Abstract
Background
Acute otitis media with perforation (AOMwiP) affects 40% of remote Indigenous children during the first 18 months of life. Streptococcus pneumoniae, Haemophilus influenzae and Moraxella catarrhalis are the primary bacterial pathogens of otitis media and their loads predict clinical ear state. Our hypothesis is that antecedent respiratory viral infection increases bacterial density and progression to perforation.
Methods
A total of 366 nasopharyngeal swabs from 114 Indigenous children were retrospectively examined. A panel of 17 respiratory viruses was screened by PCR, and densities of S. pneumoniae, H. influenzae and M. catarrhalis were estimated by quantitative real time PCR. Data are reported by clinical ear state.
Results
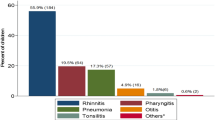
M. catarrhalis (96%), H. influenzae (91%), S. pneumoniae (89%) and respiratory viruses (59%) were common; including rhinovirus (HRV) (38%), polyomavirus (HPyV) (14%), adenovirus (HAdV) (13%), bocavirus (HBoV) (8%) and coronavirus (HCoV) (4%). Geometric mean bacterial loads were significantly higher in children with acute otitis media (AOM) compared to children without evidence of otitis media. Children infected with HAdV were 3 times more likely (p < 0.001) to have AOM with or without perforation.
Conclusion
This study confirms a positive association between nasopharyngeal bacterial load and clinical ear state, exacerbated by respiratory viruses, in Indigenous children. HAdV was independently associated with acute ear states.
Similar content being viewed by others
Background
Australian Aboriginal children have the highest published prevalence of acute otitis media (AOM), with and without perforation, in the world [1]. Infants are colonized with multiple species of respiratory bacteria within weeks of birth, including the AOM pathogens, S. pneumoniae, H. influenzae and M. catarrhalis. Early dense colonization predicts early onset of otitis media, and progression to tympanic membrane perforation (TMP) occurs in 30% of children by 6 months of age and 40% by 18 months of age [2, 3]. At any one time few as 8% of remote Indigenous children aged 6 - 30 months have bilaterally normal middle ears [1].
Understanding the burden of otitis media (OM) in remote indigenous populations is complex. Overcrowding [4], poor hygiene [5], limited formal education [6], and inadequate access to under-resourced medical services contribute to the prevalence and severity of OM in these children [2, 3]. Recurrent OM and perforation is associated with conductive hearing loss and further contributes to poor educational outcomes [7].
To date, interventions for otitis media have had modest benefits for Indigenous children in remote communities [8]. A randomised controlled trial in the Northern Territory, comparing antibiotics for the treatment of AOM, showed no difference in the risk of clinical failure between amoxicillin (54% failure rate) and azithromycin (50% failure rate) treatment groups despite significant reductions in carriage density of S. pneumoniae and H. influenzae in the azithromycin group [9].
The introduction of the seven-valent pneumococcal vaccine (7vPCV) in 2001 brought anticipation of a reduced OM burden. However a review of randomised controlled trials using PCVs to prevent AOM in children aged 12 years or younger, sourced from the Cochrane Central Register of Controlled Trials, found that the vaccine administered during infancy induced only a marginal reduction in OM disease (6-7%) [10]. In Australia a birth cohort study of Indigenous infants found a limited benefit of 7vPCV on OM compared to historic controls. No effect was observed on the proportion of children with otitis media with effusion (OME), AOM or AOMwiP at 12 months of age, however, significantly fewer vaccinees (14%) than controls (23%) experienced chronic suppurative otitis media (CSOM) and vaccinees experienced delayed onset of TMP [11].
A plausible explanation for the poor response to antibiotics and high prevalence of AOM with perforation in the Indigenous community setting is co-infection of multiple bacterial pathogens with respiratory viruses. Human rhinovirus (HRV), respiratory syncytial virus (RSV), influenza virus A and B (FLUA, FLUB), HAdV, HCoV and parainfluenza virus 1, 2 and 3 (PIV 1, PIV2 & PIV3) are commonly associated with OM [12–14]. Recent advances in molecular viral diagnostics [15] now allow a broader insight into viral OM pathogenesis.
Infection with respiratory viruses induces damage to the epithelial mucosa of airways, Eustachian tube and middle ear exposing surface elements that bacteria can adhere to [16]. The interaction between FLUA and S. pneumoniae is well noted [17, 18]. Influenza neuraminidase is demonstrated to increase pneumococcal adhesion and invasion in the Eustachian tube and middle ear [19, 20]. RSV can also facilitate bacterial adhesion. Studies reveal that in the presence of RSV, certain strains of M. catarrhalis have an enhanced ability to adhere to epithelial cells [21]. Additionally, viral mediated inflammation congests the Eustachian tube and further reduces cilial function, inhibiting bacterial clearance [22, 23], while viral immunosuppression can lead to bacterial superinfection [24]. Recent evidence suggests that pneumococcal load is driven by FLUA, HRV and RSV infection, and that enhanced transmission and acquisition occurs in the presence of FLUA [18, 25].
For young Indigenous children, the role of respiratory viruses in OM remains unclear. In the tropical North of Australia, AOM is endemic in remote communities and not subject to seasonal viral outbreaks. In this study we use molecular methods [3, 15] to investigate whether antecedent nasopharyngeal viral infection increases bacterial load and progression to AOMwiP.
Methods
Population and primary data
Samples were sourced from two previous studies based in the same remote Aboriginal communities. The first, the COMIT1 trial (1996-2001), was a birth cohort randomized controlled trial that examined the effect of long course amoxicillin versus placebo on resolution of first detected OME and the prevention of tympanic membrane perforation [8]. The second, the PRIORiTI (PRevenar Immunisation for Otitis media Reductions in Tiwi Infants) study (2001-2004), was a birth cohort observational study following the introduction of routine childhood heptavalent conjugate pneumococcal vaccination (7vPCV) [11]. In both studies examinations were undertaken at approximately one month intervals from birth till 24 months.
Sample collection and clinical definitions have been described previously [8, 11]. In summary, ear state was defined as follows: (a) Normal: normal or minor pathology (abnormal appearance or retracted drum), (b) OME: either a normal tympanic membrane (TM) or intact, non-bulging TM, plus type B tympanogram, (c) AOM: bulging TM, without perforation, and type B tympanogram, (d) AOMwiP: middle ear discharge present for less than 6 weeks and perforation covering less than 2% of the pars tensa of the TM. (e) CSOM: middle ear discharge present for longer than 6 weeks and perforation covering greater than 2% of the pars tensa of the TM. The final middle ear diagnosis reflects the child's most severely affected ear. As the focus of this study was acute ear pathology, swabs from children with CSOM were excluded.
Analysis of bacterial and viral pathogens was performed as part of a larger case-control/crossover study (Cheng et al, manuscript in preparation). Bacterial and viral DNA was extracted by defined methods [3, 15, 26, 27] and quantitative real time polymerase chain reaction (RTQ-PCR) was used to determine total and species specific pathogen loads[3]. The PCR target genes were: 16S rRNA for total bacterial load [28], autolysin (lytA) for S. pneumoniae load [29], protein D (hpd) for H. influenzae load [3], and the outer membrane protein (copB) for M. catarrhalis load [30]. Specimens were tested for FLUA and B, RSV, PIV 1, 2 and 3, HAdV, human meta-pneumovirus (HMPV), HRV, enterovirus (EV), HCoV's (HKU1, OC43, 229E and NL63), HBoV and HPyV's (WU and KI) by PCR as previously described [15, 26, 27].
Ethics
Informed written consent was obtained from the parents of all children who participated in the parent studies, COMIT and PRIORiTI. This work was performed following consultation with Indigenous community members, and was approved by the Human Research Ethics Committee of the Menzies School of Health Research and the Department of Human Services (HREC - 07/78).
Statistical Analysis
Chi-square or Fischer's exact analysis was used to compare the proportions of virus and bacteria positive swabs in each of the four ear states. Bacterial loads were log normally distributed and therefore were log transformed for analysis. Non-parametric tests for continuous variables, Kruskal-Wallis equality of populations rank test and Mann Whitney Wilcoxon ranksum tests, were used to compare bacterial loads between ear states, with or without viral co-detection. Linear regression was used to examine the relationships between the bacterial loads and interacting variables such as age and antibiotic use (<2 weeks before swab was taken) with lower limits set at lowest positive load value. The reported p values are two-tailed and significance was set at 5%. All statistical analysis was performed using STATA IC version 11 (StataCorp, Texas, USA).
Results
Bacterial and viral infection of the nasopharynx in relation to ear state
Of the 366 swabs from 114 children, 58 (16%) swabs were from children diagnosed with tympanic membrane perforation. 115 (31%) had AOM without perforation, 175 (48%) had OME, and 18 (5%) were from children with Normal ears (Table 1).
By RTQ-PCR most swabs were positive for M. catarrhalis (96%), H. influenzae (91%) and S. pneumoniae (89%). Greater than 80% had all three and all had at least one of these pathogens. Although the proportion of positive swabs was high for all ear states, more children from the AOM group were positive for each of the bacteria compared to the Normal group (Table 1).
Sixty two percent (228/366) of swabs amplified at least one virus. A total of 312 viruses were detected in the 228 positive swabs at an average of 1.37 viruses per positive swab. HRV (38%) was the most frequently encountered virus, followed by HPyV (14%), HAdV (13%), HBoV (8%) and HCoV (4%). Other viruses (FLUA and B, PIV 1,2 and 3, HMPV, RSV and EV) were only found in <9% collectively (Table 1). Further investigation of the HPyV subtypes, WU and KI, revealed the WU virus predominated (92% of HPyV positive swabs).
The proportions of HAdV detected in the nasopharyngeal swabs for each ear diagnosis were: 6% for Normal, 7% for OME, 19% for AOM and 22% for AOMwiP (Table 1) (Figure 1). No other virus revealed a positive relationship with ear state.
Bacterial load in the nasopharynx in relation to ear state
The geometric-mean bacterial loads were high throughout. Individual bacterial loads showed stepwise elevations by categorical ear state for M. catarrhalis, H. influenzae and S. pneumoniae, peaking at AOM (Figure 2). Unexpectedly, the individual bacterial loads were lower at the time of AOM with perforation than at AOM without perforation. A dichotomous ranksum test revealed a significantly higher load in the AOM group compared to Normal group for M. catarrhalis (N = 5.42 × 104, AOM = 2.19 × 106; p = 0.005); H. influenzae (N = 1.33 × 105, AOM = 1.09 × 106; p = 0.031); and S. pneumoniae (N = 2.44 × 103, AOM = 2.68 × 105; p = 0.004). The total bacterial loads were all greater than 107, showing no trend for ear state.
Bacterial load versus Clinical Ear State. The geometric mean bacterial loads are segregated by clinical ear state for each bacterial pathogen. A significantly higher load was seen for M. catarrhalis, H. influenzae and S. pneumoniae in children with AOM compared to Normal ear states (Mann Whitney Wilcoxon test). Significant differences in the loads across the ear states, for M. catarrhalis and S. pneumoniae, were confirmed by the Kruskal-Wallis equality of populations rank test. There was no significant difference in geometric mean of the Total Bacterial Load between ear states.
Bacterial load in the nasopharynx when viruses were present
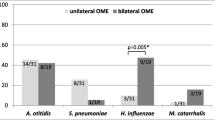
In the presence of viruses the individual loads had a general trend upwards (Figure 3). The combined load of M. catarrhalis, H. influenzae and S. pneumoniae was greater when examined by presence or absence of any study virus (p = 0.035) (data not shown). Of the individual bacterial pathogens, significantly higher loads were only seen for H. influenzae co-infected with any study virus (Virus +), HPyV and HCoV (Table 2) (Figure 3). Despite the association between HAdV and ear disease, load increases in the presence of adenovirus were insignificant.
Association between bacterial load and other known factors
The impact of sex, age, antibiotic use and number of viruses detected were assessed using linear regression (Table 2). The loads of M. catarrhalis, H. influenzae and S. pneumoniae were significantly lower in children less than 3 months of age. This was in part due to the greater proportion of children without detectable OM bacteria at this early age. In children older than 3 months the effect of age on load was neutral. Linear regression also revealed that the density of M. catarrhalis and H. influenzae was positively influenced by respiratory viral infection while the impact of sex and antibiotic use (<2 weeks before swab was taken) on load was negligible. Furthermore, comparison of total and pathogen specific loads with and without antibiotics, for each ear state, did not reveal any significant differences (data not included).
Correlation between molecular bacterial load and semi-quantitative microbiology
The positive correlation between the bacterial loads determined by RTQ-PCR and clinical ear state was consistent with original semi-quantitative microbiology (data not included). Sensitivity was enhanced using RTQ-PCR. Of the 356 swabs cultured for M. catarrhalis, 82% were culture positive and 96% were RTQ-PCR positive. Similarly for H. influenzae; 75% were culture positive and 91% were RTQ-PCR positive. All 366 swabs were cultured for S. pneumoniae of which 68% were culture positive and 89% were RTQ-PCR positive. 14%, 20% and 23% of swabs were culture negative and RTQ-PCR positive for M. catarrhalis, H. influenzae and S. pneumoniae respectively. 1%, 5% and 2% of swabs were culture positive but RTQ-PCR negative respectively.
Limitations of the study
The bacterial load estimates have limitations: The quantity of material obtained on the swab was assumed to be equal for all swabs; DNA extraction efficiency was considered to be 100%; and molecular weight calculations were based on estimates of GC content. Additionally when quantifying the total bacterial load using 16S primers, we used S. pneumoniae with 4 ribosomal operons as our control. The actual number of ribosomal operons varies greatly between species. Importantly, load values remain relative to each other throughout. It is worth noting that the hpd (protein D) target is likely to be present in closely related Haemophilus species including non-haemolytic H. haemolyticus and as such this study may over represent classical non-typeable H. influenzae.
Our swabs were collected at monthly intervals, so we have assumed that microbiology at the time of diagnosis best approximates the aetiology of that event. However our definition of the primary perforation included perforation of less than 6 weeks duration. The exact date of onset was not available. This analysis describes the nasal microbiology and ear status at the time of swab collection and presents the data for Normal, OME, AOM and AOMwiP. Furthermore, while the nasopharynx and the middle ear show concordance of infection, studies suggest a lapse of 3-10 days between upper respiratory symptoms and onset of otitis media [31, 32]. It is therefore plausible that bacterial load and viral detection will be greater in swabs collected prior to perforation, compared to swabs collected at or after the time of perforation.
Discussion
In this study the high bacterial carriage rates, high bacterial loads, and small number of children with Normal ears (5% of swabs), reflect the large burden of respiratory infection in Indigenous children who live in remote communities. Increases in individual respiratory bacterial load (and carriage) were apparent with worsening ear state with a peak at AOM, whereas the total bacterial load was high regardless of ear state.
The broad viral panel scrutinized by sensitive PCR methods in this study revealed a high proportion of virus positive nasopharyngeal swabs (62% swabs or 89% children). The proportions remained stable by ear state including the Normal group, of which 67% were positive for any study virus. HRVs, which are responsible for more cold-like illnesses than any other virus [33], were the most prevalent viruses identified in this study (38%), though no clear correlation between HRV and clinical ear state was observed.
HAdV was significantly associated with acute ear pathology (Figure 1). Children with AOM or AOMwiP were infected with HAdV at 3 times the rate of children with Normal and OME diagnosis (P < 0.001) though this was not associated with a significant increase in bacterial load. HAdV is increasingly reported as an upper respiratory pathogen with studies showing occurrence of AOM in up to 50% of those children infected [12–14, 34]. No licensed HAdV treatment is currently available [35].
Overall our data supports a potential role for viruses in AOM, particularly HAdV, encouraging prospective studies and clinical trials to better understand the role viruses might have in endemic OM.
Differences in bacterial load with viral co-infection were apparent but of limited significance. H. influenzae load was significantly elevated in the presence of HPyV, HCoV and any study virus, and linear regression revealed that the loads of M.catarrhalis and H. influenzae were higher with detection of multiple viruses (Figure 3) (Table 2). At no point were the individual bacterial loads lower in the presence of a virus. We suspect that the consistently high bacterial loads seen in these children might mask the true effect of respiratory viruses on bacterial density.
The higher individual loads seen in AOM swabs compared to AOMwiP swabs were unexpected. We suggest that this is likely due to the mismatch in timing of onset and swab collection.
Regression modelling of bacterial load by age revealed significantly lower loads for all three individual bacterial pathogens in children less than 3 months of age. In children older than 3 months, age exerted no influence on load. This finding reflects the natural acquisition of these bacteria after birth. No differences in the bacterial loads were observed between the sexes or with antibiotic use in the fortnight prior to examination and swabbing (Table 2). Treatment guidelines recommend antibiotic use upon diagnosis of AOM. The effect of antibiotic use on loads, by ear state, was also investigated (data not included). No differences in the bacterial loads were observed, with or without antibiotic use, in swabs taken at a diagnosis of AOM or AOMwiP.
Viral-bacterial co-infections are often associated with antibiotic treatment failure [36]. In the Indigenous children in our study, antibiotic therapy had no discernable effect on bacterial loads. For densely colonised children with treatment failure, more work is needed to determine whether antiviral medication might be a valid adjunct to antibiotic treatment. Recently a randomized controlled trial in the United States found that children receiving oseltamivir (Tamiflu) treatment within 48 hours of clinic presentation for laboratory confirmed influenza were considerably less likely to progress to AOM. This effect was greatest for children between the ages 1-5 years old where the incidence of AOM was the greatest [37].
The limited clinical impact of PCV7 on OM in indigenous children is largely attributed to pneumococcal serotype replacement. This study however, suggests that the influence of respiratory viruses on OM in this population may be undervalued. A review by Cripps and Otczyk in 2006 promotes a vaccine that includes both bacterial and viral antigens [38] and others have shown that the influenza vaccine can reduce overall AOM incidence by greater than 30% [39–41]. Whether vaccines for individual viruses would have an impact on endemic OM is unclear, but protection against multiple pathogens may be needed to attenuate the complex interplay between viruses and bacteria in OM in high risk populations.
Conclusions
For Indigenous children with OM this study highlights the frequency of viral infection, describes increasing bacterial density with progression to AOM, suggests a synergy of viral-bacterial co-infection, and implicates adenovirus in AOMwiP pathogenesis. Antibiotic and antiviral therapy is ever expanding and improving and it seems likely that only vaccines and/or chemotherapies targeting multiple bacterial and viral species, in combination with successful social interventions to reduce cross infection, will reduce OM prevalence, morbidity and sequelae in high risk populations. The introduction of the 10-valent pneumococcal conjugate vaccine, with broader pneumococcal serotype coverage and potential protection from the addition of the H. influenzae protein D antigen is possibly the first step in the right direction.
References
Morris PS, Leach AJ, Silberberg P, Mellon G, Wilson C, Hamilton E, Beissbarth J: Otitis media in young Aboriginal children from remote communities in Northern and Central Australia: a cross-sectional survey. BMC Pediatr. 2005, 5: 27-10.1186/1471-2431-5-27.
Leach AJ, Boswell JB, Asche V, Nienhuys TG, Mathews JD: Bacterial colonization of the nasopharynx predicts very early onset and persistence of otitis media in Australian aboriginal infants. Pediatr Infect Dis J. 1994, 13 (11): 983-989. 10.1097/00006454-199411000-00009.
Smith-Vaughan H, Byun R, Nadkarni M, Jacques NA, Hunter N, Halpin S, Morris PS, Leach AJ: Measuring nasal bacterial load and its association with otitis media. BMC ear nose and throat disorders. 2006, 6: 10-10.1186/1472-6815-6-10.
Lehmann D, Arumugaswamy A, Elsbury D, Finucane J, Stokes A, Monck R, Jeffries-Stokes C, McAullay D, Coates H, Stanley FJ: The Kalgoorlie Otitis Media Research Project: rationale, methods, population characteristics and ethical considerations. Paediatr Perinat Epidemiol. 2008, 22 (1): 60-71.
Stubbs E, Hare K, Wilson C, Morris P, Leach AJ: Streptococcus pneumoniae and noncapsular Haemophilus influenzae nasal carriage and hand contamination in children: a comparison of two populations at risk of otitis media. Pediatr Infect Dis J. 2005, 24 (5): 423-428. 10.1097/01.inf.0000160945.87356.ca.
Johnston V, Lea T, Carapetis J: Joining the dots: The links between education and health and implications for indigenous children. J Paediatr Child Health. 2009, 45 (12): 692-697. 10.1111/j.1440-1754.2009.01596.x.
Williams CJ, Jacobs AM: The impact of otitis media on cognitive and educational outcomes. Med J Aust. 2009, 191 (9 Suppl): S69-72.
Leach AJ, Morris PS, Mathews JD: Compared to placebo, long-term antibiotics resolve otitis media with effusion (OME) and prevent acute otitis media with perforation (AOMwiP) in a high-risk population: a randomized controlled trial. BMC Pediatr. 2008, 8: 23-10.1186/1471-2431-8-23.
Morris PS, Gadil G, McCallum GB, Wilson CA, Smith-Vaughan HC, Torzillo P, Leach AJ: Single-dose azithromycin versus seven days of amoxycillin in the treatment of acute otitis media in Aboriginal children (AATAAC): a double blind, randomised controlled trial. Med J Aust. 2010, 192 (1): 24-29.
Jansen AG, Hak E, Veenhoven RH, Damoiseaux RA, Schilder AG, Sanders EA: Pneumococcal conjugate vaccines for preventing otitis media. Cochrane Database Syst Rev. 2009, CD001480-2
Mackenzie GA, Carapetis JR, Leach AJ, Morris PS: Pneumococcal vaccination and otitis media in Australian Aboriginal infants: comparison of two birth cohorts before and after introduction of vaccination. BMC Pediatr. 2009, 9: 14-10.1186/1471-2431-9-14.
Alper CM, Winther B, Mandel EM, Hendley JO, Doyle WJ: Rate of concurrent otitis media in upper respiratory tract infections with specific viruses. Archives of otolaryngology--head & neck surgery. 2009, 135 (1): 17-21. 10.1001/archotol.135.1.17.
Moore HC, Jacoby P, Taylor A, Harnett G, Bowman J, Riley TV, Reuter K, Smith DW, Lehmann D: The Interaction Between Respiratory Viruses and Pathogenic Bacteria in the Upper Respiratory Tract of Asymptomatic Aboriginal and Non-Aboriginal Children. Pediatr Infect Dis J. 2010, 29 (6): 540-5.
Jacoby P, Watson K, Bowman J, Taylor A, Riley TV, Smith DW, Lehmann D: Modelling the co-occurrence of Streptococcus pneumoniae with other bacterial and viral pathogens in the upper respiratory tract. Vaccine. 2007, 25 (13): 2458-2464. 10.1016/j.vaccine.2006.09.020.
Syrmis MW, Whiley DM, Thomas M, Mackay IM, Williamson J, Siebert DJ, Nissen MD, Sloots TP: A sensitive, specific, and cost-effective multiplex reverse transcriptase-PCR assay for the detection of seven common respiratory viruses in respiratory samples. J Mol Diagn. 2004, 6 (2): 125-131. 10.1016/S1525-1578(10)60500-4.
Avadhanula V, Rodriguez CA, Devincenzo JP, Wang Y, Webby RJ, Ulett GC, Adderson EE: Respiratory viruses augment the adhesion of bacterial pathogens to respiratory epithelium in a viral species- and cell type-dependent manner. J Virol. 2006, 80 (4): 1629-1636. 10.1128/JVI.80.4.1629-1636.2006.
McCullers JA: Insights into the interaction between influenza virus and pneumococcus. Clin Microbiol Rev. 2006, 19 (3): 571-582. 10.1128/CMR.00058-05.
Diavatopoulos DA, Short KR, Price JT, Wilksch JJ, Brown LE, Briles DE, Strugnell RA, Wijburg OL: Influenza A virus facilitates Streptococcus pneumoniae transmission and disease. FASEB J. 2010, 24 (6): 1789-1798. 10.1096/fj.09-146779.
Peltola VT, McCullers JA: Respiratory viruses predisposing to bacterial infections: role of neuraminidase. Pediatr Infect Dis J. 2004, 23 (1 Suppl): S87-97.
McCullers JA, Bartmess KC: Role of neuraminidase in lethal synergism between influenza virus and Streptococcus pneumoniae. J Infect Dis. 2003, 187 (6): 1000-1009. 10.1086/368163.
El-Ahmer OR, Braun JM, Amyes SG, Weir DM, Beuth J, Blackwell CC: Comparison of Moraxella catarrhalis isolates from children and adults for growth on modified New York City medium and potential virulence factors. J Med Microbiol. 2003, 52 (Pt 10): 853-859.
Chung MH, Griffith SR, Park KH, Lim DJ, DeMaria TF: Cytological and histological changes in the middle ear after inoculation of influenza A virus. Acta Otolaryngol. 1993, 113 (1): 81-87. 10.3109/00016489309135771.
Park K, Bakaletz LO, Coticchia JM, Lim DJ: Effect of influenza A virus on ciliary activity and dye transport function in the chinchilla eustachian tube. Ann Otol Rhinol Laryngol. 1993, 102 (7): 551-558.
Bakaletz LO: Viral potentiation of bacterial superinfection of the respiratory tract. Trends Microbiol. 1995, 3 (3): 110-114. 10.1016/S0966-842X(00)88892-7.
Vu HT, Yoshida LM, Suzuki M, Nguyen HA, Nguyen CD, Nguyen AT, Oishi K, Yamamoto T, Watanabe K, Vu TD, et al: Association Between Nasopharyngeal Load of Streptococcus pneumoniae, Viral Coinfection, and Radiologically Confirmed Pneumonia in Vietnamese Children. Pediatr Infect Dis J. 2010, 30 (1): 11-8.
Bialasiewicz S, Whiley DM, Lambert SB, Gould A, Nissen MD, Sloots TP: Development and evaluation of real-time PCR assays for the detection of the newly identified KI and WU polyomaviruses. J Clin Virol. 2007, 40 (1): 9-14. 10.1016/j.jcv.2007.07.015.
Tozer SJ, Lambert SB, Whiley DM, Bialasiewicz S, Lyon MJ, Nissen MD, Sloots TP: Detection of human bocavirus in respiratory, fecal, and blood samples by real-time PCR. Journal of medical virology. 2009, 81 (3): 488-493. 10.1002/jmv.21409.
Nadkarni MA, Martin FE, Jacques NA, Hunter N: Determination of bacterial load by real-time PCR using a broad-range (universal) probe and primers set. Microbiology (Reading, England). 2002, 148 (Pt 1): 257-266.
McAvin JC, Reilly PA, Roudabush RM, Barnes WJ, Salmen A, Jackson GW, Beninga KK, Astorga A, McCleskey FK, Huff WB, et al: Sensitive and specific method for rapid identification of Streptococcus pneumoniae using real-time fluorescence PCR. J Clin Microbiol. 2001, 39 (10): 3446-3451. 10.1128/JCM.39.10.3446-3451.2001.
Greiner O, Day PJ, Altwegg M, Nadal D: Quantitative detection of Moraxella catarrhalis in nasopharyngeal secretions by real-time PCR. J Clin Microbiol. 2003, 41 (4): 1386-1390. 10.1128/JCM.41.4.1386-1390.2003.
Heikkinen T, Ruuskanen O: Temporal development of acute otitis media during upper respiratory tract infection. Pediatr Infect Dis J. 1994, 13 (7): 659-661. 10.1097/00006454-199407000-00016.
Miyamoto N, Bakaletz LO: Kinetics of the ascension of NTHi from the nasopharynx to the middle ear coincident with adenovirus-induced compromise in the chinchilla. Microb Pathog. 1997, 23 (2): 119-126. 10.1006/mpat.1997.0140.
Rotbart HA: Antiviral therapy for enteroviruses and rhinoviruses. Antivir Chem Chemother. 2000, 11 (4): 261-271.
Chonmaitree T, Revai K, Grady JJ, Clos A, Patel JA, Nair S, Fan J, Henrickson KJ: Viral upper respiratory tract infection and otitis media complication in young children. Clin Infect Dis. 2008, 46 (6): 815-823. 10.1086/528685.
Lenaerts L, De Clercq E, Naesens L: Clinical features and treatment of adenovirus infections. Rev Med Virol. 2008, 18 (6): 357-374. 10.1002/rmv.589.
Hament JM, Kimpen JL, Fleer A, Wolfs TF: Respiratory viral infection predisposing for bacterial disease: a concise review. FEMS immunology and medical microbiology. 1999, 26 (3-4): 189-195. 10.1111/j.1574-695X.1999.tb01389.x.
Winther B, Block SL, Reisinger K, Dutkowski R: Impact of oseltamivir treatment on the incidence and course of acute otitis media in children with influenza. Int J Pediatr Otorhinolaryngol. 2010, 74 (6): 684-688. 10.1016/j.ijporl.2010.03.024.
Cripps AW, Otczyk DC: Prospects for a vaccine against otitis media. Expert Rev Vaccines. 2006, 5 (4): 517-534. 10.1586/14760584.5.4.517.
Heikkinen T, Ruuskanen O, Waris M, Ziegler T, Arola M, Halonen P: Influenza vaccination in the prevention of acute otitis media in children. Am J Dis Child. 1991, 145 (4): 445-448.
Clements DA, Langdon L, Bland C, Walter E: Influenza A vaccine decreases the incidence of otitis media in 6- to 30-month-old children in day care. Arch Pediatr Adolesc Med. 1995, 149 (10): 1113-1117.
Marchisio P, Cavagna R, Maspes B, Gironi S, Esposito S, Lambertini L, Massimini A, Herzog C, Principi N: Efficacy of intranasal virosomal influenza vaccine in the prevention of recurrent acute otitis media in children. Clin Infect Dis. 2002, 35 (2): 168-174. 10.1086/341028.
Pre-publication history
The pre-publication history for this paper can be accessed here:http://www.biomedcentral.com/1471-2334/11/161/prepub
Acknowledgements
We thank the families and children that participated in this study, the Northern Territory Department of Health and Community Services, the Tiwi Health Board, and staff at the clinics involved. Without this support the study would not have been possible.
Funding
This study was supported by the NHMRC (project grant reference number 490317).
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
MB performed the DNA extractions, bacterial loads PCR's, data analysis and interpretation of results. MB is the author of this work. AC contributed to the design of the larger case control study from which these samples were drawn, and assisted with statistical analysis and editing. HSV provided expertise both to the design of the RTQ-PCR and the writing of this paper. TS's laboratory screened the swabs for the 17 respiratory viruses and TS assisted with interpretation of the virus PCR results. MN provided clinical interpretation of the virology. DW performed the DNA extractions and PCR assays to identify the viruses. JM assisted with and proofed the statistical calculations. AJL concepted this study and contributed significantly to the analysis and editing of this manuscript. All authors have read and approved the final version.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Binks, M.J., Cheng, A.C., Smith-Vaughan, H. et al. Viral-bacterial co-infection in Australian Indigenous children with acute otitis media. BMC Infect Dis 11, 161 (2011). https://doi.org/10.1186/1471-2334-11-161
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2334-11-161