Abstract
Background
The bacterial genus Listeria contains pathogenic and non-pathogenic species, including the pathogens L. monocytogenes and L. ivanovii, both of which carry homologous virulence gene clusters such as the prfA cluster and clusters of internalin genes. Initial evidence for multiple deletions of the prfA cluster during the evolution of Listeria indicates that this genus provides an interesting model for studying the evolution of virulence and also presents practical challenges with regard to definition of pathogenic strains.
Results
To better understand genome evolution and evolution of virulence characteristics in Listeria, we used a next generation sequencing approach to generate draft genomes for seven strains representing Listeria species or clades for which genome sequences were not available. Comparative analyses of these draft genomes and six publicly available genomes, which together represent the main Listeria species, showed evidence for (i) a pangenome with 2,032 core and 2,918 accessory genes identified to date, (ii) a critical role of gene loss events in transition of Listeria species from facultative pathogen to saprotroph, even though a consistent pattern of gene loss seemed to be absent, and a number of isolates representing non-pathogenic species still carried some virulence associated genes, and (iii) divergence of modern pathogenic and non-pathogenic Listeria species and strains, most likely circa 47 million years ago, from a pathogenic common ancestor that contained key virulence genes.
Conclusions
Genome evolution in Listeria involved limited gene loss and acquisition as supported by (i) a relatively high coverage of the predicted pan-genome by the observed pan-genome, (ii) conserved genome size (between 2.8 and 3.2 Mb), and (iii) a highly syntenic genome. Limited gene loss in Listeria did include loss of virulence associated genes, likely associated with multiple transitions to a saprotrophic lifestyle. The genus Listeria thus provides an example of a group of bacteria that appears to evolve through a loss of virulence rather than acquisition of virulence characteristics. While Listeria includes a number of species-like clades, many of these putative species include clades or strains with atypical virulence associated characteristics. This information will allow for the development of genetic and genomic criteria for pathogenic strains, including development of assays that specifically detect pathogenic Listeria strains.
Similar content being viewed by others
Background
The eight recognized species within the genus Listeria include L. monocytogenes, L. innocua, L. welshimeri, L. seeligeri, L. ivanovii, L. grayi, L. marthii[1] and L. rocourtiae[2], the latter two were described in 2009. L. grayi is only distantly related to the other Listeria species [1, 3] and has been proposed to represent a different genus, Murraya[4]. L. monocytogenes and L. ivanovii are pathogens of warm-blooded hosts. L monocytogenes causes a severe foodborne disease in humans as well as invasive infections in a number of other warm-blooded host species, particularly ruminants. L. ivanovii predominantly causes infections in ruminants, but has also been associated with rare infections in humans [5, 6]; this species is considered to have a narrower host range than L. monocytogenes[7]. Interestingly, each of these two pathogenic Listeria species is closely related to non-pathogenic species; L. monocytogenes is closely related to L. innocua and L. marthii[1], and L. ivanovii is closely related to L. seeligeri[3, 8], which is non-pathogenic even though many isolates contain a homologue of the main Listeria virulence gene cluster.
Genome sequencing efforts for Listeria have, so far, largely focused on L. monocytogenes; as of August 15, 2010, 25 L. monocytogenes genome sequences are publicly accessible in standard sequence databases (GenBank; EMBL). Most of these L. monocytogenes genome sequences represent strains classified into the two most common L. monocytogenes phylogenetic lineages [9] including lineage I (e.g. strains F2365, H7858 [10] ) and lineage II (e.g. strains EGD-e [11], 08-5578 and 08-5923 [12]). The other two L. monocytogenes phylogenetic lineages (III and IV) are only represented by 3 genome sequences (i.e., strains HCC23 [Genbank acc. CP001175], FSL J2-071 [Genbank acc. ARN00000000], and FSL J1-208 [Genbank acc. AARL00000000]). Publicly available genome sequences for other Listeria species include those for L. innocua CLIP11262 [11] and L. welshimeri SLCC5334 [13] as well as recently released the genome sequences for L. seeligeri SLCC3954 [14] and L. grayii DSM 20601 (Genbank acc. ACCR00000000). Knowledge of the genomic content of non-pathogenic relatives of pathogenic species is necessary though to understand the evolution of virulence associated genes and to facilitate identification of putative virulence genes [15].
The main Listeria virulence gene cluster (also known as the prfA virulence cluster or the Listeria pathogenicity island [LiPI]) encodes a number of proteins that are necessary for intracellular survival and motility [16, 17]. Specific functions encoded in this cluster include hemolysin, two phospholipases and a metalloprotease (encoded by hly, plcA, plcB, and mpl), which all contribute to escape from host cell vacuoles, an actin polymerizing protein (encoded by actA), and a global regulator of virulence gene transcription (encoded by prfA). Members of the internalin protein family, which are cell wall anchored or secreted proteins that are characterized by the presence of leucine rich repeats, are also associated with virulence in different Listeria strains. While a considerable number of genes encoding internalins have been reported in pathogenic and non-pathogenic Listeria[11, 18, 19], clear virulence related functions have only been assigned to a few internalins, including inlA and inlB, which encode proteins required for invasion of different cells types, including human intestinal epithelial cells [20], and inlC[21]. A number of atypical Listeria strains and lineages have been reported [22–24], including several putative evolutionary intermediates, which are characterized by unique virulence gene presence/absence patterns. For example, while the non-pathogenic L. innocua is typically non-hemolytic and lacks the prfA cluster, a small number of strains that contain the prfA cluster as well as inlA have been reported [22, 23]. Also, non-hemolytic L. seeligeri strains that lack the prfA cluster have been reported [24]; even though many L. seeligeri contain the prfA cluster, isolates in this species are avirulent in typically studied mammalian hosts [25].
Based on the observations outlined above, we propose that the genus Listeria represents an outstanding model system for studying the evolution of pathogenicity and the transition between pathogenic and saprotrophic lifestyles using a comparative genomics approach. We thus performed genome sequencing of (i) isolates representing Listeria species (except for L. grayi) for which no genome sequences are publicly available and (ii) atypical strains of Listeria species for which genome sequences of typical strains were already available. While L. marthii was included in our genome sequencing efforts, L. rocourtiae was not as this new species was described after completion of the work reported here.
Methods
Selection of isolates for genome sequencing
The isolates sequenced in this study (Table 1) were selected to (i) cover the full phylogenetic diversity of the genus Listeria (except for L. grayii) [9], and to (ii) represent atypical phenotypes (e.g., hemolytic L. innocua, non-hemolytic L. seeligeri) of some non-pathogenic species. L. seeligeri FSL N1-067 was selected as a typical hemolytic strain, while L. seeligeri FSL S4-171 represents an atypical non-hemolytic strain of the same species. L. marthii FSL S4-120 is the type strain of L. marthii and has recently been shown to represent the most closely related species to L. monocytogenes, however it is not pathogenic. L. ivanovii subsp. londoniensis FSL F6-596 (ATCC 49954) represents the type strain of this subspecies and represents the second pathogenic species in Listeria. L. monocytogenes FSL F2-208 represents lineage IIIC, a distinct lineage within L. monocytogenes that has not been sequenced before. L. innocua FSL J1-023 was sequenced because this strain represents a atypical hemolytic L. innocua isolate, while FSL S4-378 was sequenced as an additional typical non-hemolytic strain of L. innocua to further assess intraspecific genomic variation within L. innocua. Hemolytic activity for all strains was previously tested [26].
Genome sequencing and assembly
Genomic DNA was isolated using the UltraClean Microbial DNA Isolation Kit (MO BIO Laboratories, Carlsbad, CA) according to manufacturer's instructions. Listeria genomes were sequenced using the SOLiD™ 3 system (Applied Biosystems, Foster City, CA) following manufacturer's protocols. Mate-paired libraries with approximately 1.5 kb inserts were constructed from 20 μg of genomic DNA, and deposited on one quarter of a flow cell. Twenty-five base reads were obtained from each of the F3 and R3 tags, with 27 million to 57 million reads obtained for each of the genomes. After correcting errors in colorspace reads using a modified version of the spectral alignment tools from the EULER-USR package [27], de novo assembly was performed using the SOLiD™ System de novo Accessory Tools (http://solidsoftwaretools.com/gf/project/denovo/), which employs the Velvet assembly engine [28].
In order to identify likely misassemblies, scaffolds were aligned using MUMmer [29] to the most closely related reference genome available at the time of the analysis: L. monocytogenes scaffolds were aligned to L. monocytogenes F2365, L. innocua and L. marthii scaffolds were aligned to L. innocua Clip11262, and L. ivanovii and L. seeligeri scaffolds were aligned to L. welshimeri SLCC5334. Scaffolds were broken at points where non-contiguous regions of the reference genome were juxtaposed, and then ordered such that they were syntenic with the reference genome. All scaffolds were then concatenated into a single pseudogenome, separated by the sequence NNNNNCACACACTTAATTAATTAAGTGTGTGNNNNN, which puts stop codons in all six reading frames. Scaffolds that did not match the reference genome were concatenated in arbitrary order at the end of the pseudogenome.
The genome sequences of the seven newly sequenced strains have been deposited to GenBank as whole genome shotgun projects (see table 1 for accession numbers).
Genome annotation and whole genome alignments
Concatenated pseudogenome sequences were run through JCVI's prokaryotic annotation pipeline (http://www.jcvi.org/cms/research/projects/annotation-service/), which includes gene finding with Glimmer, Blast-extend-repraze (BER) searches to extend ORF finding beyond premature stop codons, HMM searches against Pfam [30] and TIGRFAM [31], TMHMM searches, SignalP predictions, and automatic annotations from AutoAnnotate. The manual annotation tool Manatee (downloaded from http://manatee.sourceforge.net) was used to manually review the output and aid in genome annotation and gene identification. Whole genome alignments were created in Mauve 2.3.0 [32] using the Progressive Mauve algorithm.
Orthologue analyses
Initially, orthologues found in six publicly available complete Listeria genomes (see Table 1) were identified using BLASTCLUST [33]. This analysis was limited to these six complete genomes to avoid possible problems with (sometimes incomplete) fragmented draft sequences such as the genome sequences available in the Broad Institute L. monocytogenes database (http://www.broadinstitute.org/annotation/genome/listeria_group/MultiHome.html), at the time of this analysis. Only ORFs having at least 225 nt (encoding 75 amino acids [aa]) were considered in this analysis as smaller ORFs were not annotated consistently among the genomes. A set of 3,668 unique ORFs, found among these six complete genomes by requiring 75% aa identity over at least 80%, was used to identify orthologues of these ORFs in the new genome assemblies (using TBLASTN queries with a single member of each cluster and requiring at least 65% aa identity). The 75% identity threshold was selected empirically after running blastclust with a range of percent identity values: requiring more than 75% identity resulted in too many orthologues being split into multiple clusters, and allowing lower values resulted in too many clusters containing paralogues. When an orthologue in a draft assembly was split into two or more fragments, these were considered to be a single match.
ORFs identified only in the seven newly sequenced genomes by the JCVI automated sequence annotation pipeline were split at any run of five or more ambiguous amino acids ('X'), which resulted from in-frame strings of fifteen or more 'N' between contigs in the assemblies. After splitting, all ORFs and ORF fragments with length less than 50 aa were removed from the set. Amino acid sequences of the remaining ORFs were screened against the nucleotide sequences of the six previously available finished Listeria genomes using TBLASTN with an identity threshold of 65%, and those without any hits were identified. These novel Listeria ORFs were screened against all seven draft genome assemblies in order to determine their distribution across the set of strains.
Core and pan genome analysis
The mixture model method of Snipen et al. [34] was used to estimate the number of genes in (i) the core genome (i.e., the set of genes found within every strain within the genus Listeria[34, 35]) and in (ii) the pan-genome (i.e., the core genome plus the dispensable or accessory genome, which is defined as the genes that are found in some, but not all, strains including genes that are unique to a single strain [34, 35]). To compare these estimates to another group of Gram-positive bacteria that includes closely related pathogens and non-pathogens, the same method was used to estimate the core and pan-genome of the Bacillus cereus group, a group that can be considered a single species from taxonomic point of view [36]. Only the chromosome sequences of five B. anthracis, nine B. cereus, two B. thuringiensis, and one B. weihenstephanensis strain were used for this analysis.
Cumulative pan-genome size plots for Listeria were calculated by selecting strains without replacement in random order 500 times, and then calculating the mean pan-genome size at each sampling point. Blast2GO [37] was used to perform a functional annotation of the genes found in the core and the pan-genome.
Phylogenetic analysis
Phylogenetic trees of the Listeria isolates were constructed using sequences for 100 genes in the core genome of the species studied; these genes were randomly selected form all genes that were previously shown to have no evidence for either positive selection or recombination [38] as these two processes make it problematic to infer the true organismal phylogeny. Phylogenetic trees were inferred using neighbor joining, maximum parsimony and minimum evolution phylogenetic reconstruction methods available in the MEGA package version 4.1 [39]. A Bayesian analysis was performed using MrBayes 3.12 [40] and the GTR +I+G model of nucleotide evolution. A phylogenetic reconstruction using presence or absence data of genes in the pan genome was performed using the maximum parsimony method in PAUP* version 4.010b [41].
To infer the time to the most recent common ancestor of Listeria and to infer the approximate age of the individual Listeria species, a Bayesian molecular clock analysis was performed in BEAST version 1.5.2 [42] based on the concatenated 100 core genes. One strain from each species or lineage was included in the analysis. The molecular clock analysis was performed using a GTR +I+G nucleotide substitution model, and a relaxed clock model to account for variation in substitution rates. Tracer version 1.4.1 was used to assess the proper burn-in and sampling of the model parameters. We used a mutation rate of 4.5 × 10-9 per site per year, as suggested by Ochman et al. [43] to calibrate the tree.
Evolutionary analysis of internalins
As at least some internalins have been shown to be important virulence factors in Listeria (see the Background section for more details) we decided to employ a phylogenetic approach to infer the homology and evolution of genes in the internalin gene family. This was specifically important because the JCVI automated annotation pipeline annotated most internalin genes inconsistently as either (i) internalin A, (ii) leucine-rich repeat containing protein, or (iii) cell wall anchor domain-containing protein. Amino acid sequences of ORFs predicted to encode internalins or leucine-rich repeat proteins were extracted from the annotated genomes; the newly sequenced genomes were aligned to publicly available, well-annotated genomes (e.g., EGD-e and F2365) to identify internalin genes that were missed in the initial annotation. After alignment using the EINSI strategy in the MAFFT alignment package [44], internalin aa sequences were used for phylogenetic reconstruction using a maximum parsimony heuristic search in PAUP* 4.010b [41]. Gaps identified in internalin genes were either closed by PCR and Sanger sequencing or by reassembly of the raw SOLiD™ system reads using improved versions of the de novo assembly tools. Internalins found in L. monocytogenes CLIP80459 were not included in this analysis because the majority of the internalins (26 out of 28) in this strain are identical (at the aa level) to those found in L. monocytogenes F2365.
Presence of virulence-associated genes
The presence, in the 13 genome sequences analyzed, of 78 putative virulence-associated genes previously reported by Camejo et al. [45] was assessed using whole genome alignments produced with Mauve 2.3.0. The PVclust R package [46] was used to perform a cluster analysis based on the presence/absence data for genes that showed variable presence/absence.
Horizontal gene transfer and mobile elements
SIGI-HMM [47] was used to infer if genes in the genomes were acquired through horizontal gene transfer (HGT); this program uses a codon usage based hidden Markov model to infer if genes are of alien origin (i.e., have been introduced into the genome from a divergent gene pool by HGT). Genes with more than 10% ambiguous sites were excluded from the analyses. The detection sensitivity of SIGI-HMM was set to 0.95. Prophages and prophage derived regions were identified using the online version of Prophinder [48]. The Prophinder algorithm searches for regions that are dense in phage-like proteins, using a BLASTP search [33] against all phage proteins in the ACLAME database [49]. Transposons and plasmid related genes were identified with SIGI-HMM and through examination of the initial genome annotations for transposon- or plasmid-related genes.
Identification of Restriction and Modification and CRISPR systems
Bacterial defense systems against mobile elements/foreign DNA implicated in the reduction of horizontal gene transfer include (i) restriction-modification (R-M) systems [50] and (ii) clustered, regularly interspaced, short palindromic repeat (CRISPR) systems [51]. R-M systems were identified by screening the initial annotation for genes involved in R-M systems. CRISPRfinder [52] was used to find CRISPR regions. Because the CRISPR regions consist of highly repetitive regions, which may prove problematic for short read based de novo assembly methods, the annotated genomes were also searched for CRISPR associated genes (Cas genes). When the presence of the actual CRISPR region was ambiguous, the presence of Cas genes was considered evidence for a functional CRISPR system.
Caco-2 invasion assays
The ability to invade human intestinal epithelial cells, a phenotype associated with the presence of inlA, was tested for selected strains; the L. marthii strain was not tested as its invasiveness has previously been reported [1]. The invasion assays with the human intestinal epithelial cell line Caco-2 were performed as previously described [53]. L. monocytogenes 10403S and isogenic ΔinlA (FSL K4-006) strain were included as controls in each invasion assay.
Confirmation absence virulence associated genes in non-pathogenic strains
To confirm the absence of critical virulence associated genes (i.e. inlA, inlB, inlC and the prfA cluster) in genomes where these genes were not found, we resequenced the regions where these genes are found in pathogenic strains. The absence of inlC in these non-pathogenic strains was further confirmed by PCR with degenerate primers designed to amplify inlC in L. monocytogenes and L. ivanovii. Primer sequences and additional information can be found in additional file 1.
Results
De novo assembly of short sequence reads yields high quality draft genomes for selected Listeria species
The genomes of eight Listeria strains were sequenced, using mate-paired libraries, on the SOLiD™ 3 System. The previously sequenced L. monocytogenes strain F2365 (our strain ID FSL R2-574) was included in this set to evaluate assembly of SOLiD™ system reads. Mapping of FSL R2-574 SOLiD™ system reads to the F2365 reference genome resulted in 200× median unique coverage depth (Figure 1), with unique coverage gaps only in multicopy loci (e.g., rRNA genes). The corona_lite SNP calling tool identified 21 putative SNPs and 4 ambiguities (see additional file 2 for more information); 19 of these putative SNPs appeared to be legitimate based on coverage depth. PCR amplification and Sanger sequencing confirmed that all 21 putative SNPs represent real differences between the published genome sequence of F2365 and our isolate of this strain (FSL R2-574). The four ambiguities were found to be due to single base pair insertions or deletions in FSL R2-574 relative to the F2365 genome. Three additional mutational differences (two adjacent SNPs and a single base pair insertion) were found in FSL R2-574 during Sanger sequencing-based confirmation of the SNPs identified initially. Examination of the original trace files for the F2365 genome indicated that 24 of these 28 overall differences likely represent sequence errors in the original F2365 sequence (additional file 2). However, four SNPs (see additional file 2) appear to represent real differences between the F2365 and FSL R2-574 genomes, which most likely arose during laboratory passage, as suggested previously in both L. monocytogenes[54] and Bacillus anthracis[55].
Coverage of F2365 genome by R2-574 SOLiD™ system reads. Depth of coverage of uniquely placed reads was plotted along the length of the L. monocytogenes F2365 chromosome. Gray dots indicate coverage at each base and the red line indicates the moving average with a window size of 1000. Uncovered gaps represent non-unique sequences, including the six rRNA operons.
Most of the FSL R2-574 genome was encompassed in eight large scaffolds with the largest scaffold over 1.4 Mb. Alignment of this assembly to the F2365 reference genome indicated that 98.09% of the genome was covered with identity of 99.64%, and fewer than ten misassemblies (i.e., juxtaposition in scaffolds of non-contiguous regions of the genome) were observed, indicating a high quality draft genome for FSL R2-574, according to the definition of Chain et al. [56].
Assembly of the SOLiD™ system reads resulted in 785 to 2,551 contigs per genome (Table 2); the sum of the contig lengths ranged from 2.8 to 3.2 Mb, which is comparable to genome sizes of previously sequenced Listeria genomes (Table 1). When non-contiguous genomic regions were found to be juxtaposed in the assembly, the scaffold was broken and reordered to correspond with the reference genome order. The number of potential misassemblies due to illegitimate scaffolding of contigs ranged from 68 (L. seeligeri FSL S4-171) to 500 (L. monocytogenes FSL F2-208) with a median of 152. Predicted ORF counts (based on JCVI annotations and orthology analyses) ranged from 2,724 for L. marthii FSL S4-120 to 3,017 for L. seeligeri FSL N1-067 (Table 2 and additional file 3).
The Listeria pan-genome is estimated to contain approximately 6,500 genes, including about 2,000 core genes
A total of 4,950 orthologous genes were found among the 13 genome sequences analyzed here, including the 7 new genome sequences generated. Based on these data, a mixture model approach [34] estimated the size of the actual Listeria pan-genome as 6,494 genes, suggesting that over 1,500 Listeria genes remain to be discovered by further sequencing (Figure 2a). According to this mixture model approach, the Listeria pan-genome best fitted a model with four components including (i) a component of 31% of the genes with a detection probability of 1.0 (the core-genome), (ii) a component of 7% of the genes with a detection probability of 0.82, (iii) a component of 10% of the genes with a detection probability of 0.33, and (iv) a component of 52% of the genes with a detection probability of 0.06 (Figure 2b). The lower Bayesian information criterion (BIC) of the four-component model (17,783) versus that of the three-component model (18,393) indicates a better fit. A relatively large part of the estimated pan-genome (76.2%) was covered by the actual pan-genome inferred from the 13 Listeria genome sequences. In contrast, only 42% of the estimated pan-genome was covered by the actual pan-genome inferred from the 17 genome-sequences of the B. cereus group.
Cumulative size and composition of the Listeria pan-genome. (a) Cumulative pan-genome size plots were calculated by selecting strains without replacement in random order 500 times, and then calculating the mean pan-genome size at each sampling point (solid red line). Error bars indicate one standard deviation from the mean. Estimated pan-genome size from mixture model analysis is indicated as a dotted cyan line. (b) The graphical display of the mixture model represents the four components of the pan-genome as rectangles, including (i) a component of 31% of the genes with a detection probability of 1.0 (blue: the core-genome), (ii) a component representing 7% of the genes with a detection probability of 0.82 (teal), (iii) a component of 10% of the genes with a detection probability of 0.33 (yellow), and (iv) a component of 52% of the genes with a detection probability of 0.06 (rare genes: orange).
The observed core genome shared by all 13 Listeria strains comprises 2,032 genes (shown as genes found in EGD-e and all other genomes in Figure 3), while the estimated size of the core genome is 1,994 genes, indicating that the core genome as defined by this study will change very little as more genomes are sequenced.
Comparative genome content of 13 Listeria chromosomes and L. innocua plasmid pLI100. The outermost circle indicates the source of each gene in the pan-genome with each gene represented by a constant width wedge. Starting at the top of the figure (0 Mb) and moving clockwise, all EGD-e genes are arranged in chromosomal order. Continuing clockwise, all genes not present in EGD-e are grouped by strain (as indicated by segment labels). Genes in the F2365 segment are present in F2365, but absent from EGD-e, and genes in the Clip81459 segment are present in Clip81459, but absent from F2365 and EGD-e, and so on. In this way, each gene is represented only once in the diagram. Gene order in all segments except EGD-e is monotonically increasing, but discontinuous, since shared genes may be represented in other segments. Internal circles indicate gene presence (solid color) or absence (unfilled) of each gene in each of the 13 strains examined. Circles from outer to inner are in the same order as strains on the outer circle, starting with EGD-e, followed by F2365, etc. L. monocytogenes strains are in blue; L. marthii is in green; L. innocua strains are in gold; L. welshimeri is in orange; L. seeligeri strains are in red; L. ivanovii subsp. londoniensis is in purple. The location, in the EGD-e genome, of the prfA virulence cluster, conjugative transposon tn916 and prophage A118 are specifically indicated. This figure was created using the Circos software [85].
While the number of accessory genes observed so far is 2,918, the estimated total number of accessory genes for the genus Listeria (except L. grayi) is 4,500. Only a small proportion of ORFs in the different Listeria genomes was predicted to be introduced by HGT (2.0 to 6.4%; see Table 3) and only one of these ORFs, a collagen binding protein in L. monocytogenes FSL F2-208 is potentially associated with virulence. The majority of the genes found in the core genome are involved in metabolic processes (nucleobase, nucleoside, nucleotide and nucleic acid metabolic processes [17% of GO hits], cellular macromolecule metabolic processes (14% of GO hits) and protein metabolic processes [10% of GO hits]) and transport (13% of GO hits), which is congruent with the general notion that the core genome contains genes that are essential for the survival of the organism. Genes involved in metabolic processes and transport also dominate the accessory genome (nucleobase, nucleoside, nucleotide and nucleic acid metabolic processes [21% of GO hits], cellular macromolecule metabolic processes (20% of GO hits) and transport [13% of GO hits]), which can be explained by the fact that species in Listeria have a primarily saprotrophic lifestyle and genes in the accessory genome are putatively involved in the metabolism of specific carbon sources. A large part of the accessory genome (35% of the genes), however, cannot be classified according to the Gene Ontology or is without any significant Blast hits to proteins currently in Genbank. Among these unclassified genes in the accessory genome are hypothetical proteins, proteins involved in phage resistance and prophage associated genes.
L. seeligeri genome characteristics
A total of 3,017 and 2,820 ORFs were identified in L. seeligeri FSL N1-067 and FSL S4-171, respectively (Table 2); these ORF counts are considerably higher than the 2,710 ORFs recently reported for L. seeligeri SLCC3954 [14]. In addition to 88 L. seeligeri specific ORFs (i.e., ORFs only found in both L. seeligeri FSL N1-067 and FSL S4-171), which included three ORFs encoding specific internalin-like genes, we also identified strain specific ORFs (Table 3), including seven and three genes that encode putative internalins (in FSL S4-171 and FSL N1-067, respectively). Overall, 15 and 17 internalin genes were found in L. seeligeri FSL N1-067 and FSL S4-171, respectively (Table 3); by comparison 16 internalin-like genes were reported for L. seeligeri SLCC3954 [14]. The genomic region harboring inlAB in other Listeria is completely absent from the L. seeligeri genomes (see additional file 4). The inlGHE region is absent from the FSL N1-067 genome; in FSL S4-171 this region contains ABC-transporter encoding ORFs (see additional file 5). Consistent with the absence of inlA, L. seeligeri FSL N1-067 was non-invasive in the Caco-2 cell invasion assay, with invasion efficiencies similar to the L. monocytogenes 10403S inlA null mutant (Figure 4).
Invasion efficiencies in Caco-2 cells of Listeria strains. The strains tested are shown on the x-axis and include L. seeligeri FSL N1-067, L. innocua FSL S4-378 (non-hemolytic), L. monocytogenes 10403S ΔinlA (FSL K4-006), L. innocua FSL J1-023 (hemolytic) and L. ivanovii subsp. londoniensis FSL F6-596 (ATCC 49954). Invasion efficiency (the number of recovered cells/number of cells used for inoculation) was normalized to the invasion efficiency obtained for L. monocytogenes 10403S, which was set as 100%, and was included as a control strain in each essay. Three independent invasion assays were performed for each strain tested.
The L. seeligeri FSL N1-067 prfA cluster (additional file 6) is very similar to the prfA cluster previously described [3] for L. seeligeri with the exception that instead of a duplication of plcB, two short open reading frames (encoding two proteins of 51 and 61 aa), with 62% and 41% similarity to parts of a L. seeligeri plcB sequence are found (GenBank accession AAR97365). The non-hemolytic strain FS S4-171 is characterized by the absence of the prfA cluster as also confirmed by PCR (additional file 6); however, the two ORFs (ORFX and ORFI) reported to be lost in the non-hemolytic isolates analyzed by Volokhov et al. [24] are present in the remains of the FSL S4-171 prfA cluster (additional file 6).
Evidence for the presence of a plasmid was found for the hemolytic L. seeligeri FSL N1-067; the initial annotation identified six ORFs that were annotated as two resolvases, a replication associated protein and a LtrC-like conjugative protein. BLAST searches of these plasmid-related and adjacent proteins revealed a high identity to proteins encoded on the L. innocua CLIP11262 plasmid pLI100. The sequence of this plasmid was used as a query to identify more plasmid-derived contigs in L. seeligeri FSL N1-067. P-RAST annotation [57] of these contigs and comparison to the pLI100 plasmid identified plasmid related contigs that encoded 63 putative ORFs, including 44 ORFs with homology to genes found in pLI100 (28 of these ORFs with aa identity to pLI100 ORFs of ≥ 95%). These contigs totaled 60,916 bp, representing an estimate of the minimum plasmid size (use of pLI100 as a query may not have identified non-homologous plasmid regions). Some of the predicted genes on this plasmid are involved in cadmium, arsenic and multi-drug resistance, consistent with previous reports of plasmid-associated cadmium and arsenic resistance genes in L. monocytogenes[58]. Two different R-M systems were found in the genome of L. seeligeri FSL N1-067; a MjaXIP specific Type I system and an EcoPI specific Type III system. Only one Type I restriction modification system was found in the non-hemolytic L. seeligeri strain FSL S4-171. A CRISPR system was identified in FSL N1-067, but not in FSL S4-171 (Table 3), suggesting differences in phage resistance between the two strains.
L. ivanovii subsp. londoniensis genome characteristics
A total of 2,919 ORFs were identified in the L. ivanovii subsp. londoniensis genome, which contains one prophage and two monocin-like regions (Table 3). Presence of a functional CRISPR system was inferred from the presence of Cas genes. The L. ivanovii genome included 20 genes that putatively encode internalins (Table 3); the inlAB region (additional file 4) contains seven ORFs, including five that encode internalins (i.e., one inlA and two inlB homologues, one internalin A-like protein and an internalin with distant homology to lin2724, an internalin found in L. innocua CLIP11262). Consistent with the presence of an inlA homologue, L. ivanovii subsp. londoniensis FSL F6-596 showed Caco-2 cell invasion efficiency comparable to L. monocytogenes 10403S (Figure 4).
L. innocua genome characteristics
A total of 2,855 and 2,737 ORFs were identified in the genome sequences for the non-hemolytic L. innocua FSL S4-278 and the hemolytic L. innocua FSL J1-023. Only three genes were exclusively found in all three L. innocua genomes (i.e., the two genomes sequenced here and CLIP11262); lin0464, which encodes a putative transcriptional regulator and lin1452 and lin2741, which both encode hypothetical proteins. Noteworthy among the strain specific ORFs (Table 3) are three ORFs encoding genes involved in cobalt transport systems in strain FSL J1-023. No prophage regions were identified in the genome of the hemolytic L. innocua strain FSL J1-023, while the non-hemolytic strains CLIP11262 and FSL S4-378 contain five and four prophage regions, respectively. The number of internalin genes in the L. innocua genomes ranged from 17 to 20 (Table 3). The inlAB region of the hemolytic L. innocua only contains a homologue of inlA; inlB is absent as previously reported [23]. Consistent with these findings, the hemolytic L. innocua FSL J1-023, which contains inlA, shows average Caco-2 invasion efficiencies comparable to L. monocytogenes 10403S (Figure 4), while the non-hemolytic L. innocua FSL S4-378 was non-invasive, with invasion efficiencies similar to those for the L. monocytogenes 10403S inlA null mutant (Figure 4).
Modification and restriction systems were present in all three L. innocua genomes (Table 3). L. innocua CLIP11262 and FSL S4-378 harbor a type I R-M system, while the hemolytic L. innocua FSL J1-023 has two type II R-M systems, a Sau3AI specific system and an EcoRV specific system (which is unique to this strain). CRISPR systems are present in the genomes of CLIP11262 and FSL S4-378, but were not found in FSL J1-023.
L. marthii genome characteristics
A total of 2,724 ORFs were identified in the L. marthii FSL S4-120 genome, including 74 ORFs exclusively found in this strain (Table 3). Prophinder found no evidence for the presence of prophages, however one monocin region was detected. Among the genomes examined here, the L. marthii genome has the highest percentage (6.4%) of ORFs that are introduced through HGT (Table 3). One of the regions introduced by HGT in L. marthii is a genomic island that encodes for part of a lantibiotic biosynthesis gene cluster. We also identified 23 genes that were only found in the genomes of L. marthii and the L. monocytogenes lineage IIIC strain FSL F2-208; BLAST searches against GenBank revealed that the eight of these genes were also found in other L. monocytogenes lineage III and IV genomes (2 and 6 genes in FSL J2-072 and FSL J1-208, respectively). While 19 internalin-like genes were found in the L. marthii genome, the inlAB region did not contain any internalin-like genes as confirmed by PCR and the inlGHE region contained one ORF encoding an ABC transporter and a homolog of inlC2 (additional file 5). The L. marthii genome contains a type I R-M system and a CRISPR system.
L. monocytogenes genome characteristics
A total of 2,910 ORFs were identified in L. monocytogenes lineage IIIC strain FSL F2-208 (Table 3), which is similar to the number of ORFs identified in other L. monocytogenes genomes. Only four genes were shared between all L. monocytogenes genomes but not found in the genomes of any of the other species; LMOf2365_0100 (a MerR transcriptional regulator), LMOf2365_0101 (an aldo/keto reductase family oxidoreductase), LMOf2365_0477 (a hypothetical protein) and LMOf2365_0769 (a DNA binding protein). While the number of prophages identified ranged from zero to three (Table 3), all L. monocytogenes genomes contained one monocin region, except HCC23, which seems to lack a monocin region. The chromosome of L. monocytogenes FSL F2-208 contains a region with high similarity to a putative conjugative element CTn1 found in Clostridium difficile[59]; this region contains one ORF encoding a putative virulence factor, a collagen adhesion protein. The number of internalin-like genes in the L. monocytogenes genomes ranged from 18 to 28 (Table 3). The inlAB region is highly variable, however all genomes examined here contain ORFs with homology to inlA and inlB (additional file 4). The inlGHE region is completely absent from the HCC23 genome; in FSL F2-208, this region seems to only contain an inlC2 homolog (additional file 5). All L. monocytogenes genomes, with the exception of the EGD-e genome, contain R-M systems (see Table 3).
Phylogenetic analyses identify sister groups containing pathogenic and non-pathogenic Listeria species
Alignment of the 100 concatenated genes that were used for phylogenetic reconstruction comprised 90,215 nucleotide sites, including 27,945 variable sites; the average pairwise nucleotide identity based on the 100 genes was 84.8%. All methods except maximum parsimony yielded similar topologies, placing L. seeligeri and L. ivanovii in one well-supported clade (100% bootstrap, 100% posterior probability) and L. welshimeri basal to another well supported clade containing L. innocua, L. marthii and L. monocytogenes (see Figure 5, the left tree shows the phylogeny resulting from the Bayesian analysis). Within the L. innocua/L. marthii/L. monocytogenes clade, L. innocua is the most divergent species, while L. marthii forms a sister group to L. monocytogenes. All phylogenetic relationships within this clade are well supported (bootstrap support > 98%, 100% posterior probability). The maximum parsimony tree differs from the other trees by its placement of L. welshimeri within the L. seeligeri/L. ivanovii clade. While neighbor joining, maximum likelihood, minimum evolution and parsimony methods do not find any significant bootstrap support for the placement of L. welshimeri in the phylogeny, the Bayesian analysis supports placement of L. welshimeri as a basal taxon to the L. innocua/L. marthii/L. monocytogenes clade with a highly significant posterior probability (100%).
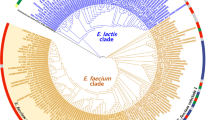
Comparison of phylogenetic trees based on Listeria core gene sequences and genomic gene content. The 100 gene sequence tree (left) was inferred using Bayesian phylogenetic inference (MrBayes v. 3.12) and the values above the branches are the posterior probabilities. The gene content tree based on the presence/absence of 4950 orthologous genes (right) was inferred using maximum parsimony and the values above the branches are bootstrap values based on 1000 bootstrap replicates. Pathogenic strains are colored red, while non-pathogenic strains are black.
A maximum parsimony phylogeny based on the presence and absence of orthologous genes yielded in a single tree (4,805 steps, consistency index 0.608, rescaled consistency index 0.446; Figure 5, right tree) with topology similar to the tree based on Bayesian, minimum evolution, maximum likelihood and neighbor joining phylogenetic inference of the 100 gene sequence dataset. The only differences are the placement of L. monocytogenes FSL F2-208 and L. innocua FSL J1-023. In the gene content-based phylogeny, the hemolytic L. innocua FSL J1-023 strain is placed is outside of the clade containing the two non-hemolytic L. innocua strains. L. monocytogenes FSL F2-208 is placed in a well-supported (97% bootstrap support) clade together with L. marthii. This placement can be mainly attributed to 23 genes that were only found in the L. monocytogenes FSL F2-208 genome and the L. marthii genome (see section 'L. marthii genome characteristics' for more information).
A molecular clock analysis of the 100-gene phylogeny, calibrated with the mutation rate proposed by Ochman et al. (Figure 6), places the time of most recent common ancestor (MRCA) of Listeria (excluding L. grayi) at 47 million years ago (mya) with a 95% highest probability density (HPD) of 58 mya to 39 mya. The time of divergence of L. welshimeri from the MRCA of L. innocua, L. marthii and L. monocytogenes was estimated at 33 mya (95% HPD: 40-27 mya). L. innocua is estimated to have diverged from the MRCA of L. monocytogenes and L. marthii at 29 mya (95% HPD: 35-24 mya). L. marthii and L. monocytogenes were estimated to have diverged from each other around the same time as the divergence of L. seeligeri and L. ivanovii (21 and 20 mya, respectively). Because the mutation rate proposed by Ochman et al. [43] is based on the divergence of Escherichia coli and Salmonella enterica, it may not be applicable to other bacteria. We therefore used the sequence divergence of 16S rRNA between L. monocytogenes and L. ivanovii to infer the putative age of the MRCA of Listeria. The divergence between L. monocytogenes and L. ivanovii is 0.9 to 1.2% (calculated based on full length 16S rDNA data previously reported [1]), which would translate to dating the MRCA at 45 to 60 mya, given the universal 16S rRNA divergence rate of 1% per 50 million years proposed by Ochmann and Wilson [60].
Maximum clade credibility tree summarizing the results of the Bayesian molecular clock analysis of the 100 concatenated core genome genes. The timeline indicates the age of the nodes when a mutation rate of 4.5 × 10-9 per year/site was used to calibrate the tree. One strain from each species or lineage was included in the analysis; L. monocytogenes lineages I (F2365), II (EGD-e) and IIIC (FSL F2-208), L. marthii (FSL S4-120), L. innocua (CLIP11262), L. welshimeri (SLCC5334), L. seeligeri (FSL N1-067), and L. ivanovii subsp. londoniensis (FSL F6-596). Values above the branches indicate posterior probability values, blue horizontal bars on the nodes show the 95% highest probability density of the inferred age of the nodes. The posterior probability of the individual trees and 95% highest probability density of the divergence time were based on 9,000,000 post burn-in generations of a 10,000,000 generation run. The values on the time lines represent ages as million years before present. Labels of pathogenic strains have been colored red.
While non-pathogenic Listeria have fewer virulence associated genes, gene presence patterns for these genes identify only internalin C as a gene that differentiates pathogenic and non-pathogenic Listeria
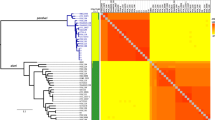
Out of 78 putative virulence-associated genes previously reported [45] (see additional file 7), 56 were found in all 13 genomes analyzed. Only 22 of these genes (not including the genes in the prfA cluster) were variable in their presence between genomes (Table 4), including three putative virulence-associated genes that were found solely in the L. monocytogenes EGD-e genome. Using a hierarchical cluster analysis based on the presence or absence of the 22 variable genes, the strains were subdivided into two clusters, including (i) a cluster composed of L. ivanovii subsp. londoniensis, all L. monocytogenes strains and the hemolytic L. innocua strain and (ii) a cluster composed of only non-pathogenic strains (the non-hemolytic L. innocua strains, L. seeligeri, L. marthii and L. welshimeri).
Other than the prfA virulence cluster, the only gene that is absent from strains in the non-pathogen cluster but present in the first cluster is inlA, encoding internalin A, a key determinant of host intestinal cell invasion. The inlC gene, encoding internalin C, is the only gene that differentiates pathogens (i.e., L. monocytogenes EGD-e, L. monocytogenes F2365, L. monocytogenes FSL F2-208 and L. ivanovii subsp. londoniensis) from non-pathogens (all other strains). There is a significant (p < 0.05, Wilcoxon test) difference between pathogenic and non-pathogenic strains in the number of variable virulence-associated genes, with 10 to 22 genes present in pathogenic strains and 5 to 12 genes present in non-pathogenic strains.
All Listeria strains contain multiple internalin-encoding genes with 16 internalin genes present in both main clades of the genus Listeria
A total of 252 putative internalin genes were identified in the genomes examined here; the number of internalin genes ranged from 9 internalins in L. welshimeri SLCC5334 to 28 internalins in L. monocytogenes CLIP81459 (Table 3). The number of internalin genes in the non-pathogenic strains (Table 1) was significantly lower as compared to the number in pathogenic strains (p < 0.05, Wilcoxon test). Within the internalin gene tree (additional file 8), three main clades can be recognized, including (i) a clade containing inlA, inlB, inlC, inlF, internalin genes found in the inlGHE region and related internalin genes, (ii) a clade containing inlI and inlJ, and (iii) a clade of genes encoding mainly uncharacterized internalin-like proteins. None of these three clades is supported by a significant posterior probability. We also identified 16 clades (marked in red in additional file 8), all with a significant posterior probability (>95%), that each contain genes from isolates of both the L. monocytogenes/L. marthii/L. innocua clade (clade A, Figure 5) as well as the L. seeligeri/L. ivanovii clade (clade B, Figure 5), supporting a MRCA that contained these internalin genes. Although some of these 16 internalin gene clades (e.g., the inlA clade) contained only a single sequence for one of the two Listeria clades, the internalin genes found in L. ivanovii and L. seeligeri (i.e., clade B) are generally highly divergent from their clade A homologues (see additional file 8), which supports presence of these genes in the MRCA as opposed to introduction by horizontal gene transfer.
Discussion
High quality draft genomes can be obtained through de novo assembly of short read sequences
The emergence and maturation of next generation sequencing (NGS) technology, driven in large part by efforts to develop approaches that allow for completion of a human genome for $1,000 [61], has made possible rapid, inexpensive, and high-throughput microbial whole-genome analysis, which promises to improve our understanding of bacterial pathogenesis, and our ability to detect and control infectious diseases. Our data show that NGS data facilitate de novo assembly and analyses of bacterial genomes. Although these draft genomes can have up to thousands of sequence gaps, the quality of the assembly is sufficient for automated annotation, and subsequent comparative genomics analyses, particularly when studying a conserved group of organisms such as the Listeria species examined here. Thus, NGS represents a feasible approach for rapid and comprehensive pathogen identification, subtyping, source tracking, and surveillance [62], and has the potential to be developed, in the long term, into routine diagnostic applications. The utility of draft genomes for identification of candidate vaccine targets has also been recently demonstrated [63].
The genus Listeria sensu stricto has a pan-genome characterized by limited introduction of new genetic material with 2,032 core and 2,918 accessory genes identified to date
Our data show that the members of the genus Listeria have a highly conserved genome with limited acquisition, from other gene pools, of homologous and non-homologous genes, even though horizontal transfer of homologous genes within and between Listeria species has clearly been shown to occur [38, 64]. Although the pangenome of the genus Listeria is not closed, there seems to be very limited on-going introduction of new genetic material from external gene pools (i.e., other genera). Data supporting this limited introduction of new genetic material into the pangenome include (i) the observation that the core and accessory genes identified among the 13 genomes analyzed represent a large proportion (i.e., 76.2%) of the predicted pan-genome, (ii) the similarity in size of observed core genome (2,032 genes) and predicted core genome (1,994 genes), suggesting limited gene loss and deletion, (iii) highly conserved estimated genome size (from 2.8 to 3.2 Mb), (iv) a relatively small fraction (4% on average) of genes that have atypical codon usage, and (v) a small number of prophages and transposons. The fact that most of these prophages have a codon usage pattern that is similar to their host indicates that they have co-evolved with their Listeria hosts [65]. A Listeria pan-genome characterized by limited introduction of genetic material is also supported by the observation that pan-genome coverage for the genus Listeria (except L. grayii) is higher than the pan-genome coverage estimates for most bacterial species, which range from 30% (for Escherichia coli, based on genomes of 22 strains) to 73% (for Francisella tularensis, based on genomes of 7 strains) [34]. A pan-genome coverage estimate performed for the Bacillus cereus group, a group of closely related pathogenic and non-pathogenic Gram-positive species, revealed a coverage of 42%, indicating pan-genome coverage of Listeria, is also high compared to Gram-positive organisms. The Bacillus cereus group, however, can be considered a single species from a taxonomic perspective [36]. In the case of Listeria this measure of shared gene content should not be interpreted to mean that Listeria species are very closely related and may in fact comprise one species. On the contrary, the Listeria species have diverged substantially in the primary sequence of their core genes with an average pair-wise nucleotide identity of 84.8%, compared to average pair-wise nucleotide identities within species of 99.2% in F. tularensis[66] and 96.7% in E. coli[67]. Phillipy et al. [68] predicted a closed pan-genome for the species L. monocytogenes, which is congruent with our observations for the complete genus.
The mechanism behind the limited occurrence of gene acquisition from outside gene pools in Listeria remains to be determined. Although several strains harbor an insertion of prophage A118 in the comK open reading frame, which encodes a transcriptional regulator of competence, comK is intact in L. marthii, L. innocua FSL J1-023 and FSL S4-378, and L. ivanovii subsp. londoniensis, as well as the previously sequenced L. monocytogenes F2365 and HCC23 genomes. While most of the competence related genes are present in all Listeria genomes [69] and while evidence for homologous recombination has been detected by multiple studies [38, 64, 70], natural competence has not yet been report for any Listeria strains [11]. Limited natural competence may thus at least partially explain the low level of gene acquisition from outside gene pools, particularly since our data suggest that most listeriophages do are part of the closed Listeria pangenome. In addition, limited gene acquisition in the genus Listeria may also be facilitated by the presence, in all genomes of the genus examined so far, of well-developed defense system against foreign DNA/mobile elements, including R-M systems [71] and/or CRISPR systems [51]. Both systems have been shown to limit or block horizontal gene transfer in Staphylococcus aureus[72, 73]. This would explain why functional transposable elements are virtually absent from Listeria, and if present (as is the case for the conjugative elements reported here) contain a putative anti-restriction gene, which protects them from the restriction modification system.
Despite the overall high conservation of genome content across different Listeria species, gene loss and deletion events, as well as introduction of genetic material through horizontal gene transfer from other gene pools occurs in this genus, often with phenotypic consequences. For example, the chromosomal region that contains inlAB in L. monocytogenes and L. ivanovii appears to be hypervariable with evidence for deletion events (e.g., in L. seeligeri) and horizontal introduction of genetic material from other genera (e.g., the presence, in the L. ivanovii inlAB region, of two ORFs with relatively high similarity to Enterococcus genes and the presence, in the inlAB region of L. marthii, of approximately 15 ORFs that were putatively introduced by horizontal gene transfer), consistent with another report [13] that also suggested putative horizontal gene transfer events in this region.
While Listeria includes a number of species-like clades, many of these putative species include subclades or strains with atypical virulence-associated characteristics and gene profiles
Generally, within the genus Listeria, only members of the species L. monocytogenes are considered to be human pathogens, while members of the species L. ivanovii are considered to be animal pathogens [7]. Key genes that clearly contribute to virulence, as supported by experimental evidence, include (i) genes located in the prfA cluster, which are critical for intracellular survival and cell to cell spread [17], (ii) inlA and inlB, which are critical for invasion of intestinal epithelial and hepatic cells, respectively [74], and (iii) inlC, which encodes a protein that is specifically required for cell-to-cell spread [21]. Strains representing L. monocytogenes lineages I, II, and III as well as the L. ivanovii subsp. londoniensis strains contained the full complement of these virulence genes (i.e., prfA cluster, inlAB, inlC), consistent with the experimentally verified virulence of these organisms [75, 76]. Our full genome analyses suggest that the evolution from a Listeria ancestor that contained all three virulence loci yielded species and strains that have lost one or more of these key virulence genes. L. welshimeri, the majority of L. innocua strains, and non-hemolytic L. seeligeri strains lack all three of these virulence loci, consistent with their observed avirulence [22, 77].
Other strains are lacking only a subset of the key virulence genes found in most L. monocytogenes and L. ivanovii. Hemolytic L. seeligeri carries the prfA cluster, but lacks inlAB and inlC, consistent with its avirulence in mammalian tissue culture and animal models [77]. Interestingly, some strains (represented by the hemolytic L. innocua strain characterized here) contain the prfA cluster as well as inlA and have the ability to invade human intestinal epithelial cells, while lacking inlC and showing avirulence in a mouse model [22]. Similarly, at least one L. monocytogenes strain (HCC23, representing lineage IIIA) contains the prfA cluster as well as inlAB, while lacking inlC and showing avirulence in mouse infection experiments [78]. These strains represent a particular challenge for virulence classification, as they would typically be classified as virulent with standard assays (as they are hemolytic and positive for some key virulence genes). Overall, our data, along with previously reported virulence characterizations of isolates representing different Listeria species as well as atypical strains (e.g., hemolytic L. innocua) [22–24], clearly indicate the need for a well-designed molecular approach to define pathogenic strains within the genus Listeria. While we hypothesize that use of multiple marker genes, e.g., genes in the prfA cluster, inlA (including identification of virulence attenuating premature stop codons [53]), inlB, and inlC is needed to identify virulent strains, further tissue culture and animal studies are needed to confirm appropriate marker genes. In addition, further comparative genomics studies of phenotypically variable Listeria will be needed to identify and validate diagnostic targets and markers.
The genus Listeria represents two main clades that diverged from a common ancestor that contained the prfA cluster and a number of internalin genes, most likely 47 million years ago
The use of 100 core genes that have been previously shown to show no evidence for positive selection nor homologous recombination resulted in a robust phylogeny dividing Listeria (except L. grayi) into two main clades; (i) a clade consisting of L. monocytogenes, L. marthii, L. innocua and L. welshimeri, and (ii) a clade consisting of L. ivanovii and L. seeligeri. The existence of two main clades has been shown in several previous studies [3, 8, 23], however the placement of L. welshimeri has always been ambiguous. While some studies placed L. welshimeri basal in the L. seeligeri/L. ivanovii clade [3], others [8, 23], like the majority of the phylogenetic reconstruction methods used here, place L. welshimeri basal in the L. monocytogenes/L. marthii/L. innocua clade. A likely explanation for this ambiguous phylogenetic placement is the "long branch attraction effect" [79] as L. welshimeri is placed on a long branch and seems to have branched off of the MRCA of the L. monocytogenes/L. marthii/L. innocua clade relatively early during the evolutionary of Listeria sensu stricto. As likelihood-based methods are less prone to long branch attraction [79], placement, by these methods, of L. welshimeri in the L. monocytogenes/L. marthii/L. innocua clade suggests that this placement is correct.
Our data also seem to support a hypothesis that the most recent common ancestor (MRCA) of Listeria possessed not only the prfA virulence cluster as indicated before [3], but also many internalins including A, B and C, which are essential for host invasion [20] and cell-to-cell spread [21]. While a few studies [19] have previously explored the evolution of internalin multigene family, including one study [19] that proposed presence of inlB in the MRCA of Listeria (except L. grayii), our analysis allowed for identification of 16 internalin genes that, like inlB, were likely present in the MRCA of Listeria sensu stricto.
Based on a Bayesian molecular clock analysis that used 100 genes of the Listeria core genome places, we propose that the MRCA of the genus Listeria (except L. grayii) can be dated to about 40 to 60 mya, similar to the date has been inferred for the most recent ancestor of S. enterica and S. bongori[80]. A plausible hypothesis for emergence of these pathogens during this time period is that a major mammalian radiation during this same epoch [81] provided strains of Listeria and Salmonella that were able to colonize mammalian hosts with a selective advantage over less adapted or environmental strains.
Loss of virulence associated genes is a recurrent evolutionary pattern in Listeria
While a number of studies have reported that gene loss and genome reduction are general patterns in the evolutionary transition from facultative pathogenic lifestyles to obligate pathogenic lifestyles in bacteria [82], our data suggest that gene loss events in multiple genomic regions and lineages coincided with multiple evolutionary transitions of Listeria from a facultative pathogenic lifestyle to an obligate saprotrophic lifestyle. The switch from a facultative pathogen to obligate saprotrophic clades seems to have occurred at least four times during the evolutionary history of Listeria sensu stricto, including (i) during the speciation event leading to L. seeligeri, which coincided with the loss of the inlAB operon and inlC, but not the prfA cluster, (ii) the speciation event leading to L. welshimeri, which coincided with loss of the prfA cluster, the inlAB operon and inlC, (iii) the speciation event leading to L. innocua, which coincided with the loss of inlB and inlC and (iv) the speciation event leading to L. marthii, which coincided with the loss of the prfA cluster, the inlAB operon and inlC. Secondary losses of additional virulence-associated genes occurred in the non-hemolytic L. seeligeri, which lost the prfA cluster, and non-hemolytic L. innocua strains, which lost the prfA cluster as well as inlA.
Despite the observation that loss of virulence genes appears to be a key event in the evolution of Listeria species, several apparently avirulent Listeria strains (hemolytic L. seeligeri strains and hemolytic L. innocua strains) have strongly conserved, and in most cases functional, homologues of key L. monocytogenes virulence genes in their genomes. For example, previous studies [83] demonstrated some functionality of different L. seeligeri virulence factors and our data suggest that the homologue of internalin A in the hemolytic L. innocua strain supports the ability to invade human intestinal epithelial cells (even though future experiments with an isogenic inlA mutant will be required to confirm this). One hypothesis is that the virulence genes in Listeria play a role in the survival of and defense against predation by protists, however this hypothesis is not supported by a recent study that demonstrates that L. monocytogenes does not survive ingestion by the amoeba Acanthamoeba polyphaga[84].
Conclusions
In order to gain an improved understanding of genome evolution in members of the genus Listeria, with a particular attention to the evolution of virulence, we generated draft genomes for seven Listeria strains focusing on species for which genome sequences were not previously available and atypical strains of species for which genome sequences were available (i.e., L. monocytogenes lineage IIIC and hemolytic L. innocua). Analysis of 13 genome sequences representing six Listeria species (including the 7 genome sequences obtained here and 6 previously reported genome sequences) suggests that (i) the genus Listeria possesses an open pan-genome with limited ongoing introduction of new genetic material, (ii) modern pathogenic and non-pathogenic Listeria species originated, approx. 40-60 mya, from a common ancestor that contained the prfA cluster and at least 16 internalin genes, and (iii) gene loss events played a key role in the evolution of Listeria. While diversification over this time period yielded a number of species-like clades in the genus Listeria, many of these putative species include clades or strains with atypical virulence characteristics and gene profiles. This information will be critical for the development of genetic and genomic criteria for pathogenic strains, including development of assays that specifically target pathogenic Listeria strains regardless of species classification.
References
Graves LM, Helsel LO, Steigerwalt AG, Morey RE, Daneshvar MI, Roof SE, Orsi RH, Fortes ED, Millilo SR, den Bakker HC, Wiedmann M, Swaminathan B, Sauders BD: Listeria marthii sp. nov., isolated from the natural environment, Finger Lakes National Forest. Int J Syst Evol Microbiol. 2010, 60: 1280-1288. 10.1099/ijs.0.014118-0.
Leclercq A, Clermont D, Bizet C, Grimont P, Le Flèche-Matéos A, Roche S, Buchrieser C, Cadet-Daniel V, Le Monnier A, Lecuit M, Allerberger F: Listeria rocourtiae sp. nov. Int J Syst Evol Microbiol. 2009, 60: 2210-2214. 10.1099/ijs.0.017376-0.
Schmid M, Ng E, Lampidis R, Emmerth M, Walcher M, Kreft J, Goebel W, Wagner M, Schleifer K: Evolutionary history of the genus Listeria and its virulence genes. Syst Appl Microbiol. 2005, 28: 1-18. 10.1016/j.syapm.2004.09.005.
Stuart S, Welshimer H: Taxonomic reexamination of Listeria Pirie and transfer of Listeria grayi and Listeria murrayi to a new genus, Murraya. Int J Syst Evol Microbiol. 1974, 24: 177-185.
Cummins AJ, Fielding AK, McLauchlin J: Listeria ivanovii infection in a patient with AIDS. J Infect. 1994, 28: 89-91. 10.1016/S0163-4453(94)94347-8.
Lessing MP, Curtis GD, Bowler IC: Listeria ivanovii infection. J Infect. 1994, 29: 230-231. 10.1016/S0163-4453(94)90914-8.
Vázquez-Boland J, Kuhn M, Berche P, Chakraborty T, Domínguez-Bernal G, Goebel W, González-Zorn B, Wehland J, Kreft J: Listeria pathogenesis and molecular virulence determinants. Clin Microbiol Rev. 2001, 14: 584-640.
Boerlin P, Rocourt J, Piffaretti JC: Taxonomy of the genus Listeria by using multilocus enzyme electrophoresis. Int J Syst Bacteriol. 1991, 41: 59-64. 10.1099/00207713-41-1-59.
Orsi RH, den Bakker HC, Wiedmann M: Listeria monocytogenes lineages: Genomics, evolution, ecology, and phenotypic characteristics. Int J Med Microbiol. 2010
Nelson KE, Fouts DE, Mongodin EF, Ravel J, DeBoy RT, Kolonay JF, Rasko DA, Angiuoli SV, Gill SR, Paulsen IT, Peterson J, White O, Nelson WC, Nierman W, Beanan MJ, Brinkac LM, Daugherty SC, Dodson RJ, Durkin AS, Madupu R, Haft DH, Selengut J, Van Aken S, Khouri H, Fedorova N, Forberger H, Tran B, Kathariou S, Wonderling LD, Uhlich GA, et al: Whole genome comparisons of serotype 4b and 1/2a strains of the food-borne pathogen Listeria monocytogenes reveal new insights into the core genome components of this species. Nucleic Acids Res. 2004, 32: 2386-2395. 10.1093/nar/gkh562.
Glaser P, Frangeul L, Buchrieser C, Rusniok C, Amend A, Baquero F, Berche P, Bloecker H, Brandt P, Chakraborty T, Charbit A, Chetouani F, Couve E, de Daruvar A, Dehoux P, Domann E, Dominguez-Bernal G, Duchaud E, Durant L, Dussurget O, Entian KD, Fsihi H, Garcia-del Portillo F, Garrido P, Gautier L, Goebel W, Gomez-Lopez N, Hain T, Hauf J, Jackson D, et al: Comparative genomics of Listeria species. Science. 2001, 294: 849-852.
Gilmour MW, Graham M, Van Domselaar G, Tyler S, Kent H, Trout-Yakel KM, Larios O, Allen V, Lee B, Nadon C: High-throughput genome sequencing of two Listeria monocytogenes clinical isolates during a large foodborne outbreak. BMC Genomics. 2010, 11: 120-
Hain T, Steinweg C, Kuenne CT, Billion A, Ghai R, Chatterjee SS, Domann E, Karst U, Goesmann A, Bekel T, Bartels D, Kaiser O, Meyer F, Puhler A, Weisshaar B, Wehland J, Liang C, Dandekar T, Lampidis R, Kreft J, Goebel W, Chakraborty T: Whole-genome sequence of Listeria welshimeri reveals common steps in genome reduction with Listeria innocua as compared to Listeria monocytogenes. J Bacteriol. 2006, 188: 7405-7415. 10.1128/JB.00758-06.
Steinweg C, Kuenne CT, Billion A, Mraheil MA, Domann E, Ghai R, Barbuddhe SB, Kärst U, Goesmann A, Pühler A, Weisshaar B, Wehland J, Lampidis R, Kreft J, Goebel W, Chakraborty T, Hain T: Complete genome sequence of Listeria seeligeri, a nonpathogenic member of the genus Listeria. J Bacteriol. 2010, 192: 1473-1474. 10.1128/JB.01415-09.
Pallen MJ, Wren BW: Bacterial pathogenomics. Nature. 2007, 449: 835-842. 10.1038/nature06248.
Vazquez-Boland JA, Dominguez-Bernal G, Gonzalez-Zorn B, Kreft J, Goebel W: Pathogenicity islands and virulence evolution in Listeria. Microbes Infect. 2001, 3: 571-584. 10.1016/S1286-4579(01)01413-7.
Freitag NE, Port GC, Miner MD: Listeria monocytogenes - from saprophyte to intracellular pathogen. Nat Rev Microbiol. 2009, 7: 623-628. 10.1038/nrmicro2171.
Hain T, Chatterjee SS, Ghai R, Kuenne CT, Billion A, Steinweg C, Domann E, Karst U, Jansch L, Wehland J, Eisenreich W, Bacher A, Joseph B, Schar J, Kreft J, Klumpp J, Loessner MJ, Dorscht J, Neuhaus K, Fuchs TM, Scherer S, Doumith M, Jacquet C, Martin P, Cossart P, Rusniock C, Glaser P, Buchrieser C, Goebel W, Chakraborty T: Pathogenomics of Listeria spp. Int J Med Microbiol. 2007, 297: 541-557. 10.1016/j.ijmm.2007.03.016.
Rooney AP, Ward TJ: Birth-and-death evolution of the internalin multigene family in Listeria. Gene. 2008, 427: 124-128. 10.1016/j.gene.2008.09.007.
Bierne H, Sabet C, Personnic N, Cossart P: Internalins: a complex family of leucine-rich repeat-containing proteins in Listeria monocytogenes. Microbes Infect. 2007, 9: 1156-1166. 10.1016/j.micinf.2007.05.003.
Rajabian T, Gavicherla B, Heisig M, Müller-Altrock S, Goebel W, Gray-Owen S, Ireton K: The bacterial virulence factor InlC perturbs apical cell junctions and promotes cell-to-cell spread of Listeria. Nat Cell Biol. 2009, 11: 1212-1218. 10.1038/ncb1964.
Johnson J, Jinneman K, Stelma G, Smith BG, Lye D, Messer J, Ulaszek J, Evsen L, Gendel S, Bennett RW, Swaminathan B, Pruckler J, Steigerwalt A, Kathariou S, Yildirim S, Volokhov D, Rasooly A, Chizhikov V, Wiedmann M, Fortes E, Duvall RE, Hitchins AD: Natural atypical Listeria innocua strains with Listeria monocytogenes pathogenicity island 1 genes. Appl Environ Microbiol. 2004, 70: 4256-4266. 10.1128/AEM.70.7.4256-4266.2004.
Volokhov D, Duperrier S, Neverov A, George J, Buchrieser C, Hitchins A: The presence of the internalin gene in natural atypically hemolytic Listeria innocua strains suggests descent from L. monocytogenes. Appl Environ Microbiol. 2007, 73: 1928-1939. 10.1128/AEM.01796-06.
Volokhov D, George J, Anderson C, Duvall R, Hitchins A: Discovery of natural atypical nonhemolytic Listeria seeligeri isolates. Appl Environ Microbiol. 2006, 72: 2439-2448. 10.1128/AEM.72.4.2439-2448.2006.
Gouin E, Mengaud J, Cossart P: The virulence gene cluster of Listeria monocytogenes is also present in Listeria ivanovii, an animal pathogen, and Listeria seeligeri, a nonpathogenic species. Infect Immun. 1994, 62: 3550-3553.
den Bakker HC, Bundrant BN, Fortes ED, Orsi RH, Wiedmann M: A population genetics-based and phylogenetic approach to understanding the evolution of virulence in the genus Listeria. Appl Environ Microbiol. 2010, 76: 6085-6100. 10.1128/AEM.00447-10.
Chaisson MJ, Brinza D, Pevzner PA: De novo fragment assembly with short mate-paired reads: Does the read length matter?. Genome Res. 2009, 19: 336-346. 10.1101/gr.079053.108.
Zerbino DR, Birney E: Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008, 18: 821-829. 10.1101/gr.074492.107.
Kurtz S, Phillippy A, Delcher AL, Smoot M, Shumway M, Antonescu C, Salzberg SL: Versatile and open software for comparing large genomes. Genome Biol. 2004, 5: R12-10.1186/gb-2004-5-2-r12.
Bateman A, Birney E, Durbin R, Eddy SR, Howe KL, Sonnhammer EL: The Pfam protein families database. Nucleic Acids Res. 2000, 28: 263-266. 10.1093/nar/28.1.263.
Haft DH, Loftus BJ, Richardson DL, Yang F, Eisen JA, Paulsen IT, White O: TIGRFAMs: a protein family resource for the functional identification of proteins. Nucleic Acids Res. 2001, 29: 41-43. 10.1093/nar/29.1.41.
Darling ACE, Mau B, Blattner FR, Perna NT: Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 2004, 14: 1394-1403. 10.1101/gr.2289704.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ: Basic local alignment search tool. J Mol Biol. 1990, 215: 403-410.
Snipen L, Almøy T, Ussery DW: Microbial comparative pan-genomics using binomial mixture models. BMC Genomics. 2009, 10: 385-10.1186/1471-2164-10-385.
Tettelin H, Masignani V, Cieslewicz MJ, Donati C, Medini D, Ward NL, Angiuoli SV, Crabtree J, Jones AL, Durkin AS, DeBoy RT, Davidsen TM, Mora M, Scarselli M, Margarit y, Ros I, Peterson JD, Hauser CR, Sundaram JP, Nelson WC, Madupu R, Brinkac LM, Dodson RJ, Rosovitz MJ, Sullivan SA, Daugherty SC, Haft DH, Selengut J, Gwinn ML, Zhou L, Zafar N, et al: Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial "pan-genome". Proc Natl Acad Sci USA. 2005, 102: 13950-13955. 10.1073/pnas.0506758102.
Didelot X, Barker M, Falush D, Priest FG: Evolution of pathogenicity in the Bacillus cereus group. Syst Appl Microbiol. 2009, 32: 81-90. 10.1016/j.syapm.2009.01.001.
Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talón M, Dopazo J, Conesa A: High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36: 3420-3435.
Orsi RH, Sun Q, Wiedmann M: Genome-wide analyses reveal lineage specific contributions of positive selection and recombination to the evolution of Listeria monocytogenes. BMC Evol Biol. 2008, 8: 233-10.1186/1471-2148-8-233.
Kumar S, Nei M, Dudley J, Tamura K: MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief Bioinformatics. 2008, 9: 299-306. 10.1093/bib/bbn017.
Ronquist F, Huelsenbeck JP: MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003, 19: 1572-1574. 10.1093/bioinformatics/btg180.
Swofford DL: PAUP*-phylogenetic analysis using parsimony (* and other methods), version 4.0. Book PAUP*-phylogenetic analysis using parsimony (* and other methods), version 4.0. 2002, City: Sinauer Associates Inc, (Editor ed.^eds.)
Drummond AJ, Rambaut A: BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol. 2007, 7: 214-10.1186/1471-2148-7-214.
Ochman H, Elwyn S, Moran NA: Calibrating bacterial evolution. Proc Natl Acad Sci USA. 1999, 96: 12638-12643. 10.1073/pnas.96.22.12638.
Katoh K, Toh H: Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinformatics. 2008, 9: 286-298. 10.1093/bib/bbn013.
Camejo A, Buchrieser C, Couvé E, Carvalho F, Reis O, Ferreira P, Sousa S, Cossart P, Cabanes D: In vivo transcriptional profiling of Listeria monocytogenes and mutagenesis identify new virulence factors involved in infection. PLoS Pathog. 2009, 5: e1000449-10.1371/journal.ppat.1000449.
Suzuki R, Shimodaira H: Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics. 2006, 22: 1540-1542. 10.1093/bioinformatics/btl117.
Waack S, Keller O, Asper R, Brodag T, Damm C, Fricke WF, Surovcik K, Meinicke P, Merkl R: Score-based prediction of genomic islands in prokaryotic genomes using hidden Markov models. BMC Bioinformatics. 2006, 7: 142-10.1186/1471-2105-7-142.
Lima-Mendez G, Van Helden J, Toussaint A, Leplae R: Prophinder: a computational tool for prophage prediction in prokaryotic genomes. Bioinformatics. 2008, 24: 863-865. 10.1093/bioinformatics/btn043.
Leplae R, Hebrant A, Wodak SJ, Toussaint A: ACLAME: a CLAssification of Mobile genetic Elements. Nucleic Acids Res. 2004, 32: D45-49. 10.1093/nar/gkh084.
Wilson GG, Murray NE: Restriction and modification systems. Annu Rev Genet. 1991, 25: 585-627. 10.1146/annurev.ge.25.120191.003101.
Horvath P, Barrangou R: CRISPR/Cas, the immune system of bacteria and archaea. Science. 2010, 327: 167-170. 10.1126/science.1179555.
Grissa I, Vergnaud G, Pourcel C: CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res. 2007, 35: W52-57. 10.1093/nar/gkm360.
Nightingale KK, Windham K, Martin KE, Yeung M, Wiedmann M: Select Listeria monocytogenes subtypes commonly found in foods carry distinct nonsense mutations in inlA, leading to expression of truncated and secreted internalin A, and are associated with a reduced invasion phenotype for human intestinal epithelial cells. Appl Environ Microbiol. 2005, 71: 8764-8772. 10.1128/AEM.71.12.8764-8772.2005.
Orsi RH, Borowsky ML, Lauer P, Young SK, Nusbaum C, Galagan JE, Birren BW, Ivy RA, Sun Q, Graves LM, Swaminathan B, Wiedmann M: Short-term genome evolution of Listeria monocytogenes in a non-controlled environment. BMC Genomics. 2008, 9: 539-10.1186/1471-2164-9-539.
Read TD, Salzberg SL, Pop M, Shumway M, Umayam L, Jiang L, Holtzapple E, Busch JD, Smith KL, Schupp JM, Solomon D, Keim P, Fraser CM: Comparative genome sequencing for discovery of novel polymorphisms in Bacillus anthracis. Science. 2002, 296: 2028-2033. 10.1126/science.1071837.
Chain PSG, Grafham DV, Fulton RS, Fitzgerald MG, Hostetler J, Muzny D, Ali J, Birren B, Bruce DC, Buhay C, Cole JR, Ding Y, Dugan S, Field D, Garrity GM, Gibbs R, Graves T, Han CS, Harrison SH, Highlander S, Hugenholtz P, Khouri HM, Kodira CD, Kolker E, Kyrpides NC, Lang D, Lapidus A, Malfatti SA, Markowitz V, Metha T, et al: Genomics. Genome project standards in a new era of sequencing. Science. 2009, 326: 236-237. 10.1126/science.1180614.
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O: The RAST Server: rapid annotations using subsystems technology. BMC Genomics. 2008, 9: 75-10.1186/1471-2164-9-75.
Lebrun M, Audurier A, Cossart P: Plasmid-borne cadmium resistance genes in Listeria monocytogenes are similar to cadA and cadC of Staphylococcus aureus and are induced by cadmium. J Bacteriol. 1994, 176: 3040-3048.
Sebaihia M, Wren BW, Mullany P, Fairweather NF, Minton N, Stabler R, Thomson NR, Roberts AP, Cerdeño-Tárraga AM, Wang H, Holden MTG, Wright A, Churcher C, Quail MA, Baker S, Bason N, Brooks K, Chillingworth T, Cronin A, Davis P, Dowd L, Fraser A, Feltwell T, Hance Z, Holroyd S, Jagels K, Moule S, Mungall K, Price C, Rabbinowitsch E, et al: The multidrug-resistant human pathogen Clostridium difficile has a highly mobile, mosaic genome. Nat Genet. 2006, 38: 779-786. 10.1038/ng1830.
Ochman H, Wilson AC: Evolution in bacteria: evidence for a universal substitution rate in cellular genomes. J Mol Evol. 1987, 26: 74-86. 10.1007/BF02111283.
Mcpherson JD: Next-generation gap. Nat Meth. 2009, 6: S2-5. 10.1038/nmeth.f.268.
Cummings CA, Chung CAB, Fang R, Barker M, Brzoska PM, Williamson P, Beaudry JA, Matthews M, Schupp JM, Wagner DM, Furtado MR, Keim P, Budowle B: Whole-genome typing of Bacillus anthracis isolates by next-generation sequencing accurately and rapidly identifies strain-specific diagnostic polymorphisms. Forensic Science International: Genetics Supplement Series. 2009, 1-2.
Greub G, Kebbi-Beghdadi C, Bertelli C, Collyn F, Riederer BM, Yersin C, Croxatto A, Raoult D: High throughput sequencing and proteomics to identify immunogenic proteins of a new pathogen: the dirty genome approach. PLoS ONE. 2009, 4: e8423-10.1371/journal.pone.0008423.
Dunn KA, Bielawski JP, Ward TJ, Urquhart C, Gu H: Reconciling ecological and genomic divergence among lineages of Listeria under an "extended mosaic genome concept". Mol Biol Evol. 2009, 26: 2605-2615. 10.1093/molbev/msp176.
Dorscht J, Klumpp J, Bielmann R, Schmelcher M, Born Y, Zimmer M, Calendar R, Loessner MJ: Comparative genome analysis of Listeria bacteriophages reveals extensive mosaicism, programmed translational frameshifting, and a novel prophage insertion site. J Bacteriol. 2009, 191: 7206-7215. 10.1128/JB.01041-09.
Larsson P, Elfsmark D, Svensson K, Wikström P, Forsman M, Brettin T, Keim P, Johansson A: Molecular evolutionary consequences of niche restriction in Francisella tularensis, a facultative intracellular pathogen. PLoS Pathog. 2009, 5: e1000472-10.1371/journal.ppat.1000472.
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM: DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol. 2007, 57: 81-91. 10.1099/ijs.0.64483-0.
Phillippy AM, Deng X, Zhang W, Salzberg SL: Efficient oligonucleotide probe selection for pan-genomic tiling arrays. BMC Bioinformatics. 2009, 10: 293-10.1186/1471-2105-10-293.
Buchrieser C: Biodiversity of the species Listeria monocytogenes and the genus Listeria. Microbes Infect. 2007, 9: 1147-1155. 10.1016/j.micinf.2007.05.002.
Nightingale K, Windham K, Wiedmann M: Evolution and molecular phylogeny of Listeria monocytogenes isolated from human and animal listeriosis cases and foods. J Bacteriol. 2005, 187: 5537-5551. 10.1128/JB.187.16.5537-5551.2005.
Tock MR, Dryden DTF: The biology of restriction and anti-restriction. Curr Opin Microbiol. 2005, 8: 466-472. 10.1016/j.mib.2005.06.003.
Waldron DE, Lindsay JA: Sau1: a novel lineage-specific type I restriction-modification system that blocks horizontal gene transfer into Staphylococcus aureus and between S. aureus isolates of different lineages. J Bacteriol. 2006, 188: 5578-5585. 10.1128/JB.00418-06.
Marraffini LA, Sontheimer EJ: CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science. 2008, 322: 1843-1845. 10.1126/science.1165771.
Braun L, Dramsi S, Dehoux P, Bierne H, Lindahl G, Cossart P: InlB: an invasion protein of Listeria monocytogenes with a novel type of surface association. Mol Microbiol. 1997, 25: 285-294. 10.1046/j.1365-2958.1997.4621825.x.
González-Zorn B, Domínguez-Bernal G, Suárez M, Ripio MT, Vega Y, Novella S, Vazquez-Boland JA: The smcL gene of Listeria ivanovii encodes a sphingomyelinase C that mediates bacterial escape from the phagocytic vacuole. Mol Microbiol. 1999, 33: 510-523.
Liu D, Ainsworth AJ, Austin FW, Lawrence ML: Characterization of virulent and avirulent Listeria monocytogenes strains by PCR amplification of putative transcriptional regulator and internalin genes. J Med Microbiol. 2003, 52: 1065-1070. 10.1099/jmm.0.05358-0.
Rocourt J, Grimont P: Listeria welshimeri sp nov and Listeria seeligeri sp. nov. Int J Syst Bacteriol. 1983, 33: 866-869. 10.1099/00207713-33-4-866.
Erdenlig S, Ainsworth AJ, Austin FW: Pathogenicity and production of virulence factors by Listeria monocytogenes isolates from channel catfish. J Food Protect. 2000, 63: 613-619.
Felsenstein J: Inferring Phylogenies. 2004, Sunderland, Massachusetts: Sinauer Associates, Inc
McQuiston JR, Herrera-Leon S, Wertheim BC, Doyle J, Fields PI, Tauxe RV, Logsdon JM: Molecular phylogeny of the salmonellae: relationships among Salmonella species and subspecies determined from four housekeeping genes and evidence of lateral gene transfer events. J Bacteriol. 2008, 190: 7060-7067. 10.1128/JB.01552-07.
Bininda-Emonds ORP, Cardillo M, Jones KE, MacPhee RDE, Beck RMD, Grenyer R, Price SA, Vos RA, Gittleman JL, Purvis A: The delayed rise of present-day mammals. Nature. 2007, 446: 507-512. 10.1038/nature05634.
Moran NA: Microbial minimalism: genome reduction in bacterial pathogens. Cell. 2002, 108: 583-586. 10.1016/S0092-8674(02)00665-7.
Karunasagar I, Lampidis R, Goebel W, Kreft J: Complementation of Listeria seeligeri with the plcA-prfA genes from L. monocytogenes activates transcription of seeligerolysin and leads to bacterial escape from the phagosome of infected mammalian cells. FEMS Microbiol Lett. 1997, 146: 303-310. 10.1111/j.1574-6968.1997.tb10209.x.
Akya A, Pointon A, Thomas C: Listeria monocytogenes does not survive ingestion by Acanthamoeba polyphaga. Microbiology. 2009,
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA: Circos: an information aesthetic for comparative genomics. Genome Res. 2009, 19: 1639-1645. 10.1101/gr.092759.109.
Machata S, Tchatalbachev S, Mohamed W, Jänsch L, Hain T, Chakraborty T: Lipoproteins of Listeria monocytogenes are critical for virulence and TLR2-mediated immune activation. J Immunol. 2008, 181: 2028-2035.
Faith N, Uhlich G, Luchansky JB, Neudeck B, Czuprynski C: A prfA transposon mutant of Listeria monocytogenes F2365, a serotype 4b strain, is able to survive in the gastrointestinal tract but does not cause systemic infection of the spleens and livers of intragastrically inoculated mice. Infect Immun. 2005, 73: 7517-7524. 10.1128/IAI.73.11.7517-7524.2005.
Acknowledgements
We would like to thank the JCVI for providing the JCVI Annotation Service, which provided us with automatic annotation data and the manual annotation tool Manatee; Barbara Bowen and Esther Fortes for their help with additional Sanger sequencing and handling of the strains; Dima Brinza for sharing de novo assembly tools prior to distribution; and Dr Joseph Peters for his helpful discussion on the possible mechanisms of deletion of the prfA cluster. Part of this work was supported by USDA Special Research Grants (2005-34459-15625, 2006-34459-16952, and 2008-34459-19043).
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
Life Technologies Corporation partially funded this study by providing sequencing reagents and instruments, and by compensating its employees (CAC, PV, LD, MB, OP, and MRF), who participated in study design, data collection and analysis, decision to publish, and preparation of the manuscript. Life Technologies Corporation also financially compensated HDB and MW for travel expenses made for a visit to Foster City.
Authors' contributions
HDB, CAC, MW, PV, OP and MRF conceived the study. PV, LD, and MB performed the genome sequencing. HDB, CAC and PV performed the genome sequence analysis. RHO helped with the genome analysis and the SNP verification. VF performed the invasion assays. HDB, CAC, and MW wrote the paper. All authors read and approved the final manuscript.
Henk C den Bakker, Craig A Cummings contributed equally to this work.
Electronic supplementary material
12864_2010_10139_MOESM1_ESM.PDF
Additional file 1:Primer sequences of primers used to confirm absence of specific virulence associated genes. (PDF 54 KB)
12864_2010_10139_MOESM2_ESM.XLS
Additional file 2:Excel file SNP differences between L. monocytogenes F2365 and its duplicate FSL R2-574 and results of Sanger sequencing SNP confirmation. (XLS 18 KB)
12864_2010_10139_MOESM5_ESM.PDF
Additional file 5:PDF file containing a graphic comparison of internalin GHE region. Gray arrows indicate conserved genes adjacent to the region. (PDF 318 KB)
12864_2010_10139_MOESM7_ESM.XLS
Additional file 7:Table in MS Excel with presence or absence of 78 virulence associated genes in the genomes of the genus Listeria. (XLS 43 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
den Bakker, H.C., Cummings, C.A., Ferreira, V. et al. Comparative genomics of the bacterial genus Listeria: Genome evolution is characterized by limited gene acquisition and limited gene loss. BMC Genomics 11, 688 (2010). https://doi.org/10.1186/1471-2164-11-688
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2164-11-688