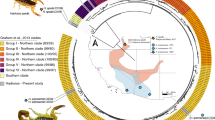
Abstract
Background
Snake venoms generally show sequence and quantitative variation within and between species, but some rattlesnakes have undergone exceptionally rapid, dramatic shifts in the composition, lethality, and pharmacological effects of their venoms. Such shifts have occurred within species, most notably in Mojave (Crotalus scutulatus), South American (C. durissus), and timber (C. horridus) rattlesnakes, resulting in some populations with extremely potent, neurotoxic venoms without the hemorrhagic effects typical of rattlesnake bites.
Results
To better understand the evolutionary changes that resulted in the potent venom of a population of C. horridus from northern Florida, we sequenced the venom-gland transcriptome of an animal from this population for comparison with the previously described transcriptome of the eastern diamondback rattlesnake (C. adamanteus), a congener with a more typical rattlesnake venom. Relative to the toxin transcription of C. adamanteus, which consisted primarily of snake-venom metalloproteinases, C-type lectins, snake-venom serine proteinases, and myotoxin-A, the toxin transcription of C. horridus was far simpler in composition and consisted almost entirely of snake-venom serine proteinases, phospholipases A2, and bradykinin-potentiating and C-type natriuretic peptides. Crotalus horridus lacked significant expression of the hemorrhagic snake-venom metalloproteinases and C-type lectins. Evolution of shared toxin families involved differential expansion and loss of toxin clades within each species and pronounced differences in the highly expressed toxin paralogs. Toxin genes showed significantly higher rates of nonsynonymous substitution than nontoxin genes. The expression patterns of nontoxin genes were conserved between species, despite the vast differences in toxin expression.
Conclusions
Our results represent the first complete, sequence-based comparison between the venoms of closely related snake species and reveal in unprecedented detail the rapid evolution of snake venoms. We found that the difference in venom properties resulted from major changes in expression levels of toxin gene families, differential gene-family expansion and loss, changes in which paralogs within gene families were expressed at high levels, and higher nonsynonymous substitution rates in the toxin genes relative to nontoxins. These massive alterations in the genetics of the venom phenotype emphasize the evolutionary lability and flexibility of this ecologically critical trait.
Similar content being viewed by others
Background
Venomous snakes rely almost entirely on their complex, largely proteinaceous venoms for feeding and defense, resulting in strong selective pressures on the genes encoding venom components [1–4] and on the ultimate repositories of the venoms, the snakes’ prey [5] and predators [6]. Although molecular signals of positive selection have been repeatedly documented for individual venom components through sequence comparisons across species [1–4, 7], such analyses characterize only minute portions of the full evolutionary stories of venoms. Proteomic approaches [8] can characterize full-venom patterns of divergence between species [9, 10], but only in broad strokes, failing to differentiate members of large venom-gene families and to provide information on sequence divergence. Even the most complex venoms are simple in terms of the number of gene families or toxin classes present; the hundreds of proteins [11] typically belong to less than 20 gene families. Proteomic approaches therefore average out many of the details of venom evolution. Venom-gland transcriptomics [12–16] have the unrealized potential to combine many, but certainly not all, of the benefits of both approaches. With adequate sequencing effort, transcriptomes can provide the full-venom information of proteomics approaches as well as the information-dense gene sequences for molecular-evolutionary analyses [17], although post-transcriptional regulation could lead to significant discrepancies between venom content and expressed toxin mRNAs [18].
Snake-venom composition can vary significantly between species [18, 19], within and between populations of a single species [18–25], and even ontogenetically within an individual [10, 26–30]. This variation is related, at least in part, to differences in diets [31]. Some general, recurrent patterns have been identified within this extensive variation, including the type I/II rattlesnake-venom classification described by Mackessy [29], which emphasizes the inverse relationship between toxicity and metalloproteinase activity seen in many rattlesnake venoms. Snake-venom metalloproteinases (SVMPs) are enzymes that break down components of the capillary basement membrane, resulting in local and systemic hemorrhage. SVMPs are more generally known to disrupt hemostasis and cause inflammation and apoptosis [32]. Type I venoms have high metalloproteinase activity and high LD50 values (>1.0 μg/g mouse body weight), whereas type II venoms have low metalloproteinase activity and low LD50 values (<1.0 μg/g mouse body weight). High metalloproteinase activity and high toxicity appear to be incompatible properties of rattlesnake venoms [33]. Type I venoms are by far the most prevalent of the two venom types, appearing in 20 out of 26 rattlesnake taxa examined by Mackessy [29]. Mackessy [29] also revealed that different subspecies can have different venom types. For example, the massasauga (Sistrurus catenatus) expresses type I venom in some subspecies (S. c. tergeminus) and type II in others (S. c. catenatus and perhaps S. c. edwardsi). Other studies have shown ontogenetic shifts between venom types, with juveniles expressing type II venom but switching to type I as adults [34, 35]. Some species, such as C. durissus, are known to express type II venom as both juveniles and adults, a pattern hypothesized to represent paedomorphism [10]. Despite the major differences in pharmacology and composition between these two venom types, evolutionary transitions between them appear to be common and to occur in parallel in several different species and perhaps even in different populations of the same species, despite the relatively short time (∼12.7 million years) since Crotalus diverged from Sistrurus[36]. The selective pressures favoring these transitions, the events triggering or enabling them, and the precise nature of any expression or genetic changes resulting in the altered venom properties are unknown. The determination of these unknowns not only has implications for our understanding of the evolution of major phenotypic innovations, but also has practical implications for snakebite treatment. Although type II venoms are the minority, bites from snakes with type II venoms show drastically different pathologies that might require unique treatment approaches.
The eastern diamondback rattlesnake (C. adamanteus) and the timber rattlesnake (C. horridus) are among the largest rattlesnake species, capable of reaching lengths of 2.4 m and 1.9 m, respectively [37]. Crotalus horridus occurs from New England and extreme southern Ontario, southward to northern Florida, and westward to eastern Texas and extreme southeastern Minnesota [38]. Crotalus adamanteus is a species of the southeastern coastal plain, ranging from extreme southeastern North Carolina, southward to the Florida Keys, and westward along the coast to extreme eastern Louisiana, and is also common on many of the Atlantic and Gulf barrier islands [38]. The two species are sympatric in parts of the Carolinas, Georgia, northern Florida, Alabama, Mississippi, and Louisiana, although they appear to be partitioned by habitat preference. Crotalus adamanteus is more often encountered in areas of high, dry, sandy soils, and C. horridus is more often found in low, wet bottomlands [37, 39]. Both species are of conservation concern because of habitat loss and human persecution [40–43]. Crotalus horridus has been extirpated from many areas, particularly in the northern part of its range, and is classified as endangered in six states and threatened in five others. Crotalus adamanteus is listed as endangered in North Carolina and is currently being reviewed for federally threatened status under the Endangered Species Act. The diets of both species are similar, consisting primarily of rabbits, squirrels, rats, mice, and occasionally birds, with rabbits being more commonly consumed by the larger C. adamanteus[37]. Crotalus horridus also occasionally consumes frogs and snakes [37]. Despite their similarities in size and diet, the venoms from some populations of the two species show dramatic differences in pharmacological properties, composition, and toxicity. Crotalus adamanteus and most populations of C. horridus express type I venom [17, 29], but at least two distinct, southern populations of C. horridus express venoms consistent with a type II classification [44].
Straight and Glenn [45] isolated an extremely lethal, heterodimeric phospholipase A2 (PLA2) presynaptic neurotoxin from C. horridus, which was homologous to Mojave toxin from C. scutulatus[46] and crotoxin from C. durissus terrificus[47]. Related toxins have also been found in C. helleri[48], C. tigris[49], neonates of C. simus[10], and rattlesnakes in the genus Sistrurus[50]. The C. horridus toxin was named canebrake toxin because it was discovered from a northern-Florida specimen belonging to the former subspecies C. h. atricaudatus, known colloquially as the canebrake rattlesnake. Glenn et al. [44] further characterized this neurotoxin, examined its geographic distribution, and found a complex pattern of venom composition in relation to the presence/absence of canebrake toxin and hemorrhagic activity. Crotalus horridus individuals fall into one of four venom types: type A venoms have canebrake toxin but no hemorrhagic activity, type B venoms lack canebrake toxin but have hemorrhagic activity, type A+B venoms have both canebrake toxin and hemorrhagic activity, and type C venoms have neither canebrake toxin nor hemorrhagic activity. Types A and B appear to be the most common types, suggesting a strong inverse relationship in venom composition between canebrake toxin and toxins such as SVMPs, which are major contributors to tissue damage and hemorrhage. Type B venom dominates throughout most of the range of C. horridus with only two known, disjunct regions where type A is common, one of which (southeastern South Carolina through eastern Georgia and northern Florida) falls in one of the regions of sympatry with C. adamanteus. Type A venom would be considered a type II rattlesnake venom under Mackessy’s classification, whereas type B would be a type I [29]. In terms of LD50 in mice, the order of decreasing toxicity for these venom types is: A > A+B > B > C. Analogous venom types, excluding type C, have been identified in C. scutulatus[46] and C. helleri[48] in relation to the presence or absence of Mojave toxin. These venom types, in particular types A and B, reflect vastly different prey incapacitation strategies and possibly different feeding ecologies, because types A and C lack predigestive effects. Low hemorrhagic activity could limit the maximum size of prey that can be consumed or prevent effective digestion at suboptimal temperatures, thereby inducing altitudinal, geographical, or seasonal limitations on foraging [33].
The venom-gland transcriptome of C. adamanteus from northern Florida has been extensively characterized by means of 454 pyrosequencing [4] and Illumina sequencing [17], and this species’ venom is clearly type I on the basis of its biochemical properties, LD50[29], and expressed venom genes [17]. The most abundant transcript in its venom gland encoded a myotoxin-A (i.e., crotamine; MYO), and SVMPs were the most abundant toxin class [17]. To compare venom-gland expression patterns between a rattlesnake with type I venom and one with type II and to elucidate the evolutionary genesis of these venom types, we sequenced the venom-gland transcriptome of C. horridus from northern Florida by means of Illumina technology, following the sequencing and de novo assembly approach used for C. adamanteus[17]. We provide the first comprehensive, sequence-based comparison of venoms between two closely related snake species and the first in-depth examination of toxin gene-family evolution, expression, and reorganization resulting from recent species divergence. We generated the first genome-scale analysis of snake molecular evolution on the basis of thousands of newly annotated nontoxins and compared the evolution of toxin sequences to these nontoxin sequences to determine whether toxins are unique in their evolutionary patterns. While a comparison between the venom-gland transcriptomes of the two venom types in C. horridus might have provided a more precise comparison of expression patterns underlying the two venom types, such a comparison would provide substantially less data on toxin and nontoxin molecular evolution and on patterns of gene-family evolution in snake venoms. By comparing the venom-gland transcriptomes of C. horridus and C. adamanteus, we provide the first transcriptome-based comparison between type I and type II rattlesnake venoms and the first genome-scale characterization of molecular divergence between two closely related venomous snake species.
Results and discussion
Crotalus horridusvenom-gland transcriptome sequencing and assembly
We generated a total of 113,344,311 pairs of 100-nucleotide (nt) raw reads, and 104,457,593 pairs passed the Illumina quality filter. We merged 64,169,665 pairs into high-quality composite reads on the basis of their 3’ overlaps as described by Rokyta et al. [17] and Rodrigue et al. [51]. These composite reads had average lengths of 133 nt with average phred scores of 46 and were the only reads used for assembly. Although we could have simply aligned our C. horridus sequencing reads to the transcripts previously identified for C. adamanteus, we performed an independent de novo assembly to provide confirmation of the C. adamanteus annotations, increase the total number of gene sequences identified for the genus Crotalus, and avoid propagating any errors that might be present in the C. adamanteus assembly. We followed the iterative assembly approach of Rokyta et al. [17]. We began by running the Extender program described by Rokyta et al. [17] on a set of 1,000 random reads to identify any extremely high-abundance transcripts. This procedure resulted in 670 contigs, 386 of which had full-length coding sequences. After eliminating duplicates, we had 39 nontoxins and 27 toxins. The high number of duplicates was indicative of extremely biased expression in the venom glands. With these 66 transcripts, we filtered ∼37% of the original reads and performed a de novo assembly with NGen on 10 million of the unfiltered reads. This assembly produced 6,112 contigs from which we annotated 24 full-length toxin and 1,479 full-length nontoxin transcripts. These 1,503 transcripts were used to filter ∼30% of the previously unfiltered reads, and 10 million of the remaining reads were used in a second de novo NGen assembly. This assembly produced 7,084 contigs, and we annotated 25 full-length toxin and 1,080 full-length nontoxin transcripts from these contigs. These 1,105 transcripts were used in a third and final filtering step, removing ∼12% of the reads from the previous set, and 10 million of the unfiltered reads were used in a final de novo NGen assembly. Of the resulting 6,825 contigs, 16 were annotated as full-length toxins and 580 as full-length nontoxins. After eliminating duplicates, our procedure generated 3,031 unique, full-length nontoxin coding sequences and 61 unique, full-length putative-toxin coding sequences.
Our 3,031 full-length, annotated nontoxin sequences accounted for 24.8% of the total sequencing reads, and our 61 toxin transcripts accounted for an additional 40.9%, illustrating clearly the extreme specialization of the venom gland. We were able to account for 65.7% of the sequencing reads (Figure 1). These percentages were similar to those of Rokyta et al. [17] who used the same overall sequencing and assembly approach for C. adamanteus (Figure 1). Our 3,092 annotated sequences (and the 3,002 sequences annotated for C. adamanteus by Rokyta et al. [17]) represent substantial fractions of the genes encoded by these rattlesnakes’ genomes; for comparison, 17,472 protein-coding genes were annotated from a full-genome assembly of the green anole lizard Anolis carolinensis[52]. The sequences generated in the present work and by Rokyta et al. [17] should facilitate genome sequencing and annotation for viperid snakes.
The venom compositions of Crotalus adamanteus and C. horridus differed drastically, consistent with their respective type I and type II classifications. The toxin-gene expression of C. adamanteus was dominated by snake-venom metalloproteinases (SVMPs), C-type lectins (CTLs), snake-venom serine proteinases (SVSPs), and a single, high-abundance myotoxin-A (MYO). These toxin classes, with the exception of SVSPs, were barely detectable in the venom-gland transcriptome of C. horridus, despite a similar number of reads mapping to toxins. The venom-gene expression of C. horridus was instead dominated by SVSPs, phospholipases A2 (PLA2s), and a single gene encoding bradykinin-potentiating and C-type natriuretic peptides (BPP). For both species, unique toxin sequences were grouped into clusters showing less than 1% nucleotide divergence. The other toxin-class abbreviations are as follows: cysteine-rich secretory protein, CRISP; and L-amino acid oxidase, LAAO.
In all that follows, we used the percentage of reads mapping to a given transcript as a proxy for its expression level. This approach was used by Rokyta et al. [17] for C. adamanteus, against whose results we will be making extensive comparisons, and matches the measures used in the many Sanger-sequencing-based venom-gland transcriptomic studies for snakes [13, 14, 53–62]. A measure such as average coverage might help correct for any correlation between the number of reads mapping to a transcript and its length, but our data showed no significant relationship between these two values (F1,3082=0.60, P=0.44). The measures of reads per kilobase of exon model per million mapped reads (RPKM) [63, 64] or fragments per kilobase of transcript per million mapped reads (FPKM) for paired-end reads [65] offer normalization for coding-RNA length and the total number of reads, providing an analog of the relative molar concentrations of transcripts. Note that our sequences, being processed mRNAs, lack introns, and this normalization for coding length is therefore less critical. More sophisticated normalization approaches have also been described [66] for RNA-seq data, but the optimal measure of transcript abundance remains under debate [67] and depends on the purpose and nature of the analyses. Generally, we were concerned with how transcriptional effort is apportioned among transcripts, which makes percentages a natural measure. In addition, percentages yield a form of compositional data [68], which has well-studied properties that enable natural comparisons among subsets. Such an approach is natural for gene-expression data [69].
Type II Crotalus horridustoxin-gene expression patterns
Some of the 61 toxin sequences were similar enough to potentially confound estimates of their individual abundances, so for the purpose of estimating transcript abundances, we clustered these sequences whenever they showed less than 1% nt divergence in their coding sequences. The clustering threshold of 1% is arbitrary but matches the approach used by Rokyta et al. [17] for C. adamanteus. Our average composite-read lengths of 133 nt and a minimum nt divergence of 1% ensure that reads can generally be mapped uniquely to contigs. Clustering the original 61 toxin sequences resulted in 53 toxin clusters (Table 1). In all clusters, the mutations responsible for the differences between the sequences were at substantial frequencies (>10%) with high coverage, suggesting that these sequences were either alleles or recent duplicates rather than simply sequencing errors. Toxins were named by a toxin-class abbreviation, a number designating the cluster (if multiple clusters were present for the class), and a lower-case letter (if the cluster included more than a single sequence). Where necessary to differentiate between species, we preceded toxin names with “Cadam” to indicate those sequences identified in the C. adamanteus transcriptome and “Chorr” to indicate those sequences identified in the C. horridus transcriptome.
Toxin sequences were vastly overrepresented in the venom-gland transcriptome of C. horridus (Figure 2), accounting for 40.9% of the total reads (Figure 1). The seven most highly expressed genes were toxins, as were 15 of the 20 most highly expressed genes. Of the 200 most highly expressed genes, 34 were toxins (Figure 2A). As has been noted previously for other species [4, 17, 60], the venom gland of C. horridus is highly specialized for the production of toxic proteins.
The venom-gland transcriptome of Crotalus horridus was dominated by toxin transcripts. The 61 unique toxin transcripts were grouped into 53 clusters with less than 1% nucleotide divergence for this analysis. Note that all vertical axes are on log scales. (A) The overall expression patterns with all of the full-length annotated sequences showed that toxins dominated the high-abundance transcripts. The inset shows a magnification of the 200 most-abundant transcripts. (B) Expression levels of the toxin sequences showed a clear dominance of snake-venom serine proteinases (SVSPs), phospholipases A2 (PLA2s), and a single gene encoding bradykinin-potentiating and C-type natriuretic peptides (BPP). Expression levels among members of toxin classes were highly variable. The remaining toxin-class abbreviations are as follows: C-type lectin, CTL; cysteine-rich secretory protein, CRISP; cysteine-rich with EGF-like domain, CREGF; glutaminyl-peptide cyclotransferase, GC; hyaluronidase, HYAL; Kunitz-type protease inhibitor, KUN; L-amino acid oxidase, LAAO; myotoxin-A, MYO; nerve growth factor, NGF; neurotrophic factor, NF; nucleotidase, NUC; phosphodiesterase, PDE; snake-venom metalloproteinase, SVMP; vascular endothelial growth factor, VEGF; venom factor, VF; and vespryn, VESP.
The venom of C. horridus, in terms of gene expression, was dominated by three classes of toxin: snake-venom serine proteinases (SVSPs), PLA2s, and bradykinin-potentiating and C-type natriuretic peptides (BPP; Figure 1 and Table 1). These three toxin classes accounted for 93.5% of the reads mapping to toxin sequences. The 11 clusters of SVSPs accounted for 58.2% of the toxin-encoding reads. These enzymatic venom components show highly variable substrate specificities but generally catalyze reactions that disrupt hemostatic mechanisms, including the coagulation cascade and the kallikrein-kinin, fibrinolytic, and complement systems [13, 70]. The nine clusters of PLA2s accounted for 22.8% of the toxin reads. PLA2s show an astounding array of toxic functions, including neurotoxic, cardiotoxic, myotoxic, hemolytic, convulsive, anticoagulant, antiplatelet, edema-inducing, and tissue-damaging effects [71, 72]. Crotalus horridus from the geographic region from which our animal was collected are known to express an extremely potent, heterodimeric, presynaptic β-neurotoxin known as canebrake toxin [44, 45, 73], which is homologous to the notoriously lethal Mojave toxin of C. scutulatus and crotoxin of C. durissus terrificus. These toxins comprise two noncovalently associated subunits, both of which are derived from PLA2 paralogs. The basic subunit is weakly toxic on its own and shows PLA2 activity. The acidic subunit is nontoxic alone and consists of three disulfide-linked polypeptide chains proteolytically derived from a PLA2-like precursor. The two most abundant PLA2 transcripts in the C. horridus venom-gland transcriptome, PLA2-2 and PLA2-1a, are most likely the basic and acidic chains of the canebrake toxin and account together for 15.2% of the toxin reads (Figure 2 and Table 1). PLA2-2 is 2.2% divergent at the nt level and 3.7% divergent at the amino-acid level from the crotoxin basic subunit (GenBank accession X12603). PLA2-1a is 3.2% divergent at the nt level and 8.4% divergent at the amino-acid level from the crotoxin acidic subunit (GenBank accession X12606). Crotalus horridus actually appeared to express four paralogous versions of the acidic subunit, including PLA2-1a, PLA2-4, PLA2-6, and PLA2-9 (Figure 3). The single BPP cluster in the C. horridus venom gland accounts for 12.5% of the toxin reads. These toxins are noted for lowering blood pressure in bite victims [13]. Together, these three major classes of toxins in high abundance suggest a unique viperid bite pathology consisting of coagulopathic, hypotensive, and neurotoxic effects.
Different clades were diversified and highly expressed in Crotalus adamanteus and C. horridus for the phospholipase A 2 (PLA2) gene family. The bars adjacent to gene names give expression levels relative to the most highly expressed family member by species. A completely colored bar indicates the highest expression for the species. Clades comprising sequences from a single species either represent gene-family expansion for that species or gene-family contraction for the other. The four-gene clade including Chorr PLA2-1a and the five-gene clade including Cadam PLA2-1a are clear examples of this phenomenon. Duplication and gene-loss events were inferred by means of a duplication-loss parsimony model. We used a homologous nontoxin-PLA2 sequence from the european rabbit (Oryctolagus cuniculus) as an outgroup to root the phylogeny. Bayesian posterior probabilities are shown for clades for which the values exceeded 50%. Numbers within circles indicate that more than one duplication event was inferred for that node.
Major classes typical of viperid venoms are notably at extremely low abundances in the C. horridus venom (Figure 1 and Table 1). We detected only three clusters of SVMPs, which accounted for just 0.11% of the toxin reads. These venom enzymes are responsible for most of the tissue damage and hemorrhage associated with most viperid bites [74] and contribute to predigestion of prey [33]. Similarly, the 11 C-type lectin (CTL) clusters accounted for just 0.22% of the toxin reads. These toxins, which typically function as multimers, are major components of hemorrhagic viperid venoms and contribute to the disruption of hemostasis by affecting plasma components and blood cells [75], ultimately leading to hemorrhage [76]. Finally, a single MYO cluster was detected, but it accounted for just 0.20% of the toxin reads. Although MYOs appear to be compatible with both type I and type II venoms [33], their primary role appears to be rapid prey incapacitation. A highly potent neurotoxin like canebrake toxin may render this particular toxin unnecessary.
We detected a number of additional low-abundance toxins in the venom-gland transcriptome of C. horridus (Figures 1 and 2 and Table 1). We identified an L-amino acid oxidase (LAAO) transcript, accounting for 1.11% of the toxin reads. LAAOs are associated with edema, apoptosis, and the inhibition of platelet aggregation [77]. A single cysteine-rich secretory protein (CRISP) sequence accounted for 0.78% of the toxin reads. CRISPs are thought to interfere with smooth-muscle contraction [78, 79]. The remaining 15 putative toxin sequences accounted for 4.08% of the toxin reads and included two vascular endothelial growth factors (VEGFs), a vespryn (VESP) [80, 81], a nerve growth factor (NGF), a nucleotidase (NUC), three phosphodiesterases (PDEs), a neurotrophic factor (NF), two Kunitz-type protease inhibitors (KUNs), a hyaluronidase (HYAL) [82], a cysteine-rich with EGF-like domain protein (CREGF), a glutaminyl-peptide cyclotransferase (GC) [83], and a venom factor (VF) [84, 85].
Type I versus type II: mRNA expression differences underlie rapid phenotypic evolution
The profiles of toxin classes expressed by type II C. horridus and its type I congener C. adamanteus were vastly different but consistent with their classifications as type I and type II venoms (Figure 1). These drastic differences were present despite the similarities in sizes, diets, and natural history of these two species and the similarity in the relative transcriptional effort expended on venom production (Figure 1). Type II venoms tend to be proteomically simpler than type I [49], and the transcriptome profiles that underlie these two venoms followed this trend. Crotalus adamanteus had 123 unique toxin sequences that fell into 78 clusters. Crotalus horridus had just 61 unique toxin sequences in 53 clusters. In terms of unique venom transcripts or toxin clusters, C. horridus had approximately 50 to 66% the complexity of C. adamanteus. In terms of major toxin classes, the simplicity of C. horridus venom was even more apparent. SVSPs, PLA2s, and, to a lesser extent, BPP made up most of the C. horridus venom transcripts (93.5% of the toxin reads), whereas the venom of C. adamanteus had a more even expression distribution over SVMPs, CTLs, SVSPs, MYO, PLA2s, and LAAO transcripts (Figure 1).
The prevalence of the more complex type I venoms in rattlesnakes [29] is difficult to reconcile with the advantages type II venoms appear to confer. The higher lethality of type II venoms implies greater efficacy in prey capture and reduced energetic costs, although toxicity has not been measured in natural, sympatric prey populations. Complex traits have been hypothesized to pay a cost in terms of their rates of adaptation [86], but this result depends on high levels of pleiotropy that do not appear to hold in most natural systems [87]. On the other hand, the higher complexity of type I venoms could lead to higher survival probability by means of functional redundancy, mutational robustness, and even increased rates of adaptation through an enlarged mutational target [88]. These potential advantages for type I venoms would provide more of a long-term evolutionary advantage, whereas the increased toxicity of type II venoms provide an immediate short-term fitness advantage. This hypothesized conflict between short-term and long-term advantages might explain both the overall prevalence of type I venoms among rattlesnakes and the fact that most species with type II venoms are still polymorphic for type I, even though type II venom dominates certain geographic regions.
Type II venoms are defined in part by their lack of hemorrhagic effects and, in particular, low SVMP activity. For the type I venom of C. adamanteus, SVMPs were the most highly expressed toxin class, accounting for 24.4% of the reads mapping to toxins. In stark contrast, SVMPs were almost undetectable in the expressed genes of the C. horridus venom-gland transcriptome, accounting for just 0.11% of the toxin reads (Figure 1). CTLs contribute to hemorrhage by either inhibiting or activating, and thereby depleting, coagulation factors [76]. They account for 22.2% of the toxin reads for C. adamanteus but just 0.2% of the toxin reads for C. horridus (Figure 1). The lack of hemorrhagic activity by C. horridus venom can therefore be explained by the lack of expression of genes responsible for this activity; we do not yet know whether all of these genes are still present in the genome but no longer expressed, or whether they have been lost from the genome. We do know, however, that some are present but expressed only in minute amounts (Figures 1 and 2).
Type II venoms are characterized by significant neurotoxic effects mediated generally by heterodimeric PLA2s homologous to crotoxin. The crotoxin homolog in C. horridus, canebrake toxin, is responsible for most of the toxicity of the type A venoms [44], and also accounts in part for the difference in PLA2 expression levels between C. adamanteus and C. horridus (Figure 1). Crotalus adamanteus expressed modest amounts of PLA2 transcripts (7.8% of its toxin reads), but, for C. horridus, PLA2s were the second most abundant class (22.8% of toxin reads). In C. scutulatus, which shows similar venom types to C. horridus, populations with predominantly type II venom show a corresponding absence of MYO [25], which causes myonecrosis and spastic hind-leg paralysis. The most abundantly expressed gene in the C. adamanteus venom-gland transcriptome was a MYO related to crotamine, but this gene was barely detectable in our type II C. horridus, accounting for just 0.2% of the toxin reads. This toxin’s probable role, prey incapacitation, is probably subsumed by the action of canebrake toxin in C. horridus.
Both species expressed high levels of SVSP transcripts, although SVSP transcripts accounted for a significantly larger portion of the toxin expression in C. horridus than in C. adamanteus (58.2% versus 20.0% of the toxin reads). Interestingly, the acidic subunit of crotoxin and its homologs are proteolytically cleaved into three peptides to produce the mature toxin; the protease responsible for the reaction is unknown, but could potentially be one or more serine proteinases, which might account for the higher expression of SVSPs in C. horridus.
A full proteomic characterization [8] and comparison will be necessary to determine whether the expression differences described above account for all of the differences in composition between the venoms of C. adamanteus and C. horridus. We have shown that dramatic changes in expression patterns for toxin gene classes underlie the correspondingly dramatic differences in venom composition and can account for the major pharmacological differences in the effects of the two venoms. In addition to the expression changes by toxin classes, changes of expression among paralogs within classes and sequence changes in individual toxins could also contribute to the different properties of the venoms (see below). Nonetheless, we have shown that extremely large and evolutionarily significant phenotypic changes between closely related species can be mediated by major changes in gene-expression patterns involving many genes, even over short evolutionary times. These dramatic changes in expression highlight a major advantage of chemical means of prey incapacitation and defense. Because venom genes, as far as is known, are expressed only in the venom glands (but see Casewell et al. [89]), major alterations in venom-gene expression can be achieved with no antagonistic pleiotropic effects. Similar large expression shifts for more typical genes would probably be strongly deleterious. Venoms are clean phenotypic modules that can undergo large changes with few, if any, deleterious pleiotropic effects, giving them potential for high evolvability.
Type II venoms differ extensively among species
Although type II venoms are unified in their broad pharmacological properties, they are far from uniform in their compositions. In those cases where it has been investigated, the neurotoxicity of these venoms was derived from the heterodimeric PLA2 crotoxin and its homologs, but the few data available suggest differences in the remainder of the expressed genes as well as in the relative amount of crotoxin homologs. A low-coverage venom-gland transcriptome for C. durissus collilineatus[60], which expresses type II venom, showed that the transcripts encoding the two subunits of crotoxin account for 88% of the toxin-encoding transcripts. In contrast, PLA2 transcripts as a class only accounted for 22.8% of the toxin reads for C. horridus. SVSP transcripts were the most abundant toxin class for C. horridus at 58.2% of the toxin reads, but they only accounted for 2.5% of the toxin sequences for C. durissus collilineatus. Proteomic data from C. simus neonates [10] and C. tigris[49], both of which express type II venoms, suggest a closer agreement with our results. In both cases, however, the SVSPs were expressed at lower levels than for C. horridus (36.0% and 26.8% compared to 58.2%), and the PLA2s were expressed at higher levels (55.9% and 66.2% compared to 22.8%). Note, however, that we are comparing transcriptomes to proteomes, which do not always show quantitative agreement [90]. Nonetheless, these differences may be responsible for the lower LD50 for the venom of C. tigris (0.07 μg/g) compared to type II C. horridus (0.22–1.0 μg/g) [29, 44].
If we assume that type I venom in adults is ancestral on the basis of its higher frequency among extant species [29], then the transition to type II venom has occured multiple times in rattlesnakes within the last ∼12.7 million years [36], a remarkable example of parallel phenotypic evolution. Calvete et al. [10] suggested that for C. durissus, type II venom represents a paedomorphic trait. If the ancestral state of rattlesnakes involved a switch between type II venom in neonates to type I as adults, such as is currently seen in C. simus[10], paedomorphism could provide a simple mechanism for frequent parallel evolution [33]. Unfortunately, the frequency of type II to type I ontogenetic shifts in rattlesnakes is unknown, although ontogenetic shifts in venom composition are not uncommon in viperids [10, 26–28, 91–93]. Of course, given the well-known among-species variation in venom composition of adult snakes, we expect similar levels of variation among juvenile venoms, thereby compounding the difficulties in elucidating the evolutionary history and patterns for snake venoms. A full investigation into the mechanisms of evolution of type II venom phenotypes in rattlesnakes, including determination of whether they represent paedomorphic traits, could provide insight into the repeatability of and constraints on large-scale phenotypic evolution.
Toxin gene-family expansion and differential paralog expression
The evolution of animal-toxin multigene families is characterized by frequent gene gain and loss and strong positive selective pressures [1]. Such patterns have been described for PLA2s [2] and three-finger toxins [7] in snakes. Unfortunately, studies of these gene families rely on sparse and unsystematic sampling of toxin sequences within species and uneven sampling across species [7] because, until recently [4, 17], complete, sequence-based characterizations of the venom components of a species were not feasible. Such sampling deficiencies probably introduce little, if any, bias into statistical tests of positive selection, but could have substantial impacts on the estimation of duplication and gene-loss rates. In particular, this bias could generate spurious signals for gene loss [7]. With our two high-coverage venom-gland transcriptomes for C. horridus and C. adamanteus, we provided the first detailed characterization of toxin gene-family evolution for snakes. Note that we could only detect sequences present in the genome and expressed in the venom glands, so gene loss in this context means that the gene was either deleted from the genome or it was no longer expressed. We only considered the evolution of the PLA2s and SVSPs, because these two families were the only two diverse gene families expressed at appreciable levels in both species.
We identified nine PLA2 transcripts for C. horridus and six for C. adamanteus (Figure 3) and used a related sequence from the european rabbit (Oryctolagus cuniculus) as our outgroup. This outgroup sequence was selected on the basis of tblastx searches of our PLA2 sequences against the NCBI nonredundant nt database, excluding results from viperids. To reconcile the PLA2 gene-family tree with the known species tree required at least 13 duplication events, one loss in C. horridus, and 4 losses in C. adamanteus (Figure 3). Crotalus adamanteus lacked orthologs for the two most highly expressed PLA2 paralogs for C. horridus: Chorr PLA2-1a and Chorr PLA2-2. Both species had bursts of species-specific gene-family expansion that involved multiple duplication events (Figure 3). The most highly expressed PLA2 for C. adamanteus, Cadam PLA2-1a, was part of a five-sequence clade unique to C. adamanteus. Similarly, the second most highly expressed PLA2 for C. horridus, Chorr PLA2-1a, was part of a four-sequence clade unique to C. horridus. These results suggest that gene-family expansion and expression levels are related, although this apparent relationship could simply reflect the use of expansion as a means of increasing expression levels. We identified 11 and 14 SVSP transcripts for C. horridus and C. adamanteus, respectively (Figure 4). The phylogeny of these sequences, with an elapid SVSP as an outgroup, showed a complex pattern of gene gain and loss, with an estimated 17 duplication events, three losses in C. horridus, and two losses in C. adamanteus (Figure 4). In constrast to the PLA2s, SVSPs showed no massive species-specific clade expansions, although species-specific duplication events were common (Figure 4). Similarly to the PLA2s, the SVSP clades expressed at high levels were different between the two species. For example, the most highly expressed SVSP for C. horridus, Chorr SVSP-1a, was paired with a low-expression C. adamanteus ortholog, Cadam SVSP-2. Our results provided the first detailed characterization of toxin gene-family evolution across closely related species. We found an overall pattern of gene-family expansion, with duplications greatly outnumbering losses. At the full-venom level, transcriptional effort among toxin classes was found to be dramatically different between C. horridus and C. adamanteus (Figure 1), and this same pattern repeated itself at a finer scale among paralogs within toxin classes (Figures 3 and 4). Even over the short time scale of divergence between congeners, expression levels for toxins were highly dynamic.
The evolution of the snake-venom serine proteinase (SVSP) gene family in Crotalus adamanteus and C. horridus was characterized by duplication and changes in expression patterns. The bars adjacent to gene names give expression levels relative to the most highly expressed family member by species. A completely colored bar indicates the highest expression for the species. Gene-family expansion and contraction appear to be less pronounced in SVSPs than phospholipases A2 (Figure 3). Duplication and gene-loss events were inferred by means of a duplication-loss parsimony model. We used a homologous SVSP from the king cobra (Ophiophagus hannah) as an outgroup to root the phylogeny. Bayesian posterior probabilities are shown for clades for which the values exceeded 50%.
Sequence divergence between Crotalus adamanteus and C. horridus
Claims abound of increased and exceptional evolutionary rates and selective pressures affecting snake venom genes [1–4], but these studies suffer from major limitations. With the exception of Gibbs and Rossiter [3], these studies average rates over the history of gene families and species and therefore capture only long-term patterns of molecular evolution. The most significant problem with these studies is the complete lack of a null expectation for molecular evolutionary patterns in snakes. We would like to know the proportion of venom genes that are evolving quickly over short time scales and whether these genes, and by implication the venom trait itself, are unique within the genome in terms of their evolutionary patterns. To address these questions, we used our annotated nontoxin sequences from C. adamanteus and C. horridus as the basis for our null expectation for molecular evolution and compared the patterns for toxins to the patterns for nontoxins. To generate our null expectations, we identified orthologous pairs of nontoxins for the two species by means of reciprocal-blast analyses. We excluded mitochondrially encoded sequences from these analyses because of their well-known high evolutionary rates. Each sequence for each species was searched against a database generated for the other species, and we performed separate searches on amino-acid sequences with blastp and nucleotide sequences with blastn. We only kept pairs of putative orthologs that were each others’ best matches for both analyses. From the 3,021 sequences from C. horridus and the 2,870 from C. adamanteus, we identified 1,903 putatively orthologous pairs. We excluded 90 pairs after alignment because their alignments contained more than 24 gapped positions, leaving 1,813 aligned pairs of orthologs. A similar treatment with the 79 toxin clusters from C. adamanteus and 53 from C. horridus yielded 30 toxin alignments.
We conducted three separate analyses of molecular evolution. For the first (Figure 5), we compared the pairwise nonsynonymous to synonymous substitution-rate ratios (d N/d S) of the toxins to the nontoxins. Because our species were closely related and an accurate estimate of d N/d S requires sufficient synonymous divergence, we first estimated pairwise d S values and excluded pairs from the analysis if d S<0.001. In addition, we excluded pairs for which d S>0.1 to avoid spurious orthologs. These constraints left 1,644 nontoxin pairs and 29 toxin pairs. The average d N/d S for nontoxins was 0.18, and the average for toxins was significantly higher at 0.63 (P≪0.001 with a Wilcoxon rank sum test). As a class, toxins have a higher d N/d S than the background nontoxins. We used the individual values from the nontoxins to generate a null distribution for genomic pairwise d N/d S and compared our toxin values to this null distribution (Figure 5). Nine of 29 toxin pairs exceeded the 95th percentile of the null distribution, but we expected fewer than two exceedances if the distributions were the same. About 30% of toxin pairs have exceptionally high d N/d S ratios. As a class of sequences and at the individual level, toxins are distinct from nontoxins in terms of their d N/d S ratios. The low level of sequence divergence between our two species suggests that estimation of the d N/d S ratio may be inaccurate, so we conducted two additional analyses that were free of the issues of estimating a ratio where the denominator is expected to be quite small. We estimated null distributions of d S and d N separately for comparison with the corresponding values from the toxins (Figures 6 and 7). If toxins are uniquely under positive selection, we would expect to see higher rates of substitution for nonsynonymous mutations, but we have no reason to expect a corresponding increase in synonymous substitution rates. Synonymous substitution rate can therefore serve as a control. For both analyses, we excluded pairs with d S>0.1 as for our d N/d S analysis. For d N, we found that the average value for toxins was 0.029, which was significantly higher than the value of 0.017 for nontoxins (P≪0.001 with a Wilcoxon rank sum test). A majority of toxin pairs (18 of 30) exceeded the 95% threshold established by the nontoxin comparisons (Figure 7). However, we also found that toxins had a higher d S than nontoxins (P<0.001 with a Wilcoxon rank sum test), and seven of 30 toxin pairs exceeded the nontoxin 95% threshold. Six of these seven pairs consisted of CTLs, which were expressed at extremely low levels in the C. horridus transcriptome. These pairs possibly represent incorrectly paired paralogs that resulted from the low CTL coverage for C. horridus. Excluding these exceptional seven from the d N analysis still left 11 of 23 toxin pairs exceeding the d N threshold established by the nontoxins. As expected for two closely related species, the distributions of pairwise d N (Figure 7) and d S (Figure 6) showed peaks at zero, corresponding to pairs with little to no sequence divergence. The distribution for d S had a second mode corresponding to approximately 1% sequence divergence.
The distribution of pairwise dN/dS ratios for toxins compared to nontoxins. The histogram shows the null distribution of d N/d S for pairs of nontoxin orthologs identified by means of a reciprocal blast search. The vertical line denotes the cutoff for the 95th percentile for the nontoxins. The d N/d S values for the toxins are plotted below the histogram, and those toxin comparisons with values exceeding the 95th percentile for nontoxins are indictated with triangles and labeled with the names of the two sequences compared. We only expected 1.45 toxins to exceed this threshold if the distributions were the same for toxins and nontoxins. The P value is based on a Wilcoxon rank sum test and shows that the average d N/d S is significantly different between toxins and nontoxins. Comparisons with d S>0.1 were excluded. We also excluded comparisons with d S<0.001 to avoid inaccurately large d N/d S estimates.
The distribution of pairwise synonymous divergence ( dS ) for toxins compared to nontoxins. The histogram shows the null distribution of d S for pairs of nontoxin orthologs identified by means of a reciprocal blast search. The vertical line denotes the cutoff for the 95th percentile for the nontoxins. The d S values for the toxins are plotted below the histogram, and those toxin comparisons with values exceeding the 95th percentile for nontoxins are indictated with triangles and labeled with the names of the two sequences compared. The P value is based on a Wilcoxon rank sum test and shows a significant difference between the average d S of toxins and nontoxins. Comparisons with d S>0.1 were excluded from the analysis. If the distributions were the same between toxins and nontoxins, we expected to see 1.5 toxins exceeding the threshold instead of the seven that were observed. Six of these seven toxin pairs were C-type lectins, a toxin class that showed a 100-fold reduction in expression in C. horridus relative to C. adamanteus. The CTLs above the threshold may represent a failure to find the true ortholog in the C. horridus transcriptome due to low coverage.
The distribution of pairwise nonsynonymous divergence ( dN ) for toxins compared to nontoxins showed an excess of outliers for toxins. The histogram shows the null distribution of d N for pairs of nontoxin orthologs identified by means of a reciprocal blast search. The vertical line denotes the cutoff for the 95th percentile for the nontoxins. The d N values for the toxins are plotted below the histogram, and those toxin comparisons with values exceeding the 95th percentile for nontoxins are indictated with triangles and labeled with the names of the two sequences compared. The P value is based on a Wilcoxon rank sum test and shows a significant difference in the mean d N between toxins and nontoxins. Comparisons with d S>0.1 were excluded, and those comparisons that also exceeded the synonymous divergence threshold (Figure 7) are labeled with white triangles instead of grey. Excluding the d S outliers, we found that almost half of the toxin comparisons exceeded the threshold established by the nontoxins, rather than the expected 5%.
We have shown that even over the short amount of time since C. horridus and C. adamanteus shared a common ancestor, many, but by no means all, venom-encoding genes have evolved in an exceptional manner compared to other coding sequences in the genome. About 30% of toxin sequences showed evidence for a higher d N/d S ratio relative to the nontoxin background sequences, a pattern consistent with those sequences having experienced stronger and/or more prolonged positive selection. Only about 20% of toxins, however, showed d N/d S>1 (Figure 5), which is definitive and extremely conservative [94] evidence for positive selection as opposed to relaxed purifying selection. We also showed that about 50% of toxin sequences had exceptionally high nonsynonymous substitution rates relative to nontoxins, which is also consistent with strong, continual positive selection acting on toxins, although relaxed purifying selection cannot be ruled out. Our null distributions for these measures were derived from evolutionary patterns for a diverse array of nontoxin sequences that were expressed in the venom glands of both species and therefore may not reflect the prevailing patterns throughout the rest of the genome. Our large sample size of nearly 2,000 nontoxin genes, however, represents a substantial fraction of the coding sequences in the genome. In addition, the extremely different venom compositions of our two species resulted in a fairly small sample size for toxins. While this small sample size was sufficient to demonstrate the molecular-evolutionary distinctiveness of toxins compared to nontoxins, future comparisons among species with the same venom types will provide more power and higher-resolution characterizations of the differences in evolutionary patterns between toxins and nontoxins.
Conserved nontoxin-expression patterns between Crotalus adamanteus and C. horridus
The differences in expression patterns between C. adamanteus and C. horridus for genes encoding putative toxins are dramatic (Figure 1) and commensurate with the phenotypic difference in their venoms’ effects, but these toxin genes are only a minute portion of the genes expressed at high levels in the venom glands (Figure 2). Rokyta et al. [17] found that for C. adamanteus, the nontoxin expression was heavily biased towards genes involved in protein production and metabolism, as expected for a tissue specialized for protein secretion. Regardless of the particular proteins secreted, the basic protein-secretory function of snake venom-glands should be consistent across species, and we therefore expected expression patterns among nontoxin sequences to be more similar than patterns for toxins. To test this hypothesis, we conducted a reciprocal-blast analysis to identify orthologous sequences between the two species and compared their expression levels. For a pair of sequences to be included in our analysis, the two sequences had to have been each other’s best blast hit for both a nucleotide-based and an animo-acid-based search with blastn and blastp, respectively. We excluded mitochondrially encoded sequences for simplicity, leaving 2,870 nontoxins for C. adamanteus and 3,021 nontoxins for C. horridus. From these sequences, we identified 1,903 reciprocal blast matches (Figure 8). For expression levels, we used the number of reads mapping to each sequence based on aligning 10 million reads from each species to their own transcript sequences. Because the same number of reads were used for each species, this procedure is effectively equivalent to using a percentage. If the overall expression levels for nontoxins were similar between the two species, which appeared to be the case (Figure 1), and the expression levels of individual nontoxins were similar, we would expect a linear relationship between expression values for the two species with a slope of one and an intercept at zero. Instead, we found a good linear fit (F1,1901=3.3×104, P≪10−10, R2=0.95), but with a slope of 0.48 and intercept of 417 reads when we used C. horridus expression values as a response variable. This lower-than-expected slope appeared to be caused by a single major outlier point representing a protein disulfide isomerase, the most highly expressed nontoxin identified for both species. Removal of this single point gave a good fit (F1,1900=1.4×104, P≪10−10, R2=0.88) with a slope of 0.95 and an intercept of 51, indicating that expression levels for nontoxin sequences present for both species are generally in close agreement.
Expression levels for nontoxins were conserved between Crotalus adamanteus and C. horridus despite major differences in toxin-expression patterns. In all plots, the diagonal line indicates equal expression levels across species. (A) The comparison between species on the basis of orthologs identified by means of a reciprocal-blast search. (B) The comparison between species on the basis of aligning 10 million reads from each species against the nontoxin sequences from C. adamanteus. (C) The comparison between species on the basis of aligning 10 million reads from each species against the nontoxin sequences from C. horridus. Note that for (B) and (C), the distortion in the pattern at lower read counts is an artifact of the minimum-read-count threshold for annotating a contig.
Our reciprocal-blast analysis indicated fairly extensive overlap between the nontoxins identified by means of independent de novo transcriptome assemblies for each species, but each species had ∼1,000 sequences without reciprocal-blast hits. This difference could reflect a difference in the identities of particular genes expressed in the two species’ venom-glands, which would represent a substantial difference in expression patterns, or it could simply represent a stochastic difference in the genes that were successfully assembled and annotated. To determine which of these two possibilities was true, we first filtered reads matching toxins for each species and then aligned 10 million of the filtered reads from each species against both sets of annotated nontoxin sequences. For the C. adamanteus nontoxins, C. horridus reads mapped to all but four template sequences (two more only had a single mapped read). For the C. horridus nontoxins, C. adamanteus reads mapped to all but a single nontoxin sequence (one more had only a single mapped read). The estimated expression levels in terms of number of mapped reads agreed well between both species for both sets of nontoxins (Figure 8). In addition, the total percentages of mapped reads were similar for both species for both sets of nontoxins. For the C. adamanteus transcripts, 42.6% and 41.2% of the reads mapped for C. adamanteus and C. horridus, respectively. For the C. horridus transcripts, 38.6% and 41.6% of the reads mapped for C. adamanteus and C. horridus, respectively. Note that in Figures 8A and 8B, read counts when mapping a species’ reads against its own transcripts showed a pattern of truncation on the lower end. This truncation reflects our procedure for selecting transcripts for annotation; we only tried to annotate contigs with at least 200 reads for each round of assembly.
Gene expression patterns generally appear to be under stabilizing selection [95–99], and the analyses above showed that nontoxin expression patterns were conserved across C. adamanteus and C. horridus despite major changes in expression patterns for toxin genes. This makes functional sense, because the same types of molecular machinery are needed to serve the secretory function of the venom-gland cells regardless of the particulars of the proteins being expressed, but it does raise questions about the regulatory control of gene expression in the venom glands. Given the large number of genes involved for both the toxins and nontoxins, and the relatively short divergence time between these species, it seems likely that toxins and nontoxins are under different regulatory control. This difference in control could contribute to the evolvability of venom by allowing large-scale changes in venom-gene expression without altering the underlying machinery for toxin production.
Sequence accession numbers
The original, unmerged sequencing reads were submitted to the National Center for Biotechnology Information (NCBI) Sequence Read Archive under accession number SRA058913. The assembled and annotated sequences were submitted to NCBI as a Transcriptome Shotgun Assembly project. This Transcriptome Shotgun Assembly project has been deposited at DDBJ/EMBL/GenBank under the accession GAAZ00000000. The version described in this paper is the first version, GAAZ01000000.
Conclusions
Rattlesnakes rely on their venoms for feeding and defense, and the ecological and evolutionary significance of these venoms ensures variation in their properties and compositions both within and between species. The most dramatic differences in rattlesnake venom properties correspond to a long-recognized dichotomy between neurotoxic and hemorrhagic venoms [29]. The timber rattlesnake (C. horridus) generally has hemorrhagic venom, but populations along the southern edge of their range express highly lethal, neurotoxic venom [44]. We sequenced the venom-gland transcriptome of an individual from one of these populations and compared the results to corresponding data from the eastern diamondback rattlesnake (C. adamanteus) [17], a congener with hemorrhagic venom.
The neurotoxic (type II) venom of C. horridus was about half as complex in terms of number of expressed genes as the hemorrhagic (type I) venom of C. adamanteus. This simplicity of type II venom might explain the general prevalence of type I venoms in rattlesnakes [29], despite the apparent advantage in potency of type II venoms. The higher complexity of type I venoms could provide evolutionary advantages in terms of functional redundancy, mutational robustness, and increased rates of adaptation through an enlarged mutational target. The primary basis for the lower complexity of the type II venom of C. horridus was the almost complete loss of expression of the two major classes of diverse hemorrhagic toxins in type I venoms, the CTLs and SVMPs. Overall, we found that the drastic difference in venom properties resulted from major changes in expression levels of toxin gene families, differential gene-family expansion and loss, changes in which paralogs within gene families were expressed at high levels, and higher d N and d N/d S values in the toxin genes relative to nontoxins. Despite the major expression differences for toxin transcripts, nontoxin expression patterns were consistent across the two species. Our work represents the first high-throughput comparative venom-gland transcriptomics study for snakes and therefore provides the first complete, in-depth look at patterns of toxin gene-family evolution, molecular evolution, and expression evolution in venomous snakes.
Methods
Venom-gland transcriptome sequencing
We followed the approach of Rokyta et al. [17] for the preparation and sequencing of the venom gland. We sequenced RNA from the venom glands of an adult female C. horridus from Bradford County, Florida. The animal weighed 1,134.7 g with a snout-to-vent length of 108 cm and a total length of 116 cm. We stimulated transcription in the glands by means of venom extraction under anesthesia [100]. The snake was anesthetized with a propofol injection (10 mg/kg) and exposure to isoflurane gas, and venom expulsion was initiated by means of electrostimulation. After allowing four days for transcription to be maximized [101], the animal was euthanized by injection of sodium pentobarbitol (100 mg/kg), and its venom glands were removed and transferred into RNAlater. The above techniques were approved by the Florida State University Institutional Animal Care and Use Committee (IACUC) under protocol #0924.
Sequencing and nonnormalized cDNA library preparation were performed by the HudsonAlpha Institute for Biotechnology Genomic Services Laboratory (http://www.hudsonalpha.org/gsl/). Transcriptome sequencing was performed essentially as described by Mortazavi et al. [63] in a modification of the standard Illumina methods described in detail in Bentley et al. [102]. Total RNA was reduced to poly-A+ RNA with oligo-dT beads. Two rounds of poly-A+ selection were performed. The purified mRNA was then subjected to a mild heat fragmentation followed by random priming for first-strand synthesis. Standard second-strand synthesis was followed by standard library preparation with the double-stranded cDNA as input material. This approach is similar to that of Illumina’s TruSeq RNA-seq library preparation kit. Sequencing was performed in one lane on the Illumina HiSeq 2000 with 100-base-pair paired-end reads.
Transcriptome assembly
We followed the iterative transcriptome assembly approach of Rokyta et al. [17]. The majority of our read pairs had overlapping 3’ ends, so we merged these pairs into longer composite reads and updated their phred quality scores accordingly [17, 51]. We also checked for and deleted any adapter sequences. Only these long, high-quality merged reads were used for assembly. We first eliminated extremely high-abundance transcripts by running the Extender program as a de novo assembler on a set of 1,000 random reads, as described by Rokyta et al. [17]. Full-length coding sequences were identified with blastx searches as described below. The resulting unique sequences were used as templates in a reference-based assembly with NGen3.1 with a 98% minimum match percentage. Ten million of the unassembled (i.e., unfiltered) reads were used in a de novo transcriptome assembly with NGen3.1 with the default settings for high-stringency, de novo transcriptome assembly for long Illumina reads, including default quality trimming. The high-stringency setting corresponded to a minimum match percentage of 93%, and we retained contigs comprising ≥200 reads. Any resulting contigs with full-length coding sequences were identified by means of blastx searches (see below). We performed two more iterative rounds of this filtering and de novo NGen assembly to yield the final set of contigs. We checked for duplicates by assembling all of the contigs with the SeqMan module of the DNAStar Lasergene software suite. We had several levels of quality control to prevent sequencing errors from being incorporated into our final sequences. We only used reads that passed Illumina’s quality filter and that were merged into overlapping, composite reads. All of our de novo assemblies with NGen used the default quality end trimming, and we only retained contigs with substantial coverage (≥200 reads).
We used blastx searches as implemented in mpiblast version 1.6.0 (http://www.mpiblast.org/) for identification and annotation of our contigs. Contig sequences were searched against the NCBI nonredundant protein database (nr) with an E-value cutoff of 10 −4, and only the best 10 matches were retained. For toxin identification, hit descriptions were searched for a set of keywords based on known snake-venom toxins and protein classes; any sequence matching these keywords was checked for a full-length putative-toxin encoding sequence. The remaining contigs were screened for those whose match lengths were ≥90% of the length of at least one of their database matches. This step was intended to eliminate fragmented or partial sequences before attempting annotation. Each annotated sequence was checked and confirmed by hand in the SeqBuilder module of the DNAStar Lasergene software suite.
We estimated transcript abundances using high-stringency reference-based assemblies in NGen3.1 with a minimum match percentage of 95. Ten million of the merged reads were mapped onto the full-length, annotated transcripts, and the percentage of reads mapping to each transcript was used as a proxy for abundance. To compare nontoxin expression levels across species, we aligned each species’ reads against both their own and the other species’ annotated nontoxin transcripts using reference-based assembly in NGen3.1 with a minimum match percentage of 95. For each species, we used 10 million reads, after first filtering reads mapping to toxin contigs.
Analysis of molecular-evolutionary patterns
Relationships among toxins within toxin families were determined by means of maximum-likelihood phylogeny estimation with PAUP*, version 4.0b10 [103] and the iterative search strategy described by Rokyta et al. [104]. All alignments were constructed with ClustalW [105]. Evolutionary models were selected using MrModelTest version 2 with Akaike Information Criterion values. Nodal support was estimated by means of posterior clade probabilities using MrBayes version 3.1.2 [106]. Markov chain Monte Carlo searches were run for 10 million generations with four chains, the temperature parameter set to 0.2, and samples taken every 1,000 generations. Samples from the first one million generations were discarded as burn-in. To infer duplication and loss events on the estimated phylogenies by reconciling them with the known three-species phylogenies, we used Notung 2.6 [107, 108].
To compare molecular-evolutionary patterns of toxins to nontoxins, we identified orthologous pairs of sequences from our two species by means of a reciprocal-blast analysis. We constructed nucleotide and amino-acid sequence databases for each species, excluding mitochondrially encoded sequences, and blasted each sequence from each species against the database generated for the other species. We performed blastn and blastp searches for each sequence with an E-value cutoff of 10−6. For blastn searches, we used the entire sequence, including untranslated regions. Putatively orthologous pairs were only retained if the two constituent sequences were each other’s best matches for both the nucleotide-based and amino-acid-sequence-based analyses. The coding sequences of retained pairs were aligned using ClustalW [105]. Alignments with more than 24 gapped positions in the coding sequences were excluded from further consideration to avoid considering potentially incorrectly annotated sequences. For the remaining orthologous pairs, we estimated the pairwise synonymous (d S) and nonsynonymous (d N) substitution rates and the pairwise ratios of nonsynonymous to synonymous substitution rates (d N/d S) with codeml from PAML version 4.4 [109, 110].
Abbreviations
- BPP:
-
Bradykinin-potentiating and C-type natriuretic peptides
- CTL:
-
C-type lectin
- CREGF:
-
Cysteine-rich with EGF-like domain
- CRISP:
-
Cysteine-rich secretory protein
- dN:
-
Nonsynonymous substitution rate
- dN/dS:
-
Ratio of nonsynonymous to synonymous substitution rates
- dS:
-
Synonymous substitution rate
- FPKM:
-
Fragments per kilobase of transcript per million mapped reads
- GC:
-
Glutaminyl-peptide cyclotransferase
- HYAL:
-
Hyaluronidase
- KUN:
-
Kunitz-type protease inhibitor
- LAAO:
-
L-amino acid oxidase
- MYO:
-
Myotoxin-A (crotamine)
- NGF:
-
Nerve growth factor
- NF:
-
Neurotrophic factor
- nt:
-
Nucleotide
- NUC:
-
Nucleotidase
- PDE:
-
Phosphodiesterase
- PLA2:
-
Phospholipase A2
- RPKM:
-
Reads per kilobase of exon model per million mapped reads
- SVMP:
-
Snake venom metalloproteinase
- SVSP:
-
Snake venom serine proteinase
- VEGF:
-
Vascular endothelial growth factor
- VESP:
-
Vespryn (ohanin-like)
References
Kordiš D, Gubenšek F: Adaptive evolution of animal toxin multigene families. Gene. 2000, 261: 43-52. 10.1016/S0378-1119(00)00490-X.
Lynch VJ: Inventing an arsenal: adaptive evolution and neofunctionalization of snake venom phospholipase A2 genes. BMC Evol Biol. 2007, 7: 2-10.1186/1471-2148-7-2.
Gibbs HL, Rossiter W: Rapid evolution by positive selection and gene gain and loss: PLA2 venom genes in closely related Sistrurus rattlesnakes with divergent diets. J Mol Evol. 2008, 66: 151-166. 10.1007/s00239-008-9067-7.
Rokyta DR, Wray KP, Lemmon AR, Lemmon EM, Caudle SB: A high-throughput venom-gland transcriptome for the eastern diamondback rattlesnake Crotalus adamanteus and evidence for pervasive positive selection across toxin classes. Toxicon. 2011, 57: 657-671. 10.1016/j.toxicon.2011.01.008.
Biardi JE, Chien DC, Coss RG: California Ground Squirrel Spermophilus beecheyi defenses against rattlesnake venom digestive and hemostatic toxins. J Chem Ecol. 2005, 31 (11): 2501-2518. 10.1007/s10886-005-7610-1.
Jansa SA, Voss RS: Adaptive evolution of the venom-targeted vWF, protein in opossums that eat pitvipers. PLoS One. 2011, 6 (6): e20997-10.1371/journal.pone.0020997.
Fry BG, Wüster W, Kini RM, Brusic V, Khan A, Venkataraman D, Rooney AP: Molecular evolution and phylogeny of elapid snake venom three-finger toxins. J Mol Evol. 2003, 57: 110-129. 10.1007/s00239-003-2461-2.
Calvete JJ, Juárez P, Sanz L: Snake venomics. Strategy and applications. J Mass Spectrom. 2007, 42: 1405-1414. 10.1002/jms.1242.
Calvete JJ, Escolano J, Sanz L: Snake venomics of Bitis species reveals large intragenus venom toxin composition variation: application to taxonomy of congeneric taxa. J Proteome Res. 2007, 6: 2732-2745. 10.1021/pr0701714.
Calvete JJ, Sanz L, Cid P, de la Torre P, Flores-Díaz M, Santos MCD, Borges A, Bremo A, Angulo Y, Lomonte B, Alape-Girón A, Gutiérrez JM: Snake venomics of the central american rattlesnake Crotalus simus and the south american Crotalus durissus complex points to neurotoxicity as an adaptive paedomorphic trend along Crotalus dispersal in South America. J Proteome Res. 2010, 9: 528-544. 10.1021/pr9008749.
Serrano SMT, Shannon JD, Wang D, Camargo ACM, Fox JW: A multifaceted analysis of viperid snake venoms by two-dimensional gel electrophoresis: an approach to understanding venom proteomics. Proteomics. 2005, 5: 501-510. 10.1002/pmic.200400931.
Francischetti IMB, My-Pham V, Harrison J, Garfield MK, Ribeiro JMC: Bitis gabonica (Gaboon viper) snake venom gland: toward a catalog for the full-length transcripts (cDNA) and proteins. Gene. 2004, 337: 55-69.
Pahari S, Mackessy SP, Kini RM: The venom gland transcriptome of the desert massasauga rattlesnake Sistrurus catenatus edwardsii: towards an understanding of venom composition among advanced snakes (Superfamily Colubroidea). BMC Mol Biol. 2007, 8: 115-10.1186/1471-2199-8-115.
Casewell NR, Harrison RA, Wüster W, Wagstaff SC: Comparative venom gland transcriptome surveys of the saw-scaled vipers Viperidae: Echis reveal substantial intra-family gene diversity and novel venom transcripts. BMC Genomics. 2009, 10: 564-10.1186/1471-2164-10-564.
Durban J, Juárez P, Angulo Y, Lomonte B, Flores-Diaz M, Alape-Girón A, Sasa M, Sanz L, Gutiérrez JM, Dopazo J, Conesa A, Calvete JJ: Profiling the venom gland transcriptomes of Costa Rican snakes by 454 pyrosequencing. BMC Genomics. 2011, 12: 259-10.1186/1471-2164-12-259.
Jiang Y, Li Y, Lee W, Xu X, Zhang Y, Zhao R, Zhang Y, Wang W: Venom gland transcriptomes of two elapid snakes Bungarus multicinctus and Naja atra and evolution of toxin genes. BMC Genomics. 2011, 12: 1-10.1186/1471-2164-12-1.
Rokyta DR, Lemmon AR, Margres MJ, Aronow K: The venom-gland transcriptome of the eastern diamondback rattlesnake Crotalus adamanteus. BMC Genomics. 2012, 13: 312-10.1186/1471-2164-13-312.
Sanz L, Escolano J, Ferritti M, Biscoglio MJ, Rivera E, Crescenti EJ, Angulo Y, Lomonte B, Gutiérrez JM, Calvete JJ: Snake venomics of the South and Central American bushmasters. Comparison of the toxin composition of Lachesis muta gathered from proteomic versus transcriptomic analysis. J Proteomics. 2008, 71: 46-60. 10.1016/j.jprot.2007.10.004.
Gibbs HL, Sanz L, Calvete JJ: Snake population venomics proteomics-based analyses of individual variation reveals significant gene regulation effects on venom protein expression in Sistrurus rattlesnakes. J Mol Evol. 2009, 68: 113-125. 10.1007/s00239-008-9186-1.
Chijiwa T, Deshimaru M, Nobuhisa I, Nakai M, Ogawa T, Oda N, ichi Nakashima K, Fukumaki Y, Shimohigashi Y, Hattori S, Ohno M: Regional evolution of venom-gland phospholipase A2 isoenzymes of Trimeresurus flavoviridis snakes in the southwestern islands of Japan. Biophys J. 2000, 347: 491-499.
Creer S, Malhotra A, Thorpe RS, Stöcklin R, Favreau P, Chou WH: Genetic and ecological correlates of intraspecific variation in pitviper venom composition detected using matrix-assisted laser desorption time-of-flight mass spectrometry (MALDI-TOF-MS) and isoelectric focusing. J Mol Evol. 2003, 56: 317-329. 10.1007/s00239-002-2403-4.
Núñez V, Cid P, Sanz L, Torre PDL, Angulo Y, Lomonte B, Gutiérrez JM, Calvete JJ: Snake venomics and antivenomics of Bothrops atrox venoms from Colombia and the Amazon regions of Brazil, Perú and Ecuador suggest the occurrence of geographic variation of venom phenotype by a trend towards paedomorphism. J Proteomics. 2009, 73: 57-78. 10.1016/j.jprot.2009.07.013.
Boldrini-França J, Corrêa-Netto C, Silva MMS, Rodrigues RS, Torre PDL, Pérez A, Soares AM, Zingali RB, Nogueira RA, Rodrigues VM, Sanz L, Calvete JJ: Snake venomics and antivenomics of Crotalus durissus subspecies from Brazil: assessment of geographic variation and its implication on snakebite management. J Proteomics. 2010, 73: 1758-1776. 10.1016/j.jprot.2010.06.001.
Calvete JJ, Sanz L, Pérez A, Borges A, Vargas AM, Lomonte B, Angulo Y, Gutiérrez JM, Chalkidis HM, Mourão RHV, Furtado MFD, da Silva AMM: Snake population venomics and antivenomics of Bothrops atrox: paedomorphism along its transamazonian dispersal and implications of geographic venom variability on snakebite management. J Proteomics. 2011, 74: 510-527. 10.1016/j.jprot.2011.01.003.
Massey DJ, Calvete JJ, Sánchez EE, Sanz L, Richards K, Curtis R, Boesen K: Venom variability and envenoming severity outcomes of the Crotalus scutulatus scutulatus (Mojave rattlesnake) from southern Arizona. J Proteomics. 2012, 75: 2576-2587. 10.1016/j.jprot.2012.02.035.
Mackessy SP: Venom ontogeny in the Pacific rattlesnakes Crotalus viridis helleri and C. v. oreganus. Copeia. 1988, 1988: 92-101. 10.2307/1445927.
Guércio RAP, Shevchenko A, Schevchenko A, López-Lozano JL, Paba J, Sousa MV, Ricart CAO: Ontogenetic variations in the venom proteome of the Amazonian snake Bothrops atrox. Proteome Sci. 2006, 4: 11-10.1186/1477-5956-4-11.
Alape-Girón A, Sanz L, Escolano J, Flores-Díaz M, Madrigal M, Sasa M, Calvete JJ: Snake venomics of the lancehead pitviper Bothrops asper geographic, individual, and ontogenetic variations. J Proteome Res. 2008, 7: 3556-3571. 10.1021/pr800332p.
Mackessy SP: Venom composition in rattlesnakes: trends and biological significance. The Biology of Rattlesnakes. Edited by: Bush S P, Cardwell M D, Beaman K R, Hayes W K, Hayes W K , Beaman K R , Cardwell M D , Bush S P . 2008, Loma Linda, CA: Loma Linda University Press, 495-510.
Zelanis A, Andrade-Silva D, Rocha MM, Furtado MF, Serrano SMT, de Azevedo ILMJ, Ho PL: A transcriptomic view of the proteome variability of newborn and adult Bothrops jararaca snake venoms. PLoS Neglect Trop D. 2012, 6 (3): e1554-10.1371/journal.pntd.0001554.
Daltry JC, Wüster W, Thorpe RS: Diet and snake venom evolution. Nature. 1996, 379: 537-540. 10.1038/379537a0.
Fox JW, Serrano SMT: Structural considerations of the snake venom metalloproteinases, key members of the M12 reprolysin family of metalloproteinases. Toxicon. 2005, 45: 969-985. 10.1016/j.toxicon.2005.02.012.
Mackessy SP: Evolutionary trends in venom composition in the western rattlesnakes (Crotalus viridis sensu lato): toxicity vs. tenderizers. Toxicon. 2010, 55: 1463-1474. 10.1016/j.toxicon.2010.02.028.
Mackessy SP: Fibrinogenolytic proteases from the venoms of juvenile and adult northern Pacific rattlesnakes (Crotalus viridis oreganus). Comp Biochem Physiol B. 1993, 106: 181-189.
Mackessy SP: Kallikrein-like and thrombin-like proteases from the venoms of juvenile and adult Northern Pacific Rattlesnakes (Crotalus viridis oreganus). J Nat Toxins. 1993, 2: 223-239.
Douglas ME, Douglas MR, Schuett GW, Porras LW: Evolution of rattlesnakes (Viperidae; Crotalus) in the warm deserts of western North America shaped by Neogene vicariance and Quaternary climate change. Mol Ecol. 2006, 15: 3353-3374. 10.1111/j.1365-294X.2006.03007.x.
Klauber LM: Rattlesnakes: Their Habits, Life Histories, and Influence on Mankind. 1997, Berkeley, California: University of California Press
McDiarmid RW, Campbell JA, Touré T: Snake Species of the World: a Taxonomic and Geographic Reference, Vol. 1. 1999, Washington, D. C.:, Herpetologists’ League
Campbell JA, Lamar WW: The Venomous Reptiles of the Western Hemisphere. 2004, Ithaca, New York: Cornell University Press
Stechert R: Historical depletion of timber rattlesnake colonies in New York state. Bull N Y Herpetol Soc. 1982, 17: 23-24.
Tyning TF: Conservation of the Timber Rattlesnake in the Northeast. 1990, Lincoln, MA: Massachusetts Audubon Society
Brown WS: Biology, status, and management of the timber rattlesnake (Crotalus horridus): a guide for conservation. Soc Study Amphibians Reptiles Cir. 1993, 22: 78-
Means DB: Effects of rattlesnake roundups on the eastern diamondback rattlesnake (Crotalus adamanteus). Herpetol Conserv Biol. 2009, 4 (2): 132-141.
Glenn JL, Straight RC, Wolf TB: Regional variation in the presence of canebrake toxin in Crotalus horridus venom. Comp Biochem Physiol C. 1994, 107 (3): 337-346.
Straight RC, Glenn JL: Isolation and characterization of basic phospholipase (PLA2) and acidic subunits of canebrake toxin from Crotalus horridus atricaudatus venom using HPLC. Toxicon. 1989, 27: 80-
Wooldridge BJ, Pineda G, Banuelas-Ornelas JJ, Dagda RK, Gasanov SE, Rael ED, Lieb CS: Mojave rattlesnakes (Crotalus scutulatus scutulatus) lacking the acidic subunit DNA sequence lack Mojave toxin in their venom. Comp Biochem Physiol B. 2001, 130: 169-179. 10.1016/S1096-4959(01)00422-5.
Hendon RA, Fraenkel-Conrat H: Biological roles of the two components of crotoxin. Proc Natl Acad Sci U S A. 1971, 68 (7): 1560-1563. 10.1073/pnas.68.7.1560.
French WJ, Hayes WK, Bush SP, Cardwell MD, Bader JO, Rael ED: Mojave toxin in venom of Crotalus helleri (southern Pacific rattlesnake) molecular and geographic characterization. Toxicon. 2004, 44: 781-791. 10.1016/j.toxicon.2004.08.008.
Calvete JJ, Pérez A, Lomonte B, Sánchez EE, Sanz L: Snake venomics of Crotalus tigris: the minimalist toxin arsenal of the deadliest neartic rattlesnake venom. Evolutionary clues for generating a pan-specific antivenom against crotalid type II venoms. J Proteome Res. 2012, 11: 1382-1390. 10.1021/pr201021d.
Calvete JJ: Proteomics in venom research: a focus on PLA2, molecules. Acta Chim Slov. 2011, 58: 629-637.
Rodrigue S, Materna AC, Timberlake SC, Blackburn MC, Malmstrom RR, Aim EJ, Chisholm SW: Unlocking short read sequencing for metagenomics. PLoS One. 2010, e11840 (7):
Alföldi J, Palma1 FD, Grabherr M, Williams C, Kong L, Mauceli E, Russell P, Lowe CB, Glor RE, Jaffe JD, Ray DA, Boissinot S, Shedlock AM, Botka C, Castoe TA, Colbourne1 JK, Fujita MK, Moreno RG, ten Hallers BF, Haussler D, Heger A, Heiman D, Janes DE, Johnson J, de Jong PJ, Koriabine MY, Lara M, Novick PA, Organ CL, Peach SE, et al: The genome of the green anole lizard and a comparative analysis with birds and mammals. Nature. 2011, 477: 587-591. 10.1038/nature10390.
de L M Junqueira-de Azevedo I, Ho PL: A survey of gene expression and diversity in the venom glands of the pitviper snake Bothrops insularis through the generation of expressed sequence tags (ESTs). Gene. 2002, 299: 279-291. 10.1016/S0378-1119(02)01080-6.
Ching ATC, Rocha MMT, Leme AFP, Pimenta DC, de Fátima D Furtado M, Serrano SMT, Ho PL deLMJunqueira-de: Some aspects of the venom proteome of the Colubridae snake Philodryas olfersii revealed from a Duvernoy’s (venom) gland transcriptome. FEBS Lett. 2006, 580: 4417-4422. 10.1016/j.febslet.2006.07.010.
Cidade DAP, Simão TA, Wagner G, de LM Junqueira-de Azevedo I, Ho PL, Bon C, Zingali RB, Albano RM, Dávila A M R: Bothrops jararaca venom gland transcriptome: analysis of the gene expression pattern. Toxicon. 2006, 48: 437-461. 10.1016/j.toxicon.2006.07.008.
de Azevedo ILMJ, Ching ATC, Carvalho E, Faria F, Nishiyama Jr MY, Diniz MRV, Ho P L: Lachesis muta (Viperidae) cDNAs reveal diverging pit viper molecules and scaffolds typical of cobra (Elapidae) venoms: implications for snake toxin repertoire evolution. Genetics. 2006, 173: 877-889. 10.1534/genetics.106.056515.
Liu Q, Zhang X, Yin W, Li C, Huang Y, Qiu P, Su X, Hu S, Yan G: A catalog for transcripts in the venom gland of the Agkistrodon acutus identification of the toxins potentially involved in coagulopathy. Biochem Biophys Res Commun. 2006, 341: 522-531. 10.1016/j.bbrc.2006.01.006.
Wagstaff SC, Harrison RA: Venom gland EST analysis of the saw-scaled viper, Echis ocellatus, reveals novel α9β1 integrin-binding motifs in venom metalloproteinases and a new group of putative toxins, renin-like aspartic proteases. Gene. 2006, 377: 21-32.
Zhang B, Liu Q, Yin W, Zhang X, Huang Y, Luo Y, Qiu P, Su X, Yu J, Hu S, Yan G: Transcriptome analysis of Deinagkistrodon acutus venomous gland focusing on cellular structure and functional aspects using expressed sequence tags. BMC Genomics. 2006, 7: 152-10.1186/1471-2164-7-152.
Boldrini-França J, Rodrigues RS, Fonseca FPP, Menaldo DL, Ferreira FB, Henrique-Silva F, Soares AM, Hamaguchi A, Rodrigues VM, Otaviano AR, Homsi-Brandeburgo MI: Crotalus durissus collilineatus venom gland transcriptome: analysis of gene expression profile. Biochimie. 2009, 91: 586-595. 10.1016/j.biochi.2009.02.001.
Leão LI, Ho PL, de LM Junqueira-de Azevedo I: Transcriptomic basis for an antiserum against Micrurus corallinus (coral snake) venom. BMC Genomics. 2009, 10: 112-10.1186/1471-2164-10-112.
Rodrigues RS, Boldrini-França J, Fonseca FPP, de la Torre P, Henrique-Silva F, Sanz L, Calvete JJ, Rodriques VM: Combined snake venomics and venom gland transcriptome analysis of Bothropoides pauloensis. J Proteomics. 2012, 75: 2707-2720. 10.1016/j.jprot.2012.03.028.
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B: Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 2008, 5 (7): 621-628. 10.1038/nmeth.1226.
Ghiselli F, Milani L, Chang PL, Hedgecock D, Davis JP, Nudzhin SV, Passamonti M: De novo assembly of the manila clam Ruditapes philippinarum transcriptome provides new insights into expression bias, mitochondrial doubly uniparental inheritance and sex determination. Mol Biol Evol. 2012, 29 (2): 771-786. 10.1093/molbev/msr248.
Garber M, Grabherr MG, Guttman M, Trapnell C: Computational methods for transcriptome annotation and quantification using RNA-seq. Nat Methods. 2011, 8 (6): 469-477. 10.1038/nmeth.1613.
Robinson MD, Oshlack A: A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010, 11: R25-10.1186/gb-2010-11-3-r25.
Bullard JH, Purdom E, Hansen KD, Dudoit S: Evaluation of statistical methods for normalization and differential expression in mRNA-Seq, experiments. Genome Biol. 2010, 11: 94-10.1186/gb-2010-11-9-r94.
Aitchison J: The Statistical Analysis of Compositional Data. 1986, : London Chapman and Hall
Vêncio RZN, Varuzza L, de B Pereira CA, Brentani H, Shmulevich I: Simcluster: clustering enumeration gene expression data on the simplex space. BMC Bioinformatics. 2007, 8: 246-10.1186/1471-2105-8-246.
Phillips DJ, Swenson SD, Francis S Markland J: Thrombin-like snake venom serine proteinases. Handbook of Venoms and Toxins of Reptiles. Edited by: Mackessy SP. 2010, Boca Raton Florida: CRC Press, 139-154.
Kini RM: Excitement ahead: structure, function and mechanism of snake venom phospholipase A2 enzymes. Toxicon. 2003, 42: 827-840. 10.1016/j.toxicon.2003.11.002.
Kini RM: Structure-function relationships and mechanism of anticoagulant phospholipase A2 enzymes from snake venoms. Toxicon. 2005, 45: 1147-1161. 10.1016/j.toxicon.2005.02.018.
Wang YM, Parmelee J, Guo YW, Tsai IH: Absence of phospholipase A2, in most Crotalus horridus venom due to translational blockage comparison with Crotalus horridus atricaudatus venom. Toxicon. 2010, 56: 93-100. 10.1016/j.toxicon.2010.03.015.
Fox JW, Serrano SMT: Snake venom metalloproteinases. Handbook of Venoms and Toxins of Reptiles. Edited by: Mackessy SP. 2010, Boca Raton Florida: CRC Press, 95-113.
Du XY, Clemetson KJ: Reptile C-type lectins. Handbook of Venoms and Toxins of Reptiles. Edited by: Mackessy SP. 2010, Boca Raton Florida: CRC Press, 359-375.
Arlinghaus FT, Eble JA: C-type lectin-like proteins from snake venoms. Toxicon. 2012, 60: 512-519. 10.1016/j.toxicon.2012.03.001.
Tan NH, Fung SY: Snake venom L-amino acid oxidases. Handbook of Venoms and Toxins of Reptiles. Edited by: Mackessy SP. 2010, Boca Raton. Florida: CRC Press, 221-235.
Yamazaki Y, Hyodo F, Morita T: Wide distribution of cysteine-rich secretory proteins in snake venoms: isolation and cloning of novel snake venom cysteine-rich secretory proteins. Arch Biochem Biophys. 2003, 412: 133-141. 10.1016/S0003-9861(03)00028-6.
Yamazaki Y, Morita T: Structure and function of snake venom cysteine-rich secretory proteins. Toxicon. 2004, 44: 227-231. 10.1016/j.toxicon.2004.05.023.
Pung YF, Wong PTH, Kumar PP, Hodgson WC, Kini RM: Ohanin, a novel protein from king cobra venom, induces hypolocomotion and hyperalgesia in mice. J Biol Chem. 2005, 280 (13): 13137-13147.
Pung YF, Kumar SV, Rajagopalan N, Fry BG, Kumar PP, Kini RM: Ohanin, a novel protein from king cobra venom: its cDNA and genomic organization. Gene. 2006, 371: 246-256. 10.1016/j.gene.2005.12.002.
Kemparaju K, Girish KS, Nagaraju S: Hyaluronidases, a neglected class of glycosidases from snake venom: beyond a spreading factor. Handbook of Venoms and Toxins of Reptiles. Edited by: Mackessy SP. 2010, Boca Raton. Florida: CRC Press, 237-258.
Pawlak J, Kini RM: Snake venom glutaminyl cyclase. Toxicon. 2006, 48: 278-286. 10.1016/j.toxicon.2006.05.013.
Eggertsen G, Lind P, Sjöquist J: Molecular characterization of the complement activating protein in the venom of the Indian cobra (Naja n. siamensis). Mol Immunol. 1981, 18 (2): 125-133. 10.1016/0161-5890(81)90078-X.
Rehana S, Kini RM: Molecular isoforms of cobra venom factor-like proteins in the venom of Austrelaps superbus. Toxicon. 2007, 50: 32-52. 10.1016/j.toxicon.2007.02.016.
Orr HA: Adaptation and the cost of complexity. Evolution. 2000, 54: 13-20.
Wagner GP, Kenney-Hunt JP, Pavlicev M, Peck JR, Waxman D, Cheverud JM: Pleiotropic scaling of gene effects and the ‘cost of complexity’. Nature. 2008, 452: 470-473. 10.1038/nature06756.
Crow KD, Wagner GP: What is the role of genome duplication in the evolution of complexity and diversity?. Mol Biol Evol. 2006, 23 (5): 887-892. 10.1093/molbev/msj083.
Casewell NR, Huttley GA, Wüster W: Dynamic evolution of venom proteins in squamate reptiles. Nat Commun. 2012, 3: 1066-
Ghazalpour A, Bennett B, Petyuk VA, Orozco L, Hagopian R, Mungrue IN, Farber CR, Sinsheimer J, Kang HM, Furlotte N, Park CC, Wen PZ, Brewer H, Wietz K, Pan C, Yordanova R, Neuhaus I, Tilford C, Siemers N, Gargalovic P, Eskin E, Kirchgessner T, Smith DJ, Smith RD, Lusis AJ, Camp II D G: Comparative analysis of proteome and transcriptome variation in mouse. PLoS Genetics. 2011, 7 (6): e1001393-10.1371/journal.pgen.1001393.
Mackessy SP, Williams K, Ashton KG: Ontogenetic variation in venom composition and diet of Crotalus oreganus concolor a case of venom paedomorphosis?. Copeia. 2003, 2003 (4): 769-782. 10.1643/HA03-037.1.
Saldarriaga MM, Otero R, Núñez V, Toro MF, Díaz A, Gutiérrez JM: Ontogenetic variability of Bothrops atrox, and Bothrops asper snake venoms from Colombia. Toxicon. 2003, 42: 405-411. 10.1016/S0041-0101(03)00171-5.
Gibbs HL, Sanz L, Chiucchi JE, Farrell TM, Calvete JJ: Proteomic analysis of ontogenetic and diet-related changes in venom composition of juvenile and adult dusky pigmy rattlesnakes (Sistrurus miliarius barbouri). J Proteomics. 2011, 74: 2169-2179. 10.1016/j.jprot.2011.06.013.
Crandall KA, Kelsey CR, Imamichi H, Lane HC, Salzman NP: Parallel evolution of drug resistance in HIV: failure of nonsynonymous/synonymous substitution rate ratio to detect selection. Mol Biol Evol. 1999, 16 (3): 372-382. 10.1093/oxfordjournals.molbev.a026118.
Denver DR, Morris K, Streelman JT, Kim SK, Lynch M, Thomas WK: The transcriptional consequences of mutation and natural selection in Caenorhabditis elegans. Nat Genet. 2005, 37 (5): 544-548. 10.1038/ng1554.
Rifkin SA, Houle D, Kim J, White KP: A mutation accumulation assay reveals a broad capacity for rapid evolution of gene expression. Nature. 2005, 438: 220-223. 10.1038/nature04114.
Lemos B, Meiklejohn CD, Cáceres M, Hartl DL: Rates of divergence in gene expression profiles of primates, mice, and flies: stabilizing selection and variability among functional categories. Evolution. 2005, 59: 126-137.
Gilad Y, Oshlack A, Rifkin SA: Natural selection on gene expression. Trends Genet. 2006, 22 (8): 456-461. 10.1016/j.tig.2006.06.002.
Bedford T, Hartl DL: Optimization of gene expression by natural selection. Proc Natl Acad Sci U S A. 2009, 106 (4): 1133-1138. 10.1073/pnas.0812009106.
McCleary RJR, Heard DJ: Venom extraction from anesthetized Florida cottonmouths, Agkistrodon piscivorus conanti, using a portable nerve stimulator. Toxicon. 2010, 55: 250-255. 10.1016/j.toxicon.2009.07.030.
Rotenberg D, Bamberger ES, Kochva E: Studies on ribonucleic acid synthesis in the venom glands of Vipera palaestinae (Ophidia, Reptilia). Biochem J. 1971, 121: 609-612.
Bentley DR, Balasubramanian S, Swerdlow HP, Smith GP, Milton J, Brown CG, Hall KP, Evers DJ, Barnes CL, Bignell HR, Boutell JM, Bryant J, Carter RJ, Keira Cheetham R, Cox AJ, Ellis DJ, Flatbush MR, Gormley NA, Humphray SJ, Irving LJ, Karbelashvili MS, Kirk SM, Li H, Liu X, Maisinger KS, Murray LJ, Obradovic B, Ost T, Parkinson ML, Pratt MR, et al: Accurate whole human genome sequencing using reversible terminator chemistry. Nature. 2008, 456 (7218): 53-59. 10.1038/nature07517.
Swofford DL: Phylogenetic Analysis Using Parsimony. 1998, Sunderland, MA: Sinauer Associates
Rokyta DR, Burch CL, Caudle SB, Wichman HA: Horizontal gene transfer and the evolution of microvirid coliphage genomes. J Bacteriol. 2006, 188 (3): 1134-1142. 10.1128/JB.188.3.1134-1142.2006.
Thompson JD, Higgins DG, Gibson TJ: CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22: 4673-4680. 10.1093/nar/22.22.4673.
Huelsenbeck JP, Ronquist F: MrBayes: Bayesian inference of phylogeny. Bioinformatics. 2001, 17: 754-755. 10.1093/bioinformatics/17.8.754.
Durand D, Halldórsson BV, Vernot B: A hybrid micro-macroevolutionary approach to gene tree reconstruction. J Comput Biol. 2006, 13 (2): 320-335. 10.1089/cmb.2006.13.320.
Vernot B, Stolzer M, Goldman A, Durand D: Reconciliation with non-binary species trees. J Comput Biol. 2008, 15 (8): 981-1006. 10.1089/cmb.2008.0092.
Yang Z: PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci. 1997, 13: 555-556.
Yang Z: PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol. 2007, 24 (8): 1586-1591. 10.1093/molbev/msm088.
Acknowledgements
The authors thank Daniel Dye for assistance in collecting the animal used in this study and S. Brian Caudle for assistance in the laboratory. Funding for this work was provided by the National Science Foundation (NSF DEB-1145978).
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
The project was conceived and planned by DRR and KPW. DRR, MJM, and KPW collected and analyzed the data. DR and KPW wrote the manuscript. All authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Rokyta, D.R., Wray, K.P. & Margres, M.J. The genesis of an exceptionally lethal venom in the timber rattlesnake (Crotalus horridus) revealed through comparative venom-gland transcriptomics. BMC Genomics 14, 394 (2013). https://doi.org/10.1186/1471-2164-14-394
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2164-14-394