Abstract
Background
Colletotrichum truncatum is a haploid, hemibiotrophic, ascomycete fungal pathogen that causes anthracnose disease on many economically important leguminous crops. This pathogen exploits sequential biotrophic- and necrotrophic- infection strategies to colonize the host. Transition from biotrophy to a destructive necrotrophic phase called the biotrophy-necrotrophy switch is critical in symptom development. C. truncatum likely secretes an arsenal of proteins that are implicated in maintaining a compatible interaction with its host. Some of them might be transition specific.
Results
A directional cDNA library was constructed from mRNA isolated from infected Lens culinaris leaflet tissues displaying the biotrophy-necrotrophy switch of C. truncatum and 5000 expressed sequence tags (ESTs) with an average read of > 600 bp from the 5-prime end were generated. Nearly 39% of the ESTs were predicted to encode proteins of fungal origin and among these, 162 ESTs were predicted to contain N-terminal signal peptides (SPs) in their deduced open reading frames (ORFs). The 162 sequences could be assembled into 122 tentative unigenes comprising 32 contigs and 90 singletons. Sequence analyses of unigenes revealed four potential groups: hydrolases, cell envelope associated proteins (CEAPs), candidate effectors and other proteins. Eleven candidate effector genes were identified based on features common to characterized fungal effectors, i.e. they encode small, soluble (lack of transmembrane domain), cysteine-rich proteins with a putative SP. For a selected subset of CEAPs and candidate effectors, semiquantitative RT-PCR showed that these transcripts were either expressed constitutively in both in vitro and in planta or induced during plant infection. Using potato virus X (PVX) based transient expression assays, we showed that one of the candidate effectors, i. e. contig 8 that encodes a cerato-platanin (CP) domain containing protein, unlike CP proteins from other fungal pathogens was unable to elicit a hypersensitive response (HR).
Conclusions
The current study catalogues proteins putatively secreted at the in planta biotrophy-necrotrophy transition of C. truncatum. Some of these proteins may have a role in establishing compatible interaction with the host plant.
Similar content being viewed by others
Background
Colletotrichum truncatum (Schwein.) Andrus W.D. Moore causes anthracnose disease of many leguminous species, including lentil (L. culinaris Medik.), soybean (Glycine max (L) Merr.), fababean (Vicia faba L.), and pea (Pisum sativum L.) [1]. This fungal pathogen employs a bi-phasic hemibiotrophic infection strategy to colonize lentil plants. The anthracnose infection is initiated by the attachment of conidia to aerial parts of the host plant. Conidia germinate immediately after adhesion and differentiate to form infection structures called appressoria devoted to mechanical penetration. After the appressorium has been formed, a thin infection or penetration peg emerging from the base of the appressorium pierces the host cuticle and cell wall, and grows in between the plant cell wall and plasma membrane. It differentiates into large, bulbous invasive primary hyphae that are biotrophic in nature. These hyphae are likely to interact with the host plasma membrane but pull away after plasmolysis (Figure 1A), indicating either initial or weak host-pathogen association. This situation differs from that of haustoria (analogous to primary hyphae) and the biotrophic invasive hyphae produced by obligate biotrophic fungi, such as powdery mildews and rusts, and other hemibiotrophs like Magnaporthe oryzae, the causal agent of rice blast, where the host plasma membrane remains adhered to fungal structures upon plasmolysis [2]. The primary hyphae of C. truncatum are entirely confined to the first infected epidermal cell throughout the biotrophic phase [3]. Thereafter, the fungus switches to the necrotrophic phase that is associated with the production of thin secondary hyphae that ramify intra- and inter-cellularly, killing and macerating host tissues by hydrolytic enzymes ahead of infection. Therefore, the biotrophy-necrotrophy switch (Figure 1B) is critical in anthracnose development. During the transition, C. truncatum probably secretes a range of proteins to establish a compatible interaction with its host, including some that may exclusively be involved in switching the pathogen to the necrotrophic phase.
Histochemical analysis of C. truncatum infected lentil leaflet tissues. (A) Plasmolyzed (0.8 M NaNO3) infected cells of lentil cultivar 'Eston' stained with Neutral Red (0.1% in water) vital dye show that the intracellular primary hyphae of C. truncatum do not form any plant-pathogen interface and remain separated during the biotrophic phase. C, conidia; A (*), appressorium; IV, infection vesicle; PH, primary hyphae; and PM, plasma membrane. (B) Infected leaflet cells of Eston display the transition from the biotrophic phase to necrotrophic phase of C. truncatum. BNS, biotrophy-necrotrophy switch; and SH, secondary hyphae. Bars = 10 μm.
In silico analyses of whole genome sequences have predicted that fungal and oomycete phytopathogens possess large repertoires of secretory proteins, which constitute 7 to 10% of predicted proteomes of M. oryzae[4], Phytophthora sojae, P. ramorum[5] and Ustilago maydis[6]. These secretory proteins play diverse roles in virulence and pathogenicity, and interactions with host cells. Secreted hydrolytic enzymes like cell wall degrading enzymes contribute to penetration of the plant cuticle and cell wall, and to tissue maceration during the onset of the necrotrophic stage [5]. Other secretory proteins, such as chitin deacetylase and proteins that bind pathogen cell wall components like chitin binding protein Avr4 of Cladosporium fulvum, modify the pathogen cell wall and are critical in evading the host defense surveillance system [7, 8]. Secreted effector proteins alter biochemical, physiological, and morphological processes in host plant cells, thereby facilitating infection (virulence factors) or triggering defense responses (avirulence factors or elicitors) [9]. Some of these proteins are active in the apoplast where they may interfere with host plant defense processes, e.g. by inhibiting plant proteases and lytic enzymes. Other fungal effector proteins enter into host cells, though the mechanism of entry of effectors is still a mystery. However, the role of RXLR (arginine, any amino acid, leucine, arginine) and dEER motifs (aspartate [less conserved], glutamate, arginine) in host cell targeting of oomycete effectors has been established [10]. Recently, Kale and colleagues [11] have shown that fungal effectors contain functional variants of RXLR motifs, and that the oomycete and fungal RXLR and dEER motifs bind to host cell surface phosphatidylinositol-3-phosphate. This binding may facilitate the uptaking of effectors through endocytosis.
One of the functions of fungal and oomycete effectors (cytoplasmic effectors) is to disrupt the host signal transduction pathways that mediate defense responses. Until now, only one effector, CgDN3 has been identified from Colletotrichum. This effector averts HR cell death during the biotrophic phase of infection and is essential for C. gloeosporioides pathogenicity on Stylosanthes guianensis[12]. Studies conducted to date have investigated the secretome at different stages of the infection process, including penetration (appressorial morphogenesis), biotrophic and necrotrophic stages. Kleeman et al. [13] identified 26 putative soluble secretory proteins from a cDNA library prepared from mature appressoria of C. higginsianum grown in vitro. Sixteen candidate effector proteins were discovered from Venturia inaequalis, the causal agent of apple scab, on the basis of their small size (< 400 amino acids in length when mature), presence of cysteine residues and of a putative SP [14]. Using a yeast signal sequence trap screen, Krijger et al. [15] identified secreted proteins from C. graminicola mycelia grown in vitro on corn cell walls and leaf extract. Recently, Takahara and colleagues [16] developed a fluorescence-activated cell sorting method to purify the intracellular biotrophic hyphae from C. higginsianum from homogenates of infected Arabidopsis leaves and constructed a biotrophy-specific cDNA library. Mosquera et al. [17] identified four biotrophy associated secreted proteins from the invasive hyphae of M. oryzae by interaction transcriptome analysis. So far, few reports have addressed the critical switch point of the hemibiotrophic infection process. Only one gene encoding a switch regulator, CLTA1 ( C . l indemuthianum t ranscriptional a ctivator 1) from C. lindemuthianum, has been identified. Mutants harboring disrupted CLTA1 were unable to switch to the necrotrophic phase. This transcriptional factor is indispensable for pathogenicity on common bean [18].
The objective of the current study was to identify proteins putatively secreted during the in planta biotrophy-necrotrophy switch of C. truncatum. To catalogue such proteins, we have generated 5000 ESTs from a directional cDNA library constructed from susceptible Canadian lentil cultivar 'Eston' inoculated with C. truncatum isolate CT-21. By biocomputational analyses of these ESTs, we identified eleven C. truncatum candidate effector genes. In planta functional analysis of three effectors was carried out using PVX-based agroinfiltration in tobacco leaves.
Results
Generation of C. truncatum ESTs
Excised leaflets of Canadian lentil cultivar 'Eston' inoculated with isolate CT-21 of C. truncatum were collected and examined microscopically to ensure that the pathogen was at the biotrophic to necrotrophic transition stage where secondary hyphae had become visible (Additional file 1). This corresponded to 48-56 hours after inoculation (hai) and we referred to this period of infection as the biotrophy-necrotrophy switch. Total RNA was extracted, and poly A+ mRNA was isolated and used for construction of a directional cDNA library. Using 5'-end single pass sequencing, 5000 ESTs were generated with an average read length of more than 600 nucleotides. BLASTX analysis with a cutoff E value ≤ 10-5 identified 57% as plant sequences, 39% as fungal sequences and the remainder (4%) as unassignable. The high abundance of fungal transcripts was further verified by semiquantitative RT-PCR using the fungal 60S ribosomal gene in an infection time-course study that included control mixtures (RNA mixtures were obtained by combining 0, 10, 20, 30, 40, 50 and 100% RNA from fungal mycelia into RNA from mock-inoculated lentil leaflets). An accumulation of around 40% of fungal 60S ribosomal transcripts, which, in turn, reflect the fungal RNA content, was predicted during the biotrophy-necrotrophy switch. The abundance of plant origin transcripts had declined with fungal proliferation in infected host tissues as shown by RT-PCR analysis of L. culinaris 60S ribosomal gene in the same infection time-course used for assessing the fungal biomass (Figure 2A).
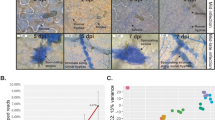
Evaluation of plant and fungal contents in infected lentil leaflet tissues, and cytological analysis of an infection time-course. (A) Semiquantitative RT-PCR amplification of the C. truncatum and L. culinaris 60S ribosomal transcripts in the appressorial penetration phase (16 hai), biotrophic phase (44 hai) and necrotrophic phase (68 hai). Fungal 60S transcripts were also qualitatively assessed in control mixtures, which were obtained by combining 0, 10, 20, 30, 40, 50, and 100% RNA from fungal mycelium into RNA from mock-inoculated leaflets. Twenty six cycles were used for amplification. (B) In planta infection time- course. C. truncatum infected lentil leaflets cultivar 'Eston' at 42-44 hai represent the biotrophic phase characterized by large intracellular primary hyphae and the necrotrophic phase at 68 hai. *Appressorium, IP, Infection peg. Bars = 10 μm.
The fungal transcriptome contains sequences encoding putative secretory proteins
Fungal sequences (1934 ESTs) were analyzed for features indicative of secreted proteins. Because of directional cloning, we could analyze the coding sequences from the 5'-end. ORF finder, and SignalP and iPSORT algorithms were exploited to deduce ORFs and SPs within the ORFs. One hundred sixty-two predicted ORFs (8.37% of the total fungal ESTs) were predicted to encode proteins with N-terminal SPs. All 162 ESTs were deposited in the NCBI GeneBank EST database (accession numbers HO663580 to HO663741). Using ContigExpress software (Invitrogen), these ORFs could be assembled into 32 contigs and 90 singletons, resulting in a total of 122 unigenes. Clone IDs belonging to various contigs are listed in the additional file 2. The average G+C content of these unigenes was around 59%. We refer to the deduced proteins encoded by these unigenes as putative secretory proteins.
We also employed BLASTX and ORF finder algorithms to investigate whether the first methionine within the amino acid translation represented the true N-terminal methionine to confirm the ORF of selected unigenes. The ORFs were then queried against the NCBI non-redundant protein database using BLASTP algorithm. Fungal effectors are most likely to be small, soluble, extracellular secreted proteins that do not become cross-linked into the fungal cell wall [13]. Therefore, predicted ORFs from these unigenes were screened for the size of the encoded polypeptide chain and the presence of transmembrane domains, cysteine residues, transmembrane domain and glycosylation sites, including glycosyl-phosphatidylinositol (GPI) modification. However, with the steady increase in the number of fungal phytopathogen genomes being sequenced, the likelihood that orthologs found in other species is increasing. Hence, some candidate effectors were identified based on orthologs in other phytopathogens. Comparison of the protein sequences encoded by the unigenes to the currently annotated databases and their sequence analyses revealed four groups. The most highly represented group comprised hydrolytic enzymes, which included 63 unigenes (52%), followed by 36 CEAPs (30%), 11 candidate effector proteins (9%) and 11 proteins (9%) classified as "other proteins". Based on these analyses, a total of 43 unigenes were predicted to encode either proteins with transmembrane domain(s) or GPI addition signal. Among them, six were grouped with hydrolases. The remaining 79 unigenes encoded putatively soluble secretory proteins, including hydrolases. A list of the clone IDs, the top hit for each sequence and the corresponding BLAST score are compiled in Table 1. An E value cutoff ≤ 10-5 was used to annotate these unigenes. Four sequences had no match at ≤ 10-5 but contained conserved signatures. Therefore, they were classified according to sequence characteristics belonging to corresponding groups and listed in Table 1.
Hydrolytic enzymes
Sixty-three unique sequences showed significant similarity to members of the enzyme classes (EC) 3 (hydrolases) and 4 (lyases). Most of them had been annotated as cell wall degrading enzymes (CWDEs). Ten unigenes encoded enzymes acting on ester bonds (EC 3 subclass 3.1); most of them were reported as pectinolytic enzymes (pectinases). These included esterases, pectinesterase, extracellular lipase, phosphoserine phosphatase and cutinase. Four depolymerase pectinolytic enzymes (polygalacturonases [EC 3 subclass 3.2] and lyases [EC 4 subclass 4.2]) were also retrieved from this library. Unigenes Ct21-1170 and Ct21-1989 were similar to exopolygalacturonase and endo-polygalacturonase, respectively. Four discrete unigenes contig 22, Ct21-1644, Ct21-1738 and Ct21-3569 encoded pectate lyase (polygalacturonate lyase) and contig 16 showed homology to pectin lyase (polymethylgalacturonate lyase).
Thirty unique sequences, including Ct21-1170 and Ct21-1989 mentioned above represented the EC 3 subclass 3.2 glycosidases (glycosyl hydrolases). Glycosidases hydrolyze the bonds between carbohydrates and release sugars from plant cell walls to provide the fungus with carbon sources and support host penetration and colonization [19]. These included xyloglucan-specific endo-β-1, 4-glucanase, cell wall glucanosyltransferase, cellulase (endoglucanases of class I, II and IV, and extracellular cellulase CelA), glycosyl hydrolase family proteins, β-glucosidase, arabinosidase, β-glucanase, mannan endo-1,4-β-mannosidase and α-galactosidase.
Sixteen unigenes belonged to the EC 3 subclass 3.4 of enzymes acting on peptide bonds (peptidases; aminopeptidase Y, serine type carboxypeptidase, glutamate carboxypeptidase II, carboxypeptidase A, serine endopeptidase, subtilisin serine protease, chymotrypsin, aspartic proteinase and metalloprotease). Correlation between peptidase activity and virulence for several phytopathogenic fungi has already been established.
Cell envelope-associated proteins (CEAPs)
Forty-three unigenes carrying SP, transmembrane helices or GPI anchor addition signal, including 14 unigenes homologous to functionally uncharacterized genes from fungi, were identified as CEAPs. Among them, six unigenes were classified under the hydrolytic enzymes category. In almost all cases except the unigene Ct21-160, the best match for these unigenes was found within the Ascomycota. Interestingly, most of the identified genes have not been previously reported in the genus Colletotrichum.
C-terminal GPI anchor sites were predicted in the translated amino acid sequences of 11 unigenes. Interestingly, we identified a member of the GPI transamidase complex (Ct21-1020) that is implicated in en bloc synthesis of GPI proteins. Three unigenes contig 1, contig 2 and Ct21-4350 encoded GPI anchored serine-threonine rich proteins. Two unigenes, contig 7 and Ct21-3268, were predicted to possess GPI anchor addition signals and cysteine rich fungal extracellular membrane (CFEM) domains (pfam05730). Unigene Ct21-4487 was identified as a hypothetical protein that possesses a GPI addition signal. Contigs 3 and 4 were matched to a putative GPI-anchored protein and GPI-anchored serine rich protein, respectively. Clones Ct21-949 and Ct21-1283 were matched to the GPI anchored cell wall organization protein Ecm33 from Pyrenophora tritici-repentis and glycolipid anchored surface protein GAS1 from C. graminicola, respectively.
The predicted products of the three discrete unigenes Ct21-2424, Ct21-2435 and Ct21-3485 resembled the biotrophy associated secreted glycoprotein Colletotrichum intracellular hypha 1 (CIH1) from C. lindemuthianum, the causal agent of common bean anthracnose [20]. The ORF of Ct21-2435 encoded a protein of 168 amino acids containing 2 lysin motif (lysM) domains. These lysMs recognize and bind N-acetyl D-glucosamine [21], and may play a role protecting fungal chitin from plant chitinases. The predicted ORFs from three discrete clones, contig 29, Ct21-1783 and Ct21-4525 encoded a protein that was similar to clock controlled protein 6 from Neurospora crassa, induced by light and during conidial development [22].
Contig 28 encoded a major facilitator superfamily (MFS) of sugar transporter that contained six transmembrane domains. MFS transporters are involved in the symport, antiport and uniport of various substrates, such as sugars, Krebs cycle intermediates, phosphate esters, oligosaccharides and antibiotics [23]. Contig 30 was similar to glucose-regulated protein 78 (GRP78) in that it is not an extracellular protein. GRP78 is a subgroup of heat shock protein 70 and located in the ER lumen where it assists in the post-translational import and folding of proteins [24] as well as the reversible binding of misfolded and underglycosylated proteins [25]. Clone Ct21-2300 was matched to a toxb precursor of P. tritici-repentis. Toxb is a host specific chlorosis toxin and was identified as a potential pathogenicity factor in P. tritici-repentis, the causal agent of tan spot of wheat [26].
Several enzymes were found to possess transmembrane domains and hence were classified as CEAPs. The Cu/Zn superoxide dismutase (Ct21-3042) is a key enzyme in the dismutation of superoxide radicals resulting from cellular oxidative metabolism into hydrogen peroxide [27]. Clone Ct21-3890 matched an enzyme called peptidyl-prolyl cis-trans isomerase B (PPIase) (EC 5.2.1.8). PPIase possesses a chaperone activity to fold proteins into active configuration by catalyzing cis/trans isomerization on proline-peptide bonds [28]. An enzyme chitin deacetylase (Ct21-4099) involved in modifying fungal cell wall was also identified. Two unigenes, Ct21-4275 and Ct21-4401, encoded metalloreductase and ascorbate peroxidase oxidoreductases, respectively. The metalloreductase contained a transmembrane domain, hence was classified as CEAP, whereas the ascorbate peroxidase lacked a transmembrane domain, and was therefore included under the category of other proteins.
Candidate effector proteins
It is assumed that fungal effectors are likely to be small [< 300 amino acids (aa)], cysteine-rich and soluble extracellular proteins that do not become cross linked into fungal cell walls [13]. With the exponential rise in the number of fungal phytopathogen genomes being sequenced, the likelihood of finding effector orthologs in other species is increasing. For example, C. fulvum lysM effector ECP6 has orthologs from 11 filamentous fungal species, including 7 plant pathogens [21]. We identified 11 secreted proteins (encoded by contigs 5, 6, 8, 10, 31, 32, Ct21-741, Ct21-1573, Ct21-1631, Ct21-2867, and Ct21-4630) that met characteristic criteria, including 8 proteins homologous to functionally uncharacterized proteins from fungal species. The predicted proteins encoded by these unigenes shared the characteristics of being small (112 to 296-aa), lacking a transmembrane domain, and being relatively rich in cysteine (3 to 16 residues). The functionally annotated homologs to two additional clones (contig 29 and Ct21-1783) also shared these features. These candidate effector proteins were predicted to possess cysteine disulfide bridges that may aid protein stability in the extracellular milieu and inside the host cell to protect them from degradation [29, 30]. Putative disulfide bond spacing was conserved between predicted ORFs and similar proteins available in the public domain. Among the functionally annotated proteins, two discrete unigenes (contig 5 and Ct21-1573) were similar to C. fulvum lysM effector protein ECP6 induced during the colonization of tomato [21]. However, unlike ECP6, contig5 and Ct21-1573 contained 1 and 2 lysM, respectively. Contig 8 exhibited similarity to an eliciting plant response-like protein that contained a CP domain. CP proteins from Ceratocystis fimbriata are surface proteins that induce necrosis and elicit phytoalexin synthesis in plants [31]. A unigene (contig 31) was matched to a small secreted protein (139-aa) from Nectria haematococca with 7 cysteine residues (form 3 disulfide bonds) in the mature protein. Contig 32 was similar to a HR-inducing protein from Ceratocystis ulmi. Homologs to 7 unigenes were hypothetical proteins.
Other proteins
The predicted peptides of a group of 11 unigenes showed similarities to diverse proteins from fungal genome sequences. The encoded peptide of clone Ct21-1408 was similar to a chloroperoxidase related protein. Chloroperoxidase catalyzes the insertion of chlorine, bromine and iodine atoms into organic acceptor molecules. EST clone Ct21-3930 was aligned with glutaminase, an enzyme catalyzing the hydrolysis of glutamine to glutamic acid. Glutaminases play a role in the acquisition of nitrogen from less preferred sources [32]. The oxidoreductase enzyme L-ascorbate peroxidase catalyzes the chemical reaction L-ascorbate + H2O2 ↔dehydroascorbate + 2 H2O. The remaining 7 unigenes could not be classified by homology search into known or putative proteins (Table 1). However, sequence analyses of predicted peptides revealed that most of these unigenes were rich in glycine or leucine. Ct21-1373 was identified as a nudix/MutT family protein nudix hydrolase (Pyrenophora tritici-repentis). Nudix hydrolases hydrolyze substrates of general structure nu cleoside di phosphate linked to another moiety, X to yield nucleotide monophosphate and P-X [33].
Time-course expression analysis of genes encoding putative effector proteins and CEAPs
Using semiquantitative RT-PCR, we examined transcript levels of genes encoding candidate effectors and CEAPs (GPI-proteins and CIH1) identified from the EST library in different cell types formed in vitro, viz. ungerminated conidia, mycelia and mycelia grown in minimal media supplemented with lentil leaflet cell walls (referred to hereafter as cell wall treated mycelia). Expression was also analyzed in planta at various stages of plant infection, namely appressorial penetration (16 hai), primary hyphae (44 hai) and secondary hyphae (68 hai) (Figure 2B).
Time-course expression profiles were obtained for 9 candidate effector encoding genes (Figure 3). When considering the in planta infection process, two kinds of expression patterns could be distinguished. EST clones contig 5, 8, 32, Ct21-1573, and Ct21-4630 were expressed at both in planta biotrophic- and necrotrophic-developmental stages. Expression at lower level was also detected in the appressorial penetration stage for two clones, contigs 8 and 32. Three clones, contigs 8, 32, and Ct21-4630 were expressed constitutively in all in vitro cell types. Clones, contig 5 and Ct21-1573 identified as homologs of lysM effector Ecp6 [21], were expressed in mycelia, whereas they were undetectable in conidia and cell wall treated mycelia, suggesting the possible involvement of these effectors genes in vegetative growth. Transcripts from four genes (contigs 6, 10, Ct21-741, and Ct21-1631) were exclusively accumulated during the necrotrophic phase of the infection process. Among them, two genes (contig 6 and Ct21-741) also showed expression in the cell wall treated mycelia, however, at a lower level than during the in planta necrotrophic phase, whereas contig 10 appeared to be expressed in mycelia and cell wall treated mycelia. Clone Ct21-1631 showed constitutive expression in all cell types tested.
Figure 3 shows expression profiles of seven (contigs 1, 2, 3, 4, Ct21-949, Ct21-1020 and Ct21-3268) out of eleven genes encoding GPI-CEAPs. All genes showed constitutive varying expression pattern in all in vitro cell types. In addition, they all were undetectable during appressorial penetration except for Ct21-3268, and during the biotrophic phase except for Contig 2 and Ct21-3268, but all were abruptly expressed during the necrotrophic phase. This was well corroborated by the abundance dynamics of the gene encoding GPI transamidase (Ct21-1020), an enzyme involved in the biosynthesis of GPI-CEAPs that was completely shut down in the biotrophic phase. These data suggest that GPI-CEAPs accumulated during the in planta necrotrophic phase and in vitro growth.
Another subclass of CEAPS tested for expression was CIH1 protein encoding unigenes (Ct21-2424, Ct21-2425 and Ct21-3485). All three were constitutively expressed in all cell types tested, indicating their role in vegetative development of the pathogen. In addition, their transcripts were also accumulated at both biotrophic- and necrotrophic- in planta developmental stages, suggesting that CIH1 glycoprotein is a basal component of the cell wall of C. truncatum as described previously [20].
Functional analysis of selected candidate effectors by PVX agroinfiltration assay
To investigate if candidate effectors have elicitor activity, we expressed 3 candidate effector proteins (encoded by unique sequences contigs 6, 8 and 32) and M. oryzae CP protein by mean of agroinfiltration in tobacco leaves.
EST analysis of contig 6 revealed a cDNA of 784-bp with an ORF of 417-bp encoding a 139-aa preprotein. A SP of 15-aa with a cleavage site in between glycine-15 and serine-16 was predicted in the preprotein, and 7 cysteine residues were found in the mature protein (124-aa). Using DISULFIND server, three cystine disulfide bridges were predicted in the mature protein at a confidence level of > 0.5. This protein showed no homology to any functionally characterized protein. The cDNA of contig 8 was 775-bp in size with an ORF of 414-bp encoding a 138-aa preprotein. A SP of 21-aa with a cleavage site between valine-21 and serine-22, and a CP domain spanned from 23-aa to 138-aa were predicted at the N- and C-termini, respectively. The CP domain contains 4 cysteine residues, which are known to form two disulphide bonds [31]. Proteins belonging to the CP family share common characteristics, such as a small size, an N-terminal SP for secretion and a conserved cysteine residue pattern [34, 35]. Protein encoded by unigene contig 8 possessed all features of a CP protein, hence we termed it CtCP1. Sequence alignment of CtCP1 with homologues from other fungal species revealed that CtCP1 showed significant similarity with fungal proteins of CP family like M. oryzae SM1 (MGG_05344), Hypocrea atroviridis eliciting plant response like protein 1 (Epl; DQ464903), Hypocrea virens snodprot1 (DQ494198) and Gibberella pulicaris snodprot-FS (AY826795) (Additional file 3). Based on sequence features, CtCP1 belongs to Epl1 cluster of CP proteins [35]. Recently, a M. oryzae CP protein MgSM1 was shown to induce HR and confer broad-spectrum disease resistance in Arabidopsis[36]. EST analysis of contig 32 revealed a cDNA of 686-bp with an ORF of 426-bp encoding a 142-aa preprotein. A SP of 17-aa with a cleavage site between alanine-17 and isolecine-18 was predicted at the N-terminus of the preprotein, and 4 cysteine residues were predicted in the mature protein (125-aa). This protein showed significant homology to the HR inducing protein of Ceratocystis ulmi and contained 4-cysteine residues.
The ORFs of three candidate effectors from C. truncatum and MgSM1 from M. oryzae with and without SP were cloned into PVX vector pGR106 [pGR106-contig 61-139, pGR106-contig 616-139, pGR106-CtCP11-138, pGR106-CtCP122-138, pGR106-contig 321-142, pGR106-contig 3216-142, pGR106-MgSM11-137 and pGR106-MgSM122-137] and transformed into Agrobacterium tumefaciens strain GV3101. We used the empty vector pGR106 as negative control, which upon transient expression showed no micro- and/or macroscopic phenotype. We also incorporated two R/Avr protein pairs as positive controls that have been shown to interact and trigger cell death in tobacco: Pto/AvrPto [37] and Cf9/Avr9 [38]. Agroinfiltration of recombinant A. tumefaciens strains carrying pGR106-MgSM11-137 in tobacco leaves resulted in macroscopic cell death 5 days after infiltration, whereas the rest of the strains were unable to cause any macroscopic phenotype (Figure 4). Transient co-expression of the genes that encode Avr/R protein pairs (Pto/AvrPto and Cf9/Avr9) elicited a spotted faint glazing on the tobacco leaf surface 19 h post infiltration, which progressed to tissue collapse and became confluent 5 days post infiltration. To investigate whether CtCP1 protein causes any microscopic cell death, we collected leaf tissues from the edges of pGR106, pGR106-CtCP11-138, pGR106-CtCP122-138, pGR106-MgSM11-137 and pGR106-MgSM122-137 infiltration zones for polyphenolic compound- associated autofluorescence under ultraviolet (UV) light. The pGR106, pGR106-CtCP11-138, pGR106-CtCP122-138 and pGR106-MgSM122-137 edges had no autofluorescence under UV light, whereas pGR106-MgSM1 edges showed strong autofluorescence under UV light (Figure 5).
In planta expression of Contigs 6, 32, CtCP1 and MgSM1 . In planta expression of Contigs 6, 32, CtCP1 and MgSM1 (A) N. tobacum leaves were challenged by agro-infiltration of A. tumefaciens strains expressing pGR106-contig 321-142 (SP), pGR106-contig 3216-142 (-SP), pGR106-contig 61-139 (SP), pGR106-contig 616-139 (-SP), pGR106-CtCP11-138 (SP), pGR106-CtCP122-138 (-SP), pGR106-MgSM11-137 (SP) and pGR106-MgSM122-137 (-SP). Empty vector pGR106 and matching pairs of R/Avr (Pto/AvrPto and Cf9/Avr9) were used as negative and positive controls, respectively. Pictures were taken 7 days after infiltration. SP, Signal peptide.
Discussion
The morphological and nutritional transition of a hemibiotrophic pathogen from biotrophy to necrotrophy plays a critical role in disease development. During this switch, fungal phytopathogens may secrete large repertoires of secretory proteins to manipulate host innate immunity in concert with a destructive arsenal of CWDEs that manifest the disease phenotype. Our strategy in this study was to catalogue genes encoding secretory proteins from C. truncatum by constructing a directional cDNA library representing the biotrophy-necrotrophy switch and then to exploit biocomputational tools to scan resulting fungal ESTs for proteins containing canonical N-terminal SPs. To date, cDNA library- and EST-based studies have been conducted to identify secretory protein encoding genes from haustoria of Puccinia striiformis f.sp. tritici[39], C. higginsianum appressoria [13], in planta infection stages of Venturia inaequalis[14] and mycelia of the corn pathogen C. graminicola grown in vitro under different secretion inducing conditions [15]. There has been no previous EST-based analysis of the biotrophy-necrotrophy switch for any fungal pathogen. Our library contained 122 fungal unigenes encoding secretory proteins. Functional classification of the secretory proteins based on homology searches and sequence analyses revealed the presence of four potential groups: hydrolytic enzymes, CEAPs, candidate effector proteins and other proteins.
Hemibiotrophs orchestrate a physiological switch from asymptomatic infection to massive cell death and tissue dissolution, presumably resulting from the coordinated secretion of cell-death elicitors and hydrolytic enzymes [40]. With 63 of 122 unigenes belonging to EC classes 3 and 4, hydrolases and lyases are a major group of enzymes identified at the biotrophy-necrotrophy switch. In general, fungal pathogens secrete a battery of CWDEs capable of hydrolyzing host cell wall components (cellulose, hemicellulose, pectin and structural proteins) to accommodate a necrotrophic life style. These CWDEs include carbohydrate esterases (pectinases), glycosyl hydrolases, polysaccharide lyases (pectinases) and peptidases, which the fungal pathogens utilize to degrade the host plant cell wall for nutrition [41]. The glycosidases like galactosidase and arabinosidase may liberate sugar moieties that can be used as a nutritional source for C. truncatum during the necrotrophic growth. Glycosidases may also have a role in decreasing the steric hinderance for other CWDEs by removing protruding side chains from the backbone of polymers and providing increased access for endo forms of CWDEs [42]. Pectinases weaken the cell wall and expose cell wall components to other enzymes, such as cellulases and hemicellulases. Genome sequencing has revealed that the hemibiotrophic phytopathogen M. oryzae is predicted to possess 121 glycosyl hydrolases, 4 polysaccharide lyases, and 13 carbohydrate esterases [41]. Our library retrieved 15 glycosyl hydrolases, 2 polysaccharide lyases, and 5 carbohydrate esterases. Another class of identified hydrolases was peptidases that may have a function in the extracellular processing of fungal secreted proteins [43], degradation of proteinaceous components involved in the plant response against pathogens [44] or degradation of host tissue [45]. Esterases, glycosyl hydrolases, lyases, and peptidases could also weaken host cell walls, allowing for an easier access to nutrients.
The interaction between a pathogen and its host is to a large extent orchestrated by the proteins that are secreted or localized to or at the cell membrane or cell wall (CEAPs). The CEAPs are secreted through the endoplasmic reticulum (ER) pathway and tethered either to the fungal cell membrane or wall. Some CEAPs lack a transmembrane domain but possess C-terminal GPI anchor addition signal. GPI anchoring is an important mechanism to tether extracellular proteins to the plasma membrane or cell wall [46]. The carboxyl termini of these proteins have a sequence motif that is recognized by a protein complex located in the ER, known as the GPI transamidase (unigene Ct21-1020). The GPI transamidase complex cleaves the protein at a position within this motif, termed the omega site, and transfers the GPI anchor en bloc to the newly generated C terminus of the protein. A genome-wide inventory of secretory proteins of P. sojae and P. ramorum revealed putative secretomes of 1,464 and 1,188 proteins, respectively [47], of which a subset of approximately 100 proteins contain putative GPI anchor addition signals. The majority of elicitins (capable of inducing an HR in tobacco) from Phytophthora possess a GPI anchor [48]. Components of the cell envelope of pathogens can be rapidly acclimatized by a changing environment. The ability to respond to a changing milieu is critical for pathogens to effectively evade, tolerate and/or suppress host immune responses. Therefore, the structure and composition of the cell envelope is dynamic and changes during different stages of in planta growth. We observed an abrogation or a reduction in transcript levels of genes encoding CEAPs during the biotrophic phase and an abrupt induction in the necrotrophic phase (Figure 3). The switch from voluminous biotrophic hyphae to thin necrotrophic hyphae involves an increased surface area to volume ratio, therefore may require elevated transcription of some of these cell surface protein encoding unigenes. This ratio and thinner cell walls of the secondary hyphae would be advantageous for both efficient nutrient uptake and secretion of wall degrading enzymes, and possibly toxins, during the necrotrophic growth of hemibiotrophic fungal pathogens [49].
The predicted products of three discrete unigenes Ct21-2424, Ct21-2435 and Ct21-3485 resembled the biotrophy associated protein CIH1 from C. lindemuthianum[24]. The CIH1 was shown to be secreted into the cell wall of invasive biotrophic hyphae and the plant-pathogen interface. In the present study, we found that the C. truncatum CIH1 homolog was expressed constitutively in all in vitro cell types tested and both during the biotrophic- and necrotrophic- phases. This result was contradictory to studies with C. higginsianum where the CIH1 protein was specifically expressed in the biotrophic phase [16]. It suggests either the transcription level of CIH1 does not reflect the protein level, or C. truncatum may employ a different approach to host invasion that is unlike that of other species of Colletotrichum.
Based on the in planta transcript profiling of nine genes encoding putative effector proteins, we identified two types of expression patterns: biotrophic/necrotrophic-specific and necrotrophic-specific. Transient expression of three candidate effectors in tobacco leaves revealed that none of them was able to elicit an HR response. Surprisingly, our data on CtCP1 were in contrast of that recently published on M. oryzae CP protein, MgSM1. MgSM1, upon transient expression, stimulated HR and conferred broad-spectrum resistance to Arabidopsis against Botrytis cinerea and Pseudomonas syringae pv. tomato[40].
The predicted products of two discrete unigenes (contig 5 and Ct21-1573) showed homology to extracellular protein 6 (ECP6) from C. fulvum. LysMs are N-acetyl D-glucosamine (chitin) binding modules that have been found in mammalian and plant pathogenic fungi as well as in saprophytes [50]. Therefore, they may also have a role in counter defense by protecting fungal cell walls against plant chitinases. Bolton et al. [21] demonstrated that ECP6 was induced by the plant pathogenic fungus C. fulvum during the initial infection of tomato and acted as a virulence factor. During infection, LysM domain-containing protein Ecp6 sequesters chitin oligosaccharides that are released from the cell walls of invading hyphae to avert the elicitation of host immunity. This sequestering may represent a common strategy of host immune suppression by fungal pathogens, because LysM effectors are widely conserved in the fungal kingdom [51].
We identified several ESTs that may encode novel effectors. Using bioinformatics tools, we identified 5 soluble secreted proteins of unknown function (contig 10, Ct21-741, Ct21-1631, Ct21-2867 and Ct21-4630). Three of them (contig 10, Ct21-741, Ct21-1631) were not expressed in the biotrophic phase while induced during the necrotrophic phase. Their expression was also detected either in some or all of the in vitro cell types. Such genes may be important for establishing fungal necrotrophy. One (Ct21-4630) was constitutively expressed during all in planta developmental stages except appressorial penetration and in all in vitro cell types (Figure 3), suggesting its general role in both vegetative development and pathogenesis.
Conclusions
We catalogued proteins putatively secreted by C. truncatum and several candidate effectors had been mined from the EST library. It was observed that the transcript levels of candidate effector genes were either induced during the necrotrophic growth of the pathogen compared with that during the biotrophic growth or elevated during both phases. Through in planta expression of candidate effectors, we demonstrated the functional role of one effector. In summary, our study suggests that suppression of host defense during the biotrophic phase and the subsequent unmasking of PAMPs or elicitors during the necrotrophic phase may be involved in pathogenesis of C. truncatum.
Methods
Plant material and cell wall extraction
Lentil plants of the Canadian cultivar 'Eston', which is fully susceptible to C. truncatum isolate CT-21, were grown at four plants per 18 cm diameter pot filled with a commercial potting soil mixture (Terralite Redi-Earth®, Scotts-Sierra Horticultural Products Co., Maryville, OH, USA). Pots were maintained in a growth chamber at approx. 25°C with a photoperiod of 16 h light/d provided by fluorescent lamps with light intensities of between 315 and 325 μmol m-2 s-1. Plants were fertilized with complete fertilizer solution (20:20:20 NPK + micronutrients) at two weeks after planting and every two weeks thereafter at a rate of 3 g l-1 and 22 ml per pot.
Cell wall was extracted from 3-week old Eston leaflets following the protocol described by Eshel et al. [52]. One hundred gram of leaflets were ground in 100 mL of acid-methanol (4% v/v HCl) and filtered through Whatman filter paper number 1 (Whatman Ltd., Maidstone, UK). Residues were collected from the filter paper and blended twice more in 100 mL of methanol. This process was repeated until no phenols could be detected in the filtrate. The residues were blended three times in 100 mL of ice chilled acetone, and air dried overnight. The residues collected were then stored at -20°C until use as sole carbon source. Two grams mycelia were incubated on a rotary shaker at 125 rpm and 240 C in 2.5 L flasks containing 200 mL minimal salt broth (20 g/L D-glucose, 1 g/L KH2PO4, 0.5 g/L MgSO4 ×7H2O, 0.5 g/L KCl, 0.15 g/L CaCl2×2H2O, 3 mg/L FeSO4× 7H2O, 3 mg/L ZnSO4×7H2O, 1.25 mg/L CuSO4×5H2O, 350 μg/L MnSO4×H2O and 250 μg/L Na2MoO4×2H2O) supplemented with 0.5% cell wall extract. After 72 hrs mycelia were collected and frozen in liquid nitrogen, and stored at -80°C until required.
Fungal material
The C. truncatum isolate CT-21 used in this study was routinely maintained in Petri dishes containing oatmeal agar (30 g oatmeal flour, Quick Oats Robin Hood, Smucker Food of Canada, Markham, ON, Canada and 8.8 g granulated agar, Difco™, Becton, Dickinson & Company, Sparks, MD, USA] to 1 L of distilled water) at 22°C. For infection assays, conidia were harvested from 10- to 14-day old cultures, washed twice by centrifugation (3000 rpm, 5 minutes), and resuspended in sterile distilled water. For isolating total conidial RNA, conidia were pelleted and kept frozen until required.
For isolating total mycelial RNA, one mL of conidial suspension (4 × 10-4 conidia mL-1) was incubated in complete medium (0.6% yeast extract, 0.3% acid casein hydrolysate, 0.3% enzymatic casein hydrolysate, 1% sucrose) at 22-25°C for 48 h with constant shaking at 150 rpm. Mycelia were harvested by filtering through nylon mesh and rinsed with distilled water twice. Collected mycelia were frozen in liquid nitrogen and stored at -80°C until required. For cell wall treated mycelia, collected mycelia were grown in vitro in minimal salt broth supplemented with Eston cell wall as sole carbon source at 22-25°C for 8 h with constant shaking at 150 rpm. Mycelia were harvested and stored as described above.
RNA isolation and cDNA library construction
For library construction, leaflets of 3-week old lentil cultivar 'Eston' were harvested and inoculated with 15 μL droplets of conidial suspension (4 × 104 conidia mL-1) of C. truncatum isolate CT-21. Three hundred 6 mm diameter infection sites were punched from the leaflets using a cork borer during 48-72 hai and microscopically assessed under a compound microscope to ensure the correct timing for the biotrophy-necrotrophy switch. Leaflet discs (~200) with biotrophic primary hyphae and first indications of secondary hyphal growth signaling the necrotrophic phase, were frozen in liquid nitrogen and stored at -80°C until required.
Total RNA was isolated using phenol/chloroform extraction and LiCl precipitation [53]. Poly(A)+ RNA (mRNA) was purified from total RNA using PolyATtract® mRNA isolation system IV (Promega, Madison, WI, USA). A directional cDNA plasmid library was generated from 5 μg poly(A)+ RNA using the pBluescript® II XR cDNA library construction kit (Stratagene, La Jolla, CA, USA). The resulting cDNA library was ligated into the pBluscript II SK (+) vector using Eco RI and Xho I restriction sites transformed into XL10-Gold ultracompetent cells (Stratagene, La Jolla, CA, USA). The 5'-end single pass sequencing of cDNA inserts using M13 reverse universal primer was carried out in the DNA Technology Unit, Plant Biotechnology Institute, Saskatoon, Canada.
EST analysis
ESTs were trimmed for vector, adaptor, and low quality sequence regions using DNAMAN software. The resulting EST sequences were queried against the NCBI non-redundant protein database using BLASTX algorithm http://www.ncbi.nlm.nih.gov/BLASTand against the Consortium for the Functional Genomics of Microbial Eukaryotes (COGEME) EST database http://cogeme.ex.ac.uk using TBLASTX algorithm [54].
An ORF finder algorithm http://www.ncbi.nlm.nih.gov/gorf was employed to predict coding region in all six frames ab initio; any sequence with a stop codon preceded by in-frame ATG codons was translated into protein sequence. All in-frame ATGs were considered as potential translation initiation sites while taking account of possible upstream ATG codons. The amino acid sequences were then scanned for potential SPs and transmembrane domain using SignalP server version 3.0 with default settings and iPSORT, and TMHMM server version 2, respectively. Proteins with predicted SPs were queried against the NCBI non-redundant protein database using BLASP algorithm with BlOSUM80 matrix without low complexity filter. Potential GPI-anchored proteins were identified using the BIG-PI Fungal Predictor http://mendel.imp.ac.at/gpi. Disulfide bonds in candidate effector proteins were predicted by SULFIND server http://disulfind.dsi.unifi.it. N-linked or O-linked glycosylation sites were predicted using NetNGlyc 1.0 and NetOGlyc 2.0 servers http://www.expasy.org.
Infection time course assays
A detached leaf assay was used for semiquantitative RT-PCR. Leaflets from 3-week old Eston plants were detached and inoculated in Petri dishes lined with wet filter paper by droplet inoculation with C. truncatum isolate CT-21 as described before. Inoculated leaflets were incubated with 12 h photoperiod. The progress of C. truncatum infection was microscopically assessed and infection sites of leaflets were harvested as described above at 3 time points: appressorial penetration phase (16 hai), biotrophic stage (44 hai), and necrotrophic stage (68 hai). These leaflet discs were then frozen in liquid nitrogen until required.
Microscopic evaluation of infection
Leaflet discs from all time points were fixed in a fixation solution (60% methanol, 30% chloroform, 10% acetic acid) until required. Fixed samples were rehydrated with decreasing ethanol gradients (100%, 80%, 70%, and 50% ethanol). Samples were then stained with 0.05% trypan blue (Harleco Parastains, Philadelphia, PA, USA) in distilled water overnight and destained in distilled water. The stained leaves were then mounted in 30% glycerol on glass slides. The developmental stages were examined and photographed using an epifluorescence microscope (Zeiss Axioplan, Jena, Germany). To investigate whether C. truncatum primary hyphae invaginate the host plasma membrane during the biotrophic phase of infection, leaflet discs from the biotrophic phase of infection were plasmolyzed and then stained with vital stain neutral red, and photographed using an epifluorescence microscope.
Semiquantitative RT-PCR
Total RNA from each time point sample (conidia, mycelia, and cell wall treated mycelia [in vitro grown], and appressorial penetration, primary hyphae and secondary hyphae [in planta]) was isolated as described above. The genomic DNA was eliminated using RNase-free amplification grade DNase I (Invitrogen, Carlsbad, USA) from isolated total RNA and stored frozen at -80°C until required. Two micrograms of total RNA was reverse transcribed in a 20 μL reaction volume using 200 U SuperScript reverse transcriptase (Invitrogen, Carlsbad, CA, USA) following the protocol of the supplier. The resulting cDNA was diluted 20 times in sterilized ultrapure water. PCR amplification was performed with one microliter of cDNA from each sample as template using 28 cycles in an Eppendorf Mastercycler (Eppendorf, Hamburg, Germany). The PCR products were separated on 1% agarose gel and stained with ethidium bromide. The gene encoding 60S subunit of ribosmal protein (identified in the library) was used as control. All gene-specific primers used for RT-PCR analysis are listed in the additional file 4.
Binary constructs and agroinfiltration
All primers and plasmids used in this study are described in additional file 5. For in planta expression of selected candidate effectors, including M. oryzae MgSM1, binary PVX constructs were made in vector pGR106 [55]. ORFs with and without SPs were cloned into pCR2.1 (Invitrogen) and sequenced. Confirmed fragments were digested with Cla I and Not I and ligated into pGR106. The constructs were then transformed to A. tumefaciens GV3101 carrying pSoup helper plasmid.
Infiltration assays with 0.3 OD600 of recombinant A. tumefaciens strains were performed on 4-6 weeks-old Nicotiana tobacum plants as described previously [38]. Responses were monitored from 2-10 days after infiltration. For autofluorescence detection, edges of lesions/infiltration zone were observed under UV light (480/40 nm excitation filter; 510 nm barrier) and photographed.
References
Sutton BC: The Coelomycetes. 1980, Wallingford: CAB International Press
Kankanala P, Czymmek K, Valenta B: Roles for rice membrane dynamics and plasmodesmata during biotrophic invasion by the blast fungus. Plant cell. 2007, 19: 706-724. 10.1105/tpc.106.046300.
O'Connell RJ, Uronu AB, Waksman G, Nash C, Keon JPR, Bailey JA: Hemibiotrophic infection of Pisum sativum by Colletotrichum truncatum. Plant Pathol. 1993, 42: 774-783. 10.1111/j.1365-3059.1993.tb01564.x.
Dean RA, Talbot NJ, Ebbole DJ, Farman ML, Mitchell TK, Orbach MJ, Thon M, Kulkarni R, Xu JR, Pan H, et al: The genome sequence of the rice blast fungus Magnaporthe grisea. Nature. 2005, 434: 980-986. 10.1038/nature03449.
Tyler BM, Tripathy S, Zhang X, Dehal P, Jiang RHY, Aerts A, Arredondo FD, Baxter L, Bensasson D, Beynon JL, et al: Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science. 2006, 313: 1261-1266. 10.1126/science.1128796.
Kämper J, Kahmann R, Bölker M, Ma LJ, Brefort T, Saville BJ, Banuett F, Kronstad JW, Gold SE, Müller O, et al: Insights from the genome of the biotrophic fungal plant pathogen Ustilago maydis. Nature. 2006, 444: 97-101. 10.1038/nature05248.
van den Burg HA, Harrison SJ, Joosten M, Vervoort J, de Wit P: Cladosporium fulvum Avr4 protects fungal cell walls against hydrolysis by plant chitinases accumulating during infection. Mol Plant-Microbe Interact. 2006, 19: 1420-1430. 10.1094/MPMI-19-1420.
El Gueddari NE, Rauchhaus U, Moerschbacher BM, Deising HB: Developmentally regulated conversion of surface-exposed chitin to chitosan in cell walls of plant pathogenic fungi. New Phytol. 2002, 156: 103-112. 10.1046/j.1469-8137.2002.00487.x.
Bent AF, Mackey D: Elicitors, Effectors and R Genes. Ann Rev Phytopathol. 2007, 45: 399-436. 10.1146/annurev.phyto.45.062806.094427.
Whisson SC, Boevink PC, Moleleki L, Avrova AO, Morales JG, Gilroy EM, Armstrong MR, Grouffaud S, van West P, Chapman S, Hein I, Toth IK, Pritchard L, Birch PRJ: A translocation signal for delivery of oomycete effector proteins into host plant cells. Nature. 2007, 450: 115-118. 10.1038/nature06203.
Kale SD, Gu B, Capelluto DGS, Dou D, Feldman E, Rumore A, Arredondo FD, Hanlon R, Fudal I, Rouxel T, Lawrence CB, Shan W, Tyler BM: External lipid PI3P mediates entry of eukaryotic pathogen effectors into plant and animal host cells. Cell. 2010, 142: 284-295. 10.1016/j.cell.2010.06.008.
Stephenson SA, Hatfield J, Rusu AG, Maclean DJ, Manners JM: CgDN3: an essential pathogenicity gene of Colletotrichum gloeosporioides necessary to avert a hypersensitive-like response in the host Stylosanthes guianensis. Mol Plant-Microbe Interact. 2000, 13: 929-941. 10.1094/MPMI.2000.13.9.929.
Kleemann J, Takahara H, Stüber K, O'Connell R: Identification of soluble secreted proteins from appressoria of Colletotrichum higginsianum by analysis of expressed sequence tags. Microbiology. 2008, 154: 1204-1217. 10.1099/mic.0.2007/014944-0.
Bowen JK, Mesarich CH, Rees-George J, Cui W, Fitzgerald A, Win J, Plummer KM, Templeton MD: Candidate effector gene identification in the ascomycete fungal phytopathogen Venturia inaequalis by expressed sequence tag analysis. Mol Plant Pathol. 2009, 10 (3): 431-48. 10.1111/j.1364-3703.2009.00543.x.
Krijger JJ, Horbach R, Behr M, Schweizer P, Deising HB, Wirsel SGR: The yeast signal sequence trap identifies secreted proteins of the hemibiotrophic corn pathogen Colletotrichum graminicola. Mol Plant-Microbe Interact. 2008, 21 (10): 1325-1336. 10.1094/MPMI-21-10-1325.
Takahara H, Dolf A, Endl E, O'Connel R: Flow cytometric purification of Colletotrichum higginsianum biotrophic hyphae from Arabidopsis leaves for stage-specific transcriptome analysis. Plant J. 2009, 59: 672-683. 10.1111/j.1365-313X.2009.03896.x.
Mosquera G, Giraldo MC, Khang CH, Coughlan S, Valent B: Interaction transcriptome analysis identifies Magnaporthe oryzae BAS1-4 as biotrophy-associated secreted proteins in rice blast disease. Plant Cell. 2009, 21: 1273-1290. 10.1105/tpc.107.055228.
Dufresne M, Perfect S, Pellier AL, Bailey JA, Langin T: A GAL4-like protein is involved in the switch between biotrophic and necrotrophic phases of the infection process of Collectotrichum lindemuthianum on common bean. Plant Cell. 2000, 12: 1579-1590.
You BJ, Chung KR: Phenotypic characterization of mutants of the citrus pathogen Colletotrichum acutatum defective in a PacC-mediated pH regulatory pathway. FEMS Microbiol Lett. 2007, 277: 107-114. 10.1111/j.1574-6968.2007.00951.x.
Perfect SE, O'Connell RJ, Green EF, Doering-Saad C, Green JR: Expression cloning of a fungal proline-rich glycoprotein specific to the biotrophic interface formed in the Colletotrichum-bean interaction. Plant J. 1998, 15: 273-279. 10.1046/j.1365-313X.1998.00196.x.
Bolton MD, van Esse HP, Vossen JH, de Jonge R, Stergiopoulos I, Stulemeijer IJE, van den Berg GCM, Borrás-Hidalgo O, Dekker HL, de Koster CG, de Wit PJGM, Joosten MHAJ, Thomma BPHJ: The novel Cladosporium fulvum lysin motif effector Ecp6 is a virulence factor with orthologues in other fungal species. Mol Microbiol. 2008, 69: 119-136. 10.1111/j.1365-2958.2008.06270.x.
Bell-Pedersen D, Shinohara ML, Loros JJ, Dunlap JC: Circadian clock-controlled genes isolated from Neurospora crassa are late night- to early morning-specific. Proc Natl Acad Sci USA. 1996, 93: 13096-13101. 10.1073/pnas.93.23.13096.
Marger MD, Saier MH: A major superfamily of transmembrane facilitators that catalyse uniport, symport and antiport. Trends Biochem Sci. 1993, 18: 13-20. 10.1016/0968-0004(93)90081-W.
Corsi AK, Scheckman R: Mechanism of polypeptide translocation into the endoplasmic reticulum. J Biol Chem. 1996, 271: 30299-30302. 10.1074/jbc.271.48.30299.
Pelham HR: Speculations on the functions of the major heat shock and glucose-regulated proteins. Cell. 1986, 46: 959-961. 10.1016/0092-8674(86)90693-8.
Orolaza NP, Lamari L, Ballance GM: Evidence of a host-specific chlorosis toxin from Pyrenophora tritici-repentis, the causal agent of tan spot of wheat. Phytopathology. 1995, 85: 1282-1287. 10.1094/Phyto-85-1282.
Fridovich I: The biology of oxygen radicals. Science. 1978, 201: 875-880. 10.1126/science.210504.
Kromina KA, Ignatov AN, Abdeeva IA: Role of peptidyl-prolyl cis/trans-isomerases in pathologic processes. Biochemistry. 2008, 2 (3): 195-202.
Rep M: Small proteins of plant-pathogenic fungi secreted during host colonization. FEMS Microbiol Lett. 2005, 253: 19-27. 10.1016/j.femsle.2005.09.014.
Templeton MD, Rikkerink EHA, Beever RE: Small, cysteine-rich proteins and recognition in fungal-plant interactions. Mol Plant-Microbe Interact. 1994, 7: 320-325. 10.1094/MPMI-7-0320.
Pazzagli L, Cappugi G, Manao G, Camici G, Santini A, Scala A: Purification, characterization, and amino acid sequence of cerato-platanin, a new phytotoxic protein from Ceratocystis fimbriata f. sp. Platani. J Biol Chem. 1999, 274: 24959-24964. 10.1074/jbc.274.35.24959.
Shaffer PM, Arst HN, Estberg L, Fernando L, Ly T, Sitter M: An asparaginase of Aspergillus nidulans is subject to oxygen repression in addition to nitrogen metabolite repression. Mol Gen Genet. 1988, 212: 337-341. 10.1007/BF00334704.
Bessman MJ, Frick DN, O'Handley SF: The MutT proteins or 'nudix' hydrolases, a family of versatile, widely distributed, 'housecleaning' enzymes. J Biol Chem. 1996, 271: 25059-25062. 10.1074/jbc.271.41.25059.
Wösten HAB: Hydrophobins: multipurpose proteins. Annu Rev Microbiol. 2001, 55: 625-646. 10.1146/annurev.micro.55.1.625.
Seidl V, Marchetti M, Schandl R, Allmaier G, Kubicek CP: Epl1, the major secreted protein of Hypocrea atroviridis on glucose, is a member of a strongly conserved protein family comprising plant defense response elicitors. FEBS J. 2006, 273: 4346-4359. 10.1111/j.1742-4658.2006.05435.x.
Yang Y, Zhang H, Li G, Li W, Wang X, Song F: Ectopic expression of MgSM1, a Cerato-platanin family protein from Magnaporthe grisea, confers broad-spectrum disease resistance in Arabidopsis. Plant Biotech J. 2009, 7 (8): 763-777. 10.1111/j.1467-7652.2009.00442.x.
Abramovitch RB, Kim YJ, Chen SR, Dickman MB, Martin GB: Pseudomonas type III effector AvrPtoB induces plant disease susceptibility by inhibition of host programmed cell death. EMBO J. 2003, 22: 60-69. 10.1093/emboj/cdg006.
Van der Hoorn RAL, Laurent F, Roth R, De Wit PJGM: Agroinfiltration is a versatile tool that facilitates comparative analyses of Avr9/Cf-9-induced and Avr4/Cf-4-induced necrosis. Mol Plant Microbe Interact. 2000, 13: 439-446. 10.1094/MPMI.2000.13.4.439.
Yin C, Chen X, Wang X, Han Q, Kang Z, Hulbert SH: Generation and analysis of expression sequence tags from haustoria of the wheat stripe rust fungus Puccinia striiformis f. sp. Tritici. BMC Genom. 2009, 10: 626-10.1186/1471-2164-10-626.
Kelley BS, Lee SJ, Damasceno CMB, Chakravarthy S, Kim BD, Martin GB, Rose JKC: A secreted effector protein (SNE1) from Phytophthora infestans is a broadly acting suppressor of programmed cell death. Plant J. 2010, 62: 357-366. 10.1111/j.1365-313X.2010.04160.x.
Soanes DM, Richards TA, Talbot NJ: Insights from sequencing fungal and oomycete genomes: what can we learn about plant disease and the evolution of pathogenicity?. Plant Cell. 2007, 19: 3318-3326. 10.1105/tpc.107.056663.
Keon JPR, Byrde RJW, Cooper RM: Some aspects of fungal enzymes that degrade plant cell walls. Fungal Infection of Plants. Edited by: Pegg GF and Ayres PG. 1987, Cambridge: University Press, 133-157.
Martinez-Ruiz A, Martinez del Pozo A, Lacadena J, Mancheno JM, Onaderra M, Gavilanes JG: Characterization of a natural larger form of the antifungal protein (AFP) from Aspergillus giganteus. Biochim Biophys Acta. 1997, 1340: 81-87. 10.1016/S0167-4838(97)00038-1.
Olivieri F, Zanetti ME, Oliva CR, Covarrubias AA, Casalongue CA: Characterization of an extracellular serine protease of Fusarium eumartii and its action on pathogenesis related proteins. Eur J Plant Pathol. 2002, 108: 63-72. 10.1023/A:1013920929965.
Carlile AJ, Bindschedler LV, Bailey AM, Bowyer P, Clarkson JM, Cooper RM: Characterization of SNP1, a cell wall degrading trypsin, produced during infection by Stagonospora nodorum. Mol Plant-Microbe Interact. 2000, 13: 538-550. 10.1094/MPMI.2000.13.5.538.
Eisenhaber B, Wildpaner M, Schultz CJ, Borner GH, Dupree P, Eisenhaber F: Glycosylphosphatidylinositol lipid anchoring of plant proteins. Sensitive prediction from sequence- and genome-wide studies for Arabidopsis and rice. Plant Physiol. 2003, 133: 1691-1701. 10.1104/pp.103.023580.
Jiang RHY: Footprints of evolution: The dynamics of effector genes in the Phytophthora genome. PhD thesis. 2006, Wageningen University, Laboratorium voor Fytopathologie
Jiang RHY, Tyler BM, Whisson SC, Hardham AR, Govers F: Ancient origin of elicitin gene clusters in Phytophthora genomes. Mol Biol Evol. 2006, 23: 338-351.
Perfect SE, Green JR, O'Connell RJ: Surface characteristics of necrotrophic secondary hyphae produced by the bean anthracnose fungus, Colletotrichum lindemuthianum. Eur J Plant Pathol. 2001, 107: 813-819. 10.1023/A:1012473823851.
de Jonge R, Thomma BPHJ: Fungal LysM effectors: extinguishers of host immunity?. Trends Microbiol. 2009, 17: 151-157. 10.1016/j.tim.2009.01.002.
de Jonge R, van Esse HP, Kombrink A, Shinya T, Desaki Y, Bours R, van der Krol S, Shibuya N, Joosten MHAJ, Thomma BPHJ: Conserved fungal LysM effector Ecp6 prevents chitin-triggered immunity in plants. Science. 2010, 329: 953-955. 10.1126/science.1190859.
Eshel D, Miyara I, Ailing T, Dinoor A, Prusky D: pH regulates endoglucanase expression and virulence of Alternaria alternata in persimmon fruits. Mol Plant-Microbe Interact. 2002, 15: 774-779. 10.1094/MPMI.2002.15.8.774.
Wilkins TA, Smart LB: Isolation of RNA from plant tissue. A Laboratory Guide to RNA: Isolation, Analysis, and Synthesis. Edited by: Krieg PA. 1996, New York: Wiley-Liss Inc, 21-42.
Altschul SF, Warren G, Webb M, Eugene WM, David JL: Basic local alignment search tool. J Mol Biol. 1990, 215: 403-10.
Jones L, Hamilton AJ, Voinnet O, Thomas CL, Maule AJ, Baulcombe DC: RNA-DNA interactions and DNA methylation in post-transcriptional gene silencing. Plant 6/24/2011Cell. 1999, 11: 2291-2302.
Acknowledgements and funding
The authors thank Dr. D. Baulcombe for providing pGR106, Dr. G.B. Martin for Cf9/Avr9 and Pto/AvrPto, and Cheryl Cho and Candace Knihniski for their technical assistance. This work was supported by NSERC-CRD and the Saskatchewan Pulse Growers grants to Drs SB, AV and YW.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Authors' contributions
VB constructed cDNA library and conducted EST analysis, expression profiling and functional analysis of candidate effectors, and drafted the manuscript. SB, AV and YW conceived the study, participated in its design and coordination, and helped to draft the manuscript. GS coordinated the sequencing aspects of the project. All authors read and approved the final manuscript.
Electronic supplementary material
12864_2011_3500_MOESM1_ESM.DOC
Additional file 1:Fungal developmental stages. Per cent of in planta fungal developmental stages in excised infected leaflet discs used for cDNA library construction. (DOC 28 KB)
12864_2011_3500_MOESM2_ESM.DOC
Additional file 2:Clone IDs under various contigs. NCBI GeneBank EST database ID to individual ESTs in each contig. (DOC 43 KB)
12864_2011_3500_MOESM3_ESM.DOC
Additional file 3:Sequence alignment of CtCP1 with its orthologs. ClustalW program was used to align C. truncatum CtCP1 with CP proteins present in other fungal species. (DOC 32 KB)
12864_2011_3500_MOESM4_ESM.DOC
Additional file 4:List of primers (F, Forward; R, Reverse). Gene-specific primer sets were used to catalogue the expression profiles of CEAPs and candidate effectors. (DOC 35 KB)
12864_2011_3500_MOESM5_ESM.DOC
Additional file 5:List of primers (F, Forward; R, Reverse). Primers were used to clone candidate effectors in the binary PVX-based expression vector pGR106. (DOC 36 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Bhadauria, V., Banniza, S., Vandenberg, A. et al. EST mining identifies proteins putatively secreted by the anthracnose pathogen Colletotrichum truncatum. BMC Genomics 12, 327 (2011). https://doi.org/10.1186/1471-2164-12-327
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2164-12-327