A, Sato K, Kimura H: Binding of human minichromosome maintenance proteins with histone H3. J Biol Chem. 1996, 271: 24115-24122.
Ishimi Y, Komamura-Kohno Y, You Z, Omori A, Kitagawa M: Inhibition of Mcm4,6,7 helicase activity by phosphorylation with cyclin A/Cdk2. J Biol Chem. 2000, 275: 16235-16241.
Jiang CL, Jin SG, Pfeifer GP: MBD3L1 is a transcriptional repressor that interacts with methyl-CpG-binding protein 2 (MBD2) and components of the NuRD complex. J Biol Chem. 2004, 279: 52456-52464.
Johnson CA, Padget K, Austin CA, Turner BM: Deacetylase activity associates with topoisomerase II and is necessary for etoposide-induced apoptosis. J Biol Chem. 2001, 276: 4539-4542.
Johnson CA, White DA, Lavender JS, O'Neill LP, Turner BM: Human class I histone deacetylase complexes show enhanced catalytic activity in the presence of ATP and co-immunoprecipitate with the ATP-dependent chaperone protein Hsp70. J Biol Chem. 2002, 277: 9590-9597.
Joseph J, Tan SH, Karpova TS, McNally JG, Dasso M: SUMO-1 targets RanGAP1 to kinetochores and mitotic spindles. J Cell Biol. 2002, 156: 595-602.
Kato H, Tjernberg A, Zhang W, Krutchinsky AN, An W, Takeuchi T, Ohtsuki Y, Sugano S, de Bruijn DR, Chait BT, Roeder RG: SYT associates with human SNF/SWI complexes and the C-terminal region of its fusion partner SSX1 targets histones. J Biol Chem. 2002, 277: 5498-5505.
Kehlenbach RH, Gerace L: Phosphorylation of the nuclear transport machinery down-regulates nuclear protein import in vitro. J Biol Chem. 2000, 275: 17848-17856.
Kitagawa H, Fujiki R, Yoshimura K, Mezaki Y, Uematsu Y, Matsui D, Ogawa S, Unno K, Okubo M, Tokita A, et al: The chromatin-remodeling complex WINAC targets a nuclear receptor to promoters and is impaired in Williams syndrome. Cell. 2003, 113: 905-917.
Kneissl M, Putter V, Szalay AA, Grummt F: Interaction and assembly of murine pre-replicative complex proteins in yeast and mouse cells. J Mol Biol. 2003, 327: 111-128.
Kohler M, Speck C, Christiansen M, Bischoff FR, Prehn S, Haller H, Gorlich D, Hartmann E: Evidence for distinct substrate specificities of importin alpha family members in nuclear protein import. Mol Cell Biol. 1999, 19: 7782-7791.
Koipally J, Georgopoulos K: A molecular dissection of the repression circuitry of Ikaros. J Biol Chem. 2002, 277: 27697-27705.
Koipally J, Georgopoulos K: Ikaros-CtIP interactions do not require C-terminal binding protein and participate in a deacetylase-independent mode of repression. J Biol Chem. 2002, 277: 23143-23149.
Koipally J, Renold A, Kim J, Georgopoulos K: Repression by Ikaros and Aiolos is mediated through histone deacetylase complexes. Embo J. 1999, 18: 3090-3100.
Kutay U, Izaurralde E, Bischoff FR, Mattaj IW, Gorlich D: Dominant-negative mutants of importin-beta block multiple pathways of import and export through the nuclear pore complex. Embo J. 1997, 16: 1153-1163.
Kuzmichev A, Zhang Y, Erdjument-Bromage H, Tempst P, Reinberg D: Role of the Sin3-histone deacetylase complex in growth regulation by the candidate tumor suppressor p33(ING1). Mol Cell Biol. 2002, 22: 835-848.
Laherty CD, Yang WM, Sun JM, Davie JR, Seto E, Eisenman RN: Histone deacetylases associated with the mSin3 corepressor mediate mad transcriptional repression. Cell. 1997, 89: 349-356.
Lattanzi G, Cenni V, Marmiroli S, Capanni C, Mattioli E, Merlini L, Squarzoni S, Maraldi NM: Association of emerin with nuclear and cytoplasmic actin is regulated in differentiating myoblasts. Biochem Biophys Res Commun. 2003, 303: 764-770.
Lee CG, Hague LK, Li H, Donnelly R: Identification of toposome, a novel multisubunit complex containing topoisomerase IIalpha. Cell Cycle. 2004, 3: 638-647.
Lee JH, Skalnik DG: CpG-binding protein (CXXC finger protein 1) is a component of the mammalian Set1 histone H3-Lys4 methyltransferase complex, the analogue of the yeast Set1/COMPASS complex. J Biol Chem. 2005, 280: 41725-41731.
Li YP, Busch RK, Valdez BC, Busch H: C23 interacts with B23, a putative nucleolar-localization-signal-binding protein. Eur J Biochem. 1996, 237: 153-158.
Liu JH, Wei S, Burnette PK, Gamero AM, Hutton M, Djeu JY: Functional association of TGF-beta receptor II with cyclin B. Oncogene. 1999, 18: 269-275.
Lounsbury KM, Richards SA, Perlungher RR, Macara IG: Ran binding domains promote the interaction of Ran with p97/beta-karyopherin, linking the docking and translocation steps of nuclear import. J Biol Chem. 1996, 271: 2357-2360.
Mahajan R, Delphin C, Guan T, Gerace L, Melchior F: A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell. 1997, 88: 97-107.
Makar AB, McMartin KE, Palese M, Tephly TR: Formate assay in body fluids: application in methanol poisoning. Biochem Med. 1975, 13: 117-126.
Marango J, Shimoyama M, Nishio H, Meyer JA, Min DJ, Sirulnik A, Martinez-Martinez Y, Chesi M, Bergsagel PL, Zhou MM, et al: The MMSET protein is a histone methyltransferase with characteristics of a transcriptional corepressor. Blood. 2008, 111: 3145-3154.
Mayeur GL, Fraser CS, Peiretti F, Block KL, Hershey JW: Characterization of eIF3k: a newly discovered subunit of mammalian translation initiation factor elF3. Eur J Biochem. 2003, 270: 4133-4139.
Mazumdar A, Wang RA, Mishra SK, Adam L, Bagheri-Yarmand R, Mandal M, Vadlamudi RK, Kumar R: Transcriptional repression of oestrogen receptor by metastasis-associated protein 1 corepressor. Nat Cell Biol. 2001, 3: 30-37.
McKeegan KS, Debieux CM, Boulon S, Bertrand E, Watkins NJ: A dynamic scaffold of pre-snoRNP factors facilitates human box C/D snoRNP assembly. Mol Cell Biol. 2007, 27: 6782-6793.
Methot N, Rom E, Olsen H, Sonenberg N: The human homologue of the yeast Prt1 protein is an integral part of the eukaryotic initiation factor 3 complex and interacts with p170. J Biol Chem. 1997, 272: 1110-1116.
Meurer M, Degitz K: [Antiphospholipid antibodies]. Hautarzt. 1992, 43: 111-113.
Moraes KC, Quaresma AJ, Maehnss K, Kobarg J: Identification and characterization of proteins that selectively interact with isoforms of the mRNA binding protein AUF1 (hnRNP D). Biol Chem. 2003, 384: 25-37.
Moroianu J, Blobel G, Radu A: RanGTP-mediated nuclear export of karyopherin alpha involves its interaction with the nucleoporin Nup153. Proc Natl Acad Sci USA. 1997, 94: 9699-9704.
Morris-Desbois C, Rety S, Ferro M, Garin J, Jalinot P: The human protein HSPC021 interacts with Int-6 and is associated with eukaryotic translation initiation factor 3. J Biol Chem. 2001, 276: 45988-45995.
Munnia A, Schutz N, Romeike BF, Maldener E, Glass B, Maas R, Nastainczyk W, Feiden W, Fischer U, Meese E: Expression, cellular distribution and protein binding of the glioma amplified sequence (GAS41), a highly conserved putative transcription factor. Oncogene. 2001, 20: 4853-4863.
Musahl C, Schulte D, Burkhart R, Knippers R: A human homologue of the yeast replication protein Cdc21. Interactions with other Mcm proteins. Eur J Biochem. 1995, 230: 1096-1101.
Nagoshi E, Yoneda Y: Dimerization of sterol regulatory element-binding protein 2 via the helix-loop-helix-leucine zipper domain is a prerequisite for its nuclear localization mediated by importin beta. Mol Cell Biol. 2001, 21: 2779-2789.
Nakielny S, Shaikh S, Burke B, Dreyfuss G: Nup153 is an M9-containing mobile nucleoporin with a novel Ran-binding domain. Embo J. 1999, 18: 1982-1995.
Natsiuk MV, Chekman IS: [Level of nicotinamide coenzymes in the liver and myocardium of rats poisoned with dichlorethane]. Biull Eksp Biol Med. 1975, 79: 58-60.
Neish AS, Anderson SF, Schlegel BP, Wei W, Parvin JD: Factors associated with the mammalian RNA polymerase II holoenzyme. Nucleic Acids Res. 1998, 26: 847-853.
Ng HH, Zhang Y, Hendrich B, Johnson CA, Turner BM, Erdjument-Bromage H, Tempst P, Reinberg D, Bird A: MBD2 is a transcriptional repressor belonging to the MeCP1 histone deacetylase complex. Nat Genet. 1999, 23: 58-61.
Nicol SM, Causevic M, Prescott AR, Fuller-Pace FV: The nuclear DEAD box RNA helicase p68 interacts with the nucleolar protein fibrillarin and colocalizes specifically in nascent nucleoli during telophase. Exp Cell Res. 2000, 257: 272-280.
Nicolas E, Ait-Si-Ali S, Trouche D: The histone deacetylase HDAC3 targets RbAp48 to the retinoblastoma protein. Nucleic Acids Res. 2001, 29: 3131-3136.
Nicolas E, Morales V, Magnaghi-Jaulin L, Harel-Bellan A, Richard-Foy H, Trouche D: RbAp48 belongs to the histone deacetylase complex that associates with the retinoblastoma protein. J Biol Chem. 2000, 275: 9797-9804.
Nie Z, Yan Z, Chen EH, Sechi S, Ling C, Zhou S, Xue Y, Yang D, Murray D, Kanakubo E, et al: Novel SWI/SNF chromatin-remodeling complexes contain a mixed-lineage leukemia chromosomal translocation partner. Mol Cell Biol. 2003, 23: 2942-2952.
Nikolaev AY, Papanikolaou NA, Li M, Qin J, Gu W: Identification of a novel BRMS1-homologue protein p40 as a component of the mSin3A/p33(ING1b)/HDAC1 deacetylase complex. Biochem Biophys Res Commun. 2004, 323: 1216-1222.
Ohnishi M, Tanaka Y, Tutui T, Bann S: Extensive malignant schwannoma of the mandibular nerve. Case report. Int J Oral Maxillofac Surg. 1992, 21: 280-281.
Olave I, Wang W, Xue Y, Kuo A, Crabtree GR: Identification of a polymorphic, neuron-specific chromatin remodeling complex. Genes Dev. 2002, 16: 2509-2517.
Otsuki T, Furukawa Y, Ikeda K, Endo H, Yamashita T, Shinohara A, Iwamatsu A, Ozawa K, Liu JM: Fanconi anemia protein, FANCA, associates with BRG1, a component of the human SWI/SNF complex. Hum Mol Genet. 2001, 10: 2651-2660.
Park J, Wood MA, Cole MD: BAF53 forms distinct nuclear complexes and functions as a critical c-Myc-interacting nuclear cofactor for oncogenic transformation. Mol Cell Biol. 2002, 22: 1307-1316.
Phelan ML, Sif S, Narlikar GJ, Kingston RE: Reconstitution of a core chromatin remodeling complex from SWI/SNF subunits. Mol Cell. 1999, 3: 247-253.
Quintana DG, Thome KC, Hou ZH, Ligon AH, Morton CC, Dutta A: ORC5L, a new member of the human origin recognition complex, is deleted in uterine leiomyomas and malignant myeloid diseases. J Biol Chem. 1998, 273: 27137-27145.
Reichman TW, Muniz LC, Mathews MB: The RNA binding protein nuclear factor 90 functions as both a positive and negative regulator of gene expression in mammalian cells. Mol Cell Biol. 2002, 22: 343-356.
Rountree MR, Bachman KE, Baylin SB: DNMT1 binds HDAC2 and a new co-repressor, DMAP1, to form a complex at replication foci. Nat Genet. 2000, 25: 269-277.
Rozenblatt-Rosen O, Hughes CM, Nannepaga SJ, Shanmugam KS, Copeland TD, Guszczynski T, Resau JH, Meyerson M: The parafibromin tumor suppressor protein is part of a human Paf1 complex. Mol Cell Biol. 2005, 25: 612-620.
Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, Berriz GF, Gibbons FD, Dreze M, Ayivi-Guedehoussou N, et al: Towards a proteome-scale map of the human protein-protein interaction network. Nature. 2005, 437: 1173-1178.
Saito M, Ishikawa F: The mCpG-binding domain of human MBD3 does not bind to mCpG but interacts with NuRD/Mi2 components HDAC1 and MTA2. J Biol Chem. 2002, 277: 35434-35439.
Sakai H, Urano T, Ookata K, Kim MH, Hirai Y, Saito M, Nojima Y, Ishikawa F: MBD3 and HDAC1, two components of the NuRD complex, are localized at Aurora-A-positive centrosomes in M phase. J Biol Chem. 2002, 277: 48714-48723.
Scheffzek K, Ahmadian MR, Kabsch W, Wiesmuller L, Lautwein A, Schmitz F, Wittinghofer A: The Ras-RasGAP complex: structural basis for GTPase activation and its loss in oncogenic Ras mutants. Science. 1997, 277: 333-338.
Schmidt DR, Schreiber SL: Molecular association between ATR and two components of the nucleosome remodeling and deacetylating complex, HDAC2 and CHD4. Biochemistry. 1999, 38: 14711-14717.
Schmiesing JA, Ball AR, Gregson HC, Alderton JM, Zhou S, Yokomori K: Identification of two distinct human SMC protein complexes involved in mitotic chromosome dynamics. Proc Natl Acad Sci USA. 1998, 95: 12906-12911.
Schmiesing JA, Gregson HC, Zhou S, Yokomori K: A human condensin complex containing hCAP-C-hCAP-E and CNAP1, a homolog of Xenopus XCAP-D2, colocalizes with phosphorylated histone H3 during the early stage of mitotic chromosome condensation. Mol Cell Biol. 2000, 20: 6996-7006.
Schulte D, Richter A, Burkhart R, Musahl C, Knippers R: Properties of the human nuclear protein p85Mcm. Expression, nuclear localization and interaction with other Mcm proteins. Eur J Biochem. 1996, 235: 144-151.
Schwerk C, Prasad J, Degenhardt K, Erdjument-Bromage H, White E, Tempst P, Kidd VJ, Manley JL, Lahti JM, Reinberg D: ASAP, a novel protein complex involved in RNA processing and apoptosis. Mol Cell Biol. 2003, 23: 2981-2990.
Shi Y, Downes M, Xie W, Kao HY, Ordentlich P, Tsai CC, Hon M, Evans RM: Sharp, an inducible cofactor that integrates nuclear receptor repression and activation. Genes Dev. 2001, 15: 1140-1151.
Sif S, Saurin AJ, Imbalzano AN, Kingston RE: Purification and characterization of mSin3A-containing Brg1 and hBrm chromatin remodeling complexes. Genes Dev. 2001, 15: 603-618.
Singh BB, Patel HH, Roepman R, Schick D, Ferreira PA: The zinc finger cluster domain of RanBP2 is a specific docking site for the nuclear export factor, exportin-1. J Biol Chem. 1999, 274: 37370-37378.
Stelzl U, Worm U, Lalowski M, Haenig C, Brembeck FH, Goehler H, Stroedicke M, Zenkner M, Schoenherr A, Koeppen S, et al: A human protein-protein interaction network: a resource for annotating the proteome. Cell. 2005, 122: 957-968.
Sullivan EK, Weirich CS, Guyon JR, Sif S, Kingston RE: Transcriptional activation domains of human heat shock factor 1 recruit human SWI/SNF. Mol Cell Biol. 2001, 21: 5826-5837.
Takezawa S, Yokoyama A, Okada M, Fujiki R, Iriyama A, Yanagi Y, Ito H, Takada I, Kishimoto M, Miyajima A, et al: A cell cycle-dependent co-repressor mediates photoreceptor cell-specific nuclear receptor function. Embo J. 2007, 26: 764-774.
Taplick J, Kurtev V, Kroboth K, Posch M, Lechner T, Seiser C: Homo-oligomerisation and nuclear localisation of mouse histone deacetylase 1. J Mol Biol. 2001, 308: 27-38.
Tatematsu KI, Yamazaki T, Ishikawa F: MBD2–MBD3 complex binds to hemi-methylated DNA and forms a complex containing DNMT1 at the replication foci in late S phase. Genes Cells. 2000, 5: 677-688.
Tickenbrock L, Cramer J, Vetter IR, Muller O: The coiled coil region (amino acids 129–250) of the tumor suppressor protein adenomatous polyposis coli (APC). Its structure and its interaction with chromosome maintenance region 1 (Crm-1). J Biol Chem. 2002, 277: 32332-32338.
Tong JK, Hassig CA, Schnitzler GR, Kingston RE, Schreiber SL: Chromatin deacetylation by an ATP-dependent nucleosome remodelling complex. Nature. 1998, 395: 917-921.
Tsai SC, Valkov N, Yang WM, Gump J, Sullivan D, Seto E: Histone deacetylase interacts directly with DNA topoisomerase II. Nat Genet. 2000, 26: 349-353.
Underhill C, Qutob MS, Yee SP, Torchia J: A novel nuclear receptor corepressor complex, N-CoR, contains components of the mammalian SWI/SNF complex and the corepressor KAP-1. J Biol Chem. 2000, 275: 40463-40470.
Vlag van der J, Otte AP: Transcriptional repression mediated by the human polycomb-group protein EED involves histone deacetylation. Nat Genet. 1999, 23: 474-478.
Vashee S, Simancek P, Challberg MD, Kelly TJ: Assembly of the human origin recognition complex. J Biol Chem. 2001, 276: 26666-26673.
Wang W, Cote J, Xue Y, Zhou S, Khavari PA, Biggar SR, Muchardt C, Kalpana GV, Goff SP, Yaniv M, et al: Purification and biochemical heterogeneity of the mammalian SWI-SNF complex. Embo J. 1996, 15: 5370-5382.
Watkins NJ, Dickmanns A, Luhrmann R: Conserved stem II of the box C/D motif is essential for nucleolar localization and is required, along with the 15.5 K protein, for the hierarchical assembly of the box C/D snoRNP. Mol Cell Biol. 2002, 22: 8342-8352.
Will CL, Urlaub H, Achsel T, Gentzel M, Wilm M, Luhrmann R: Characterization of novel SF3b and 17S U2 snRNP proteins, including a human Prp5p homologue and an SF3b DEAD-box protein. Embo J. 2002, 21: 4978-4988.
Wilson BJ, Bates GJ, Nicol SM, Gregory DJ, Perkins ND, Fuller-Pace FV: The p68 and p72 DEAD box RNA helicases interact with HDAC1 and repress transcription in a promoter-specific manner. BMC Mol Biol. 2004, 5: 11-
Xia ZB, Anderson M, Diaz MO, Zeleznik-Le NJ: MLL repression domain interacts with histone deacetylases, the polycomb group proteins HPC2 and BMI-1, and the corepressor C-terminal-binding protein. Proc Natl Acad Sci USA. 2003, 100: 8342-8347.
Xue Y, Canman JC, Lee CS, Nie Z, Yang D, Moreno GT, Young MK, Salmon ED, Wang W: The human SWI/SNF-B chromatin-remodeling complex is related to yeast rsc and localizes at kinetochores of mitotic chromosomes. Proc Natl Acad Sci USA. 2000, 97: 13015-13020.
Yabuta N, Kajimura N, Mayanagi K, Sato M, Gotow T, Uchiyama Y, Ishimi Y, Nojima H: Mammalian Mcm2/4/6/7 complex forms a toroidal structure. Genes Cells. 2003, 8: 413-421.
Yanagida M, Hayano T, Yamauchi Y, Shinkawa T, Natsume T, Isobe T, Takahashi N: Human fibrillarin forms a sub-complex with splicing factor 2-associated p32, protein arginine methyltransferases, and tubulins alpha 3 and beta 1 that is independent of its association with preribosomal ribonucleoprotein complexes. J Biol Chem. 2004, 279: 1607-1614.
Yang L, Mei Q, Zielinska-Kwiatkowska A, Matsui Y, Blackburn ML, Benedetti D, Krumm AA, Taborsky GJ, Chansky HA: An ERG (ets-related gene)-associated histone methyltransferase interacts with histone deacetylases 1/2 and transcription co-repressors mSin3A/B. Biochem J. 2003, 369: 651-657.
Yang WM, Yao YL, Sun JM, Davie JR, Seto E: Isolation and characterization of cDNAs corresponding to an additional member of the human histone deacetylase gene family. J Biol Chem. 1997, 272: 28001-28007.
Yao YL, Yang WM: The metastasis-associated proteins 1 and 2 form distinct protein complexes with histone deacetylase activity. J Biol Chem. 2003, 278: 42560-42568.
Yao YL, Yang WM, Seto E: Regulation of transcription factor YY1 by acetylation and deacetylation. Mol Cell Biol. 2001, 21: 5979-5991.
Yaseen NR, Blobel G: Two distinct classes of Ran-binding sites on the nucleoporin Nup-358. Proc Natl Acad Sci USA. 1999, 96: 5516-5521.
Yaseen NR, Blobel G: GTP hydrolysis links initiation and termination of nuclear import on the nucleoporin nup358. J Biol Chem. 1999, 274: 26493-26502.
Yasui D, Miyano M, Cai S, Varga-Weisz P, Kohwi-Shigematsu T: SATB1 targets chromatin remodelling to regulate genes over long distances. Nature. 2002, 419: 641-645.
Yoshida T, Kokura K, Makino Y, Ossipow V, Tamura T: Heterogeneous nuclear RNA-ribonucleoprotein F binds to DNA via an oligo(dG)-motif and is associated with RNA polymerase II. Genes Cells. 1999, 4: 707-719.
You A, Tong JK, Grozinger CM, Schreiber SL: CoREST is an integral component of the CoREST- human histone deacetylase complex. Proc Natl Acad Sci USA. 2001, 98: 1454-1458.
You Z, Ishimi Y, Masai H, Hanaoka F: Roles of Mcm7 and Mcm4 subunits in the DNA helicase activity of the mouse Mcm4/6/7 complex. J Biol Chem. 2002, 277: 42471-42479.
You Z, Komamura Y, Ishimi Y: Biochemical analysis of the intrinsic Mcm4–Mcm6–mcm7 DNA helicase activity. Mol Cell Biol. 1999, 19: 8003-8015.
Young PG, Attardi G: Characterization of double-stranded RNA from HeLa cell mitochondria. Biochem Biophys Res Commun. 1975, 65: 1201-1207.
Zhang H, Saitoh H, Matunis MJ: Enzymes of the SUMO modification pathway localize to filaments of the nuclear pore complex. Mol Cell Biol. 2002, 22: 6498-6508.
Zhang H, Shi X, Paddon H, Hampong M, Dai W, Pelech S: B23/nucleophosmin serine 4 phosphorylation mediates mitotic functions of polo-like kinase 1. J Biol Chem. 2004, 279: 35726-35734.
Zhang Y, Dufau ML: Silencing of transcription of the human luteinizing hormone receptor gene by histone deacetylase-mSin3A complex. J Biol Chem. 2002, 277: 33431-33438.
Zhang Y, Dufau ML: Dual mechanisms of regulation of transcription of luteinizing hormone receptor gene by nuclear orphan receptors and histone deacetylase complexes. J Steroid Biochem Mol Biol. 2003, 85: 401-414.
Zhang Y, Iratni R, Erdjument-Bromage H, Tempst P, Reinberg D: Histone deacetylases and SAP18, a novel polypeptide, are components of a human Sin3 complex. Cell. 1997, 89: 357-364.
Zhang Y, LeRoy G, Seelig HP, Lane WS, Reinberg D: The dermatomyositis-specific autoantigen Mi2 is a component of a complex containing histone deacetylase and nucleosome remodeling activities. Cell. 1998, 95: 279-289.
Zhang Y, Ng HH, Erdjument-Bromage H, Tempst P, Bird A, Reinberg D: Analysis of the NuRD subunits reveals a histone deacetylase core complex and a connection with DNA methylation. Genes Dev. 1999, 13: 1924-1935.
Zhang Y, Sun ZW, Iratni R, Erdjument-Bromage H, Tempst P, Hampsey M, Reinberg D: SAP30, a novel protein conserved between human and yeast, is a component of a histone deacetylase complex. Mol Cell. 1998, 1: 1021-1031.
Zhao K, Wang W, Rando OJ, Xue Y, Swiderek K, Kuo A, Crabtree GR: Rapid and phosphoinositol-dependent binding of the SWI/SNF-like BAF complex to chromatin after T lymphocyte receptor signaling. Cell. 1998, 95: 625-636.
Zhou J, Chau CM, Deng Z, Shiekhattar R, Spindler MP, Schepers A, Lieberman PM: Cell cycle regulation of chromatin at an origin of DNA replication. Embo J. 2005, 24: 1406-1417.
Zhou K, Choe KT, Zaidi Z, Wang Q, Mathews MB, Lee CG: RNA helicase A interacts with dsDNA and topoisomerase IIalpha. Nucleic Acids Res. 2003, 31: 2253-2260.
Zhupan VF: [Causes of inflammation of an artificial esophagus and its treatment]. Vestn Khir Im I I Grek. 1975, 114: 83-87.
Dehe PM, Dichtl B, Schaft D, Roguev A, Pamblanco M, Lebrun R, Rodriguez-Gil A, Mkandawire M, Landsberg K, Shevchenko A, et al: Protein interactions within the Set1 complex and their roles in the regulation of histone 3 lysine 4 methylation. J Biol Chem. 2006, 281: 35404-35412.
Dehe PM, Geli V: The multiple faces of Set1. Biochem Cell Biol. 2006, 84: 536-548.
Reinberg D, Sims RJ: de FACTo nucleosome dynamics. J Biol Chem. 2006, 281: 23297-23301.
Murr R, Vaissiere T, Sawan C, Shukla V, Herceg Z: Orchestration of chromatin-based processes: mind the TRRAP. Oncogene. 2007, 26: 5358-5372.
Mohrmann L, Verrijzer CP: Composition and functional specificity of SWI2/SNF2 class chromatin remodeling complexes. Biochim Biophys Acta. 2005, 1681: 59-73.
Cavellan E, Asp P, Percipalle P, Farrants AK: The WSTF-SNF2h chromatin remodeling complex interacts with several nuclear proteins in transcription. J Biol Chem. 2006, 281: 16264-16271.
Silverstein RA, Ekwall K: Sin3: a flexible regulator of global gene expression and genome stability. Curr Genet. 2005, 47: 1-17.
Tyagi M, Karn J: CBF-1 promotes transcriptional silencing during the establishment of HIV-1 latency. Embo J. 2007, 26: 4985-4995.
Denslow SA, Wade PA: The human Mi-2/NuRD complex and gene regulation. Oncogene. 2007, 26: 5433-5438.
Cismasiu VB, Paskaleva E, Suman Daya S, Canki M, Duus K, Avram D: BCL11B is a general transcriptional repressor of the HIV-1 long terminal repeat in T lymphocytes through recruitment of the NuRD complex. Virology. 2008, 380: 173-181.
Boeke J, Ammerpohl O, Kegel S, Moehren U, Renkawitz R: The minimal repression domain of MBD2b overlaps with the methyl-CpG-binding domain and binds directly to Sin3A. J Biol Chem. 2000, 275: 34963-34967.
Brackertz M, Gong Z, Leers J, Renkawitz R: p66alpha and p66beta of the Mi-2/NuRD complex mediate MBD2 and histone interaction. Nucleic Acids Res. 2006, 34: 397-406.
Dobosy JR, Selker EU: Emerging connections between DNA methylation and histone acetylation. Cell Mol Life Sci. 2001, 58: 721-727.
Kransdorf EP, Wang SZ, Zhu SZ, Langston TB, Rupon JW, Ginder GD: MBD2 is a critical component of a methyl cytosine-binding protein complex isolated from primary erythroid cells. Blood. 2006, 108: 2836-2845.
Kuzmichev A, Nishioka K, Erdjument-Bromage H, Tempst P, Reinberg D: Histone methyltransferase activity associated with a human multiprotein complex containing the Enhancer of Zeste protein. Genes Dev. 2002, 16: 2893-2905.
Hirano T: At the heart of the chromosome: SMC proteins in action. Nat Rev Mol Cell Biol. 2006, 7: 311-322.
Bhat MA, Philp AV, Glover DM, Bellen HJ: Chromatid segregation at anaphase requires the barren product, a novel chromosome-associated protein that interacts with Topoisomerase II. Cell. 1996, 87: 1103-1114.
Forsburg SL: Eukaryotic MCM proteins: beyond replication initiation. Microbiol Mol Biol Rev. 2004, 68: 109-131.
Sasaki T, Gilbert DM: The many faces of the origin recognition complex. Curr Opin Cell Biol. 2007, 19: 337-343.
Antonin W, Ellenberg J, Dultz E: Nuclear pore complex assembly through the cell cycle: regulation and membrane organization. FEBS Lett. 2008, 582: 2004-2016.
Wagner N, Krohne G: LEM-Domain proteins: new insights into lamin-interacting proteins. Int Rev Cytol. 2007, 261: 1-46.
Bridger JM, Foeger N, Kill IR, Herrmann H: The nuclear lamina. Both a structural framework and a platform for genome organization. Febs J. 2007, 274: 1354-1361.
Kiernan RE, Vanhulle C, Schiltz L, Adam E, Xiao H, Maudoux F, Calomme C, Burny A, Nakatani Y, Jeang KT, et al: HIV-1 tat transcriptional activity is regulated by acetylation. Embo J. 1999, 18: 6106-6118.
Bres V, Kiernan R, Emiliani S, Benkirane M: Tat acetyl-acceptor lysines are important for human immunodeficiency virus type-1 replication. J Biol Chem. 2002, 277: 22215-22221.
Bukrinsky M: SNFing HIV transcription. Retrovirology. 2006, 3: 49-
Pumfery A, Deng L, Maddukuri A, de la Fuente C, Li H, Wade JD, Lambert P, Kumar A, Kashanchi F: Chromatin remodeling and modification during HIV-1 Tat-activated transcription. Curr HIV Res. 2003, 1: 343-362.
Coull JJ, Romerio F, Sun JM, Volker JL, Galvin KM, Davie JR, Shi Y, Hansen U, Margolis DM: The human factors YY1 and LSF repress the human immunodeficiency virus type 1 long terminal repeat via recruitment of histone deacetylase 1. J Virol. 2000, 74: 6790-6799.
Critchfield JW, Ho O, Roberts BD, Van Lint C, Verdin E, Butera ST: Isoquinolinesulphonamide derivatives inhibit transcriptional elongation of human immunodeficiency virus type 1 RNA in a promyelocytic model of latency. Antivir Chem Chemother. 1999, 10: 275-284.
He G, Margolis DM: Counterregulation of chromatin deacetylation and histone deacetylase occupancy at the integrated promoter of human immunodeficiency virus type 1 (HIV-1) by the HIV-1 repressor YY1 and HIV-1 activator Tat. Mol Cell Biol. 2002, 22: 2965-2973.
Imai K, Okamoto T: Transcriptional repression of human immunodeficiency virus type 1 by AP-4. J Biol Chem. 2006, 281: 12495-12505.
Jiang G, Espeseth A, Hazuda DJ, Margolis DM: c-Myc and Sp1 contribute to proviral latency by recruiting histone deacetylase 1 to the human immunodeficiency virus type 1 promoter. J Virol. 2007, 81: 10914-10923.
Kiefer HL, Hanley TM, Marcello JE, Karthik AG, Viglianti GA: Retinoic acid inhibition of chromatin remodeling at the human immunodeficiency virus type 1 promoter. Uncoupling of histone acetylation and chromatin remodeling. J Biol Chem. 2004, 279: 43604-43613.
Williams SA, Chen LF, Kwon H, Ruiz-Jarabo CM, Verdin E, Greene WC: NF-kappaB p50 promotes HIV latency through HDAC recruitment and repression of transcriptional initiation. Embo J. 2006, 25: 139-149.
Deng L, de la Fuente C, Fu P, Wang L, Donnelly R, Wade JD, Lambert P, Li H, Lee CG, Kashanchi F: Acetylation of HIV-1 Tat by CBP/P300 increases transcription of integrated HIV-1 genome and enhances binding to core histones. Virology. 2000, 277: 278-295.
Ott M, Schnolzer M, Garnica J, Fischle W, Emiliani S, Rackwitz HR, Verdin E: Acetylation of the HIV-1 Tat protein by p300 is important for its transcriptional activity. Curr Biol. 1999, 9: 1489-1492.
Kalverda B, Roling MD, Fornerod M: Chromatin organization in relation to the nuclear periphery. FEBS Lett. 2008, 582: 2017-2022.
Shaklai S, Amariglio N, Rechavi G, Simon AJ: Gene silencing at the nuclear periphery. Febs J. 2007, 274: 1383-1392.
Ye Q, Callebaut I, Pezhman A, Courvalin JC, Worman HJ: Domain-specific interactions of human HP1-type chromodomain proteins and inner nuclear membrane protein LBR. J Biol Chem. 1997, 272: 14983-14989.
Dorner D, Gotzmann J, Foisner R: Nucleoplasmic lamins and their interaction partners, LAP2alpha, Rb, and BAF, in transcriptional regulation. Febs J. 2007, 274: 1362-1373.
Gautier VW, Sheehy N, Duffy M, Hashimoto K, Hall WW: Direct interaction of the human I-mfa domain-containing protein, HIC, with HIV-1 Tat results in cytoplasmic sequestration and control of Tat activity. Proc Natl Acad Sci USA. 2005, 102: 16362-16367.
Acknowledgements
This work was supported by the National Virus Reference Laboratory (NVRL), University College Dublin, Dublin, Ireland and by the UCD-SMMS Research Support Scheme (2007), University College Dublin, Dublin, Ireland.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
VG conceived and designed the study, planned and coordinated its execution, conducted experimental procedures, data interpretation and in silico analysis, and drafted the manuscript. LG performed GST-pull down/Western blotting experiments. NOD participated in the experimental design and performed LC MS/MS and peptide and protein identification. SP participated in the experimental design and supervised the proteomic analysis. NS participated in the interpretation of results and final editing of the manuscript. WWH supervised the study design, execution, analysis and revised the manuscript critically.
Electronic supplementary material
12977_2008_1052_MOESM1_ESM.xls
Additional file 1: List of the 183 components of the Tat nuclear interactome in Jurkat identified by GST pull-down combined with LC-MS/MS. Amino acid coverage (Coverage%), number of MS/MS peptides used for the identification (MS/MS peptide no), TurboSEQUEST score, Genebank accession number and gene ontology (GO) analysis (cellular process) for each identification are indicated. (XLS 96 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Gautier, V.W., Gu, L., O'Donoghue, N. et al. In vitro nuclear interactome of the HIV-1 Tat protein. Retrovirology 6, 47 (2009). https://doi.org/10.1186/1742-4690-6-47
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1742-4690-6-47