Abstract
Background
Rat oligonucleotide microarrays were used to detect changes in gene expression in the dorsal root ganglion (DRG) 3 days following sciatic nerve transection (axotomy). Two comparisons were made using two sets of triplicate microarrays, naïve versus naïve and naïve versus axotomy.
Results
Microarray variability was assessed using the naïve versus naïve comparison. These results support use of a P < 0.05 significance threshold for detecting regulated genes, despite the large number of hypothesis tests required. For the naïve versus axotomy comparison, a 2-fold cut off alone led to an estimated error rate of 16%; combining a >1.5-fold expression change and P < 0.05 significance reduced the estimated error to 5%. The 2-fold cut off identified 178 genes while the combined >1.5-fold and P < 0.05 criteria generated 240 putatively regulated genes, which we have listed. Many of these have not been described as regulated in the DRG by axotomy. Northern blot, quantitative slot blots and in situ hybridization verified the expression of 24 transcripts. These data showed an 83% concordance rate with the arrays; most mismatches represent genes with low expression levels reflecting limits of array sensitivity. A significant correlation was found between actual mRNA differences and relative changes between microarrays (r2 = 0.8567). Temporal patterns of individual genes regulation varied.
Conclusions
We identify parameters for microarray analysis which reduce error while identifying many putatively regulated genes. Functional classification of these genes suggest reorganization of cell structural components, activation of genes expressed by immune and inflammatory cells and down-regulation of genes involved in neurotransmission.
Similar content being viewed by others
Background
Transection of the peripheral axons of primary sensory neurons results in profound alterations in their metabolism, regenerative capacity, survival, excitability, transmitter function and sensitivity to diverse extrinsic and intrinsic signals [1, 2]. These changes are contributed to by transcriptional alterations triggered by a loss of trophic support from peripheral target organs, and by novel signals generated at the injury site. The transcriptional changes lead both to adaptive responses, such as the capacity to survive the injury and re-grow the injured axon, and maladaptive responses that can lead to a change in sensation, including the generation of neuropathic pain [3, 4].
Measurements of mRNA and protein made in many laboratories using diverse methodologies have identified approximately 40 genes that are up-regulated and 25 down-regulated after peripheral nerve injury [3, 5, 6]. These regulated genes include members of several classes: G protein coupled receptors, ligand- and voltage-gated ion channels, receptor tyrosine kinases, growth factors, cytokines, neuropeptides, cell cytoskeletal genes, cell surface/extracellular matrix genes, and a miscellaneous group of enzymes.
We have now used high-density rat genome oligonucleotide microarrays to analyze changes in gene expression at an early time point (3 days) following a peripheral nerve injury in adult rats. Oligonucleotide microarrays provide the capacity to analyze parallel changes in many thousands of genes, and have been used successfully to examine expression profile changes in many neuronal and non-neuronal systems [7] including DRG neurons [8, 9]. Studying the effect of peripheral nerve injury on gene expression profiles in the DRG offers several advantages. The DRG represents a dense collection of cell bodies of one general class of neuron, the primary sensory neuron. The lesion has a uniform impact on the cells, and the existence of a large pool of genes with known regulation allows for quality controls for changes identified by the microarrays [3, 5, 6].
Although microarrray technology offers enormous potential advantages, there remain important concerns about representation, sensitivity, reproducibility, variability and the false positive and negative detection rates [10, 11]. What, if any, is the threshold fold-difference between probe sets that reflects real regulation? Is fold-difference the most sensitive measure for detecting changes? Are genes with a low expression levels in all conditions detected? What is the concordance rate between genes identified as regulated by independent methods and genes identified by the microarray analysis? What criteria need to be used to jointly minimize the false negative and the false positive rates? How many arrays need to be analyzed?
We now show that fold-difference alone results in a high degree of error whilst detecting regulated genes from microarrays, replicate arrays with statistical analysis reduces false positives and negatives. The oligonucleotide microarray screen indicates that hundreds of genes are regulated by neuronal injury, and that this technique is a powerful primary screen for such changes.
Results
Array sensitivity and variability
Affymetrix rat U34A oligonucleotide arrays were used to screen for changes in gene expression in DRG neurons three days following a peripheral nerve transection (axotomy, Ax) by comparing expression levels with non-injured DRGs (naïve, N). Nine biologically independent array hybridizations were performed (six naïve and three axotomy). DRG tissue (L4 and L5 from the left or ipsilateral side to the injury) from 5 male Sprague-Dawley rats were pooled for each RNA population. Each RNA sample was labeled separately and hybridized to a separate array. Genes were defined as detected if they received a present or marginal call in at least one of the arrays within each comparison. Of the 8799 annotated genes and expressed sequence tags (ESTs) represented on the rat U34A array, 52% (n= 6, naïve arrays) were detected in arrays hybridized with naïve rat DRG cRNA and 54% (n = 3, axotomy arrays) genes in the axotomy comparisons.
To assess the degree of variability inherent in the system, the mean probe set intensity values of the two independent sets of three naïve samples were compared on a scatter plot (Figure 1A). The majority of probe sets from the two control groups (N vs. N) lie on or very close to the identity line, with a linear regression of r2 = 0.9898 (for all detected genes in the N vs. N comparison, Figure 1A). Figure 1A shows that most of the variation between the two naïve groups exists at the lower end of the gene expression levels. This suggests that this variability is in part due to array sensitivity. In consequence, differences in low intensity genes may be unreliable.
Intensity plots of average values of all present genes for the naïve group 1 versus naïve group 2. (A) and the naïve group 1 versus the 3 day axotomy group (B). Intensity plots of average values of all genes deemed to vary significantly using an unpaired two-tailed t-test, for the naïve group 1 versus naïve group 2 comparison (C) and the naïve group 1 versus the 3 day axotomy group comparison (D). Plots are color coded so that P < 0.05 (blue points), P < 0.01 (green points), P < 0.001 (red points). Tabulated in (E) are the numbers of genes detected (grey) and those genes achieving P < 0.05 (blue), P < 0.01 (green), P < 0.001 (red) as a function of fold change.
A plot of the mean intensity from triplicate naives against triplicate 3 day axotomy (N vs. Ax) arrays (Figure 1B) show many data points lying both above and below the identity line, suggesting regulation. Since each individual sample was pooled from 5 male Sprague-Dawley animals of a similar age and from a single supplier (Charles River), biological variation is likely to be minimal.
Defining regulated genes
Many of the detected genes in both the N vs. N and the N vs. Ax array comparison groups show no difference, defined as fold-difference up or down less than 1.15 (69%, N vs. N and 47%, N vs. Ax) (Figure 1E). Further, many show small fold-differences of 1.15 to 1.5 (27%, N vs. N and 41%, N vs. Ax) (Figure 1E). In total, genes displaying a fold-difference below 1.5-fold accounted for 97% of the N vs. N arrays and 88% of the N vs. Ax comparison.
When statistical significance is not taken into account, 122 genes display a difference of 1.5–2.0 fold and 28 a greater than 2-fold difference in the N vs. N group (2.6% and 0.6% of present genes respectively). In the N vs. Ax group 369 genes show a fold-difference of 1.5–2.0 and 178 greater than 2-fold (7.8% and 3.8% of present genes respectively). Although a greater than 2-fold change cut off eliminates most false positives (estimated error rate, 16%), this specificity is at the expense of many putatively regulated genes. In the 1.5–2.0 fold range in the N vs. Ax group, there are still three times as many genes as are in the N vs. N group. (369 genes [N vs. Ax] vs. 122 genes [N vs. N]; Figure 1E).
In order to distinguish systematic from random differences in the array intensity levels, a two-tailed t-test was performed for both comparison groups, which was possible since each group consisted of three independent experiments. One of the fundamental problems in analyzing microarray data is the risk of false positive results due to multiple hypothesis testing. The most conservative approach to this problem, a Bonferroni-corrected significance threshold, would not result in any significantly regulated genes, as the adjusted threshold would be 1.06 × 10-5, while the lowest P value attained in our data set for the N vs. Ax. comparison was 1.51 × 10-5. This would also rule out identification of genes via step-down methods based on the Bonferroni correction. Therefore, we attempted to assess the accumulation of false positives empirically, by testing multiple hypotheses in our system on a sample not expected to include any true positives, i.e. the N. vs. N. comparison. Those probe sets with a unadjusted significance of P < 0.05 (blue), P < 0.01 (green) or P < 0.001 (red) are indicated on the scatter plots in Figures 1C and 1D, while the distribution of significant differences at different fold-changes is illustrated in the table in Figure 1E. Of the 4599 detected genes in the naïve samples, 109 show a significant difference between the two sets (2.4%, P < 0.05). The vast majority of these genes, however, exhibit low fold changes (<1.5-fold, 89%, Figure 1E).
Of the 369 genes that show a fold-difference of 1.5–2.0 and the 178 that were above 2-fold in the N vs. Ax comparison, 223 (60%) and 84 (57%) respectively were not significant (Figure 1E, P < 0.05). These data indicate that using a fold-difference of greater than 2 from triplicate samples to select regulated genes will include many false positives. However, combining fold difference with significant difference in a two dimensional matrix can reduce the false positive error rate drastically. The combination of a >1.5-fold and significant difference (P < 0.05) identifies 12 genes in the N vs. N comparison compared with 240 genes for N vs. Ax, a 20-fold difference (Figure 1E). The estimated error of 5% using these two criteria is much less than that obtained using just above 2-fold (16%) or significance alone (20%) and prevents many genes regulated below 2-fold being excluded from consideration of regulation.
The inherent array error rate (determined by the significant difference rate in the N vs. N analysis) is clustered at low fold changes (Figure 1E). However, many more genes achieve statistical significance at these small fold changes in the N vs. Ax comparison than in the N vs. N group (1.15–1.5 fold: 6.2% [N vs. Ax] and 1.9% [N vs. N]; Figure 1E) indicating that some low fold differences may be real. The degree of significance (P < 0.001; 0.01, 0.05) helps distinguish erroneous changes at these low fold levels (Figure 1E).
Comparison of array data with the literature
Table 1 (see additional file 1) lists those genes previously shown to be regulated in various sciatic nerve injury models using a variety techniques, as well as the expression intensity, fold change and P value derived for these genes from the arrays. Of 69 genes reported in the literature to be expressed in the DRG (Table 1, see additional file 1) 16 are not detected by the arrays indicating that array sensitivity is an issue. Amongst these are particular functional classes, i.e. such as G protein coupled receptors, which are expressed at low levels [12].
25 of the 53 detected genes matched the criteria (>1.5-fold, P < 0.05) for defining regulated genes across the N vs. Ax comparison, and all of these were concordant with the changes reported in the literature (Table 1, see additional file 1). In addition 4 of the 53 detected genes whose levels had previously been described as unchanged by axotomy also did not vary on the arrays. Mismatches between changes expected from the literature and the arrays may be due to a failure of small differences to achieve statistical significance when using a triplicate analysis. Furthermore, differences in the timing, the nature of the injury models used and detection methods employed between this and earlier studies make detailed analysis of these data unfeasible. In order to detect true false positive and negative rates a direct comparison needs to be made between the array data set and transcript levels measured in samples equivalent to those used for array hybridization.
Northern blotting validation of transcriptional regulation
Twenty four genes detected by the arrays as being present in the DRG and displaying up-, down- or no regulation three days after sciatic nerve axotomy were analyzed by replicate quantitative slot blot (Figure 2) and Northern blot analysis (Figure 3). The genes were chosen in a non-random fashion to encompass the range of hybridization intensities seen on the arrays, with expression levels less than 1000 to intensity levels greater than 10,000. Genes were also chosen which displayed a wide range of fold-changes and varying p values and possible functions were taken into account (Table 2).
Triplicate Northern slot blots for three genes with altered expression, 2 showing an increase after axotomy (A and B), one a decrease (C). Cyclophilin a gene known not to alter following axotomy shows equal loading of the slot blots and acts as a normalization control (see methods) (D). The histograms indicate the Northern slot blot data expressed as percentage of naïve expression levels or above the graphs as fold change. Below the blots is the gene expression fold change calculated from the triplicate microarray data. ** P < 0.01, * P < 0.05 (two-tailed unpaired t-test). E. Correlation between 19 separate mRNA fold-differences calculated by triplicate slot blot and those calculated from the triplicate arrays. Linear regression line of best fit is also plotted, scale does not represent values between -1 and 1 as fold change values within this range do not exist due to the method of calculation (see methods).
Northern blots showing differences in expression of 24 genes in naïve/control total DRG RNA (left lane) versus 3 day sciatic nerve axotomy total DRG RNA (right lane). Gene descriptions and accession numbers are shown above each blot. The relative positions of 28S and 18S ribosomal RNA are indicated which migrate at 4.7 kb and 1.9 kb respectively. Abbreviations: GTPcyclo GTP cyclohydrolase-1; Endo-1 Endothelin-1; CB-1 Cannabinoid receptor-1; CCK-B Cholecystokinin-B receptor; 53 Kd Stromelysin-1 (matrix metalloproteinase-3); SproR Small proline rich protein-1A; ET-B Endothelin receptor-B; MSS4 Guanine nucleotide releasing protein; IES-JE Monocyte chemoattractant protein-1; PKBS Peripheral benzodiazepine receptor; 5HT3 5HT3A receptor; GADD45 Growth arrest and DNA damage inducible gene alpha; PACAP Pituitary adenylate cyclase activating peptide; Pho Cl - Phospholemman chloride channel; VGF nerve growth factor inducible protein VGF ; HFH-2 HNF-3/fork-head homolog-2; GFRα-1 GDNF receptor alpha-1 subtype; CLP36 PDZ and LIM domain protein-1 (ELFIN); MET-1 Metallothionein-1L; CCHL2A Calcium channel α-2 subunit; α-Macro alpha(2)-macroglobulin; Giα sub Guanine nucleotide-binding protein G-I, α subunit; SNAP-25 Synaptosomal associated protein 25A & B.
Figure 2 illustrates representative triplicate slot blots for three genes with altered expression, 2 showing an increase after axotomy (Figure 2A and 2B) and one a decrease (Figure 2C). Cyclophilin, a gene known not to alter following axotomy[13], was used to correct for loading (Figure 2D). Each blot was prepared from independent L4 and L5 DRG RNA samples extracted from different groups of animals than those used for the arrays.
Triplicate slot blots were produced for all 24 genes present in Table 2. The concordance rate between the microarray and slot blot fold changes for the 24 genes was 83% overall but concordance depended on gene expression level (Table 2). Genes expressed at low levels in the arrays tended to display fold changes that varied from that found by the slot blots. It is likely that this is a consequence of the unreliable average intensity estimation for low level genes due to limited array sensitivity (Figure 1A). If genes with a mean intensity expression level of less than 1000 are removed from the analysis (top five rows, Table 2), and the array fold changes of the remaining 19 genes are plotted against the slot blot fold changes, a very strong correlation is evident (linear regression r2 = 0.8567) (Figure 2E). The concordance rate of this group is 95% (using >1.5-fold, P < 0.05 to predict regulation). This shows that within the bounds of array sensitivity, average fold changes for genes found to differ significantly between triplicate control and experimental arrays represent a reasonable estimate of the relative changes in gene expression.
Northern blot analysis, although requiring more tissue, both confirms that the correct transcript size is recognized by the probe and enables splice variants to be identified (Figure 3). All 24 genes in Table 2 were analyzed in this way and alternate splice forms were evident for 5 genes.
A list of all the putatively regulated genes within the DRG detected by the arrays (>1.5-fold and P < 0.05) 3 days following a sciatic nerve transection are given as additional data in table 3 (see additional file 2).
In situHybridization
Dorsal root ganglia as well as containing the cell bodies of primary sensory neurons also contain satellite glial cells, Schwann cells and immune cells and it is important, therefore to determine which of these cell types express the regulated genes. Here we use isotopic- in situ hybridization (ISH) to determine the cellular localization of 6 transcripts (Figure 4). All genes selected are neuronal. GTPcylo, IES-JE, CCHL2A and VGF all show a marked increase in the number of labeled neurons while SNAP25 and the 5HT3 receptor show a decrease in numbers and intensity of labeled DRG neurons following axotomy (Figure 4).
Darkfield photomicrographs of 6 μm thick L4 DRG sections processed for in situ hybridization with probes indicated. GTPcyclo, IES-JE, CCHL2A and VGF transcripts show a strong upregulation following injury compared to the naïve. SNAP25, and 5HT3 receptor mRNA show a downregulation following injury to the peripheral neurons. Scale bar = 200 μm.
Time dependent changes in transcription
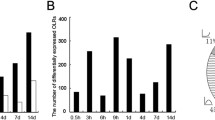
The pattern of expression of four genes within the DRG (Lysozyme, PACAP, MET-1 and SNAP25) were further quantified across a time course of 1, 3, 7 and 14 days post axotomy by triplicate northern slot blot (Figure 5). These genes show diverse patterns of regulation. While lysozyme, PACAP, MET-1 are all up-regulated, for MET1 this is only transient. Identical results were found by in situ hybridization (Fig. 4). Profiling changes in transcription over time is therefore, important.
Axotomy induced changes in expression of Lysozyme, PACAP, MET-1 and SNAP25 transcript levels over time. The histograms indicate the triplicate Northern slot blot data expressed as percentage of naïve expression levels 1, 3, 7 and 14 days post axotomy. ***P < 0.001 ** P < 0.01, * P < 0.05 (two-tailed unpaired t-test).
Potential roles of injury-regulated genes
Table 4 (see additional file 3) indicates the pattern of expression, class, action, and functional role for 15 previously uncharacterized, and 9 novel injury-regulated genes, amongst the 24 validated genes. Five of these genes have been reported in DRG neurons previously and six of the genes are expressed in PC12 cells, a pheochromocytoma-derived cell line that shares neural crest origin with neurons of the DRG. Two genes are expressed in macrophages and 6 in glial cells. Three genes have been reported to show axotomy-induced regulation in non-DRG neurons (sympathetic and motor neurons). These genes belong to several different functional classes and from what is known about their functional role in other cells, may have a role in regeneration, cell survival, or alterations in sensory processing after nerve injury (Table 4, see additional file 3). Once a change in expression of the 240 genes in Table 3 have been validated, similar analyses of their putative function need to be made in order to begin to address what role the genes play in the adaptive and maladaptive response to nerve injury.
Global changes in transcription
The 240 genes defined as putatively regulated (>1.5-fold, P < 0.05) within the DRG 3 days after sciatic nerve section represent 5.1% of the detected genes, and of these 2.8% are up-regulated and 2.3% down-regulated (Figure 6A). These genes were assigned into functional classes and characterized as up- and down-regulated. Some gene classes are selectively regulated after axotomy (Figure 6B). Genes associated with the cell cytoskeleton, positive and negative regulators of apoptosis as well as immune markers tend to be up-regulated, whereas genes involved in membrane excitability (ion channels) or neurotransmission (neurotransmitters and vesicle trafficking genes) tend to be down-regulated.
Percentage of detected genes regulated above 1.5-fold (P < 0.05) with the proportion up and down regulated also shown (A). Genes regulated above 1.5-fold (P < 0.05) were classified into functional classes, plotted are the percent of up and down regulated genes respectively in each functional class (B).
Discussion
Although many studies have used microarray technology for expression profiling, the criteria for determining putatively regulated genes from such analyses remain poorly defined, the current standard being a >2-fold difference in replicate arrays. Using Affymetrix high-density rat oligonucleotide-array data from control (naïve, non-injured) and experimental (axotomy, 3 day post peripheral nerve injury) DRGs, and comparing the differences detected in control vs. control and control vs. experimental arrays, we have explored the optimal criteria for minimizing the false positive error rate. A >2-fold criterion alone was found to generate a substantial proportion of potential false positives, while also excluding many putatively regulated genes. Based on significant difference for each probe set (P < 0.05), 109 genes differed in the control vs. control group, and 556 in the control vs. experimental group, again a prohibitively high false positive rate (20%). However, a combination of fold and significant difference was found to maximize the difference between control vs. control and control vs. experimental array comparisons. Using >1.5-fold and P < 0.05, 240 putatively regulated genes were detected in the experimental group compared with only 12 in the control group, an estimated error rate of 5%. If fold differences of 1.25 to 1.5 are included, many more significantly different genes are recruited (219 in the experimental vs. 41 in the control comparison) reducing the potential false negative rate. At the same time, the estimated false positive error rate rises to 11.5 %, still lower than the 2-fold alone error. Producing a reduction in false positive without also producing false negative differences requires the use both of fold and significant difference. The established 2-fold criterion is much too conservative; when using replicate arrays with pooled samples from multiple animals to reduce biological variation, it loses over 50% of putatively regulated genes and fails to eliminate false positives. The very high concordance rate found between genes predicted from the array analysis to be regulated and measurements of mRNA for 24 genes by quantitative Northern slot blots and in situ hybridization, validates utilization of a two dimensional matrix of fold and significant difference.
One problem with oligonucleotide arrays is sensitivity. Some groups of transcripts, particularly GPCRs, including the opiate receptors, which are expressed in the DRG, [14] fall below the microarray detection threshold. Lack of sensitivity may result from technical issues such as poor probe performance [11] or low copy number transcripts. Tissue heterogeneity and neuronal subpopulation-restricted gene expression may lower the concentration of a transcript in the total RNA sample to below detection threshold [10].
Recently Xiao et al [9] published a study using 7.5 K cDNA arrays to analyze genes regulated in the DRG 2, 7, 14 and 28 days following sciatic nerve axotomy. Of the genes listed in Table 4, seven were found in both studies (Peripheral Benzodiazepine receptor, 5HT3, Gadd45, CLP36, Lysozyme, SNAP25 and VGF). Of those genes whose expression was analyzed across time in this study (Figure 5) two, Lysozyme and SNAP25, were also detected by Xiao et al [9]. Both genes show very similar expression patterns over time in the two studies.
In a different cDNA array study, also recently published, 16 genes were identified to be regulated by at least 2-fold one week after sciatic axotomy [8]. Of these, seven were also identified by our study. Five of the non-concordant genes were not represented by known probe sets on the array used here. Others changed, but did not reach statistical significance, and likely represented differences in regulation levels relative to the time points used (1 week as opposed to 3 days). One gene verified in this study, the small proline-rich protein 1A (Table 2), was characterized in detail by Bonilla et al, who show that it colocalizes with filamentous actin in membrane ruffles and augments axonal outgrowth [8].
Comparison of the global lists of regulated genes between this study and other reports are complicated by the differing criteria used to identify regulated genes in the different studies. However, it is encouraging to note that a reasonable number of genes have been cross identified in three separate array studies looking at the effects of peripheral nerve injury on dorsal root ganglion gene expression, pointing to the reliability of array technologies across differing platforms.
Why is the number of genes regulated in sensory neurons after peripheral nerve injury so large? The injury constitutes an enormous stress to the cell and also deprives it from contact with its normal target-derived neurotrophic support. Both of these events represent perturbations from normal cellular functioning that require a variety of compensatory responses. A major issue is cell survival. Injured adult DRG neurons do not die after axonal injury [15] as a result of the upregulation of cell survival factors such as the small heat shock protein HSP27 [16]. We now find several genes that could have a survival-promoting role, including the peripheral benzodiazepine receptor, whose agonists are potent antiapoptotic compounds [17], GADD45A, which maintains genomic stability and appears to be anti-apoptotic in neurons after ischemic damage [18], metallothionein 1L, which is a free radical scavenger [19], GTP cyclohydrolase 1, which is obligatory for the actions of NGF on PC12 cells [20], and IES-JE (MCP-1) which attracts macrophages, which in turn have been reported to have a role in neuronal survival [21].
Neurons with axons in the peripheral nervous system can successfully regenerate when injured [22]. This is in part due to an environment permissive for axonal growth but also due to the upregulation of regeneration-associated genes (RAGs) which increase intrinsic growth capacity [23]. Three examples of RAGs are GAP-43 and CAP-23 [24] and sprp1A [8]. We now find several other potential candidates including; α2 macroglobulin, CLP36, and VGF all of which have been described to promote neurite outgrowth or interact with the cytoskeleton in a way that may promote regeneration (see Table 4 [additional file 3] for references). MMP3 also known as 53 kD polypeptide, transin or stromelysin-1, a secreted protease, may facilitate neurite growth by dissolving the extracellular matrix of the basal lamina at the growing tip of the axon (see Table 4 for references).
Sensory neurons react to peripheral nerve injury by increasing their excitability, changing their constitutive synaptic transmitter profile and even their synaptic contacts in the spinal cord. Interestingly, an analysis of the functional classes of the genes differentially expressed (Figure 6) reveals the contribution that alterations in transcription make to these processes. Several of these changes are maladaptive in the sense that they contribute to the generation of the abnormal sensations that constitute neuropathic pain, by producing ectopic spontaneous activity into the CNS, altering synaptic drive, increasing excitability and diminishing inhibitory action in the dorsal horn of the spinal cord, and by making novel synaptic contacts with inappropriate neurons [1, 4]. A number of the injury-regulated genes we describe here may contribute directly to this altered sensory processing; CB1 receptor, VGF, the phospholemman ion channel, SNAP25 A, endothelin-1 and the ligand-gated 5-HT3 receptor (Table 4, see additional file 3). We have only validated a small fraction of the putatively regulated genes detected by the arrays. Our analysis indicates that at a conservative estimate over two hundred genes (240 genes, >1.5-fold, P < 0.05) are regulated by the injury. These include genes of a number of distinct functional classes, some of which appear to change in a uniform way. Ion channels and neurotransmitter related genes tend to decrease their expression while those associated with the cytoskeleton increase. This raises the possibility that numbers of different genes may be regulated together by common changes in signal transduction and transcription induced by the injury, and contribute in this way to the coordinated changes that constitute the axotomy response.
Discovering and validating regulated genes by array analysis is clearly only the first step in elucidating their functional role. An evaluation of the cellular localization and distribution of the gene product as well as loss and gain of function manipulations in vitro and in vivo are required. Functional studies cannot yet be performed at a high throughput level and this represents one of the major limiting factors in analyzing the large numbers of candidate genes identified by microarray analysis. Elucidating the specific role of genes regulated in sensory neurons after peripheral nerve injury will provide insight though into many major biological issues including cell survival, growth, intercellular communication and the factors that contribute to sensory abnormalities. Microarray technology provides a powerful tool for beginning this analysis in a high throughput mode by revealing the extent of change in neuronal gene expression. Such analysis requires replicate measures to minimize variability, calculation of fold and significant differences to detect regulated genes with minimal false positive and negative confounders and validation of individual genes.
Conclusions
1: A comparison of two naïve (control) triplicate data sets allowed determination of the inherent variation present in the oligonucleotide array data. Analysis revealed a relatively high false positive error rate in the naïve versus axotomy (experimental) comparisons (16%) when using only a 2-fold criterion for regulated gene detection.
2: When statistical significance (P < 0.05) was combined with a fold difference of greater than 1.5 as the criteria for detecting putatively regulated genes, not only was the estimated false positive error substantially reduced (to 5%), but also many more genes were identified compared with the established 2-fold criterion, in the naïve versus axotomy comparison (240 as opposed to 178 genes).
3: All 240 genes displaying a fold change of greater than 1.5 and a P-value of less than 0.05 are listed.
4: To validate these findings we performed quantitative triplicate slot blots for 24 genes, enabling a direct comparison between the statistically analyzed array data and the mRNA measurements. These comparisons show strong concordance between putative and actual regulation as well as a direct relationship between array fold change and mRNA fold change.
5: Some genes were further quantified across a time course of 1, 3, 7 and 14 days post axotomy. These genes show diverse patterns of regulation in response to nerve injury and underlie the importance of including time as a dimension in any expression profile study.
6: Genes detected from the arrays, as being likely to be regulated, were assigned to functional groups. These data suggest the reorganization of cell structural components following peripheral nerve injury, activation of genes expressed by immune and inflammatory cells, and a down-regulation of genes involved in neurotransmission.
Methods
Surgical procedures
All procedures were performed in accordance with Massachusetts General Hospital animal care regulations. Adult male Sprague Dawley rats (200–300 g) were anesthetized with halothane. For the sciatic nerve transection (axotomy), the left sciatic nerve was exposed at the mid thigh level, ligated with 3/0 silk and sectioned distally. The wound was sutured in two layers, and the animals were allowed to recover.
Tissue and RNA preparation
Animals were terminally anesthetized with CO2, the L4 and L5 DRGs rapidly removed, and stored at -80°C. Total RNA was extracted from homogenized DRG samples using acid phenol extraction (TRIzol reagent, Gibco-BRL). RNA concentration was evaluated by A260 measurement and quality assessed by electrophoresis on a 1.5% agarose gel. Each RNA sample used for hybridization of each array was extracted from rat L4 and L5 DRGs (10 ganglia pooled from 5 animals, per sample).
Microarray Analysis
Affymetrix rat genome U34A oligonucleotide microarrays, representing 8799 known transcripts and expressed sequence tags (ESTs), were used (Santa Clara, CA http://www.affymetrix.com). Oligonucleotides are arranged in pairs corresponding to different regions of the target mRNA with multiple probe pairs. Each probe pair consists of a 25 nucleotide perfect match (PM) to the target region coupled with a 25-mer with a single mismatch (MM) at the 13th nucleotide. Transcript abundance is estimated by analysis of signal intensity of the PM/MM pairs. The arrays are hybridized with biotin-labeled cRNA, prepared as per standard Affymetrix protocol. Briefly, total RNA (8 μg) from DRGs was reverse transcribed using an oligo-dT primer coupled to a T7 RNA polymerase binding site. Double-stranded cDNA was made and biotinylated-cRNA synthesized using T7 polymerase. The cRNA was hybridized for 16 hours to an array, followed by binding with a streptavidin-conjugated fluorescent marker, and then incubated with a polyclonal anti-streptavidin antibody coupled to phycoerythrin as an amplification step. Following washing, the chips were scanned with a Hewlett-Packard GeneArray laser scanner and data analyzed using GeneChip software. External standards were included to control for hybridization efficiency and sensitivity.
Hybridization levels for each species of mRNA detected on the arrays are expressed by intensity (signal) and as present (P), marginal (M) or absent (A) calls, calculated by Affymetrix software (MAS 5.0, α1 = 0.04 α2 = 0.06). To normalize the array data standard Affymetrix protocols were employed, each array was scaled to a target signal of 2500 across all probe sets (MAS 5.0).
The arrays were grouped for two comparisons: two triplicate sets of naïve data compared with one another, and one triplicate naïve set compared with one triplicate post-axotomy set. The individual naïve arrays included in each triplicate set were picked randomly. A probe set was determined undetected if it received an A call in all of the six arrays involved in the comparison. Detected were Present or Marginal by MAS5.0 in at least one array for each analysis. Mean signal and standard deviation were calculated for each detected probe set. The p-value for rejecting the null hypothesis that the mean signals were equal between the two triplicate sets was calculated using an unpaired, two-tailed t-test for independent samples with unequal variance (Satterthwaite's method). Fold-differences between the mean signals (A and B) in the two triplicate sets were calculated as max(A, B) / min(A, B) with down regulation relative to naïve expressed as negative.
cDNA Probe production
To generate specific probes for Northern blot hybridization experiments, primers based on the rat accession number provided by Affymetrix were designed, primer pairs were chosen using the Primer3 software http://www-genome.wi.mit.edu/ from the 1000 most 3' nucleotides within each accession sequence. PCR was performed on cDNA reverse transcribed from total RNA, extracted from lumbar DRGs, using poly-dT as a primer to obtain cDNA fragments (141 to 596 bp). These fragments were subsequently cloned into the PCRII vector (TA cloning Kit, Invitrogen) and the identity of each was confirmed by sequencing in both directions. These cDNAs were gel-purified and used to produce 32P-labeled cDNA probes (Prime-It kit, Stratagene).
Northern blot analysis
Total RNA was size separated by electrophoresis on a 1.5% agarose/formaldehyde gel (10 μg of total RNA per lane) and transferred to a Hybond N+ nylon membrane. Membranes were hybridized with labeled-probes (see above) in ExpressHyb (Clontech) overnight at 65°C, washed and exposed to X-ray film with an intensifying screen at -80°C.
Slot Blots
Total RNA (1.25 μg) was directly transferred to Hybond N+ nylon membrane under vacuum using a Hoefer PR648 slot blot apparatus (Amersham Pharmacia Biotech) as described in [25]. The slot blots were produced in batches of ten from a relevant master mix and probed with the same cDNA under the same conditions as described for Northern blot analysis. Levels of hybridization were quantified using the 24450 phosphorimager system (Molecular Dynamics, Sunnyvale CA.). One of the blots in each batch was probed for cyclophilin to act as a loading control. The other nine blots were used to measure the expression levels of individual genes. Loading levels between samples on each blot were normalized using the cyclophilin levels from the control blot.
Isotopic in situHybridization
DRGs were rapidly removed, embedded in OCT (Tissue Tek) and frozen. Sections were cut serially at 6 μm. Isotopic-in situ hybridization was carried out using forty-eight base pair oligonucleotide probes, designed to have 50% G-C content and be complementary to the mRNAs whose accession numbers were provided by Affymetrix. Probes were 3'-end labeled with 35S or 33P-dATP using a terminal transferase reaction and hybridization carried out [26]. Autoradiograms were generated by dipping slides in NTB2 nuclear track emulsion and storing in the dark at 4°C. Sections were exposed for 1–8 weeks (depending on the abundance of transcript), developed, fixed and viewed under darkfield using a fiber-optic darkfield stage adapter (MVI). Controls to confirm specificity of oligonucleotide probes included hybridization of sections with labeled probe with a 1,000-fold excess of cold probe or labeled probe with a 1,000-fold excess of another, dissimilar cold probe of the same length and similar G-C content.
References
Devor M, Seltzer Z: Pathophysiology of damaged nerves in realtion to chronic pain. In Textbook of Pain. Edited by: Wall PD, Melzack R. 1999, Edinburgh: Churchill Livingstone, 129-164.
Kerr BJ, Thompson SW, Wynick D, McMahon SB: Endogenous galanin is required for the full expression of central sensitization following peripheral nerve injury. Neuroreport. 2001, 12: 3331-3334. 10.1097/00001756-200110290-00037.
Hokfelt T, Zhang X, Wiesenfeld-Hallin Z: Messenger plasticity in primary sensory neurons following axotomy and its functional implications. TINS. 1994, 17 (1): 22-30. 10.1016/0166-2236(94)90031-0.
Woolf CJ, Mannion RJ: Neuropathic pain: aetiology, symptoms, mechanisms, and management. Lancet. 1999, 353: 1959-1964. 10.1016/S0140-6736(99)01307-0.
Wisden W, Gundlach AL, Barnard EA, Seeburg PH, Hunt SP: Distribution of GABAA receptor subunit mRNAs in rat lumbar spinal cord. Brain Res Mol Brain Res. 1991, 10: 179-183. 10.1016/0169-328X(91)90109-B.
Fernandes KJL, Tetzlaff W: Gene Expression in Axotomized Neurons: Identifying the Intrinsic Determinants of Axonal Growth. In Axonal Regeneration in the Central Nervous System. Edited by: Ingoglia NA, Murray M. 2001, New York: Marcel Dekker, 219-266.
van Berkum NL, Holstege FC: DNA microarrays: raising the profile. Curr Opin Biotechnol. 2001, 12: 48-52. 10.1016/S0958-1669(00)00173-7.
Bonilla IE, Tanabe K, Strittmatter SM: Small proline-rich repeat protein 1A is expressed by axotomized neurons and promotes axonal outgrowth. J Neurosci. 2002, 22: 1303-1315.
Xiao HS, Huang QH, Zhang FX, Bao L, Lu YJ, Guo C, Yang L, Huang WJ, Fu G, Xu SH, et al: Identification of gene expression profile of dorsal root ganglion in the rat peripheral axotomy model of neuropathic pain. Proc Natl Acad Sci U S A. 2002, 99: 8360-8365. 10.1073/pnas.122231899.
Mirnics K, Middleton FA, Lewis DA, Levitt P: Analysis of complex brain disorders with gene expression microarrays: schizophrenia as a disease of the synapse. Trends Neurosci. 2001, 24: 479-486. 10.1016/S0166-2236(00)01862-2.
Schadt EE, Li C, Su C, Wong WH: Analyzing high-density oligonucleotide gene expression array data. J Cell Biochem. 2000, 80: 192-202. 10.1002/1097-4644(20010201)80:2<192::AID-JCB50>3.3.CO;2-N.
Kieffer BL: Molecular aspects of opioid receptors. In Handbook of experimental pharmacology; The pharmacology of pain. Edited by: Dickenson AH, Besson JM. 1997, 281-303.
Costigan M, Mannion RJ, Kendall G, Lewis SE, Campagna JA, Coggeshall RE, Meredith-Middleton J, Tate S, Woolf CJ: Heat Shock Protein 27: developmental regulation and expression after peripheral nerve injury. J Neurosci. 1998, 18: 5891-5900.
Mansour A, Fox CA, Akil H, Watson SJ: Opioid-receptor mRNA expression in the rat CNS: anatomical and functional implications. Trends Neurosci. 1995, 18: 22-29. 10.1016/0166-2236(95)93946-U.
Tandrup T, Woolf CJ, Coggeshall RE: Delayed loss of small dorsal root ganglion cells after transection of the rat sciatic nerve. J Comp Neurol. 2000, 422: 172-180. 10.1002/(SICI)1096-9861(20000626)422:2<172::AID-CNE2>3.0.CO;2-H.
Lewis SE, Mannion RJ, White FA, Coggeshall RE, Beggs S, Costigan M, Martin JL, Dillmann WH, Woolf CJ: A role for HSP27 in sensory neuron survival. J Neurosci. 1999, 19: 8945-8953.
Bono F, Lamarche I, Prabonnaud V, Le Fur G, Herbert JM: Peripheral benzodiazepine receptor agonists exhibit potent antiapoptotic activities. Biochem Biophys Res Commun. 1999, 265: 457-461. 10.1006/bbrc.1999.1683.
Chen J, Uchimura K, Stetler RA, Zhu RL, Nakayama M, Jin K, Graham SH, Simon RP: Transient global ischemia triggers expression of the DNA damage-inducible gene GADD45 in the rat brain. J Cereb Blood Flow Metab. 1998, 18: 646-657. 10.1097/00004647-199806000-00007.
Campagne MV, Thibodeaux H, van Bruggen N, Cairns B, Lowe DG: Increased binding activity at an antioxidant-responsive element in the metallothionein-1 promoter and rapid induction of metallothionein-1 and -2 in response to cerebral ischemia and reperfusion. J Neurosci. 2000, 20: 5200-5207.
Anastasiadis PZ, Bezin L, Imerman BA, Kuhn DM, Louie MC, Levine RA: Tetrahydrobiopterin as a mediator of PC12 cell proliferation induced by EGF and NGF. Eur J Neurosci. 1997, 9: 1831-1837.
Subang MC, Richardson PM: Influence of injury and cytokines on synthesis of monocyte chemoattractant protein-1 mRNA in peripheral nervous tissue. Eur J Neurosci. 2001, 13: 521-528. 10.1046/j.1460-9568.2001.01425.x.
Fawcett JW, Keynes RJ: Peripheral nerve regeneration. Ann Rev Neurosci. 1990, 13: 43-60. 10.1146/annurev.neuro.13.1.43.
Skene JHP: Axonal growth associated proteins. Ann Rev Neurosci. 1989, 12: 127-156. 10.1146/annurev.ne.12.030189.001015.
Bomze HM, Bulsara KR, Iskandar BJ, Caroni P, Pate Skene JH: Spinal axon regeneration evoked by replacing two growth cone proteins in adult neurons. Nat Neurosci. 2001, 4: 38-43. 10.1038/82881.
Sambrook J, Russell DW: Molecular cloning : a laboratory manual. New York, Cold Spring Harbor Laboratory Press. 2001, Ref Type: Serial (Book, Monograph), 3rd
Karchewski LA, Kim FA, Johnston J, McKnight RM, Verge VM: Anatomical evidence supporting the potential for modulation by multiple neurotrophins in the majority of adult lumbar sensory neurons. J Comp Neurol. 1999, 413: 327-341. 10.1002/(SICI)1096-9861(19991018)413:2<327::AID-CNE11>3.0.CO;2-3.
Shi TS, Winzer-Serhan U, Leslie F, Hokfelt T: Distribution and regulation of alpha(2)-adrenoceptors in rat dorsal root ganglia. Pain. 2000, 84: 319-330. 10.1016/S0304-3959(99)00224-9.
Birder LA, Perl ER: Expression of alpha2-adrenergic receptors in rat primary afferent neurones after peripheral nerve injury or inflammation. J Physiol. 1999, 515 (Pt 2): 533-542.
Gallinat S, Yu M, Dorst A, Unger T, Herdegen T: Sciatic nerve transection evokes lasting up-regulation of angiotensin AT2 and AT1 receptor RNA min adult rat dorsal root ganglia and sciatic nerves. Brain Res Mol Brain Res. 1998, 57: 111-122. 10.1016/S0169-328X(98)00079-5.
Levy D, Zochodne DW: Increased mRNA expression of the B1 and B2 bradykinin receptors and antinociceptive effects of their antagonists in an animal model of neuropathic pain. Pain. 2000, 86: 265-271. 10.1016/S0304-3959(00)00256-6.
Zhang X, Dagerlind A, Elde RP, Castel MN, Broberger C, Wiesenfeld-Hallin Z, Hokfelt T: Marked increase in cholecystokinin B receptor messenger RNA levels in rat dorsal root ganglia after peripheral axotomy. Neurosci. 1993, 57: 227-233. 10.1016/0306-4522(93)90057-M.
Antunes Bras JM, Laporte AM, Benoliel JJ, Bourgoin S, Mauborgne A, Hamon M, Cesselin F, Pohl M: Effects of peripheral axotomy on cholecystokinin neurotransmission in the rat spinal cord. J Neurochem. 1999, 72: 858-867. 10.1046/j.1471-4159.1999.720858.x.
Xu ZQ, Shi TJ, Landry M, Hokfelt T: Evidence for galanin receptors in primary sensory neurones and effect of axotomy and inflammation. Neurorep. 1996, 8: 237-242.
Zhang X, Xu ZO, Shi TJ, Landry M, Holmberg K, Ju G, Tong YG, Bao L, Cheng XP, Wiesenfeld-Hallin Z, et al: Regulation of expression of galanin and galanin receptors in dorsal root ganglia and spinal cord after axotomy and inflammation. Ann N Y Acad Sci. 1998, 863: 402-413.
Sten Shi TJ, Zhang X, Holmberg K, Xu ZQ, Hokfelt T: Expression and regulation of galanin-R2 receptors in rat primary sensory neurons: effect of axotomy and inflammation. Neurosci Lett. 1997, 237: 57-60. 10.1016/S0304-3940(97)00805-7.
Zhang X, Wiesenfeld Z-Hallin, Hokfelt T: Effect of peripheral axotomy on expression of neuropeptide Y receptor mRNA in rat lumbar dorsal root ganglia. Eur J Neurosci. 1994, 6: 43-57.
Landry M, Holmberg K, Zhang X, Hokfelt T: Effect of axotomy on expression of NPY, galanin, and NPY Y1 and Y2 receptors in dorsal root ganglia and the superior cervical ganglion studied with double-labeling in situ hybridization and immunohistochemistry. Exp Neurol. 2000, 162: 361-384. 10.1006/exnr.1999.7329.
Zhang X, Bao L, Shi TJ, Ju G, Elde R, Hokfelt T: Down-regulation of mu-opioid receptors in rat and monkey dorsal root ganglion neurons and spinal cord after peripheral axotomy. Neurosci. 1998, 82: 223-240. 10.1016/S0306-4522(97)00240-6.
Zhang X, Bao L, Arvidsson U, Elde R, Hokfelt T: Localization and regulation of the delta-opioid receptor in dorsal root ganglia and spinal cord of the rat and monkey: evidence for association with the membrane of large dense-core vesicles. Neurosci. 1998, 82: 1225-1242. 10.1016/S0306-4522(97)00341-2.
Fukuoka T, Tokunaga A, Kondo E, Miki K, Tachibana T, Noguchi K: Change in mRNAs for neuropeptides and the GABA(A) receptor in dorsal root ganglion neurons in a rat experimental neuropathic pain model. Pain. 1998, 78: 13-26. 10.1016/S0304-3959(98)00111-0.
Bradbury EJ, Burnstock G, McMahon SB: The Expression of P2X3 Purinoreceptors in Sensory Neurons: Effects of Axotomy and Glial-Derived Neurotrophic Factor. Mol Cell Neurosci. 1998, 12: 256-268. 10.1006/mcne.1998.0719.
Yiangou Y, Facer P, Birch R, Sangameswaran L, Eglen R, Anand P: P2X3 receptor in injured human sensory neurons. Neurorep. 2000, 11: 993-996.
Michael GJ, Priestley JV: Differential expression of the mRNA for the vanilloid receptor subtype 1 in cells of the adult rat dorsal root and nodose ganglia and its downregulation by axotomy. J Neurosci. 1999, 19: 1844-1854.
Krekoski CA, Parhad IM, Clark AW: Attenuation and recovery of nerve growth factor receptor mRNA in dorsal root ganglion neurons following axotomy. J Neurosci Res. 1996, 43: 1-11.
Bennett DL, Boucher TJ, Armanini MP, Poulsen KT, Michael GJ, Priestley JV, Phillips HS, McMahon SB, Shelton DL: The glial cell line-derived neurotrophic factor family receptor components are differentially regulated within sensory neurons after nerve injury. J Neurosci. 2000, 20: 427-437.
Kashiba H, Hyon B, Senba E: Glial cell line-derived neurotrophic factor and nerve growth factor receptor mRNAs are expressed in distinct subgroups of dorsal root ganglion neurons and are differentially regulated by peripheral axotomy in the rat. Neurosci Lett. 1998, 252: 107-110. 10.1016/S0304-3940(98)00558-8.
Bergman E, Fundin BT, Ulfhake B: Effects of aging and axotomy on the expression of neurotrophin receptors in primary sensory neurons. J Comp Neurol. 1999, 410: 368-386. 10.1002/(SICI)1096-9861(19990802)410:3<368::AID-CNE2>3.0.CO;2-I.
Kashiba H, Senba E: Up- and down-regulation of BDNF mRNA in distinct subgroups of rat sensory neurons after axotomy. Neurorep. 1999, 10: 3561-3565.
Cho HJ, Kim JK, Park HC, Kim JK, Kim DS, Ha SO, Hong HS: Changes in brain-derived neurotrophic factor immunoreactivity in rat dorsal root ganglia, spinal cord, and gracile nuclei following cut or crush injuries. Exp Neurol. 1998, 154: 224-230. 10.1006/exnr.1998.6936.
Noguchi K, Senba E, Morita Y, Sato M, Tohyama M: Alpha-CGRP and beta-CGRP mRNAs are differentially regulated in the rat spinal cord and dorsal root ganglion. Brain Res Mol Brain Res. 1990, 7: 299-304. 10.1016/0169-328X(90)90080-W.
Verge VM, Wiesenfeld-Hallin Z, Hokfelt T: Cholecystokinin in mammalian primary sensory neurons and spinal cord: in situ hybridization studies in rat and monkey. Eur J Neurosci. 1993, 5: 240-250.
Ji RR, Zhang Q, Zhang X, Piehl F, Reilly T, Pettersson RF, Hokfelt T: Prominent expression of bFGF in dorsal root ganglia after axotomy. Eur J Neurosci. 1995, 7: 2458-2468.
Hokfelt T, Zhang X, Wiesenfeld-Hallin Z: Messenger plasticity in primary sensory neurons following axotomy and its functional implications. Trends Neurosci. 1994, 17: 22-30. 10.1016/0166-2236(94)90031-0.
Murphy PG, Grondin J, Altares M, Richardson PM: Induction of interleukin-6 in axotomized sensory neurons. J Neurosci. 1995, 15: 5130-5138.
Jongsma H, Danielsen N, Sundler F, Kanje M: Alteration of PACAP distribution and PACAP receptor binding in the rat sensory nervous system following sciatic nerve transection. Brain Res. 2000, 853: 186-196. 10.1016/S0006-8993(99)02233-7.
Alvares D, Fitzgerald M: Building blocks of pain: the regulation of key molecules in spinal sensory neurones during development and following peripheral axotomy. Pain. 1999, Suppl 6: S71-S85. 10.1016/S0304-3959(99)00140-2.
Zhang Q, Ji RR, Lindsay R, Hokfelt T: Effect of growth factors on substance P mRNA expression in axotomized dorsal root ganglia. Neurorep. 1995, 6: 1309-1312.
Mulder H, Zhang Y, Danielsen N, Sundler F: Islet amyloid polypeptide and calcitonin gene-related peptide expression are down-regulated in dorsal root ganglia upon sciatic nerve transection. Brain Res Mol Brain Res. 1997, 47: 322-330. 10.1016/S0169-328X(97)00060-0.
Livesey FJ, O'Brien JA, Li M, Smith AG, Murphy LJ, Hunt SP: A Schwann cell mitogen accompanying regeneration of motor neurons. Nature. 1997, 390: 614-618. 10.1038/37615.
Waxman SG, Kocsis JD, Black JA: Type III sodium channel mRNA is expressed in embryonic but not adult spinal sensory neurons, and is reexpressed following axotomy. J Neurophysiol. 1994, 72: 466-470.
Dib-Hajj S, Black JA, Felts P, Waxman SG: Down-regulation of transcripts for Na channel alpha-SNS in spinal sensory neurons following axotomy. Proc Natl Acad Sci U S A. 1996, 93: 14950-14954. 10.1073/pnas.93.25.14950.
Ishikawa K, Tanaka M, Black JA, Waxman SG: Changes in expression of voltage-gated potassium channels in dorsal root ganglion neurons following axotomy. Muscle Nerve. 1999, 22: 502-507. 10.1002/(SICI)1097-4598(199904)22:4<502::AID-MUS12>3.0.CO;2-K.
Okuse K, Chaplan SR, McMahon SB, Luo ZD, Calcutt NA, Scott BP, AN Akopian, Wood JN: Regulation of Expression of the Sensory Neuron-Specific Sodium Channel SNS in Inflammatory and Neuropathic Pain. Mol Cell Neurosci. 1997, 10: 196-207. 10.1006/mcne.1997.0657.
Sleeper AA, Cummins TR, Dib SD-Hajj, Hormuzdiar W, Tyrrell L, Waxman SG, Black JA: Changes in expression of two tetrodotoxin-resistant sodium channels and their currents in dorsal root ganglion neurons after sciatic nerve injury but not rhizotomy. J Neurosci. 2000, 20: 7279-7289.
Luo ZD, Chaplan SR, Higuera ES, Sorkin LS, Stauderman KA, Williams ME, Yaksh TL: Upregulation of dorsal root ganglion (alpha)2(delta) calcium channel subunit and its correlation with allodynia in spinal nerve-injured rats. J Neurosci. 2001, 21: 1868-1875.
Dib SD-Hajj, Tyrrell L, Black JA, Waxman SG: NaN, a novel voltage-gated Na channel, is expressed preferentially in peripheral sensory neurons and down-regulated after axotomy. Proc Natl Acad Sci U S A. 1998, 95: 8963-8968. 10.1073/pnas.95.15.8963.
Lund LM, McQuarrie IG: Axonal regrowth upregulates beta-actin and Jun D mRNA expression. J Neurobiol. 1996, 31: 476-486. 10.1002/(SICI)1097-4695(199612)31:4<476::AID-NEU7>3.3.CO;2-8.
Chong MS, Reynolds ML, Irwin N, Coggeshall RE, Emson PC, Benowitz LI, Woolf CJ: GAP-43 expression in primary sensory neurons following central axotomy. J Neurosci. 1994, 14: 4375-4384.
Woodham P, Anderson PN, Nadim W, Turmaine M: Satellite cells surrounding axotomised rat dorsal root ganglion cells increase expression of a GFAP-like protein. Neurosci Lett. 1989, 98: 8-12. 10.1016/0304-3940(89)90364-9.
Wong J, Oblinger MM: Differential regulation of peripherin and neurofilament gene expression in regenerating rat DRG neurons. J Neurosci Res. 1990, 27: 332-341.
Oblinger MM, Wong J, Parysek LM: Axotomy-induced changes in the expression of a type III neuronal intermediate filament gene. J Neurosci. 1989, 9: 3766-3775.
Chadan S, Moya KL, Portier MM, Filliatreau G: Identification of a peripherin dimer: changes during axonal development and regeneration of the rat sciatic nerve. J Neurochem. 1994, 62: 1894-1905.
Jiang YQ, Pickett J, Oblinger MM: Comparison of changes in beta-tubulin and NF gene expression in rat DRG neurons under regeneration-permissive and regeneration-prohibitive conditions. Brain Res. 1994, 637: 233-241. 10.1016/0006-8993(94)91238-6.
Moskowitz PF, Smith R, Pickett J, Frankfurter A, Oblinger MM: Expression of the class III beta-tubulin gene during axonal regeneration of rat dorsal root ganglion neurons. J Neurosci Res. 1993, 34: 129-134.
Newton RA, Bingham S, Davey PD, Medhurst AD, Piercy V, Raval P, Parsons AA, Sanger GJ, Case CP, Lawson SN: Identification of differentially expressed genes in dorsal root ganglia following partial sciatic nerve injury. Neurosci. 2000, 95: 1111-1120. 10.1016/S0306-4522(99)00515-1.
Tsujino H, Kondo E, Fukuoka T, Dai Y, Tokunaga A, Miki K, Yonenobu K, Ochi T, Noguchi K: Activating transcription factor 3 (ATF3) induction by axotomy in sensory and motoneurons: A novel neuronal marker of nerve injury. Mol Cell Neurosci. 2000, 15: 170-182. 10.1006/mcne.1999.0814.
Herdegen T, Fiallos-Estrada CE, Schmid W, Bravo R, Zimmermann M: The transcription factors c-JUN, JUN D and CREB, but not FOS and KROX-24, are differentially regulated in axotomized neurons following transection of rat sciatic nerve. Brain Res Mol Brain Res. 1992, 14: 155-165. 10.1016/0169-328X(92)90170-G.
De Leon M, Nahin RL, Molina CA, De Leon DD, Ruda MA: Comparison of c-jun, junB, and junD mRNA expression and protein in the rat dorsal root ganglia following sciatic nerve transection. J Neurosci Res. 1995, 42: 391-401.
Kenney AM, Kocsis JD: Temporal variability of jun family transcription factor levels in peripherally or centrally transected adult rat dorsal root ganglia. Brain Res Mol Brain Res. 1997, 52: 53-61. 10.1016/S0169-328X(97)00211-8.
Werner A, Willem M, Jones LL, Kreutzberg GW, Mayer U, Raivich G: Impaired axonal regeneration in alpha7 integrin-deficient mice. J Neurosci. 2000, 20: 1822-1830.
Daniloff JK, Levi G, Grumet M, Rieger F, Edelman GM: Altered expression of neuronal cell adhesion molecules induced by nerve injury and repair. J Cell Biol. 1986, 103: 929-945. 10.1083/jcb.103.3.929.
Zhang Y, Roslan R, Lang D, Schachner M, Lieberman AR, Anderson PN: Expression of CHL1 and L1 by neurons and glia following sciatic nerve and dorsal root injury. Mol Cell Neurosci. 2000, 16: 71-86. 10.1006/mcne.2000.0852.
Gavazzi I, Stonehouse J, Sandvig A, Reza JN, Appiah-Kubi LS, Keynes R, Cohen J: Peripheral, but not central, axotomy induces neuropilin-1 mRNA expression in adult large diameter primary sensory neurons. J Comp Neurol. 2000, 423: 492-499. 10.1002/1096-9861(20000731)423:3<492::AID-CNE11>3.3.CO;2-C.
Araki T, Milbrandt J: Ninjurin, a novel adhesion molecule, is induced by nerve injury and promotes axonal growth. Neuron. 1996, 17: 353-361. 10.1016/S0896-6273(00)80166-X.
Shi TJ, Holmberg K, Xu ZQ, Steinbusch H, de Vente J, Hokfelt T: Effect of peripheral nerve injury on cGMP and nitric oxide synthase levels in rat dorsal root ganglia: time course and coexistence. Pain. 1998, 78: 171-180. 10.1016/S0304-3959(98)00124-9.
Gonzalez-Hernandez T, Rustioni A: Expression of three forms of nitric oxide synthase in peripheral nerve regeneration. J Neurosci Res. 1999, 55: 198-207. 10.1002/(SICI)1097-4547(19990115)55:2<198::AID-JNR7>3.3.CO;2-D.
Gillardon F, Wickert H, Zimmermann M: Differential expression of bcl-2 and bax mRNA in axotomized dorsal root ganglia of young and adult rats. Eur J Neurosci. 1994, 6: 1641-1644.
Gillardon F, Klimaschewski L, Wickert H, Krajewski S, Reed JC, Zimmermann M: Expression pattern of candidate cell death effector proteins Bax, Bcl-2, Bcl-X, and c-Jun in sensory and motor neurons following sciatic nerve transection in the rat. Brain Res. 1996, 739: 244-250. 10.1016/S0006-8993(96)00829-3.
Alberi S, Raggenbass M, de Bilbao F, Dubois-Dauphin M: Axotomized neonatal motoneurons overexpressing the bcl2 proto-oncogene retain functional electrophysiological properties. Proc Natl Acad Sci U S A. 1996, 93: 3978-3983. 10.1073/pnas.93.9.3978.
De Leon M, Welcher AA, Nahin RH, Liu Y, Ruda MA, Shooter EM, Molina CA: Fatty acid binding protein is induced in neurons of the dorsal root ganglia after peripheral nerve injury. J Neurosci Res. 1996, 44: 283-292. 10.1002/(SICI)1097-4547(19960501)44:3<283::AID-JNR9>3.3.CO;2-R.
Cho S, Volpe BT, Bae Y, Hwang O, Choi HJ, Gal J, Park LC, Chu CK, Du J, Joh TH: Blockade of tetrahydrobiopterin synthesis protects neurons after transient forebrain ischemia in rat: a novel role for the cofactor. J Neurosci. 1999, 19: 878-889.
Stegenga SL, Hirayama K, Kapatos G: Regulation of GTP cyclohydrolase I gene expression and tetrahydrobiopterin content in cultured sympathetic neurons by leukemia inhibitory factor and ciliary neurotrophic factor. J Neurochem. 1996, 66: 2541-2545.
Gokin AP, Fareed MU, Pan HL, Hans G, Strichartz GR, Davar G: Local injection of endothelin-1 produces pain-like behavior and excitation of nociceptors in rats. J Neurosci. 2001, 21: 5358-5366.
Milner P, Loesch A, Burnstock G: Endothelin immunoreactivity and mRNA expression in sensory and sympathetic neurones following selective denervation. Int J Dev Neurosci. 2000, 18: 727-734. 10.1016/S0736-5748(00)00054-X.
Giaid A, Gibson SJ, Ibrahim BN, Legon S, Bloom SR, Yanagisawa M, Masaki T, Varndell IM, Polak JM: Endothelin 1, an endothelium-derived peptide, is expressed in neurons of the human spinal cord and dorsal root ganglia. Proc Natl Acad Sci U S A. 1989, 86: 7634-7638.
Millns PJ, Chapman V, Kendall DA: Cannabinoid inhibition of the capsaicin-induced calcium response in rat dorsal root ganglion neurones. Br J Pharmacol. 2001, 132: 969-971.
Hohmann AG, Herkenham M: Localization of central cannabinoid CB1 receptor messenger RNA in neuronal subpopulations of rat dorsal root ganglia: a double-label in situ hybridization study. Neurosci. 1999, 90: 923-931. 10.1016/S0306-4522(98)00524-7.
Chapman V: Functional changes in the inhibitory effect of spinal cannabinoid (CB) receptor activation in nerve injured rats. Neuropharmacol. 2001, 41: 870-877. 10.1016/S0028-3908(01)00125-3.
Ahluwalia J, Urban L, Capogna M, Bevan S, Nagy I: Cannabinoid 1 receptors are expressed in nociceptive primary sensory neurons. Neurosci. 2000, 100: 685-688. 10.1016/S0306-4522(00)00389-4.
Ferguson TA, Muir D: MMP-2 and MMP-9 increase the neurite-promoting potential of schwann cell basal laminae and are upregulated in degenerated nerve. Mol Cell Neurosci. 2000, 16: 157-167. 10.1006/mcne.2000.0859.
Vaillant C, Didier-Bazes M, Hutter A, Belin MF, Thomasset N: Spatiotemporal expression patterns of metalloproteinases and their inhibitors in the postnatal developing rat cerebellum. J Neurosci. 1999, 19: 4994-5004.
Nordstrom LA, Lochner J, Yeung W, Ciment G: The metalloproteinase stromelysin-1 (transin) mediates PC12 cell growth cone invasiveness through basal laminae. Mol Cell Neurosci. 1995, 6: 56-68. 10.1006/mcne.1995.1006.
Cosgaya JM, Recio JA, Aranda A: Influence of Ras and retinoic acid on nerve growth factor induction of transin gene expression in PC12 cells. Oncogene. 1997, 14: 1687-1696. 10.1038/sj.onc.1200997.
Schreiber RC, Krivacic K, Kirby B, Vaccariello SA, Wei T, Ransohoff RM, Zigmond RE: Monocyte chemoattractant protein (MCP)-1 is rapidly expressed by sympathetic ganglion neurons following axonal injury. Neurorep. 2001, 12: 601-606. 10.1097/00001756-200103050-00034.
Carroll SL, Frohnert PW: Expression of JE (monocyte chemoattractant protein-1) is induced by sciatic axotomy in wild type rodents but not in C57BL/Wld(s) mice. J Neuropathol Exp Neurol. 1998, 57: 915-930.
Siebert H, Sachse A, Kuziel WA, Maeda N, Bruck W: The chemokine receptor CCR2 is involved in macrophage recruitment to the injured peripheral nervous system. J Neuroimmunol. 2000, 110: 177-185. 10.1016/S0165-5728(00)00343-X.
Flugel A, Hager G, Horvat A, Spitzer C, Singer GM, Graeber MB, Kreutzberg GW, Schwaiger FW: Neuronal MCP-1 expression in response to remote nerve injury. J Cereb Blood Flow Metab. 2001, 21: 69-76. 10.1097/00004647-200101000-00009.
Lacor P, Benavides J, Ferzaz B: Enhanced expression of the peripheral benzodiazepine receptor (PBR) and its endogenous ligand octadecaneuropeptide (ODN) in the regenerating adult rat sciatic nerve. Neurosci Lett. 1996, 220: 61-65. 10.1016/S0304-3940(96)13187-6.
Lacor P, Gandolfo P, Tonon MC, Brault E, Dalibert I, Schumacher M, Benavides J, Ferzaz B: Regulation of the expression of peripheral benzodiazepine receptors and their endogenous ligands during rat sciatic nerve degeneration and regeneration: a role for PBR in neurosteroidogenesis. Brain Res. 1999, 815: 70-80. 10.1016/S0006-8993(98)01105-6.
Gehlert DR, Stephenson DT, Schober DA, Rash K, Clemens JA: Increased expression of peripheral benzodiazepine receptors in the facial nucleus following motor neuron axotomy. Neurochem Int. 1997, 31: 705-713. 10.1016/S0197-0186(97)00007-7.
Rosenberg M, Pie B, Cooper E: Developing neonatal rat sympathetic and sensory neurons differ in their regulation of 5-HT3 receptor expression. J Neurosci. 1997, 17: 6629-6638.
Scroggs RS, Cardenas CG, Whittaker JA, Kitai ST: Muscarine Reduces Calcium-Dependent Electrical Activity in Substantia Nigra Dopaminergic Neurons. J Neurophysiol. 2001, 86: 2966-2972.
Ali Z, Wu G, Kozlov A, Barasi S: The role of 5HT3 in nociceptive processing in the rat spinal cord: results from behavioural and electrophysiological studies. Neurosci Lett. 1996, 208: 203-207. 10.1016/0304-3940(95)12600-7.
Morales M, McCollum N, Kirkness EF: 5-HT(3)-receptor subunits A and B are co-expressed in neurons of the dorsal root ganglion. J Comp Neurol. 2001, 438: 163-172. 10.1002/cne.1307.
Chou TT, Trojanowski JQ, Lee VM: p38 mitogen-activated protein kinase-independent induction of gadd45 expression in nerve growth factor-induced apoptosis in medulloblastomas. J Biol Chem. 2001, 276: 41120-41127. 10.1074/jbc.M102832200.
Kojima S, Mayumi-Matsuda K, Suzuki H, Sakata T: Molecular cloning of rat GADD45gamma, gene induction and its role during neuronal cell death. FEBS Lett. 1999, 446: 313-317. 10.1016/S0014-5793(99)00234-3.
Torp R, Su JH, Deng G, Cotman CW: GADD45 is induced in Alzheimer's disease, and protects against apoptosis in vitro. Neurobiol Dis. 1998, 5: 245-252. 10.1006/nbdi.1998.0201.
Bogaev RC, Jia LG, Kobayashi YM, Palmer CJ, Mounsey JP, Moorman JR, Jones LR, Tucker AL: Gene structure and expression of phospholemman in mouse. Gene. 2001, 271: 69-79. 10.1016/S0378-1119(01)00497-8.
Moorman JR, Palmer CJ, John JE, Durieux ME, Jones LR: Phospholemman expression induces a hyperpolarization-activated chloride current in Xenopus oocytes. J Biol Chem. 1992, 267: 14551-14554.
Sweadner KJ, Rael E: The FXYD gene family of small ion transport regulators or channels: cDNA sequence, protein signature sequence, and expression. Genomics. 2000, 68: 41-56. 10.1006/geno.2000.6274.
Moorman JR, Ackerman SJ, Kowdley GC, Griffin MP, Mounsey JP, Chen Z, Cala SE, O'Brian JJ, Szabo G, Jones LR: Unitary anion currents through phospholemman channel molecules. Nature. 1995, 377: 737-740. 10.1038/377737a0.
Snyder SE, Pintar JE, Salton SR: Developmental expression of VGF mRNA in the prenatal and postnatal rat. J Comp Neurol. 1998, 394: 64-90. 10.1002/(SICI)1096-9861(19980427)394:1<64::AID-CNE6>3.3.CO;2-8.
Snyder SE, Cheng HW, Murray KD, Isackson PJ, McNeill TH, Salton SR: The messenger RNA encoding VGF, a neuronal peptide precursor, is rapidly regulated in the rat central nervous system by neuronal activity, seizure and lesion. Neurosci. 1998, 82: 7-19. 10.1016/S0306-4522(97)00280-7.
Angelastro JM, Klimaschewski L, Tang S, Vitolo OV, Weissman TA, Donlin LT, Shelanski ML, Greene LA: Identification of diverse nerve growth factor-regulated genes by serial analysis of gene expression (SAGE) profiling. Proc Natl Acad Sci U S A. 2000, 97: 10424-10429. 10.1073/pnas.97.19.10424.
Bauer K, Kratzer M, Otte M, de Quintana KL, Hagmann J, Arnold GJ, Eckerskorn C, Lottspeich F, Siess W: Human CLP36, a PDZ-domain and LIM-domain protein, binds to alpha-actinin-1 and associates with actin filaments and stress fibers in activated platelets and endothelial cells. Blood. 2000, 96: 4236-4245.
Nakagawa N, Hoshijima M, Oyasu M, Saito N, Tanizawa K, Kuroda S: ENH, containing PDZ and LIM domains, heart/skeletal muscle-specific protein, associates with cytoskeletal proteins through the PDZ domain. Biochem Biophys Res Commun. 2000, 272: 505-512. 10.1006/bbrc.2000.2787.
Pomies P, Louis HA, Beckerle MC: CRP1, a LIM domain protein implicated in muscle differentiation, interacts with alpha-actinin. J Cell Biol. 1997, 139: 157-168. 10.1083/jcb.139.1.157.
Levadoux-Martin M, Hesketh JE, Beattie JH, Wallace HM: Influence of metallothionein-1 localization on its function. Biochem J. 2001, 355: 473-479. 10.1042/0264-6021:3550473.
van Lookeren CM, Thibodeaux H, van Bruggen N, Cairns B, Gerlai R, Palmer JT, Williams SP, Lowe DG: Evidence for a protective role of metallothionein-1 in focal cerebral ischemia. Proc Natl Acad Sci U S A. 1999, 96: 12870-12875. 10.1073/pnas.96.22.12870.
Lee PG, Koo PH: Rat alpha(2)-macroglobulin inhibits NGF-promoted neurite outgrowth, TrK phosphorylation, and gene expression of pheochromocytoma PC12 cells. J Neurosci Res. 1999, 57: 872-883. 10.1002/(SICI)1097-4547(19990915)57:6<872::AID-JNR13>3.3.CO;2-9.
Koo PH, Liebl DJ: Inhibition of nerve growth factor-stimulated neurite outgrowth by methylamine-modified alpha 2-macroglobulin. J Neurosci Res. 1992, 31: 678-692.
Koo PH, Stach RW: Interaction of nerve growth factor with murine alpha-macroglobulin. J Neurosci Res. 1989, 22: 247-261.
Marzolo MP, von Bernhardi R, Bu G, Inestrosa NC: Expression of alpha(2)-macroglobulin receptor/low density lipoprotein receptor-related protein (LRP) in rat microglial cells. J Neurosci Res. 2000, 60: 401-411. 10.1002/(SICI)1097-4547(20000501)60:3<401::AID-JNR15>3.3.CO;2-C.
Fenzi F, Benedetti MD, Moretto G, Rizzuto N: Glial cell and macrophage reactions in rat spinal ganglion after peripheral nerve lesions: an immunocytochemical and morphometric study. Arch Ital Biol. 2001, 139: 357-365.
Lu X, Richardson PM: Responses of macrophages in rat dorsal root ganglia following peripheral nerve injury. J Neurocytol. 1993, 22: 334-341.
Hikawa N, Horie H, Takenaka T: Macrophage-enhanced neurite regeneration of adult dorsal root ganglia neurones in culture. Neurorep. 1993, 5: 41-44.
Jacobsson G, Piehl F, Bark IC, Zhang X, Meister B: Differential subcellular localization of SNAP-25a and SNAP-25b RNA transcripts in spinal motoneurons and plasticity in expression after nerve injury. Brain Res Mol Brain Res. 1996, 37: 49-62. 10.1016/0169-328X(95)00272-T.
Hepp R, Dupont JL, Aunis D, Langley K, Grant NJ: NGF enhances depolarization effects on SNAP-25 expression: induction of SNAP-25b isoform. Neurorep. 2001, 12: 673-677. 10.1097/00001756-200103260-00011.
Shirasu M, Kimura K, Kataoka M, Takahashi M, Okajima S, Kawaguchi S, Hirasawa Y, Ide C, Mizoguchi A: VAMP-2 promotes neurite elongation and SNAP-25A increases neurite sprouting in PC12 cells. Neurosci Res. 2000, 37: 265-275. 10.1016/S0168-0102(00)00125-5.
Acknowledgements
We thank Bayer AG and the NIH (NS38253, HD38533) for financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
MC carried out array probe preparation, slot blot analysis and northern analysis, participated in the design of the study and drafted the manuscript. KB carried out northern analysis and data preparation, participated in the design of the study and drafted the manuscript. LK carried out in situ hybridization and participated in the design of the study. RSG carried out statistical data analysis and functional group analysis and participated in the design of the study. DD'U participated in data preparation and in the design of the study. AA and JS carried out tissue preparation. JWM participated in data preparation. REP participated in the design of the study. CJW conceived the study and participated in its design and coordination.
Michael Costigan, Katia Befort contributed equally to this work.
Electronic supplementary material
12868_2002_38_MOESM1_ESM.pdf
Additional File 1: Comparison of microarray and known regulation data for genes whose expression pattern within the DRG following nerve injury has been studied previously. (PDF 58 KB)
12868_2002_38_MOESM2_ESM.pdf
Additional File 2: Summary of genes identified as regulated >1.5-fold (P < 0.05) in the DRG 3 days after sciatic nerve axotomy. (PDF 68 KB)
12868_2002_38_MOESM3_ESM.pdf
Additional File 3: Summary of expression patterns and functional roles of uncharacterized nerve-injury regulated genes. (PDF 92 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Costigan, M., Befort, K., Karchewski, L. et al. Replicate high-density rat genome oligonucleotide microarrays reveal hundreds of regulated genes in the dorsal root ganglion after peripheral nerve injury.. BMC Neurosci 3, 16 (2002). https://doi.org/10.1186/1471-2202-3-16
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2202-3-16