Abstract
While the use of high-resolution molecular typing in routine matching of h uman l eukocyte a ntigens (HLA) is expected to improve unrelated donor selection and transplant outcome, the genetic complexity of HLA still makes the current methodology limited and laborious. Pyrosequencing™ is a gel-free, sequencing-by-synthesis method. In a Pyrosequencing reaction, nucleotide incorporation proceeds sequentially along each DNA template at a given n ucleotide d ispensation o rder (NDO) that is programmed into a pyrosequencer. Here we describe the design of a NDO that generates a pyrogram unique for any given allele or combination of alleles. We present examples of unique pyrograms generated from each of two heterozygous HLA templates, which would otherwise remain cis/trans ambiguous using standard s equencing b ased t yping (SBT) method. In addition, we display representative data that demonstrate long read and linear signal generation. These features are prerequisite of high-resolution typing and automated data analysis. In conclusion Pyrosequencing is a one-step method for high resolution DNA typing.
Similar content being viewed by others
Introduction
Solid organ transplantation and allogeneic stem cell transplantation currently represent a common treatment for end-stage organ failure and several hematological and non-hematological malignances. Matching of patient and unrelated donor for h uman l eukocyte a ntigen (HLA) molecules significantly decreases the probability of graft rejection, graft vs. host disease and transplant-related mortality [1]. However, the extensive diversity of the HLA genes makes the identification of matched donors extremely challenging. Although, in several instances it might not be feasible to identify perfect matches, algorithms have been developed that allow identification of likely histocompatibility based on the molecular definition of individual alleles [2, 3]. This algorithm grades mismatches according to the number of variant epitopes present between donor and recipient. As histocompatibility is inversely correlated with number of mismatches it is likely that sequence-based information that provides the definitive information about HLA allele identity will become increasingly important in the future. High-resolution information about HLA alleles identity is best achieved using sequencing-based methodology that could be performed using high-throughput automated systems [4]. Although significant advancement has been made in resolution, automation, throughput and data analysis in DNA sequencing and other polymorphism analysis techniques, the search continues for more efficient methods that could resolve cis/trans ambiguities in highly polymorphic genetic systems such as HLA genes. Currently, commonly used HLA molecular typing methods include s equence s pecific o ligonucleotide p robes (SSOP), p olymerase c hain r eaction (PCR) using s equence s pecific p rimers (SSP) and sequence based typing (SBT) [5]. Among them, SSOP solely exploits DNA hybridization and, therefore, results in the most cis/trans ambiguities. SSP can solve ambiguous combinations if primers are designed to cover the geneomic region where the ambiguity is present. In this case, amplification of the genomic region framed by two primers assures the occurrence in cis of these two regions. This strategy, however, requires a large number of primers to reach a desired resolution and cover various combinations of ambiguous sites within HLA loci. SBT provides by far the highest resolution and currently represents the golden standard for high resolution DNA typing and novel allele discovery. In addition, recent advances made possible to perform SBT at a high throughput level in routine HLA typing laboratories [4]. The biggest challenge that SBT of HLA alleles incurs is the resolution of intrinsic cis/trans ambiguities that cannot be solved by SBT unless time consuming cloning of individual genes is performed [6]. This is because nucleotide incorporation proceeds simultaneously along all DNA templates in a SBT reaction [7].
Pyrosequencing™ [9–11] is a real-time, sequencing by synthesis method catalyzed by four kinetically well-balanced enzymes, DNA polymerase, ATP sulfurylase, luciferase, and apyrase. It fundamentally differs from Sanger's sequencing method in the order of nucleotide incorporation. Each nucleotide is dispensed and tested individually for its incorporation into a nascent DNA template. Each incorporation event is accompanied by release of pyrophosphate (PPi) in a quantity equimolar to the amount of nucleotide incorporated. ATP sulfurylase quantitatively converts PPi to ATP in the presence of adenosine 5' phosphosulfate. ATP then drives the luciferase-mediated conversion of luciferin to oxyluciferin that generates visible light in amounts that are proportional to the amount of ATP. The light is detected by a charge coupled device (CCD) camera and displayed as a peak in a pyrogram™. Each peak height is proportional to the number of nucleotides incorporated. Unincorporated dNTP and excess ATP are continuously degraded by Apyrase. After the degradation is completed, the next dNTP is added and a new Pyrosequencing cycle is started. As the process continues, the complementary DNA strand is built up. To pyrosequence an unknown DNA sequence, a cyclic n ucleotide d ispensation o rder (NDO) is generally used. As a result of each cycle of dATP, dGTP, dCTP and dTTP dispensation, one of the four dNTPs is incorporated into the DNA template while the other dNTPs are degraded by Apyrase. When a DNA sequence is known, non-cyclic NDOs can be programmed with predictable pyrograms. Nucleotide sequence is determined from the order of nucleotide dispensation and peak height in the pyrogram.
Based on the programmable nucleotide incorporation feature of Pyrosequencing, we set out to optimize Pyrosequencing for high resolution HLA DNA typing. Here we describe the design of NDO that generates a pyrogram that is unique for any given allele or combination of alleles. We present unique pyrograms generated from each of the heterozygous HLA templates that would otherwise be cis/trans ambiguous using s equencing b ased t yping (SBT) methods. We also present representative data that demonstrate long read and linear signal generation. These features are prerequisite of high-resolution typing and automated data analysis. In conclusion, Pyrosequencing can be used as a one-step method for high resolution DNA typing and could be applied in several settings spanning from HLA typing in support of donor/recipient selection to become a complement to comprehensive immunogenetic profiling in several clinical setting where other aspects of immune polymorphism need to be explored [8].
Results
Design of n ucleotide d ispensation o rder (NDO) that generates unique pyrogram for any allele or combination of alleles
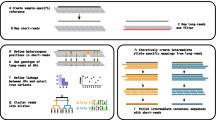
Two types of nucleotide dispensation can be used to pyrosequence a homozygous HLA template. An in-phase dispensation results in incorporation of nucleotides into all templates at the same base pair position(s). A negative dispensation results in no incorporation of any nucleotide, generating background signal (zero peak) only. Introducing negative dispensations at different positions results in different pyrograms from the same homozygous template [11]. In addition to in-phase and negative dispensations, it is possible to exploit out-of-phase dispensation to pyrosequence heterozygous DNA templates. Out-of-phase dispensation results in nucleotide incorporation along one allele, which put the sequencing reaction ahead of the other allele. Nucleotide incorporation can become in-phase again at various downstream positions, which can be controlled by NDO. Figure 1 shows five NDOs designed to sequence heterozygous genomic regions of the HLA-class II locus, DRB1. In this case, the goal is to the differentiate DRB1*11011, 13011 (black bars) combination from the DRB1*0319, 1320 (Red bars) whose sequences are only different at positions 5'-298-299-3'. All NDOs start with nucleotide incorporation at position 5'-286-3' and end at or after nucleotide incorporation at position 5'-299-3' of both alleles. Each pyrogram peak represents the sum of nucleotide incorporation at each nucleotide dispensation step, into all DNA templates in the same reaction mixture in either in-phase or out-of-phase fashion. NDO 1 requires the least number of nucleotide dispensations but it generates the same theoretical pyrogram from both templates. NDO 2, which is a typical cyclic NDO, generates unique theoretical pyrograms from each template but it requires more nucleotide dispensations than the other four NDOs, partly due to the inclusion of four negative dispensations. NDO 3 generates unique theoretical pyrograms from both templates at dispensations 5, 18 and 19 and, requires less nucleotide dispensations than NDO 2 because of the lack of negative dispensations. NDO 4 also generates unique theoretical pyrograms at three dispensations (dispensations 14, 15 and 16). In addition, it requires less nucleotide dispensations as compared to NDO 3 (18 Vs. 20). NDO 5 is the most effective. It generates the most number of differential theoretical peak heights at positions 12, 14, 16 and 18, and requires only 18 dispensations as well. Using this technique, on one hand, each DNA template sequence can generate different pyrograms. On the other hand, different DNA template sequences can generate identical pyrograms. Our NDO design software automatically compares a theoretical pyrogram generated by a given NDO from any homozygous or heterozygous HLA template sequence against that from all other homozygous or heterozygous HLA template sequences in the database (Shi et al, unpublished results). Among the NDOs that result in unique theoretical pyrogram, NDOs that produce that shorter theoretical number of dispensations are chosen.
NDO optimization. Sequences of DRB1*11011, 13011 and DRB1*1111, 1306 are shown on the top (5'-286-305-3'). NDO 1 to NDO 5 are designed to pyrosequence these two pairs of heterozygous templates. X-axis represents the NDO. G, A, T and C represent the dNTP that is dispensed at each. Alphabetical numbers represent the dispensation step. E.g. the first step dispenses dATP. Y-axis represents theoretical peak height, shown as number of nucleotide incorporated into the two molecules of template alleles at each nucleotide dispensation step. Black peaks represent signals generated from DRB1*11011, 13011. Red peaks represent signals generated from DRB1*1111, 1306.
Pyrosequencing resolves intrinsic s equencing b ased t yping (SBT) cis/trans ambiguity
Although high-resolution SBT of HLA allleles provides the highest resolution, it cannot effectively solve many intrinsic cis/trans ambiguities unless coupled with time consuming cloning of sequencing of individual clones. The sequence difference between the two heterozygous templates at position 5'-298-299-3' as described in Figure 1, for example, is a commonly encountered SBT ambiguous example. In an effort to solve this SBT ambiguity, we tested whether or not experimentally obtained pyrogram matched the theoretical pyrogram predicted by NDO 5. Figure 2 further illustrates NDO 5 step-wise. We chose to place the 3' end of the Pyrosequencing primer just upstream of another polymorphic site at 5'-286-3', designated reference polymorphic site. Out-of-phase NDO is designed at the very first nucleotide dispensation. T is incorporated in to template DRB1*110101 but not DRB1*130101. The pyrogram output, as shown in Figure 3, demonstrates differential peak heights at all four theoretically different positions. Using the 11th peak adjacent and upstream of the first differential peak (the 12th peak) as a normalizer, we could observe that peak height ratios at peaks 12, 14, 16 and 18 closely correlated with theoretical peak height ratios proposed in Figure 1 (IDO 5). The 12th peak deviates from the prediction by 18%. The 14th peak deviates by 13.5%. while both 16th and 18th peaks demonstrated deviations from prediction close to 0%. As an average the deviation from theoretical prediction was 7.9%. Figures 4 demonstrates another HLA-DRB1 SBT ambiguity, DRB1*030101, 130101 Vs. DRB1*0319, 1320, that Pyrosequencing can solve. Using the upstream adjacent peak (the 7th peak) as a normalizer, calculated peak height ratios at peaks 8, 11, 14, 15 and 16 also closely correlated with theoretical ratios with a deviation range between 0% to 16.7% and an average deviation of 6.7%. These two examples demonstrate how Pyrosequencing can be used to quantify differences and therefore identify the cis/trans conformation of ambiguous HLA heterozygous pairs that cannot be resolved by SBT.
The same NDO can generate unique pyrogram from DRB1*11011, 13011 and DRB1*1111, 1306. The two heterozygous templates used are DRB1*11011, 13011 and DRB1*1111, 1306. The SBT ambiguous positions are at 5'-298-299-3'. The Pyrosequencing primer is high lighted in purple color with its 3' end located at 5'-285-3'. Nucleotide incorporation starts at position 5'-286-3'. NDO is indicated on the left, from top to bottom. The nucleotides incorporated into each allele of template DRB1*11011, 13011 and each allele of template DRB1*11011, 13011 are highlighted in yellow at each nucleotide dispensation step except for the nucleotides involve in the ambiguities that are with the color red for  , green for
, green for  and black for
and black for  .
.
Pyrograms generated from two heterozygous DNA templates using the same NDO. Top pyrogram is generated from Pyrosequencing reaction in one well of a 96 well plate. Bottom pyrogram is generated from Pyrosequencing reaction in a separate well of a 96 well plate. The X axis of each pyrogram, from left to right, indicates the order of reagent addition. E represents enzyme. S represents substrate. The expected number of nucleotides incorporation into each pair of heterozygous DNA template and actual peak height are indicated for the normalizer peaks and differential peaks below each pyrogram. At the bottom of the figure is shown expected ratio of peak heights and the ratio of normalized peak heights (in shadowed area).
The same NDO can generate unique pyrogram from DRB1*030101, 130101 Vs. DRB1*0319, 1320. Top pyrogram is generated from Pyrosequencing reaction in one well of a 96 well plate. Bottom pyrogram is generated from Pyrosequencing reaction in a separate well of a 96 well plate. The X axis of each pyrogram, from left to right, indicates the order of reagent addition. E represents enzyme. S represents substrate. The expected number of nucleotides incorporation into each pair of heterozygous DNA template and actual peak height are indicated for the normalizer peaks and differential peaks below each Pyrogram. At the bottom of the figure is shown expected ratio of peak heights and the ratio of normalized peak heights (in shadowed area).
The general principles for the design of NDO can be summarized as follows: a primer is usually placed in proximity upstream of the reference polymorphic site chosen to be the one closest to the ambiguous polymorphic site to be investigated. The first nucleotide dispensation is usually out-of-phase. As a result, SBT ambiguity at one position is generally magnified into pyrograms differences at multiple peaks. This greatly enhances sensitivity and accuracy in detection of peak height differences. In our experience, ambiguities that cannot be solved by SBT within the HLA-DRB1 locus can be consistently solved by unique Pyrosequencing NDO (Wang et al, unpublished results).
Long read and linear signal generation facilitates automated data analysis
The ability to perform long Pyrosequencing reads (length of the genomic region investigated) is often necessary for reasonable throughput. It is essential for achieving high resolution when the reference polymorphic site downstream of the Pyrosequencing primer is distant from the ambiguous site. In addition to the optimized NDO, the PCR amplicons are designed to prevent background generation that could occur during a long Pyrosequencing reaction. The pyrogram shown in Figure 5 is an example of a linear and predictable reduction in signal generation with low background signal generation through 72 nucleotide dispensations. The background signal ranges from 2% to 11% with an average of 6% of the signals immediately upstream and downstream (Figure 6, bottom panel). The low background signal makes possible the discrimination of linear sequence-specific signals. One trend line is plotted against the signals generated from dATP (Figure 6, Top panel). A similar trend line is plotted against the signals generated from dGTP, dCTP and dTTP (Figure 6, middle panel). The dATP trend line is plotted separately because of its kinetics slightly faster than the other three dNTPs. Note that both trend lines indicate high confidence level with R2 greater than 95%. This linearity allows the extrapolation of the actual peak height relative to the dispensation point. Combining the two trend lines, the actual peak height can be extrapolated using the formula: "Extrapolated peak = [Split Height + (Slope × Disp#)] × Nuc#". The extrapolated peak heights only vary from the theoretical peak heights from 0% to 20%, averaging at 4.3%. This algorithm offers powerful aid to automated data analysis of Pyrosequencing results.
Linear signal generation through a 72 nucleotide dispensation Pyrosequencing run. Shown on the top in red is the Pyrosequencing primer sequence from 5' end to 3' end in the direction the red arrow points to. In this Pyrosequencing reaction, nucleotide incorporation into DRB1*1201, 1302 starts immediately downstream of the 3' end of Pyrosequencing primer and ends at the 3' end template sequence shown. The polymorphic positions are shown and underlined in blue. Pyrogram is shown below template sequence. Y-axis represents peak heights. X-axis represents NDO.
The percentage of each background peak over sequence-specific signal is calculated by dividing each background peak height with the average peak height of the proximal upstream and downstream sequence-specific peak height. Top panel depicts a trend line generated from all the "A" peak heights of the pyrogram shown in Figure 5. Middle panel depicts a trend line generated from all the "G, C and T" peak heights of the pyrogram shown in Figure 5. R2 are shown on the upper right corners of both panels. Bottom depicts theoretical and extrapolated peak heights generated from the NDO and peak heights shown in top panel. The formula that is used is: Extrapolated peak= [Split Height + (Slope × Disp#)] × Nuc#. Average background is the average of all background signals. MAX background is the highest background signal over sequence specific signal. MIN background is the lowest background signal over sequence specific signal.
Discussion
Pyrosequencing offers a new approach to data acquisition, analysis and identification of known and unknown (new) alleles, in particular in heterozygous conditions. This method may represent a useful tool to the screening and characterization of polymorphic genetic markers in several clinical or experimental settings [12–24]. In addition, Pyrosequencing has been applied for the study of gene expression [23] and could be a usefull complement to high throughput single nucleotide polymorphism identification system as a substitute to SBT [8, 24]. Here we propose that Pyrosequencing may confront the most challenging task of solving ambiguities in HLA typing by SBT in heterozygous conditions. Although its reading length is currently shorter than that routinely covered by SBT, automated dNTP dispensation could compensate for this limitation by controling simultaneous reactions in multiple wells using primers that anneal to different locations of the template DNA. In fact, a reading length of 70 to 100 nucleotides allows the high-resolution genotyping of Exon II of HLA-DRB1 (Wang et al, unpublished results). NDOs can also be designed to achieve higher throughput and lower genotyping resolution by introducing fewer numbers of out-of-phase dispensations (Wang et al, unpublished results). Without automatization, it is possible to process 96 to 384 wells PCR product by Pyrosequencing within 4 hours. Constant improvements in the chemistry for sample preparation for Pyrosequencing and Pyrosequencing [25–34] and the implementation of automation devices http://www.pyrosequencing.com it may be possible in the future to apply this technology directly for routine typing of HLA and other immune related genes characterized by extensive polymorphisms [8].
Materials and Methods
DNA samples
Genomic DNA samples were locally available or obtained from the International Histocompatibility Workshops (IHW) cell lines panel, UCLA interchange panel and samples.
PCR amplification
Each PCR amplification mixture of 50 μl contains 1 × PCR buffer (made in house), 2 mM MgCl2, 0.2 mM of each dNTP (purchased from Amersham Biosciences Inc.), 0.2 mM PCR primers, 2 U Taq DNA polymerase, and 250 ng genomic DNA. Either forward or reverse primer is biotinylated. PCR reaction starts with a 95°C denaturation for 5 minutes. This is followed with a 50-cycle thermal cycling. Each cycle is programmed to include 30 seconds denaturation at 95°C, 60 seconds annealing at appropriate temperature, and a 10 seconds final extention at 72°C. The PCR amplicon produced is enough for more 8 pyrosequencing reactions. The PCR amplicons used in this work is 286 bp containing Exon II and the flanking intron sequences.
Sample Preparation
Biotinylated PCR products are immobilized on streptavidin-coated Sepharose beads (Amersham Biosciences). 50 ul of Binding buffer (PyrosequencingAB) was added to the 50 ul of PCR product. Then 4 ul of streptavidin-coated Sepharose beads was added and the mixture was vigorously mixed at room temperature for 10 minutes. The streptavidin-coated Sepharose bead and PCR mixture is transferred to a filter plate (Amersham Biosciences) and the Binding buffer is removed by vacuum. The biotinylated DNA attached to the streptavidin-coated Sepharose beads was denaturated in 50 ul of Denaturation buffer (PyrosequencingAB) for 1 minute. The Denaturation buffer was removed by vacuum and DNA was washed twice in 150 ul of Wash Buffer (PyrosequencingAB). The DNA is resuspended in 50 ul of Annealing buffer (PyrosequencingAB).
Pyrosequencing Reaction
40 ul of well mixed DNA was transferred to a 96-well PSQ96 plate (PyrosequencingAB). The appropriate sequencing primer was added in a volume of 5 ul using a 3 uM stock solution, resulting in 45 ul reaction volume. The sequencing primer is allowed to anneal on a heat plate set for 80°C for 2 minutes. Samples are allowed to cool for 5 minutes at room temperature. Once samples have cooled down the plate in placed on the Pyrosequencer and the PSQ96 reagents are added to the SQA cartridge (PyrosequencingAB). NDO is automatically designed using software developed at Pel-Freez Clinical Systems. Pyrosequencing data output is quantified using Peak Height Determination Software v1.1 (PyrosequencingAB).
References
Petersdorf EW, Mickelson EM, Anasetti C, Martin PJ, Wool-frey AE, Hansen JA: Effect of HLA mismatches on the outcome of hematopoietic transplants. Curr Opin Immunol. 1999, 11: 521-526. 10.1016/S0952-7915(99)00016-3.
Duquesnoy RJ: HLA matchmaker: a molecularly based algorithm for histocompatibility determination. I Description of the algorithm. Hum Immunol. 2002, 63: 339-352. 10.1016/S0198-8859(02)00382-8.
Duquesnoy RJ, Marrari M: HLA matchmaker: a molecularly based algorithm for histocompatibility determination. II. Verification of the algorithm and determination of the relative immunogenicity of amino acid triplet-defined epitopes. Hum Immunol. 2002, 63: 353-363. 10.1016/S0198-8859(02)00381-6.
Adams SD, Krausa P, McGinnis M, Simonis TB, Stein J, Marincola FM: Practicality of high-throughput HLA sequencing based typing. ASHI Quarterly. 2001, 25: 54-57.
Erlich HA, Opelz G, Hansen JA: HLA DNA typing and transplantation. Immunity. 2001, 14: 347-356. 10.1016/S1074-7613(01)00115-7.
Adams SD, Barracchini KC, Chen D, Robbins F-M, Stroncek D, Marincola FM: Ambiguous allele combinations in sequence-based typing. ASHI Quarterly. 2003,
Sanger F, Nickels S, Coulson AR: DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977, 74: 5463-5467.
Jin P, Wang E: Polymorphism in clinical immunology. From HLA typing to immunogenetic profiling. J Transl Med. 2003, 1 (18):
Ronaghi M, Uhlen M, Nyrén P: A sequencing method based on real-time pyrophosphate. Science. 1998, 281: 363-365. 10.1126/science.281.5375.363.
Ronaghi M: Pyrosequencing sheds light on DNA sequencing. Genome Res. 2001, 11: 3-11. 10.1101/gr.11.1.3.
Wang L, Marincola FM: Applaying Pyrosequencing to HLA-typing. ASHI Quarterly. 2003, 27: 16-18.
Ahmadian A, Gharizadeh B, Gustafsson AC, Sterky F, Nyren P, Uhlen M, Lundeberg J: Single-nucleotide polymorphism analysis by Pyrosequencing. Anal Biochem. 2000, 280: 103-10. 10.1006/abio.2000.4493.
Sun YQ, Monstein HJ, Nilsson LE, Petersson F, Borch K: Profiling and identification of eubacteria in the stomach of Mongolian gerbils with and without Helicobacter pylory infection. Helicobacter. 2003, 8: 149-157.
Vozarova B, Fernandez-Real JM, Knowler WC, Gallart L, Hanson RL, Gruber JD, Ricart W, Vendrell J, Richart C, Tataranni PA, Wolford JK.: The interleukin-6 (-174) G/C promoter polymorphism is associated with type-2 diabetes mellitus in Native Americans and Caucasians. Hum Genet. 2003, 112: 409-413.
Hefler LA, Ludwig E, Lampe D, Zeillinger R, Leodolter S, Gitsch G, Koelbl H, Tempfer CB: Polymorphisms of the endothelial nitric oxide synthase gene in ovarian cancer. Gynecol Oncol. 2002, 86: 134-137. 10.1006/gyno.2002.6749.
Elahi E, Pourmand N, Chaung R, Rofoogaran A, Boisver J, Samimi-Rad K, Davis RW, Ronaghi M: Determination of hepatitis C virus genotype by Pyrosequencing. J Virol Methods. 2003, 109: 171-176. 10.1016/S0166-0934(03)00068-5.
Pacey-Miller T, Henry R: Single-nucleotide polymorphism detection in plants using a single stranded Pyrosequencing protocol with a universal biotinylated primer. Analyt Biochem. 2003, 317: 165-170.
De Vivo I, Huggins GS, Hankinson SE, Lescault PJ, Boezen M, Colditz GA, Hunter DJ: A functional polymorphism in the promoter of the progesterone receptor gene associated with endometrial cancer risk. Proc Natl Acad Sci U S A. 2002, 99: 12263-12268. 10.1073/pnas.192172299.
Wolford JK, Gruber JD, Ossowski VM, Vozarova B, Antonio Tataranni P, Bogardus C, Hanson RL: A C-reactive protein promoter polymorphism is associated with type 2 diabeter mellitus in Pima Indians. Molec Genet Metab. 2003, 78: 136-144. 10.1016/S1096-7192(02)00230-5.
Barber RC, O'Keefe GE: Characterization of a single nucleotide polymorphism in the lipopolysaccaride binding protein and its association with sepsis. Am J Respir Crit Care Med. 2003, 167: 1316-1320. 10.1164/rccm.200209-1064OC.
Robin C, Lyman RF, Long AD, Langley CH, Mackay TFC: Hairy: a quantitative trait locus for Drosophila Sensory Bristle Number. Genetics. 2002, 162: 155-164.
Olofsson P, Holmberg J, Tordsson J, Lu S, Akerstrom B, Holmdahl R: Positional identification of Ncf1 as a gene that regulates arthritis severity in rats. Nature Genetics. 2003, 33: 25-32. 10.1038/ng1058.
Nordstrom T, Nourizad K, Ronaghi M, Nyren P: Method enabling Pyrosequencing on double-stranded DNA. Anal Biochem. 2000, 282: 186-93. 10.1006/abio.2000.4603.
Sundstrom M, Vliagoftis H, Karlberg P, Butterfield JH, Nilsson K, Metcalfe DD, Nilsson G: Functional and phenotypic studies of two variants of a human mast cell line with a distinct set of mutations in the c-kit proto-oncogene. Immunology. 2003, 108: 89-97. 10.1046/j.1365-2567.2003.01559.x.
Ronaghi M, Karamohamed S, Pettersson B, Uhlen M, Nyren P: Real-time DNA sequencing using detection of pyrophosphate release. Anal Biochem. 1996, 242: 84-9. 10.1006/abio.1996.0432.
Wang E, Adams S, Zhao Y, Panelli MC, Simon R, Klein H, Marincola FM: A strategy for detection of known and unknown SNP using a minimum number of oligonucleotides. J Transl Med. 2003, 1: 4-10.1186/1479-5876-1-4.
Eriksson S, Berg LM, Wadelius M, Aldeborn A: Cttochrome P450 genotyping by multiplex real-time DNA sequencing with Pyrosequencing™ technology. Assay Drug Develop Technol. 2002, 1: 49-59. 10.1089/154065802761001301.
Ringquist S, Alexander AM, Rudert WA, Styche A, Trucco M: Pyrosequencing-based typing of alleles of the HLA-DQB1 gene. Biotechniques. 2002, 33: 166-175.
Gharizadeh B, Nordstrom T, Ahmadian A, Ronaghi M, Nyren P: Long-read Pyrosequencing using pure 2'-deoxyadenosine-5'-O'-(1-thiotriphosphate) Sp-isomer. Analyt Biochem. 2002, 301: 82-90. 10.1006/abio.2001.5494.
Gharizadeh B, Ghaderi M, Donnelly D, Amini B, Wallin KL, Nyren P: Multiple-primer DNA sequencing method. Electrophoresis. 2003, 24: 1145-1151. 10.1002/elps.200390147.
Lahser FC, Wright-Minogue J, Skelton A, Molcolm BA: Quantitative estimation of viral fitness using Pyrosequencing. Biotechniques. 2003, 34: 26-28.
Ronaghi M, Uhlen M, Nyren P: A sequencing method based on real-time pyrophosphate. Science. 1998, 281: 363-365. 10.1126/science.281.5375.363.
Hochberg EP, Miklos DB, Neuberg D, Eichner DA, McLaughlin SF, Mattes-Ritz A, Alyea EP, Antin JH, Soiffer RJ, Ritz J: A novel rapid single nucleotide polymorphism (SNP)-based method for assessment of hematopoietic chimerism after allogeneic stem cell transplantation. Blood. 2003, 101: 363-369. 10.1182/blood-2002-05-1365.
Fakhrai-Rad H, Pourmand N, Ronaghi M: Pyrosequencing: an accurate detection platform for single nucleotide polymorphisms. Hum Mutat. 2002, 19: 479-85. 10.1002/humu.10078.
Acknowledgements
We wish to acknowledge PyrosequencingAB for their outstanding technical support. We are grateful to Mostafa Ronaghi for the valuable discussions. We thank Yunxia Wang, Joel Shi, Dina Berchanskiy and Xiang Jun Liu for their assistance and helpful discussions. Daniel Ramon is a PhD candidate in the field of Biochemistry at Universidad Nacional de San Luis (Argentina).
Author information
Authors and Affiliations
Corresponding author
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Ramon, D., Braden, M., Adams, S. et al. Pyrosequencing™ : A one-step method for high resolution HLA typing. J Transl Med 1, 9 (2003). https://doi.org/10.1186/1479-5876-1-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1479-5876-1-9