Abstract
Background
Antimicrobial resistance is under-documented and commensal Escherichia coli can be used as indicator organisms to study the resistance in the community. We sought to determine the prevalence of resistance to broad-spectrum antimicrobials with particular focus on the quinolones, which have recently been introduced in parts of Africa, including Ghana.
Results
Forty (13.7%) of 293 E. coli isolates evaluated were nalidixic acid-resistant. Thirteen (52%) of 2006 and 2007 isolates and 10 (66.7%) of 2008 isolates were also resistant to ciprofloxacin. All but one of the quinolone-resistant isolates were resistant to three or more other antimicrobial classes. Sequencing the quinolone-resistance determining regions of gyrA and parC, which encode quinolone targets, revealed that 28 quinolone-resistant E. coli harboured a substitution at position 83 of the gyrA gene product and 20 of these isolates had other gyrA and/or parC substitutions. Horizontally-acquired quinolone-resistance genes qnrB1, qnrB2, qnrS1 or qepA were detected in 12 of the isolates. In spite of considerable overall diversity among E. coli from Ghana, as evaluated by multilocus sequence typing, 15 quinolone-resistant E. coli belonged to sequence type complex 10. Five of these isolates carried qnrS1 alleles.
Conclusions
Quinolone-resistant E. coli are commonly present in the faecal flora of Accra residents. The isolates have evolved resistance through multiple mechanisms and belong to very few lineages, suggesting clonal expansion. Containment strategies to limit the spread of quinolone-resistant E. coli need to be deployed to conserve quinolone effectiveness and promote alternatives to their use.
Similar content being viewed by others
Background
Following emergence of resistance to inexpensive broad-spectrum antimicrobials across much of Africa, quinolone antibacterials have recently been introduced and are widely used. West African studies that sought quinolone resistance in commensal or diarrhoeagenic Escherichia coli before 2004 reported no or very low incidences of resistance to nalidixic acid and the fluoroquinolones [1–4]. Thus, available data suggests that resistance to the quinolones was rare in West Africa until the first decade of the 21st century. More recent anecdotal reports and surveillance studies point to emergence of quinolone resistance among enteric pathogens and faecal enteric bacteria in Ghana and elsewhere in West Africa [5–8]. In a study by Nys et al. (2004) faecal isolates of adult volunteers in eight different countries were assessed for susceptibility to antimicrobials in the same laboratory [8]. Resistance to broad spectrum first-generation antibiotics was common and ciprofloxacin resistance was found to be slowly emerging in Asian, South American and African countries, including Ghana [8]. Newman et al. (2004) collected 5099 clinical bacterial isolates (1105 of which were E. coli) from nine of the ten regions in Ghana and tested them for antimicrobial susceptibility. They found that over 70% of the isolates were resistant to tetracycline, trimethoprim-sulphamethoxazole, ampicillin and chloramphenicol and reported that 11% of the isolates were ciprofloxacin-resistant [7].
Quinolones inhibit the activity of bacterial DNA gyrase and DNA topoisomerase enzymes, which are essential for replication. Single nucleotide polymorphisms (SNPs) in the quinolone resistance determining regions (QRDR) of gyrA and parC, the two genes that encode DNA gyrase and topoisomerase IV respectively, can lead to conformational changes in these enzymes that cause them to block quinolones from binding to the DNA- substrate complex, yet still preserve their enzymatic function [9]. In Escherichia coli and related Gram-negative bacteria, DNA gyrase is the first target for fluoroquinolones. If gyrA has resistance-conferring mutations, the primary target of fluoroquinolone switches from DNA gyrase to topoisomerase IV [10, 11]. Studies from other parts of the world have found that resistance-conferring mutations are typically selected in gyrA first, and then parC.
Although mutations in the QRDR of gyrA and parC are the most commonly documented resistance mechanisms, resistance has also been known to be conferred by mutations in the second topoisomerase gene, parE. Another mechanism of quinolone resistance relies on upregulation of efflux pumps, which export quinolones and other antimicrobials out of the bacterial cell. For example, mutations in the gene encoding a repressor of the acrAB pump genes, acrR, are associated with quinolone resistance [12]. Quinolone resistance can also be acquired horizontally through transferable quinolone resistance (qnr) or other DNA. The Qnr gene product inhibits quinolones binding to target proteins [13]. Other horizontally acquired quinolone resistance genes include aac(6')-Ib , encoding a fluroquinolone acetylating enzyme, as well as qepA and oqxAB, which encode horizontally transmitted efflux pumps [14–16].
Resistance to the quinolones often emerges at low-levels by acquisition of an initial resistance-conferring mutation or gene. Acquisition of subsequent mutations leads to higher levels of resistance to the first-generation quinolone, nalidixic acid and a broadening of the resistance spectrum to include second-generation quinolones (first-generation fluoroquinolones) such as ciprofloxacin, followed by newer second- and third-generation fluoroquinolones [17].
Although multiple mechanisms of quinolone resistance have been reported from other continents, there are few data from sub-Saharan Africa on the molecular basis for quinolone resistance. We performed antimicrobial susceptibility testing on fecal E. coli isolates from Accra, Ghana in 2006, 2007 and 2008. We identified isolates that were resistant to nalidixic acid and screened these strains for mutations in the QRDR of gyrA and parC as well as horizontally-acquired quinolone-resistance genes. In order to gain some insight into resistance dissemination, we also studied inter-strain relatedness among quinolone-resistant E. coli isolates by multilocus sequence typing.
Results
Resistance to commonly used antimicrobials is high and resistance to the quinolones was detected
In 2006, 2007 and 2008 respectively, 156, 78 and 101 stool specimens were collected. A total of 293 Escherichia coli isolates were recovered from culture of the 335 stool specimens. Consistent with the results of recent studies from West African countries, including Ghana [1, 7, 8], 50-90% of the E. coli isolates were resistant to the broad-spectrum antimicrobials ampicillin, streptomycin, sulphonamides, tetracycline and trimethoprim (Figure 1). Resistance to chloramphenicol was less common but was seen in 30-41% of the isolates. The proportions of isolates resistant to most agents were comparable between 2006 and 2007. However, the proportion of isolates resistant to each antimicrobial in 2008 was significantly greater than those seen in 2006, for all agents (p < 0.05) (Figure 1).
As illustrated in Figure 1 in 2006 and 2007, we recorded resistance rates to nalidixic acid of 12.3% but by 2008, 15 (18.2%) of isolates were nalidixic acid resistant. Ciprofloxacin-resistant isolates represented 7 (5.4%) and 6 (7.7%) of the total number of isolates in 2006 and 2007 respectively. In 2008, 10 (9.9%) of the isolates were fluoroquinolone resistant. Thus, in 2006 and 2007, 13 (52%) quinolone-resistant E. coli isolates were ciprofloxacin resistant but in 2008, 10 (67%) of the quinolone-resistant E. coli were resistant to ciprofloxacin (Figure 1). Although the numbers were too small to attain statistical significance, organisms with higher nalidixic acid MICs were recovered more commonly in 2008 than in 2006 (Table 1).
Quinolone resistance was almost always seen in multiply-resistant E. coli. As shown in Table 1, all quinolone-resistant E. coli (QREC) were resistant to at least one other antimicrobial and all but three of the QREC isolates were resistant to four or more non-quinolone antibacterials. Most QREC demonstrated high-level resistance to nalidixic acid with 21 of 40 of the QREC isolates showing a nalidixic acid MIC that exceeded 1024 mg/L. Among 2006 isolates, low-level resistance was more common, with the MIC50 in that year being 128 mg/L. In both 2007 and 2008, the MIC50 was >1024 mg/L.
Quinolone resistant E. coli predominantly harbour mutations in gyrA, parCor both
Increasing nalidixic acid MICs, accompanied by resistance to fluoroquinolones is often due to the acquisition of multiple mutations in quinolone targets. We sequenced the quinolone-resistance determining regions (QRDRs) of gyrA and parC in the 40 QREC isolates. As shown in Table 1, 28 (70%) of the quinolone-resistant isolates had at least one non-synonymous substitution in the QRDR of gyrA and 18 of these isolates also had one or more non-synonymous mutations in parC. Twenty-seven of the 28 isolates with at least one mutation in gyrA had a serine to leucine substitution at position 83, one of the most commonly documented resistance conferring mutations [10]. Twenty of these isolates also harboured the frequently documented aspartic acid to asparagine substitution at position 87 and all of these isolates had a nalidixic acid MIC of at least 256 mg/L. Eighteen of them were resistant to ciprofloxacin as well as nalidixic acid.
Eighteen QREC isolates had non-synonymous mutations in the QRDR of parC with a serine to isoleucine substitution at position 80, present in 16 strains, being the most common substitution (Table 1). The 2007 isolate with a Thr66Ile substitution in ParC had a single GyrA substitution, Ser83Leu. All other isolates with ParC substitutions also had Ser83Leu and Asp87Asn substitutions in GyrA. Five isolates had more than one ParC substitution. Thr66Ile and Asn105Ser substitutions in ParC, seen in two isolates in this study, have not previously been described in E. coli but Thr66Ile has been seen in Salmonella enterica serovars Heidelberg and Mbandaka [18](Table 1). Both substitutions occur in strains with other previously described non-synonymous polymorphisms in parC and gyrA. In each case, the level and spectrum of resistance seen is not significantly greater than that for isolates that lack the novel substitution. Therefore, it is not possible to conclude that either substitution confers additional resistance although confirmation of this assumption, or otherwise, can only be made in isogenic strains.
Multiple horizontally-transmitted quinlone resistance genes were detected among E. colifrom Accra
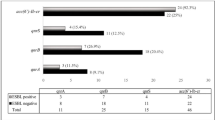
We used PCR to screen for qnrA, qnrB, qnrS and qepA genes and confirmed all amplicons by sequencing. Of the 40 strains evaluated twelve carried one horizontally acquired quinolone resistance gene. These were qnrB1 (2 isolates), qnrB2 (1 isolate), qnrS1 (7 isolates) and qepA (2 isolates). In two isolates, without mutations in gyrA and parC QRDRs, horizontally-acquired resistance genes could account for the resistance seen. However, in the vast majority of cases, horizontally acquired resistance was seen in combination with QRDR mutations.
Quinolone-resistant E. colifrom Accra are over-represented among multi-locus sequence type 10
We hypothesized that clonal expansion might account, at least in part, for the rise in resistance seen in the course of the study. To test this hypothesis, we subjected all the 40 QREC isolates to multi-locus sequence typing by the scheme of Wirth et al [19] and deposited their allelic profiles in the database at http://www.mlst.net. We identified 30 Sequence Types (STs) among 40 QREC isolates from Ghana (0.75 STs per strain). As shown in Figure 2, quinolone resistance is seen in diverse lineages that have been detected in Ghana. STs that were recovered more than once among the QREC included ST10 (9 isolates) as well as STs101, 156, 227, 648 and 1466 (2 isolates each) (Table 1). Although there were 10 QREC STs that were identified for the first time in this study (reflecting the low proportion of strains from West Africa in the database), only one of these (1466) was seen more than once among QREC (Figure 2, Table 1). Three others were related to STs that were also seen among QREC - ST1471 was a single-locus variant of ST206, and STs1286 and 1467 were respectively single- and double-locus variants of ST10. Horizontally-transmitted quinlone resistance determinants were expectedly detected in strains belonging to multiple STs. However qnrS1 alleles were in all but two cases detected among strains belonging to the ST10 complex.
eBURST output for 165 E. coli isolates in the http://www.mlst.net database that were isolated in Ghana, including 48 isolates sequence-typed in this study. Each ST is marked as a dot or node. The size of the node is proportional to the number of isolates contained in that ST. Blue nodes represent predicted founder STs and sub-founders are indicated in yellow. All other STs marked as black dots. STs annotated in green are comprised of quinolone-resistant strains only and those written in pink contain quinolone-sensitive and quinolone-resistant isolates.
Nine of the 40 QREC isolates obtained in this study belonged to ST10, in contrast to 10 of 125 other E. coli from Ghana in the database (p = 0.02, Fisher's exact test). Moreover six other QREC isolates were single- or double- locus variants of ST10. Thus ST10 appeared to be over-represented among QREC isolates from Ghana. Because most of the isolates from Ghana were deposited in the database two to four years in advance of our own study, we sequence typed eight non-QREC isolates selected at random from our 2008 isolates. All eight belonged to different sequence types (10, 349, 541, 1474, 1475, 1476, 1477 and 1478), five of the eight sequence types were novel, and only one 2008 non-QREC strain was an ST10 isolate. Therefore our data suggest that ST10-complex QREC may represent a successful quinolone-resistant lineage.
Discussion
Evolution of reduced susceptibility to the quinolones is causing concern following rapidly rising rates of fluoroquinolone-resistant E. coli in many parts of the world [20]. In African countries with a high infectious disease burden, formal and informal health systems depend heavily on broad spectrum orally-administrable antibacterials. In this study, we found that most commensal E. coli isolates are resistant to ampicillin, sulphonamides, tetracycline and trimethoprim, as well as streptomycin, which have been used to treat actual and supposed bacterial infections in Ghana for over four decades, and that resistance to these agents is increasing with time. We also found that about a third of isolates were resistant to chloramphenicol. Fluoroquinolone antimicrobials have been recently introduced as an effective alternative to older antibacterials that have been compromised by resistance. However, although resistance rates were markedly lower for this class of drugs, we also found that quinolone resistance was increasingly common among fecal E. coli in this study.
We determined that 12-18% of fecal E. coli isolated from healthy individuals in Accra in 2006, 2007 and 2008 are quinolone resistant. Twenty-three of the 40 QREC isolated were resistant to the fluoroquinolone ciprofloxacin. Ciprofloxacin-resistant QREC, showing high-level nalidixic acid resistance, were more commonly isolated in 2008 than in 2006 and 2007. Strains with one or no mutations in gyrA were typically ciprofloxacin sensitive. However most isolates had accumulated a second gyrA mutation and/or mutations in parC and were fluoroquinolone resistant. The QRDR polymorphisms most commonly detected in this study are those most frequently reported in the literature [10]. As has been validated experimentally in isogenic strains, high-level nalidixic acid resistance and fluoroquinolone resistance in isolates in this study was associated with parC substitutions in strains also harbouring substitutions in gyrA [17]. However, gyrA and parC mutations did not absolutely correlate with nalidixic acid MICs, partly due to horizontally-acquired quinolone-resistance genes. We sought qnrA, qnrB, qnrS and qepA genes by PCR and confirmed all amplicons by sequencing. We found that two isolates without mutations in the QRDRs of gyrA and parC, as well as ten isolates with QRDR mutations carried a qnrS1, a qnrB or a qepA allele. The presence of horizontally-acquired genes accounted in part for elevated nalidixic acid MICs in strains that harboured these genes, but not completely. It is therefore possible that other resistance mechanisms, such as ParE polymorphisms, other horizontally acquired resistance genes (such as oqxAB and aac(6')-Ib for example), over-active efflux, or even novel mechanisms are present in some of the isolates.
Resistance patterns in pathogens often mirror those in commensals. This is borne out by our recent documentation of quinolone resistance in Vibrio cholerae isolates recovered in the same time frame as the E. coli strains presented in this report [21]. Fifteen of the 40 QREC isolates identified in this study belonged to ST10, or were single- or double-locus variants of this ST, pointing to the possibility of clonal expansion. ST10-complex strains were isolated in all three years and therefore over-representation of these STs in our sample cannot be explained by short-term, localized clustering. There are four major E. coli phylogenetic clades: ECOR A, B1, B2 and D. Few studies have looked at the geographical variance in the distribution of these groups but overall, QREC from Ghana were predominantly drawn from ECOR group A. Of the STs identified in this study that are classified into ECOR clades at the E. coli MLST database, ST10 complex (14 isolates) belong to ECOR group A, ST131 (1 isolate) to ECOR B2, STs101 and 410 (3 isolates) to ECOR B1 and STs 156, 206 and 210 (4 isolates) are hybrids of ECOR A and B1, that is AxB1. Available data appear to suggest that ECOR A strains are highly prevalent in Africa, compared to some other world regions [22]. However, when we compared the sequence types of quinolone-resistant and -susceptible strains from Ghana only, we still found that resistant strains were over-represented in the ST10 complex. Pandemic clonal expansion of some QREC lineages has been reported in the literature [23–28]. For example, ST131 is a globally disseminated multi-resistant clone and was detected once among the QREC in this study. Recent reports suggest that isolates from Europe and North America that belong to ST10- or ST131- clonal complexes may be less likely to carry virulence factors for invasive disease, but more likely to be fluoroquinolone resistant [24–28]. However it is equally likely that mutations to fluoroquinolone resistance are more likely to be stably inherited in a specific genetic background. Our own data also appear to suggest that, although horizontally acquired, qnrS1 is associated with ST10 complex.
A recent paper by Davidson et al suggests that the antimalarial chloroquine may select for fluoroquinolone-resistant fecal bacteria in malaria endemic areas and proposes that chloroquine-mediated selection accounts for high levels of QREC in fecal flora in villages in South America [29]. However, data from within Africa (where chloroquine has seen heaviest use) to support this hypothesis are currently lacking and evidence from elsewhere supports a link between quinolone resistance rates in E. coli and fluoroquinolone consumption at population levels [30, 31]. Our own data suggest that even if resistance was present at low levels prior to the introduction of the quinolones, an upsurge in resistance may reflect a selective advantage that came with quinolone introduction. This is particularly likely for Ghana were artemisinin combination therapies have recently been introduced to replace chloroquine. Furthermore, because almost all quinolone resistant-E. coli were multiply resistant, selective pressure from other, even more commonly applied, antimicrobials will help to maintain the quinolone-resistant clonal groups we identified in this study.
Conclusion
Fluoroquinolones, largely ciprofloxacin, were introduced very recently to Ghana with high expectations. This study demonstrates that resistance to these drugs is already common and occurs through multiple mechanisms, suggesting that heavy use of these valuable drugs may rapidly obliterate their usefulness. In addition to the impact that the emergence and dissemination of quinolone resistant bacteria may have on the use of fluoroquinolone antibacterials, we found that QREC were almost invariably resistant to multiple antimicrobials. This is worrisome because it means that, if the commensal flora is reflective of resistance profiles in pathogens, there may be few low-cost alternatives for managing infections due to Gram-negative enteric organisms. Additionally, horizontally-acquired resistance to the quinolones, and presumably other agents may be present on mobile elements that could be transmitted to pathogens. Recent calls for antimicrobial development have spotlighted hospital pathogens and Gram-positive community-acquired pathogens such as Staphylococci [32]. Our data suggest that there is also a pressing need for orally administrable drugs with activity against Gram-negative organisms, which can be used to manage community enteric infections in Ghana and other parts of Africa. Additionally, known strategies for containing antimicrobial resistance need to be more rigorously applied [33–35].
Methods
Strains
This study was approved by the Institutional Review Board of the University of Ghana Medical School. E. coli isolates were recovered from stool specimens collected from consenting, apparently healthy individuals who presented for medical check-ups at the Korle-Bu Teaching Hospital and the Microbiology Department of the University of Ghana Medical School. Colonies with a typical E. coli morphology on MacConkey agar were subjected to biochemical testing, and where this was inconclusive, by 16 S amplification and sequencing [36]. Colonies from the same specimen with identical biochemical and susceptibility profiles were treated as identical strains.
Antimicrobial susceptibility testing
Each isolate was tested for susceptibility to eight antimicrobials using the Clinical and Laboratory Standards Institute (CLSI, formerly NCCLS) disc diffusion method [37]. Antimicrobial discs and control strain E. coli ATCC 35218 were obtained from Remel. The antimicrobial discs used contained ampicillin (10 μg), streptomycin (10 μg), trimethoprim (5 μg), tetracycline (30 μg), nalidixic acid (30 μg), chloramphenicol (30 μg), ciprofloxacin (5 μg) and sulphonamide (300 μg). Inhibition zone diameters were interpreted in accordance with CLSI guidelines with WHONET software version 5.3 [38]. Minimum inhibitory concentrations (MICs) to nalidixic acid were measured using the agar dilution technique on Mueller-Hinton agar as recommended by the CLSI and using E. coli ATCC 35218 as control [39].
Mutational analysis of the Quinolone-Resistance Determining Regions of gyrA and parC
DNA was extracted from each quinolone-resistant isolate, using the Promega Wizard genomic extraction kit. The QRDR of the gyrA and parC genes were amplified from DNA templates by PCR using Platinum PCR supermix (Invitrogen) and the primer pairs listed in Table 2. PCR reactions began with a two-minute hot start at 94°C followed by 30 cycles of 94°C for 30 s, annealing temperature, 30 s and 72°C for 30 s. gyrA amplifications were annealed at 58°C and parC reactions were annealed at 52°C. E. coli K-12 MG1655 [40] was used as a control. Amplicons were sequenced on both strands and predicted peptide sequences were compared to the corresponding gene from the MG1655 genome [40] by pair-wise FASTA alignments.
Identification of horizontally-acquired quinolone-resistance genes
Horizontally-acquired quinolone-resistance genes were identified by PCR. The primers of Liu et al [41] were used to screen for the qepA gene and qnrA, qnrB, and qnrS were identified with PCR using the primer pairs published by Wu et al [42] (Table 2). Amplicons were sequence-verified.
Multi-locus sequence typing
Gene fragments from the adk, fumC, gyrB, icd, mdh, purA and recA were amplified using primers listed in Table 2, as described by Wirth et al [19]. Amplified DNA products were sequenced from both ends. Allele assignments were made at the publicly accessible E. coli MLST database at http://www.mlst.net. Phylogenetic inferences about ancestral allelic profiles and strain interrelatedness were made using eBURSTv3 at http://eburst.mlst.net/ defining clonal complexes based on groups sharing five identical alleles and bootstrapping with 1000 samplings.
Statistical analysis
Proportions were compared using the χ2 or Fisher's exact test with p-values less than 0.05 being considered significant.
Funding
This work was supported by a Branco Weiss Fellowship from the Society in Science, ETHZ, Zürich to INO. SSN and RSL were HHMI-supported undergraduate researchers, and RSL was also an Arnold and Mabel Beckman Scholar, at Haverford College.
References
Okeke IN, Fayinka ST, Lamikanra A: Antibiotic resistance trends in Escherichia coli from apparently healthy Nigerian students (1986-1998). Emerg Infect Dis. 2000, 6 (4): 393-396. 10.3201/eid0604.000413.
Mendez Arancibia E, Pitart C, Ruiz J, Marco F, Gascon J, Vila J: Evolution of antimicrobial resistance in enteroaggregative Escherichia coli and enterotoxigenic Escherichia coli causing traveller's diarrhoea. J Antimicrob Chemother. 2009, 64 (2): 343-347. 10.1093/jac/dkp178.
Okeke IN, Lamikanra A, Czeczulin J, Dubovsky F, Kaper JB, Nataro JP: Heterogeneous virulence of enteroaggregative Escherchia coli strains isolated from children in Southwest Nigeria. J Infect Dis. 2000, 181: 252-260. 10.1086/315204.
Okeke IN, Steinruck H, Kanack KJ, Elliott SJ, Sundstrom L, Kaper JB, Lamikanra A: Antibiotic-resistant cell-detaching Escherichia coli strains from Nigerian children. J Clin Microbiol. 2002, 40 (1): 301-305. 10.1128/JCM.40.1.301-305.2002.
Soge OO, Adeniyi BA, Roberts MC: New antibiotic resistance genes associated with CTX-M plasmids from uropathogenic Nigerian Klebsiella pneumoniae. J Antimicrob Chemother. 2006, 58 (5): 1048-1053. 10.1093/jac/dkl370.
Nabeth P, Perrier-Gros-Claude J-D, Juergens-Behr A, Dromigny J-A: In vitro susceptibility of quinolone-resistant Enterobacteriaceae uropathogens to fosfomycin trometamol, in Dakar, Senegal. Scand J Infect Dis. 2005, 37 (6): 497-499. 10.1080/00365540510038956.
Newman MJ, Frimpong E, Asamoah-Adu A, Sampane-Donkor E, Opintan JA: Resistance to antimicrobial drugs in Ghana. The Ghanaian-Dutch Collaboration for Health Research and Development. 2004
Nys S, Okeke IN, Kariuki S, Dinant GJ, Driessen C, Stobberingh EE: Antibiotic resistance of faecal Escherichia coli from healthy volunteers from eight developing countries. J Antimicrob Chemother. 2004, 54 (5): 952-955. 10.1093/jac/dkh448.
Hawkey PM: Mechanisms of quinolone action and microbial response. J Antimicrob Chemother. 2003, 29-35. 10.1093/jac/dkg207. 51 Suppl 1
Hopkins KL, Davies RH, Threlfall EJ: Mechanisms of quinolone resistance in Escherichia coli and Salmonella: recent developments. Int J Antimicrob Agents. 2005, 25 (5): 358-373. 10.1016/j.ijantimicag.2005.02.006.
Hooper DC: Mechanisms of action of antimicrobials: focus on fluoroquinolones. Clin Infect Dis. 2001, 32 (Suppl 1): S9-S15. 10.1086/319370.
Wang H, Dzink-Fox JL, Chen M, Levy SB: Genetic characterization of highly fluoroquinolone-resistant clinical Escherichia coli strains from China: role of acrR mutations. Antimicrob Agents Chemother. 2001, 45 (5): 1515-1521. 10.1128/AAC.45.5.1515-1521.2001.
Tran JH, Jacoby GA: Mechanism of plasmid-mediated quinolone resistance. Proc Natl Acad Sci USA. 2002, 99 (8): 5638-5642. 10.1073/pnas.082092899.
Hansen LH, Jensen LB, Sørensen HI, Sørensen SJ: Substrate specificity of the OqxAB multidrug resistance pump in Escherichia coli and selected enteric bacteria. J Antimicrob Chemother. 2007, 60 (1): 145-147. 10.1093/jac/dkm167.
Yamane K, Wachino J-i, Suzuki S, Kimura K, Shibata N, Kato H, Shibayama K, Konda T, Arakawa Y: New plasmid-mediated fluoroquinolone efflux pump, QepA, found in an Escherichia coli clinical isolate. Antimicrob Agents Chemother. 2007, 51 (9): 3354-3360. 10.1128/AAC.00339-07.
Strahilevitz J, Jacoby GA, Hooper DC, Robicsek A: Plasmid-mediated quinolone resistance: a multifaceted threat. Clin Microbiol Rev. 2009, 22 (4): 664-689. 10.1128/CMR.00016-09.
Morgan-Linnell SK, Zechiedrich L: Contributions of the combined effects of topoisomerase mutations toward fluoroquinolone resistance in Escherichia coli. Antimicrob Agents Chemother. 2007, 51 (11): 4205-4208. 10.1128/AAC.00647-07.
Eaves DJ, Randall L, Gray DT, Buckley A, Woodward MJ, White AP, Piddock LJV: Prevalence of mutations within the quinolone resistance-determining region of gyrA, gyrB, parC, and parE and association with antibiotic resistance in quinolone-resistant Salmonella enterica. Antimicrob Agents Chemother. 2004, 48 (10): 4012-4015. 10.1128/AAC.48.10.4012-4015.2004.
Wirth T, Falush D, Lan R, Colles F, Mensa P, Wieler LH, Karch H, Reeves PR, Maiden MC, Ochman H, et al: Sex and virulence in Escherichia coli: an evolutionary perspective. Mol Microbiol. 2006, 60 (5): 1136-1151. 10.1111/j.1365-2958.2006.05172.x.
Livermore DM: Has the era of untreatable infections arrived?. J Antimicrob Chemother. 2009, 64 (Suppl 1): i29-36. 10.1093/jac/dkp255.
Opintan JA, Newman MJ, Nsiah-Poodoh OA, Okeke IN: Vibrio cholerae O1 from Accra, Ghana carrying a class 2 integron and the SXT element. J Antimicrob Chemother. 2008, 62 (5): 929-933. 10.1093/jac/dkn334.
Robinson DA, Falush D, Feil EJ: Bacterial population genetics in infectious disease. 2010, Hoboken, N.J.: J. Wiley
Uchida Y, Mochimaru T, Morokuma Y, Kiyosuke M, Fujise M, Eto F, Eriguchi Y, Nagasaki Y, Shimono N, Kang D: Clonal spread in Eastern Asia of ciprofloxacin-resistant Escherichia coli serogroup O25 strains, and associated virulence factors. Int J Antimicrob Ag. 2010, 35 (5): 444-450. 10.1016/j.ijantimicag.2009.12.012.
Johnson James R, Kuskowski Michael A, Owens K, Gajewski A, Winokur Patricia L: Phylogenetic origin and virulence genotype in relation to resistance to fluoroquinolones and/or extended spectrum cephalosporins and cephamycins among Escherichia coli isolates from animals and humans. J Infect Dis. 2003, 188 (5): 759-768. 10.1086/377455.
Moreno E, Prats G, Sabate M, Perez T, Johnson JR, Andreu A: Quinolone, fluoroquinolone and trimethoprim/sulfamethoxazole resistance in relation to virulence determinants and phylogenetic background among uropathogenic Escherichia coli. J Antimicrob Chemother. 2006, 57 (2): 204-211. 10.1093/jac/dki468.
Johnson JR, Menard M, Johnston B, Kuskowski MA, Nichol K, Zhanel GG: Epidemic clonal groups of Escherichia coli as a cause of antimicrobial-resistant urinary tract infections in Canada, 2002 to 2004. Antimicrob Agents Chemother. 2009, 53 (7): 2733-2739. 10.1128/AAC.00297-09.
Grude N, Strand L, Mykland H, Nowrouzian FL, Nyhus J, Jenkins A, Kristiansen BE: Fluoroquinolone-resistant uropathogenic Escherichia coli in Norway: evidence of clonal spread. Clin Microbiol Infect. 2008, 14 (5): 498-500. 10.1111/j.1469-0691.2008.01952.x.
Boyd L, Atmar R, Randall G, Hamill R, Steffen D, Zechiedrich L: Increased fluoroquinolone resistance with time in Escherichia coli from >17,000 patients at a large county hospital as a function of culture site, age, sex, and location. BMC Infectious Diseases. 2008, 8 (1): 4-10.1186/1471-2334-8-4.
Davidson RJ, Davis I, Willey BM, Rizg K, Bolotin S, Porter V, Polsky J, Daneman N, McGeer A, Yang P, et al: Antimalarial therapy selection for quinolone resistance among Escherichia coli in the absence of quinolone exposure, in tropical South America. PLoS ONE. 2008, 3 (7): e2727-10.1371/journal.pone.0002727.
van de Sande-Bruinsma N, Grundmann H, Verloo D, Tiemersma E, Monen J, Goossens H, Ferech M: Antimicrobial drug use and resistance in Europe. Emerg Infect Dis. 2008, 14 (11): 1722-1730. 10.3201/eid1411.070467.
Gottesman Bat S, Carmeli Y, Shitrit P, Chowers M: Impact of quinolone restriction on resistance patterns of Escherichia coli isolated from urine by culture in a community setting. Clin Infect Dis. 2009, 49 (6): 869-875.
Talbot GH, Bradley J, Edwards JE, Gilbert D, Scheld M, Bartlett JG: Bad bugs need drugs: An update on the development pipeline from the antimicrobial availability task force of the Infectious Diseases Society of America. Clin Infect Dis. 2006, 42 (5): 657-668. 10.1086/499819.
Okeke IN, Klugman KP, Bhutta ZA, Duse AG, Jenkins P, O'Brien TF, Pablos-Mendez A, Laxminarayan R: Antimicrobial resistance in developing countries. Part II: strategies for containment. Lancet Infect Dis. 2005, 5 (9): 568-580. 10.1016/S1473-3099(05)70217-6.
Okeke IN, Aboderin AO, Byarugaba DK, Ojo O, Opintan JA: Growing problem of multidrug-resistant enteric pathogens in Africa. Emerg Infect Dis. 2007, 13 (11): 1640-1646.
Nugent R, Okeke IN: When medicines fail: recommendations for curbing antibiotic resistance. J Infect Dev Ctries. 2010, 4 (6): 355-356.
Lane DJ: 16S/23 S rRNA sequencing. Nucleic Acid Techniques in Bacterial Systematics. Edited by: Stackebrandt E, Goodfellow M. 1991, New York: John Wiley and Sons, 115-175.
NCCLS: Performance standards for antimicrobial disk susceptibility tests, 8th Edition; Approved standard. 2003, Villanova, PA: National Committee for Clinical Laboratory Standards, 130-
O'Brien TF, Stelling JM: WHONET: an information system for monitoring antimicrobial resistance. Emerg Infect Dis. 1995, 1 (2): 66-
CLSI: Methods for dilution antimicrobial susceptiblity tests for bacteria that grow aerobically, 7th Edition; Approved standard. 2006, Wayne, PA: Clinical and Laboratory Standards Institute
Blattner FR, Plunkett G, Bloch CA, Perna NT, Burland V, Riley M, Collado-Vides J, Glasner JD, Rode CK, Mayhew GF, et al: The complete genome sequence of Escherichia coli K-12. Science. 1997, 277 (5331): 1453-1474. 10.1126/science.277.5331.1453.
Liu J-H, Deng Y-T, Zeng Z-L, Gao J-H, Chen L, Arakawa Y, Chen Z-L: Coprevalence of plasmid-mediated quinolone resistance determinants QepA, Qnr, and AAC(6')-Ib-cr among 16 S rRNA methylase RmtB-producing Escherichia coli isolates from pigs. Antimicrob Agents Chemother. 2008, 52 (8): 2992-2993. 10.1128/AAC.01686-07.
Wu J-J, Ko W-C, Tsai S-H, Yan J-J: Prevalence of plasmid-mediated quinolone resistance determinants QnrA, QnrB, and QnrS among clinical isolates of Enterobacter cloacae in a Taiwanese hospital. Antimicrob Agents Chemother. 2007, 51 (4): 1223-1227. 10.1128/AAC.01195-06.
Deguchi T, Yasuda M, Nakano M, Ozeki S, Kanematsu E, Nishino Y, Ishihara S, Kawada Y: Detection of mutations in the gyrA and parC genes in quinolone-resistant clinical isolates of Enterobacter cloacae. J Antimicrob Chemother. 1997, 40 (4): 543-549. 10.1093/jac/40.4.543.
Acknowledgements
We thank Owusu Agyemang Nsiah-Poodoh, Jessica Glaubman, Cindy Manu and Bing Dao Zhang for technical assistance, as well as John Wain and Jennifer Crowe for helpful comments. This study was dependent on the E. coli MLST database curated by Mark Achtman and made publicly available from http://www.mlst.net.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
SSN performed molecular experiments, analysed and interpreted data, and contributed to writing the paper. JAO collected isolates and performed microbiology experiments. RSL designed and performed molecular experiments. MJN co-conceived the study and collected isolates. INO co-conceived the study, performed microbiology and molecular experiments, analysed and interpreted data and wrote the manuscript. All authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Namboodiri, S.S., Opintan, J.A., Lijek, R.S. et al. Quinolone resistance in Escherichia coli from Accra, Ghana. BMC Microbiol 11, 44 (2011). https://doi.org/10.1186/1471-2180-11-44
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2180-11-44